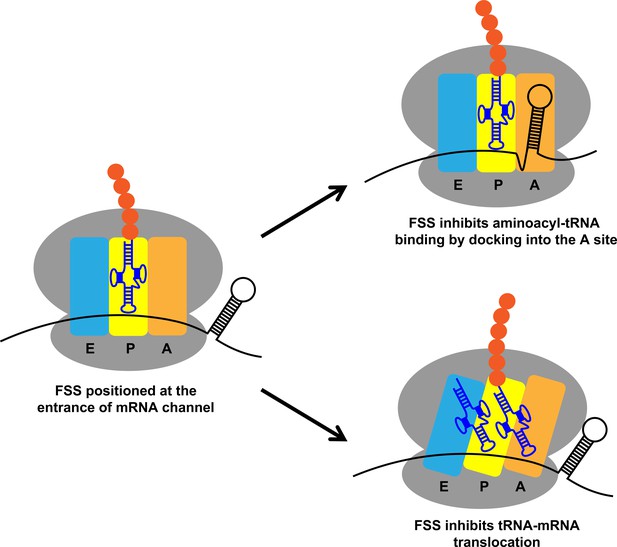
mRNA stem-loops can pause the ribosome by hindering A-site tRNA binding
Figures
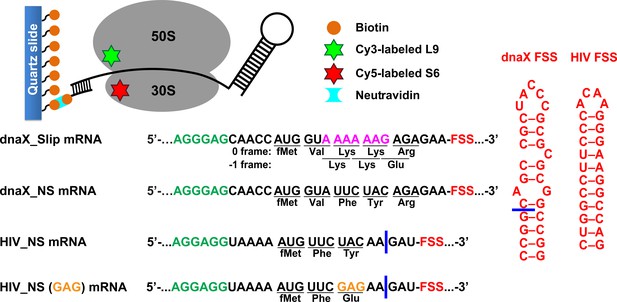
Experimental design.
The effect of frameshift-inducing mRNA stem-loops on translation elongation was studied using FRET between cy5 (red) and cy3 (green) attached to 30S protein S6 and 50S protein L9, respectively. S6-cy5/L9-cy3 ribosomes were immobilized on quartz slides using neutravidin and biotinylated DNA oligomers annealed to the mRNA. dnaX_Slip mRNA contains an internal SD sequence (green), a slippery sequence (magenta) and an FSS (red). In the non-slippery (NS) dnaX and HIV mRNAs, the slippery sequences were replaced by non-slippery codons. Two different HIV_NS mRNAs contain either a UAC or a GAG (orange) codon. Corresponding polypeptide sequences are shown below each mRNA. The ΔFSS mRNAs are truncated as indicated by blue bars.
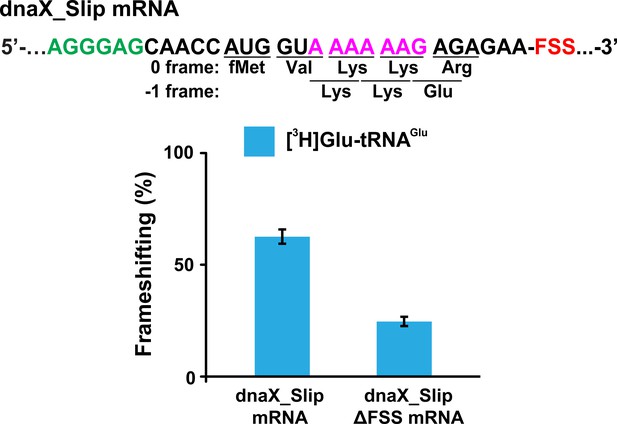
FSS and slippery sequence of dnaX mRNA stimulate −1 RPF.
Ribosomes containing P-site N-Ac-Val-tRNAVal were programmed with either dnaX_Slip or dnaX_Slip ΔFSS mRNA. The ribosomes were incubated with EF-G•GTP, EF-Tu•GTP, Lys-tRNALys, Arg-tRNAArg (binds in 0 frame) and [3H]Glu-tRNAGlu (binds in −1 frame) for 6 min. [3H]Glu-tRNAGlu binding was measured by filter-binding assay. Frameshifting efficiency (ribosome A-site occupancy by [3H]Glu-tRNAGlu) was normalized by the P-site occupancy of N-Ac-[3H]Glu-tRNAGlu non-enzymatically bound to the ribosome programmed with dnaX_Slip ΔFSS mRNA. Error bars show standard deviations of triplicated measurements.
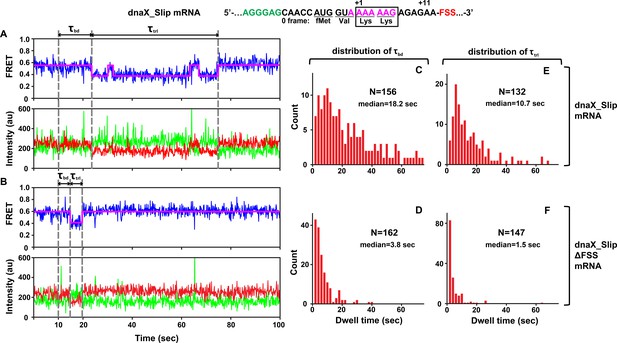
DnaX FSS slows ribosome intersubunit rotation.
