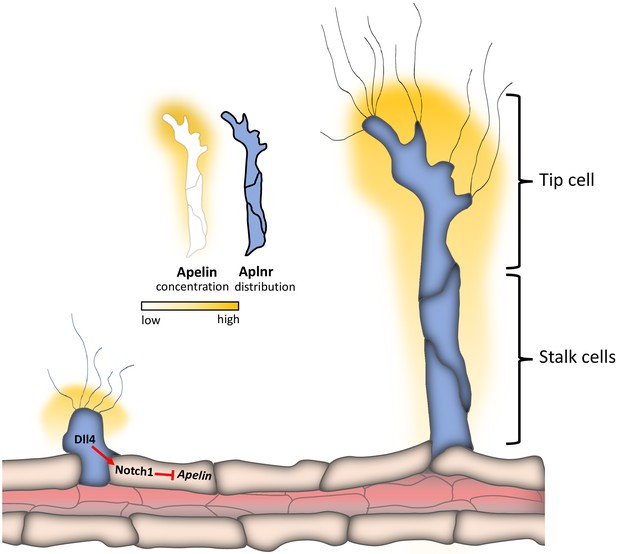
Apelin signaling drives vascular endothelial cells toward a pro-angiogenic state
Figures
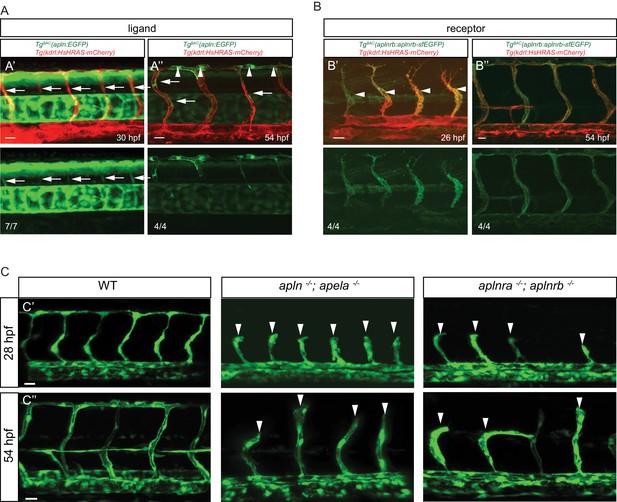
Apelin signaling promotes endothelial sprouting.
Visualization of apelin and apelin receptor b expression using transgenic reporter lines. Confocal projection images of the trunk region of zebrafish embryos. (A) TgBAC(apln:EGFP) expression is detectable in growing ISVs at 30 and 54 hpf. Arrowheads point to strong apln expression in tip cells, while arrows point to weak apln expression in stalk cells. (B) TgBAC(aplnrb:aplnrb-EGFP) expression is detectable in sprouting ECs (arrowheads) at 26 hpf and is clearly present in the ISVs and DLAV at 54 hpf. (C) Inactivation of Apelin ligand and receptor genes impairs angiogenesis. Confocal projection images of the blood vasculature in the trunk region of Tg(fli1a:EGFP) embryos. apln -/-; apela -/- as well as aplnra -/-; aplnrb -/- embryos exhibit a reduction in vascular sprouting at 28 and 54 hpf. Arrowheads point to stalled ISVs. Scale bars: A’, 10 µm; A’’, B’, C’, C’’, 20 µm; B’’, 15 µm.
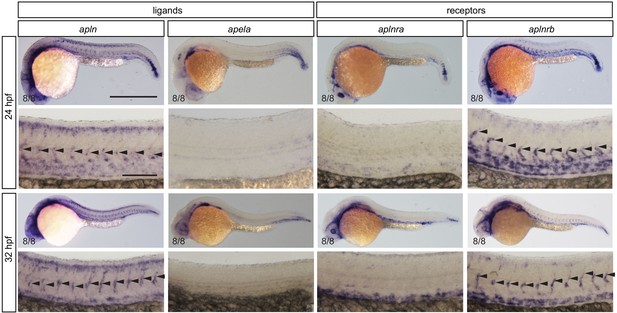
Expression of apelin ligand and receptor genes by in situ hybridization.
Expression of apelin ligand and receptor genes at 24 and 32 hpf. apln is expressed in ISVs (arrowheads). apela is not detected in blood vessels. aplnra is expressed in the posterior cardinal vein. aplnrb is expressed in the posterior cardinal vein and ISVs (arrowheads). Scale bars: 500 µm; insets, 100 µm.
