The His-Gly motif of acid-sensing ion channels resides in a reentrant ‘loop’ implicated in gating and ion selectivity
Figures
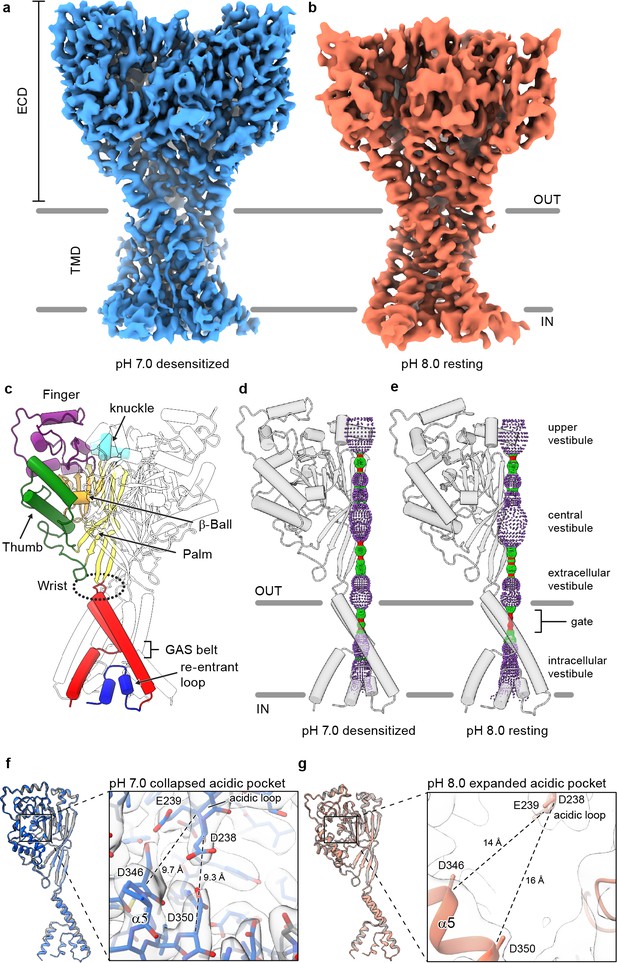
Structures of cASIC1-SMA.
(a-b) Cryo-EM maps of cASIC1-SMA at pH 7.0 (a) and pH 8.0 (b). (c) Cartoon diagram of cASIC1 with single subunit shown colored by domain. (d-e) Pore profiles for cASIC1-SMA in a desensitized (d) and resting (e) state calculated with HOLE software (pore radius: red <1.15 Å<green < 2.3 Å<purple). (g) Single subunit superposition of cASIC1-SMA in a desensitized state (blue) and the desensitized state X-ray structure (Baconguis et al., 2014; Gonzales et al., 2009) (PDB 4NYK, grey). Detailed view of the collapsed acidic pocket is shown in the inset. (h) Single subunit superposition of cASIC1-SMA in a resting state (salmon) and the resting state X-ray structure (Yoder et al., 2018a) (PDB 5WKU, grey). Detailed view of the expanded acidic pocket is shown in the inset.
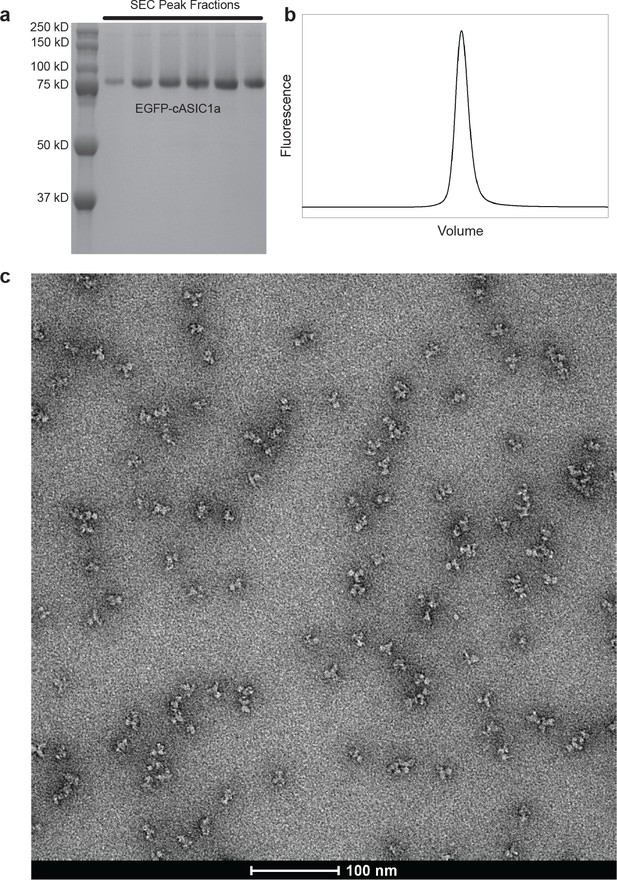
Purification of cASIC1-SMA.
(a-b) SDS-PAGE (a) and FSEC (b) analysis of SMA-solubilized EGFP-cASIC1 particles after size-exclusion chromatography. (c) Negative stain transmission electron microscopy of SMA-solubilized EGFP-cASIC1 particles.