S6-cy5/L9-cy3 ribosomes containing P-site N-Ac-Val-Lys-tRNALys were programmed with either dnaX_Slip (A, C, E) or dnaX_Slip ΔFSS (B, D, F) mRNAs. After 10 s of imaging, EF-Tu•GTP•Lys-tRNALys and EF-G•GTP were co-injected into the flow-through chamber. (A–B) Representative smFRET traces show cy3 fluorescence (green), cy5 fluorescence (red), FRET efficiency (blue) and the HHM fit of FRET efficiency (magenta). τbd is the dwell time between the injection and Lys-tRNALys binding to the A site, which corresponds to the transition from NR (0.6 FRET) to R (0.4 FRET) state of the ribosome. τtrl is the dwell time between A-site binding of Lys-tRNALys and EF-G-catalyzed tRNA translocation, which corresponds to the transition from R to the stable (i.e. lasting over 4 s) NR state of the ribosome. The full-length views of smFRET traces are shown in Figure 2—figure supplement 1A,B. (C–F) Histograms (2 s binning size) compiled from over 100 traces show the distributions and median values of τbd and τtrl. N indicates the number of FRET traces assembled into each histogram.
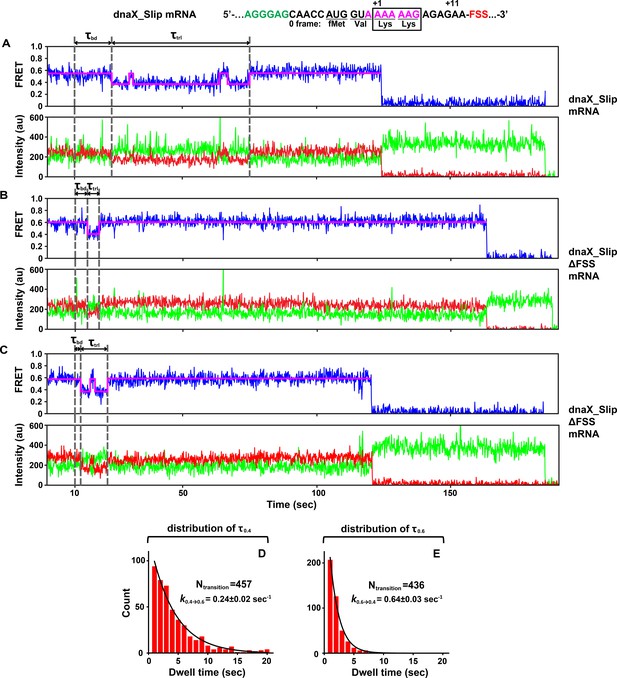
Both A-site tRNA binding and translocation are hindered by the dnaX FSS positioned at the entrance of mRNA channel.
(A–B) The full-length view of smFRET traces shown in Figure 2A–B with a single-step photobleaching of both Cy3 and Cy5 fluorophores. (C) Ribosome programmed with dnaX_Slip ΔFSS mRNA showed excursion from 0.4 to 0.6 FRET in pre-translocation state. (D–E) In the ribosome containing deacylated tRNAPhe in the P site, distributions of dwell time in 0.4 (τ0.4, (D) and 0.6 (τ0.6, (E) FRET state are fitted to single exponential decay shown as solid black line. N indicates the total number of 0.4-to-0.6 (D) and 0.6-to-0.4 (E) transitions, respectively.
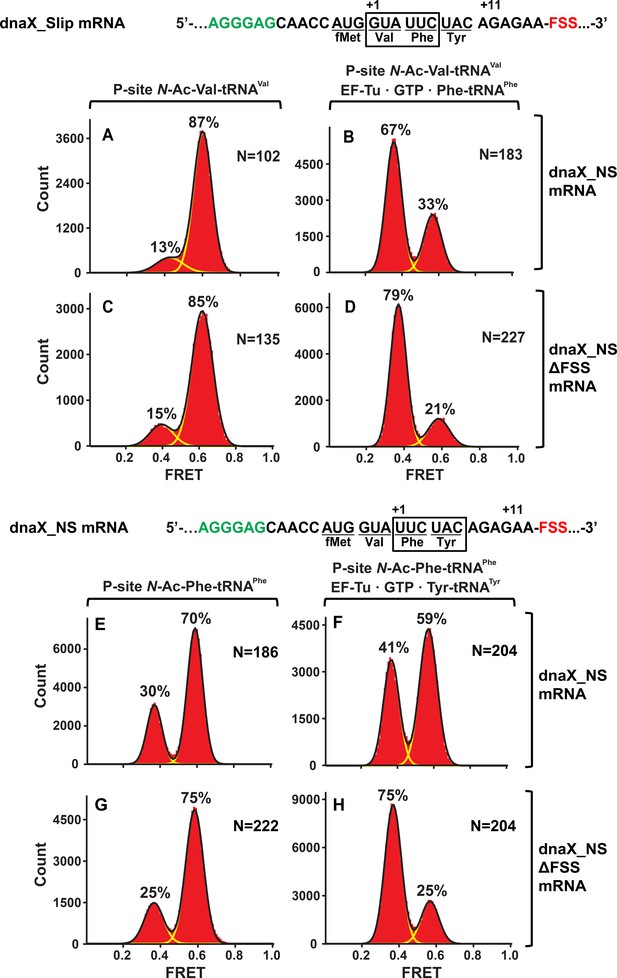
In the context of non-slippery codons, the dnaX FSS stalls the ribosome in the NR conformation.