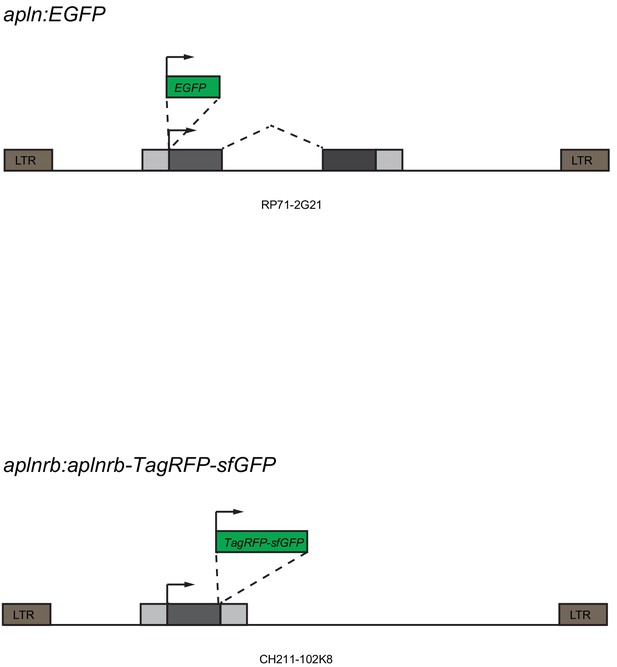
Generation of the TgBAC(apln:EGFP) and Tg(aplnrb:aplnrb-TagRFP-sfGFP) reporter lines.
BAC engineering was used to generate the apln:EGFP and aplnrb:aplnrb-TagRFP-sfGFP constructs. The clone RP71-2G21 was engineered to replace the ATG of apln with an EGFP cassette. The clone CH211-102K8 was engineered to replace the stop codon of aplnrb with a TagRFP-sfGFP cassette. To enhance the frequency of insertion into the genome, tol2 arms were added to the backbone of the BAC.
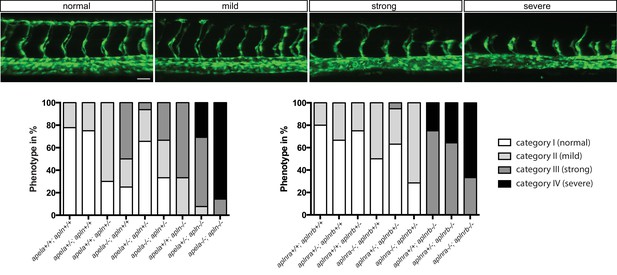
Quantification of angiogenic defects.
Confocal projection images of the trunk region of Tg(fli1a:EGFP) embryos at 30 hpf. Analysis of the offspring of apln +/-; apela+/- (n = 85) and aplnra +/-; aplnrb +/- (n = 71) fish resulted in different phenotypic categories according to the extent of ISV development. Genotyping of single embryos revealed an additive effect for apln and apela ligands and aplnra and aplnrb receptors. Mild phenotypes correlated with loss of single ligand or receptor genes, while deficiency in apln resulted in stronger phenotypes than deficiency in apela. aplnrb deficiency resulted in stronger phenotypes than deficiency in aplnra. Deficiencies in both ligands or both receptors resulted in more severe phenotypes. Scale bar: 40 µm.
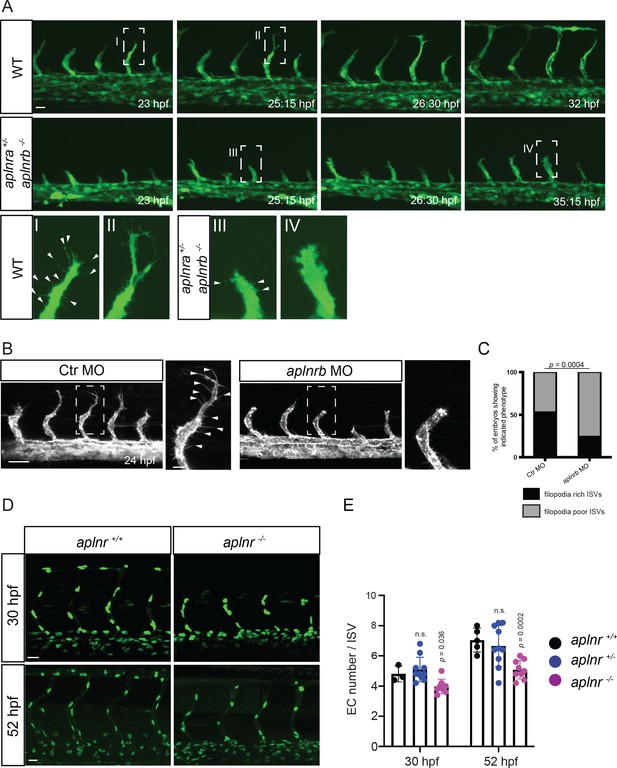
Apelin signaling regulates endothelial filopodia formation and endothelial cell numbers.
(A) Still images from confocal time-lapse movies of vascular development in wild-type and aplnra +/-; aplnrb -/- embryos. During sprouting, wild-type tip cells send out filopodia (arrowheads). aplnra +/-; aplnrb -/- embryos exhibit smaller sprouts and fail to form filopodia. (B) Confocal images of the blood vasculature in 24 hpf Tg(kdrl:HsHRAS-EGFP) embryos injected with Ctr MO and aplnrb MO. aplnrb morphant embryos exhibit smaller sprouts and fail to form filopodia (arrowheads). (C) aplnrb morphant embryos exhibit a reduction in the number of endothelial filopodia (Ctr MO, n = 10; aplnrb MO, n = 15). (D) Confocal images of the blood vasculature of 30 and 52 hpf Tg(fli1a:nEGFP) wild-type and aplnra +/-; aplnrb -/- embryos showing EC cell nuclei. (E) aplnra +/-; aplnrb -/- embryos exhibit reduced EC numbers in the ISVs (30 hpf: aplnr +/+, n = 3; aplnr +/-, n = 10; aplnr -/-, n = 8; 52 hpf: aplnr +/+, n = 5; aplnr +/-, n = 10; aplnr -/-, n = 9). n.s. not significant (two-tailed t-test). Scale bars: A, D, 20 µm; B, 40 µm; B, inset 10 µm.
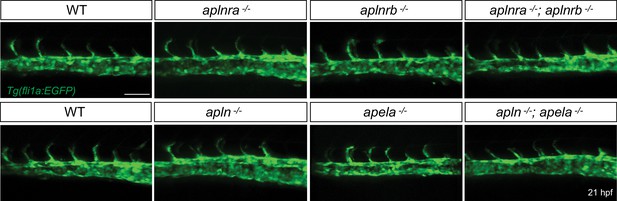
No obvious defects during initiation of EC sprouting in Apelin signaling-deficient embryos.
Confocal projection images of the trunk region of Tg(fli1a:EGFP) embryos at 21 hpf. Sprouting of ECs from the DA does not appear to be altered in Apelin signaling-deficient embryos. Scale bar: 50 µm.
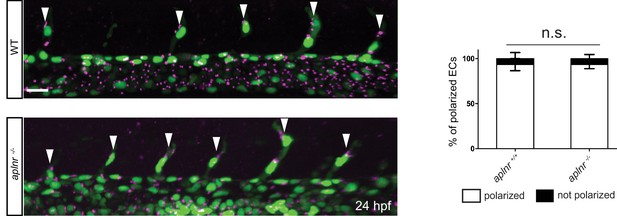
No obvious defects in EC polarity in Apelin deficient embryos.
Confocal projection images of the trunk region of Tg(kdrl:NLS-EGFP); Tg(fli1a:B4GALT1-mCherry) embryos at 24 hpf. Apelin deficiency does not cause obvious defects in EC polarity (aplnr +/+, n = 6; aplnr -/-, n = 8). Arrowheads point to polarized tip cells in the ISVs. Scale bar: 20 µm.
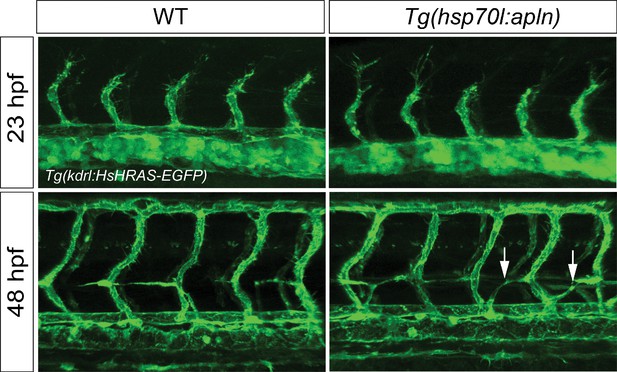
Overexpression of apln does not cause ectopic EC sprouting.