Histograms show FRET distributions in S6-cy5/L9-cy3 ribosomes programmed with dnaX_NS (A–B, E–G) or dnaX_NS ∆FSS (C–D, H–J) mRNA, respectively. Ribosomes were bound with P-site peptidyl tRNA analogs, N-Ac-Val-tRNAVal (A, C) or N-Ac-Phe-tRNAPhe (E, G). The ribosomes were then incubated with either EF-Tu•GTP•Phe-tRNAPhe (B, D) or EF-Tu•GTP•Tyr-tRNATyr (F, H) for 5 min and imaged after removal of unbound aminoacyl-tRNAs. Yellow lines show individual Gaussian fits of FRET distributions. Black lines indicate the sum of Gaussian fits. N indicates the number of FRET traces compiled into each histogram. The fractions of the ribosome in R and NR conformations are shown above the corresponding 0.4 and 0.6 Gaussian peaks, respectively.
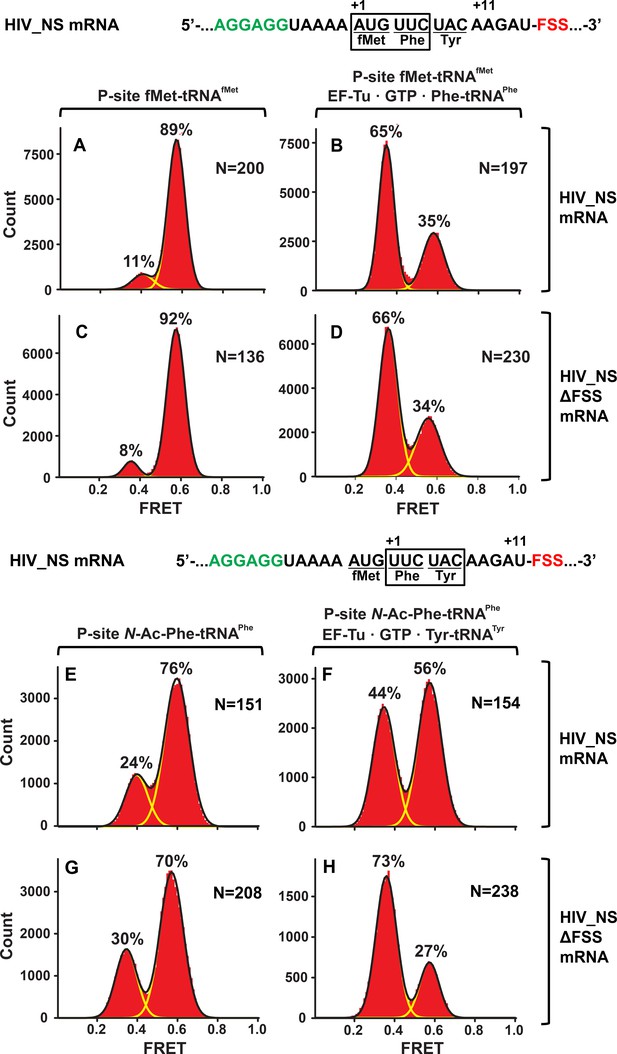
In the context of non-slippery codons, the HIV FSS stalls the ribosome in NR conformation.
Histograms show FRET distributions in S6-cy5/L9-cy3 ribosomes programmed with HIV_NS (A–B, E–G) or HIV_NS ∆FSS (C–D, H–J) mRNA, respectively. Ribosomes contained fMet-tRNAfMet (A, C) or N-Ac-Phe-tRNAPhe (E, G) in the P site. The ribosomes were then incubated with either EF-Tu•GTP•Phe-tRNAPhe (B, D) or EF-Tu•GTP•Tyr-tRNATyr (F, H) for 5 min and imaged after removal of unbound aminoacyl-tRNAs. Yellow lines show individual Gaussian fits of FRET distributions. Black lines indicate the sum of Gaussian fits. N indicates the number of FRET traces compiled into each histogram. The fractions of the ribosome in R and NR conformations are shown above the corresponding 0.4 and 0.6 Gaussian peaks, respectively.
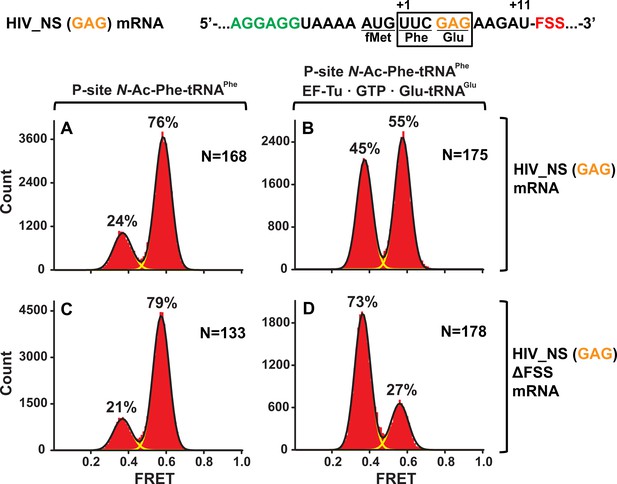
The FSS-induced ribosome stalling in NR conformation is independent of A-site codon identity.
Histograms show FRET distributions in S6-cy5/L9-cy3 ribosomes programmed with HIV_NS (GAG) (A–B) or HIV_NS (GAG) ∆FSS (C–D) mRNA, respectively. Ribosomes containing P-site N-Ac-Phe-tRNAPhe (A, C) were incubated with EF-Tu•GTP•Tyr-tRNATyr (B, D) for 5 min and imaged after removal of unbound aminoacyl-tRNAs. Yellow lines show individual Gaussian fits of FRET distributions. Black lines indicate the sum of Gaussian fits. N indicates the number of FRET traces compiled into each histogram. The fractions of the ribosome in R and NR conformations are shown above the corresponding 0.4 and 0.6 Gaussian peaks, respectively.
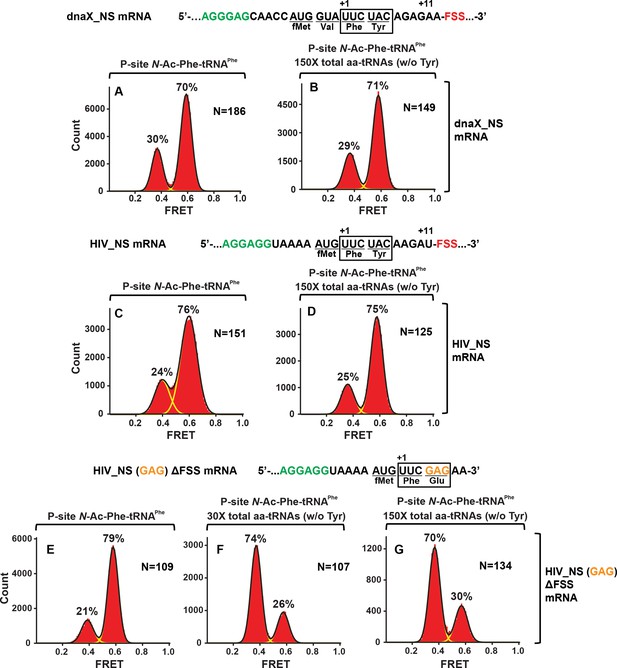
In the absence of slippery sequence, levels of the FSS-induced frameshifting are negligible.
Histograms show FRET distributions in S6-cy5/L9-cy3 ribosomes programmed with the dnaX_NS (A, B), HIV_NS (C, D), or HIV_NS (GAG) ∆FSS (E–G) mRNA, respectively. Ribosomes containing P-site N-Ac-Phe-tRNAPhe (A, C, E) were incubated with 150- (B, D, G) or 30- (F) fold molar excess of total aminoacyl-tRNAs•EF-Tu•GTP (mixture of all aminoacyl-tRNAs except Tyr-tRNATyr) for 5 min and imaged after removal of unbound aminoacyl-tRNA.
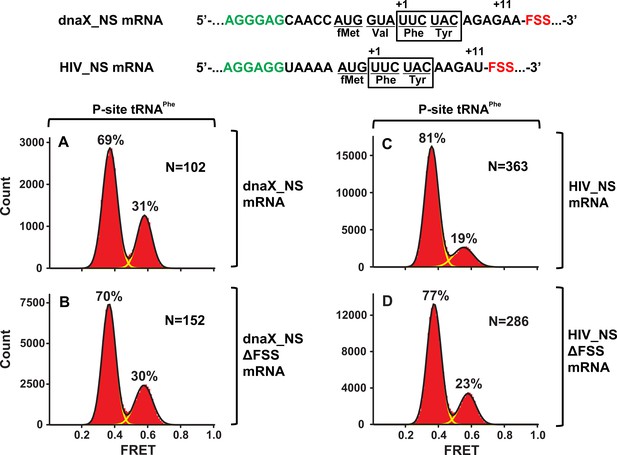
The FSSs do not stabilize the pretranslocation ribosome in the NR conformation.
Histograms show FRET distributions in S6-cy5/L9-cy3 ribosomes, which contained deacylated tRNAPhe in the P site and were programmed with the dnaX_NS (A), dnaX_NS ∆FSS (B), HIV_NS (C), or HIV_NS ∆FSS (D) mRNA, respectively.
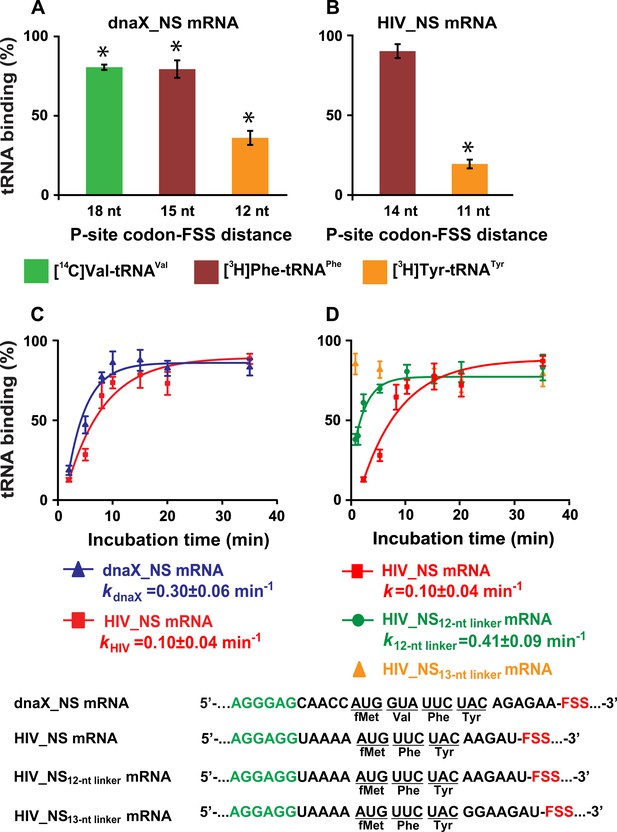
The dnaX and HIV FSSs inhibit A-site tRNA binding.