Confocal projection images of the trunk region of Tg(fli1a:LIFEACT-GFP) embryos at 23 and 48 hpf. apln expression was induced by a single heatshock at 19.5 hpf for 1 hr at 39°C and analyzed at 23 hpf or heatshocked at 42 and 45 hpf for 1 hr at 39°C and analyzed at 48 hpf. Global overexpression of apln does not lead to ectopic vascular sprouting but mispatterning of the string of parachordal lymphangioblasts (arrows).
Confocal time-lapse-imaging of a Tg(fli1a:EGFP) wild-type embryo from 23 to 32 hpf.
During sprouting, wild-type tip cells send out filopodia. Related to Figure 2A.
Confocal time-lapse-imaging of a Tg(fli1a:EGFP) aplnra +/-; aplnrb -/- embryos from 23 to 35 hpf.
During sprouting, aplnra +/-; aplnrb -/- embryos exhibit smaller sprouts and fail to form filopodia. Related to Figure 2A.
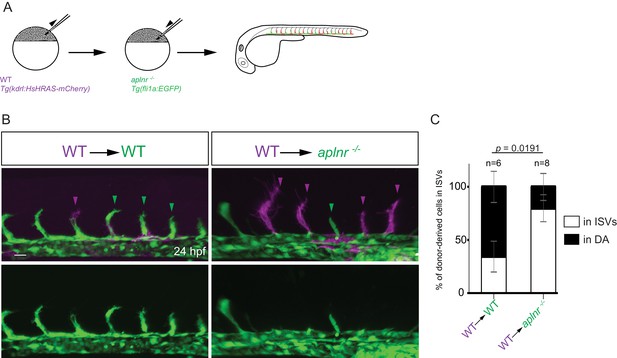
Apelin signaling promotes the sprouting behavior of ECs.
(A) Experimental design: At the blastula stage, cells from Tg(kdrl:HsHRAS-mCherry) embryos were transplanted into host embryos obtained from Tg(fli1a:EGFP) aplnra +/-; aplnrb +/- incrosses. At 24 hpf, the mosaic embryos were imaged and the donor-derived ECs scored for their position. (B, C) 34,5% of wild-type donor-derived ECs in wild-type hosts were found within the ISVs. 80% of wild-type donor-derived ECs in aplnra +/-; aplnrb -/- hosts were found within the ISVs. Notably, wild-type ECs transplanted into aplnr- deficient embryos completely substituted for the lack of cells in the dorsal part of the vasculature at 54 hpf (Figure 3—figure supplement 2). Scale bars: B, 20 µm.
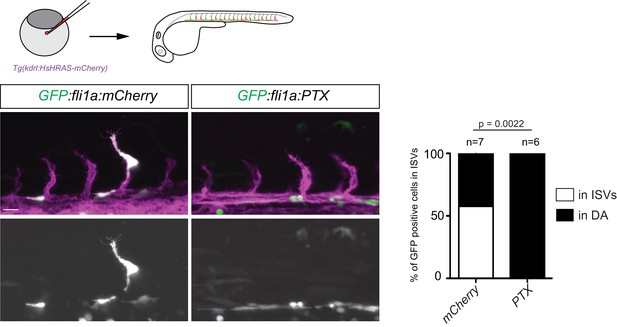
Overexpression of PTX phenocopies loss of Apelin signaling.
Confocal projection images of the trunk region of Tg(kdrl:HsHRAS-mCherry) embryos at 24 hpf. PTX was overexpressed by injecting a bidirectional fli1a promoter driving PTX and GFP at the same time. ECs expressing PTX were detected only in the DA while mCherry expressing control ECs were also detected in the ISVs. Scale bar: 20 µm.