(A–B) Incorporation of radio-labeled amino acids during translation through first four codons of dnaX_NS mRNA (A) or first three codons of HIV_NS mRNA (B) were measured by filter-binding assays. (C) Kinetics of EF-Tu-catalyzed [3H]Tyr-tRNATyr binding to the A site of ribosomes containing N-Ac-Phe-tRNAPhe in the P site. Ribosomes were programmed with dnaX_NS mRNA (blue) or HIV_NS mRNA (red). Single exponential fits are shown as line graphs. (D) Kinetics of EF-Tu-catalyzed [3H]Tyr-tRNATyr binding to the A site of ribosomes containing N-Ac-Phe-tRNAPhe in the P site. Ribosomes were programmed with HIV_NS mRNA (red), HIV_NS12-nt linker mRNA (green), and HIV_NS13-nt linker mRNA (yellow), respectively. The binding of radio-labeled amino acids to ribosomes programmed with FSS-containing mRNA is shown relative to that observed in ribosomes programmed with corresponding ∆FSS mRNA (A–D). Asterisks indicate that amino acid incorporation into ribosomes programmed with FSS-containing mRNA was significantly different from that in ribosomes programmed with ∆FSS mRNA, as p-values determined by the Student t-test were below 0.05. Error bars in each panel show standard deviations of triplicated measurements.
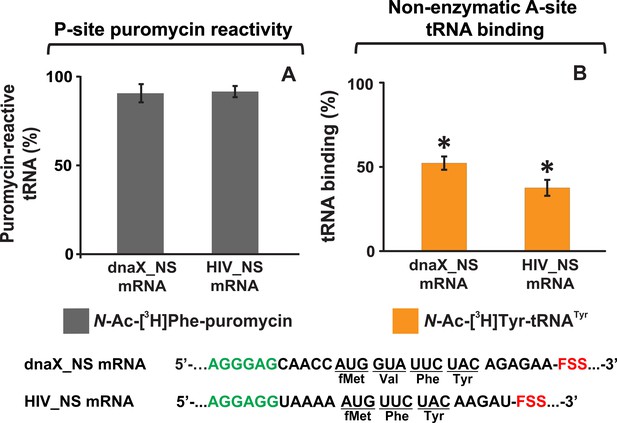
The FSS inhibits A-site tRNA binding but not ribosome-catalyzed transpeptidation reaction.
(A) Ribosomes bound with P-site N-Ac-[3H]Phe-tRNAPhe and FSS-containing mRNA were incubated with puromycin. The amount of N-Ac-[3H]Phe-puromycin extracted from the ribosomes programmed with dnaX_NS or HIV_NS mRNA was normalized by the amount of N-Ac-[3H]Phe-puromycin extracted from the ribosomes programmed with corresponding ∆FSS mRNAs. (B) The extent of N-Ac-[3H]Tyr-tRNATyr binding to ribosomes bound with P-site deacylated tRNAPhe and programmed with dnaX or HIV_NS mRNA after 10 min incubation. The binding of N-Ac-[3H]Tyr-tRNATyr determined by filter-binding assay is shown relative to that measured in ribosomes programmed with corresponding ∆FSS mRNAs. Asterisks indicate that amino acid incorporation into ribosomes programmed with FSS-containing mRNA was significantly different from that in ribosomes programmed with ∆FSS mRNA, as p-values determined by the Student t-test were below 0.05. Error bars in each panel show standard deviations of triplicated measurements.
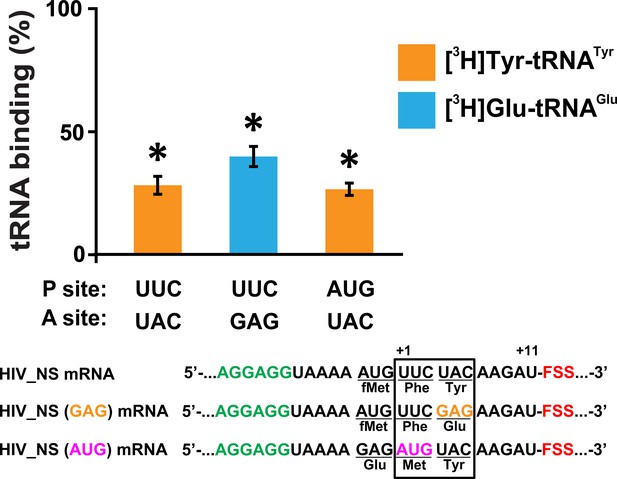
The FSS-mediated inhibition of A-site tRNA binding is independent of P-site and A-site codon identities.
The extent of EF-Tu-catalyzed cognate aminoacyl-tRNA binding after a 5 min incubation with ribosomes programmed with HIV_NS, HIV_NS (GAG) or HIV_NS (AUG) mRNAs. The P site of the ribosome was bound with N-Ac-Phe-tRNAPhe (in the presence of HIV_NS and HIV_NS (GAG) mRNAs) or N-Ac-Met-tRNAMet (in the presence of HIV_NS (AUG) mRNA). The binding of radio-labeled amino acids to ribosomes programmed with FSS-containing mRNA is shown relative to that observed in ribosomes programmed with corresponding ∆FSS mRNA. Asterisks indicate that amino acid incorporation into ribosomes programmed with FSS-containing mRNA was significantly different from that in ribosomes programmed with ∆FSS mRNA, as p-values determined by the Student t-test were below 0.05. Error bars in each panel show standard deviations of triplicated measurements.
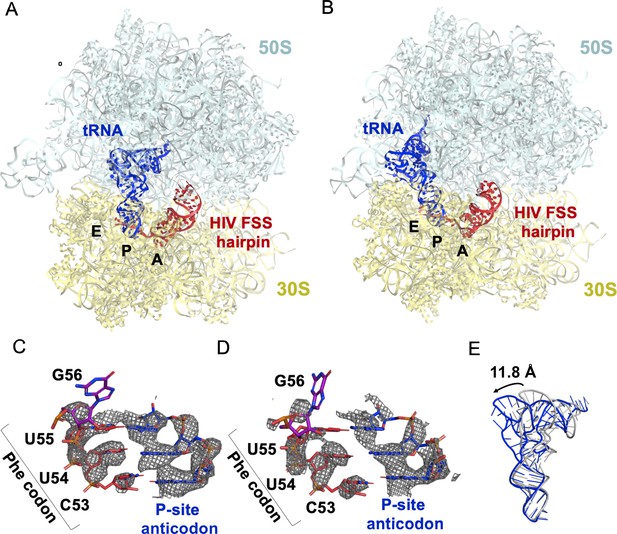
The HIV FSS hairpin occupies the ribosomal A site.
(A) Cryo-EM structures of the 70S ribosome in non-rotated (NR-I) and (B) rotated (R–I) conformations. The large subunits (50S) are shown in aqua, the small subunits (30S) in yellow, P-site tRNA in blue, and HIV FSS hairpin in red. (C and D) Close-up views of the codon and anti-codon basepairs of the NR-I (C) and R-I (D) states illustrating in-frame basepairing of the HIV_NS(GAG) mRNA (red) with the P-site tRNA (blue). The first position of the GAG A-site codon is shown in purple. The cryo-EM map (gray mesh) was sharpened by applying a B-factor of −50 Å2. (E) Overlay of NR-I P-site tRNA with P-site tRNA bound in the P/P classical site (PDB ID: 4V5D) shows a 11.8 Å rotation of the tRNA elbow towards the E site.
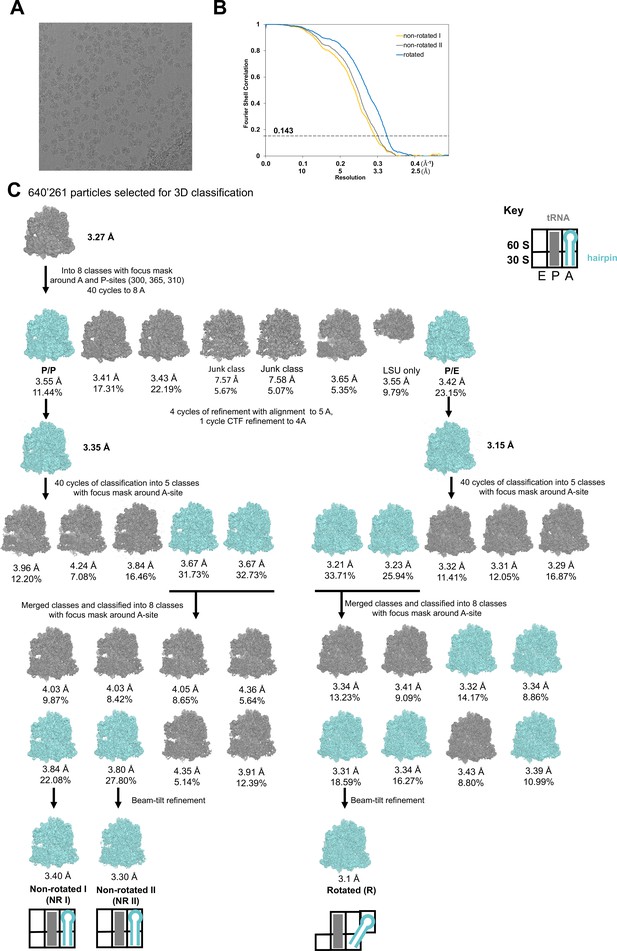
Data and schematic of cryo-EM refinement and classification.