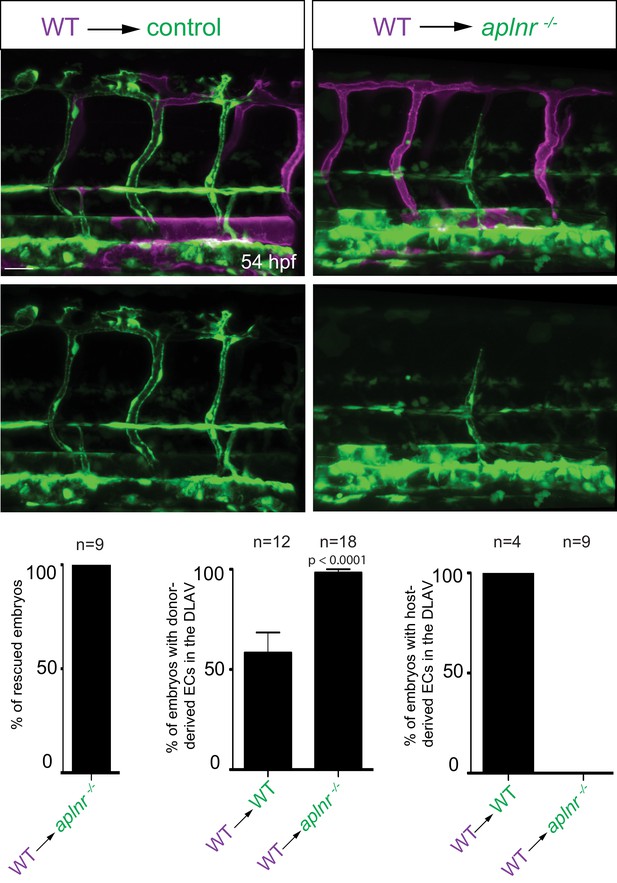
Apelin signaling functions cell-autonomously in ECs.
Confocal projection images of the trunk region of Tg(fli1a:EGFP) embryos at 54 hpf. At the blastula stage, cells from Tg(kdrl:HsHRAS-mCherry) embryos were transplanted into host embryos obtained from Tg(fli1a:EGFP) aplnra +/-; aplnrb +/- incrosses. At 54 hpf, the mosaic embryos were imaged and the donor-derived ECs scored for their position. Notably, wild-type ECs transplanted into aplnr deficient embryos completely substituted for the lack of cells in the dorsal part of the vasculature at 54 hpf (Figure 3—figure supplement 1). Scale bar: 30 µm.
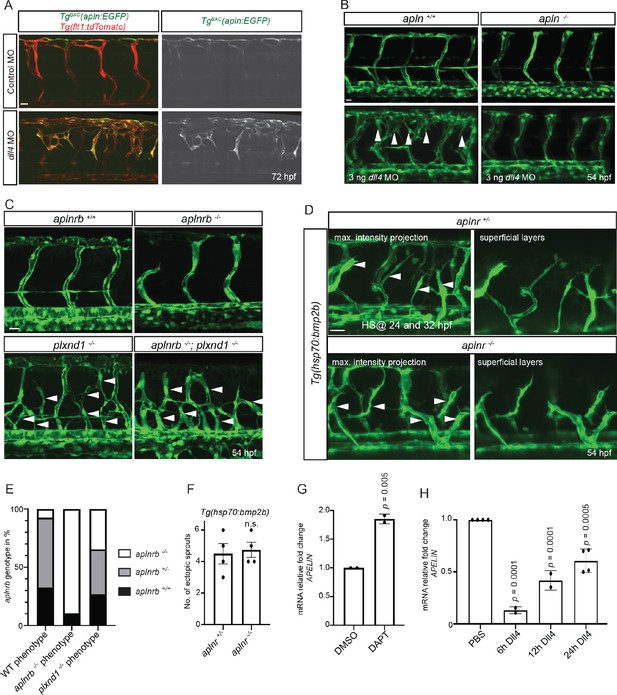
Apelin signaling functions downstream of Notch signaling in endothelial cells.
(A - D) Confocal projection images of the blood vasculature in the trunk region of Tg(flt1:tdTomato) (A) and Tg(fli1a:EGFP) (B–D) animals at 54 (B–D) and 72 (A) hpf. (A) Injection of a dll4 morpholino leads to an increase in TgBAC(apln:EGFP) expression. (B) Loss of Apelin function can block excessive endothelial sprouting in dll4 morphants. (C, E) Angiogenic response in aplnrb -/-, plxnd1 -/-, and aplnrb -/-; plxnd1 -/- embryos (arrowheads) (n = 95). (D, F) Angiogenic response to bmp2b overexpression in aplnr +/- and aplnr -/- embryos (arrowheads). (E) Genotype of embryos for aplnrb after sorting them according to phenotype. (G) RT-qPCR analysis of APELIN mRNA levels in HUVECs treated with DAPT for 24 hr. Blocking Notch signaling with DAPT induces APELIN expression. (H) RT-qPCR analysis of APELIN mRNA levels in HUVECs cultured on DLL4 to activate Notch signaling. Activating Notch signaling represses APELIN expression. Arrowheads point to ectopic sprouts. n.s. not significant (two-tailed t-test). Ct values can be found in Figure 4—source data 1. Scale bars: A, C, 20 µm; B, 15 µm; D, 30 µm.
-
Figure 4—source data 1
Ct values of RT-qPCR.
- https://cdn.elifesciences.org/articles/55589/elife-55589-fig4-data1-v2.docx
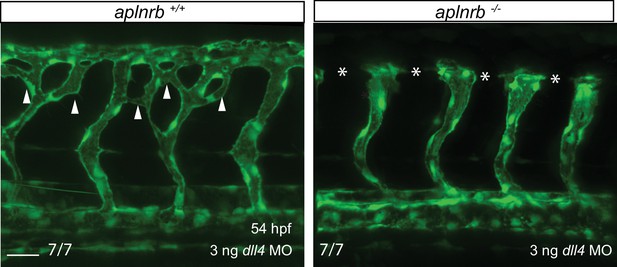
Apelin signaling functions downstream of Notch signaling in ECs.
Confocal projection images of the trunk region of Tg(fli1a:EGFP) embryos at 54 hpf. Loss of aplnrb function can rescue excessive endothelial sprouting in dll4 morphants. Arrowheads point to ectopic sprouts and asterisks indicate missing DLAV fragments. Scale bar: 30 µm.
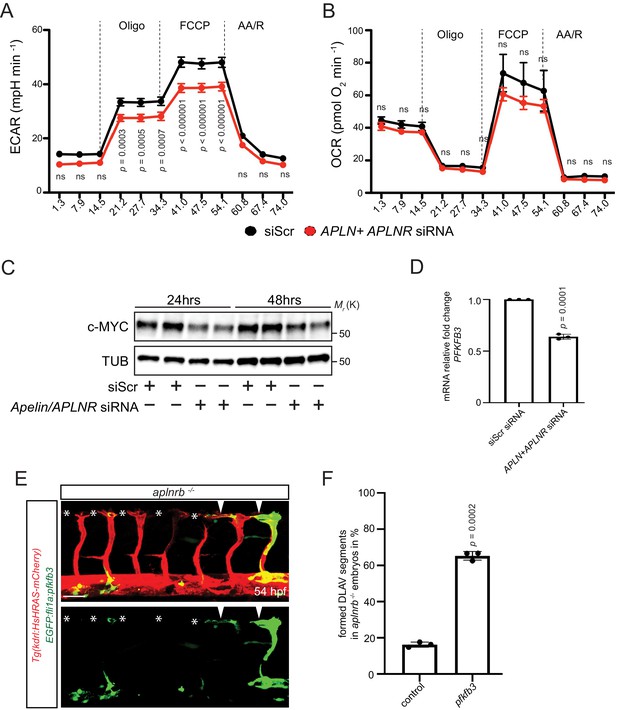
Apelin signaling positively regulates EC metabolism.