(A) Representative micrograph. (B) Fourier shell correlation as a function of resolution for the NR (I and II) and R structures. (C) 640,261 particles were aligned to a single model. Focused 3D classification using a spherical mask around the A and P sites yielded one class of R and one class of NR ribosomes. Each class was extracted and refined separately at 5 Å. Sub-classification of each class with a spherical mask around the A site yielded two classes of R and NR ribosomes with weak density for the HIV FSS hairpin in the A site. Particles were extracted based on HIV FSS hairpin occupancy and further sub-classified with a mask as described for the previous step.
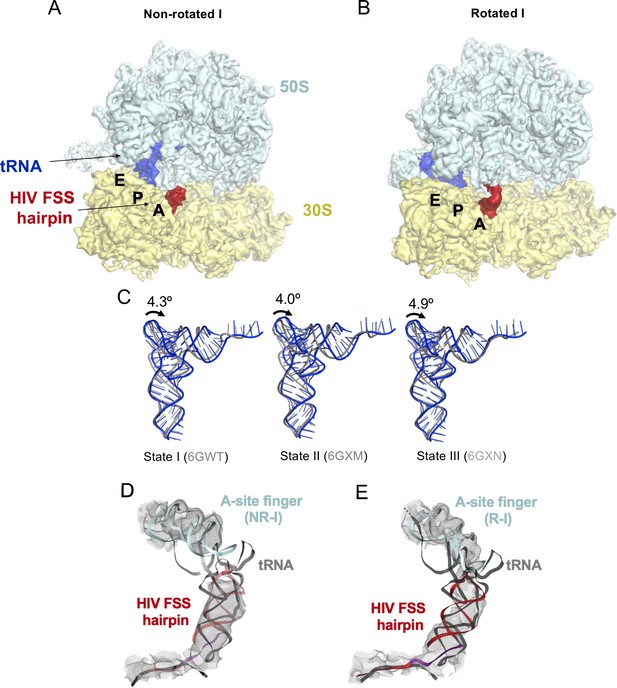
Density maps of (A) NR-I and (B) R-I filtered to 5 Å resolution.
The cryo-EM maps are colored as in Figure 5. (C) Comparison of P-site tRNA of the NR-I structure from this work (blue) with tRNA from states I-III of recycling factor 1 and 3 (RF1 and RF3) bound ribosome structures stalled with Apidaecin 137 (gray) (Graf et al., 2018). (D–E) Fitting of a tRNA (gray) into the hairpin density of the (C) NR-I and (D) R-I states shows that tRNA cannot be accommodated without steric clashes with the 50S A-site finger (aqua) in either of the observed states. This rules out that the density corresponds to a tRNA.
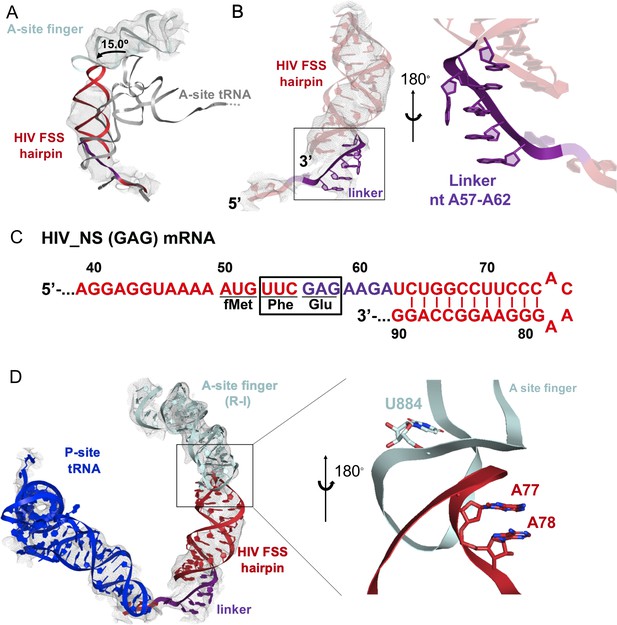
Stacking interactions of linker nucleotides stabilize the HIV FSS in the A site.