(A - B) Extracellular acidification aate (ECAR) (A) and oxygen consumption rates (OCR) (B) in siScr and APLN+APLNR siRNA-treated HUVECs under basal conditions and in response to oligomycin, fluoro-carbonyl cyanide phenylhydrazone (FCCP) and antimycin A (AA)/rotenone. (A) Reduced basal and maximal glycolytic activity in APLN+APLNR siRNA-treated compared to siScr-treated HUVECs. (B) No significant difference in oxygen consumption in APLN+APLNR siRNA-treated compared to siScr-treated HUVECs. (C) Reduced c-MYC levels in APLN+APLNR siRNA-treated compared to siScr-treated HUVECs. (D) RT-qPCR analysis of PFKFB3 mRNA levels in APLN+APLNR siRNA-treated compared to siScr-treated HUVECs. (E) Confocal projection images of the blood vasculature in the trunk region of a 54 hpf Tg(kdrl:HsHRAS-mCherry) animal injected with an EGFP:fli1a:pfkfb3 plasmid. Arrowheads point to formed DLAV fragments while asterisks indicate missing DLAV fragments. (F) Quantification of the rescue of the DLAV fragment by mosaic pfkfb3 overexpression in aplnrb -/- embryos. n.s. not significant (two-tailed t-test). Scale bar: E, 50 µm.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| genetic reagent (D. rerio) | Tg(fli1a:EGFP)y1 | Lawson and Weinstein, 2002 | ZFIN: y1 | |
| genetic reagent (D. rerio) | Tg(fli1a:nEGFP)y7 | Roman et al., 2002, | ZFIN: y7 | |
| genetic reagent (D. rerio) | Tg(kdrl:HsHRAS-mCherry)s896 | Chi et al., 2008 | ZFIN: s896 | |
| genetic reagent (D. rerio) | aplnramu296 | Helker et al., 2015 | ZFIN: mu296 | |
| genetic reagent (D. rerio) | aplnrbmu281 | codes for another allele of aplnrb from Helker et al., 2015 | ZFIN: mu281 | |
| genetic reagent (D. rerio) | aplnmu267 | Helker et al., 2015 | ZFIN: mu267 | |
| genetic reagent (D. rerio) | Tg(hsp70:bmp2b)fr13 | Chocron et al., 2007 | ZFIN: fr13 | |
| genetic reagent (D. rerio) | apelabr13 | Chng et al., 2013, | ZFIN: br13 | |
| genetic reagent (D. rerio) | Tg(fli:lifeact-GFP)mu240 | Hamm et al., 2016 | ZFIN: mu240 | |
| genetic reagent (D. rerio) | Tg(fli1a:Hsa.B4GALT1-mCherry)bns9 | Kwon et al., 2016 | ZFIN: bns9 | |
| genetic reagent (D. rerio) | Tg(hsp70:apln)mu269 | This manuscript | ZFIN: mu269 | |
| genetic reagent (D. rerio) | Tg(kdrl:HsHRAS-EGFP)mu280 | This manuscript | ZFIN: mu280 | |
| genetic reagent (D. rerio) | Tg(apln:EGFP)bns157 | This manuscript | ZFIN: bns157 | |
| genetic reagent (D. rerio) | Tg(aplnrb:aplnrb-TagRFP- sfGFP)bns309 | This manuscript | ZFIN: bns309 | |
| antibody | anti-FOXO1 (rabbit monoclonal) | Cell Signaling Technology | Cat#2880 | (1:1000) |
| antibody | anti-pThr24FOXO1/pThr32FOXO3a (rabbit monoclonal) | Cell Signaling Technology | Cat#9464 | (1:1000) |
| antibody | anti-c-MYC (rabbit polyclonal) | Cell Signaling Technology | Cat#9402 | (1:1000) |
| antibody | anti-Tubulin (rabbit polyclonal) | Cell Signaling Technology | Cat#2148 | (1:1000) |
| other | Taqman probe APLN | Thermo Fisher Scientific | Hs00175572_m1 | |
| other | Taqman probe APLNR | Thermo Fisher Scientific | Hs00270873_s1 | |
| other | Taqman probe PFKFB3 | Thermo Fisher Scientific | Hs00270873_s1 | |
| other | Taqman probe ACTB | Thermo Fisher Scientific | Hs01060665_g1 | |
| commercial assay or kit | In-Fusion HD Cloning Plus | Takara Bio | Cat# 638910 | |
| transfected construct (human) | APLN | Dharmacon | Cat# L-017023-01-0005 | 50 nM |
| transfected construct (human) | APLNR | Dharmacon | Cat# L-005430-00-0005 | 50 nM |
| transfected construct (human) | ON-TARGETplus Non-targeting Pool | Dharmacon | Cat# D-001810-10-05 | 50 nM |
| commercial assay or kit | mMessage mMachine SP6 Transcription Kit | Thermo Fisher Scientific | Cat# AM1340 | |
| commercial assay or kit | DIG RNA labelling kit | Roche | Cat# 11277073910 | |
| commercial assay or kit | SuperScript III First-Strand Synthesis System | Thermo Fisher Scientific | Cat#18080051 | |
| commercial assay or kit | RNA Clean and Concentrator Kit | Zymo Research | Cat# R1013 | |
| software, algorithm | ZEN Blue 2012 | Zeiss, Germany | ||
| software, algorithm | ZEN Black 2012 | Zeiss, Germany | ||
| software, algorithm | Imaris - Version 8.4.0 | Bitplane, UK | ||
| software, algorithm | GraphPad Prism 6 | GraphPad Software, USA |