(A) Overlay of the NR-I HIV FSS hairpin from this work with A-site tRNA (gray) accommodated in a ribosome in the classical state (PDB ID: 4V5D). The hairpin density is shown after filtering to 8 Å. (B) View of the R-I HIV FSS hairpin model (red, linker in purple) in cryo-EM density filtered to 5 Å (gray mesh) and close-up of the purine stack (shown in purple) after 180° rotation. (C) Primary sequence and secondary structure of the HIV_NS(GAG) mRNA. The linker sequence is highlighted in purple. (D) In the rotated state, the HIV FSS hairpin (red) contacts the A-site finger (aqua) of the large ribosomal subunit. The hairpin density allows to clearly identify helical pitch, major and minor grooves of the A-form RNA. The close-up after 180° rotation shows that the only complementary bases within the two loop regions point away from each other and likely do not contribute to A-site finger/hairpin binding. P-site tRNA is blue.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Escherichia coli) | dnaX | doi: 10.1093/nar/14.20.8091 | Uniprot ID: P06710 | |
| Gene (Human Immunodeficiency Virus Type 1) | gag-pol | doi: 10.1089/aid.1987.3.57 | Uniprot ID: P04585 | |
| Strain, strain background (Escherichia coli) | MRE600 | ATCC | ATCC #29417, (NCTC #8164, NCIB #10115) | E. coli strain K-12 that lacks the RNase I activity |
| Strain, strain background (Escherichia coli) | DH5α competent cell | Thermo Fisher Scientific | Catalog #: 18265017 | |
| Genetic reagent (Escherichia coli) | tRNA | Chemical Block | tRNAPhe tRNATyrtRNAfMet tRNAMettRNAGlu tRNAValtRNALys tRNAArg | |
| Genetic reagent (Escherichia coli) | Total tRNA from E. coli MRE600 | Sigma-Aldrich | Catalog #: 10109541001 | |
| Transfected construct (Escherichia coli) | pSP64 poly (A) | Promega | Catalog #: P1241 | |
| Biological sample (Escherichia coli) | ribosome (30S, 50S and 70S) | doi: 10.1016/j.jmb.2007.04.042 doi: 10.1073/pnas.1520337112 | ||
| Recombinant DNA reagent | SacI-HF | New England Biolabs | Catalog #: R3156 | |
| Recombinant DNA reagent | BglII | New England Biolabs | Catalog #: R0144 | |
| Recombinant DNA reagent | HindIII-HF | New England Biolabs | Catalog #: R3104 | |
| Recombinant DNA reagent | T4 DNA ligase | New England Biolabs | Catalog #: M0202 | |
| Peptide, recombinant protein | elongation factor Tu (EF-Tu) | doi: 10.1016/j.jmb.2007.04.042 | ||
| Peptide, recombinant protein | elongation factor G (EF-G) | doi: 10.1016/j.jmb.2007.04.042 | ||
| Peptide, recombinant protein | T7 polymerase | doi: 10.1073/pnas.95.2.515 | ||
| Commercial assay or kit | Plasmid Miniprep System | Promega | Catalog #: A1223 | |
| Commercial assay or kit | Gel and PCR Clean-Up System | Promega | Catalog #: A9281 | |
| Commercial assay or kit | DNA oligo synthesis | INTEGRATED DNA TECHNOLOGIES (IDT) | ||
| Commercial assay or kit | DNA sequencing | ACGT, INC | ||
| Chemical compound, drug | puromycin | Sigma-Aldrich | Catalog #: P8833 | |
| Chemical compound, drug | cy3 maleimide | Click Chemistry Tools | Catalog #: 1009 | |
| Chemical compound, drug | cy5 maleimide | Click Chemistry Tools | Catalog #: 1004 | |
| Chemical compound, drug | Phenylalanine, L -[2,3,4,5,6-3H]- | PerkinElmer | Catalog #: NET112201MC | |
| Chemical compound, drug | Valine, L-[U-14C]- | PerkinElmer | Catalog #: NEC291EU050UC | |
| Chemical compound, drug | Methionine, L-[35S]- | PerkinElmer | Catalog #: NEG009T001MC | |
| Chemical compound, drug | Glutamic Acid, L-[3,4–3H]- | PerkinElmer | Catalog #: NET490001MC | |
| Chemical compound, drug | Tyrosine, L -[ring-3,5 | PerkinElmer | Catalog #: NET127001MC | |
| Software, algorithm | smFRET data acquisition and analysis package | Taekjip Ha’s laboratory website at Johns Hopkins University (http://ha.med.jhmi.edu/resources/) | ||
| Software, algorithm | IDL | ITT, INC. (https://www.harrisgeospatial.com/Software-Technology/IDL) | ||
| Software, algorithm | HaMMy | Taekjip Ha’s laboratory website at Johns Hopkins University (http://ha.med.jhmi.edu/resources/) doi: 10.1529/biophysj.106.082487 | ||
| Software, algorithm | SerialEM | (https://bio3d.colorado.edu/SerialEM/) doi: 10.1016/j.jsb.2005.07.007 | ||
| Software, algorithm | cisTEM | (https://cistem.org/) doi: 10.7554/eLife.35383 | ||
| Software, algorithm | Phenix-1.17.1–3660 | (https://www.phenix-online.org/) doi:10.1107/S2059798319011471 | ||
| Software, algorithm | Coot v0.9 pre-EL | Part of CCPEM 1.3.0 suite (https://www.ccpem.ac.uk/index.php) doi:10.1107/S2059798317007859 | ||
| Software, algorithm | PyMol 2.3.2 | Schrödinger, LLC (https://pymol.org) |
Additional files
-
Supplementary file 1
Model mRNA sequences.
The SD sequence is shown in green, slippery sequence in magenta, FSS in red, and handle sequence, which is complementary to biotinylated DNA oligo, underlined. To change the linker length between the HIV FSS and the P-site codon, HIV_NS12-nt linker mRNA and HIV_NS13-nt linker mRNA were prepared by extending the linker from native 11 nucleotides to 12 and 13 nucleotides, respectively. To alter codon identities, a UAC-to-GAG mutation was made on HIV_NS mRNA to generate HIV_NS (GAG) mRNA. Similarly, AUG-to-GAG and UUC-to-AUG mutations were made to generate HIV_NS (AUG) mRNA. The codon replacements are colored orange. Vertical black bars indicate the 3’ ends of ΔFSS mRNAs.
- https://cdn.elifesciences.org/articles/55799/elife-55799-supp1-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/55799/elife-55799-transrepform-v2.pdf