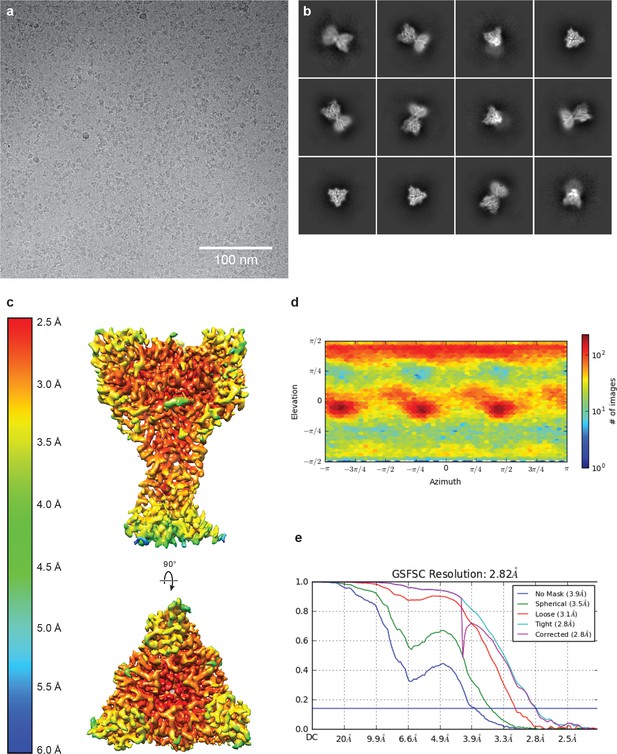
Cryo-EM of cASIC1-SMA at pH 7.0.
(a-b) Representative micrograph (a) and 2D classes (b) of cASIC1-SMA at pH 7.0. (c–e) Local resolution estimation (c) angular distribution (d) and gold standard FSC resolution estimation (e) from final non-uniform refinement in cryoSPARC V2.
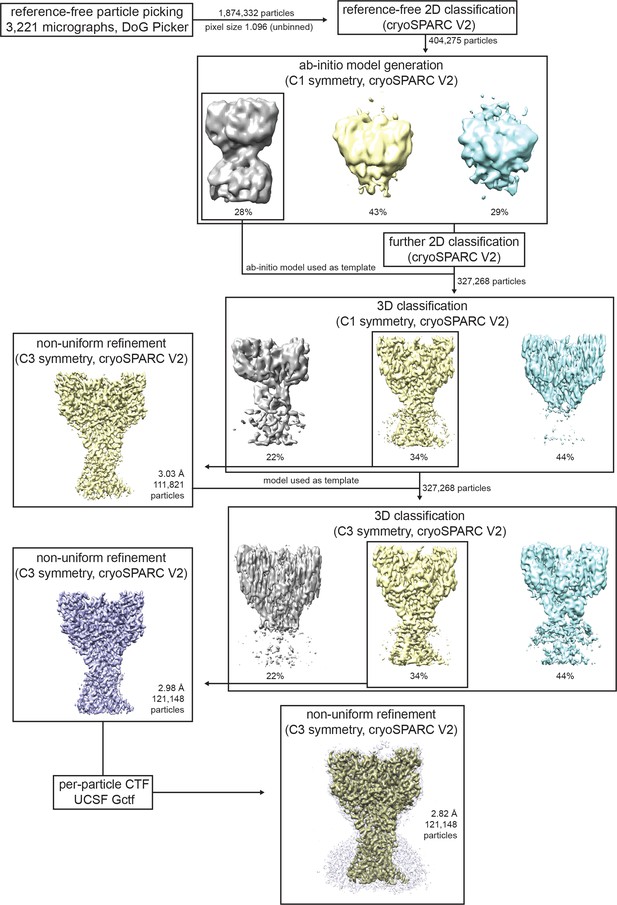
Cryo-EM data processing for cASIC1-SMA at pH 7.0.
Data processing strategy for cASIC1-SMA pH 7.0 dataset.
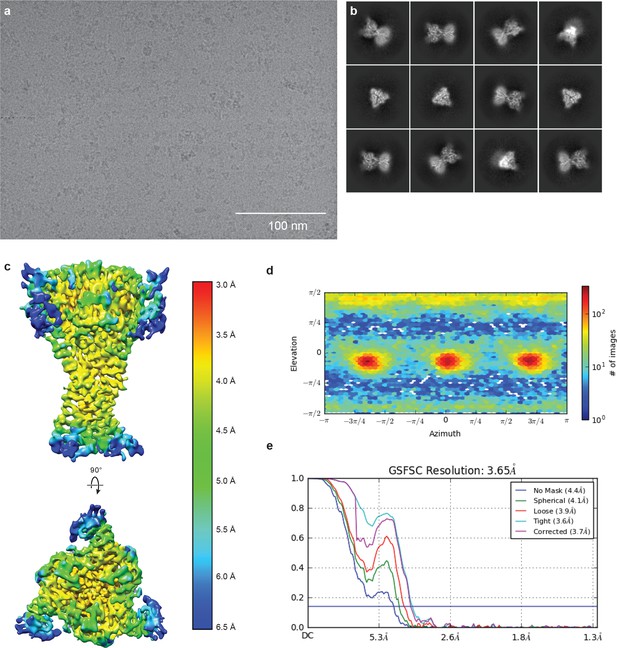
Cryo-EM of cASIC1-SMA at pH 8.0.
(a-b) Representative micrograph (a) and 2D classes (b) of cASIC1-SMA at pH 8.0. (c–e) Local resolution estimation (c) angular distribution (d) and gold standard FSC resolution estimation (e) from final non-uniform refinement in cryoSPARC V2.
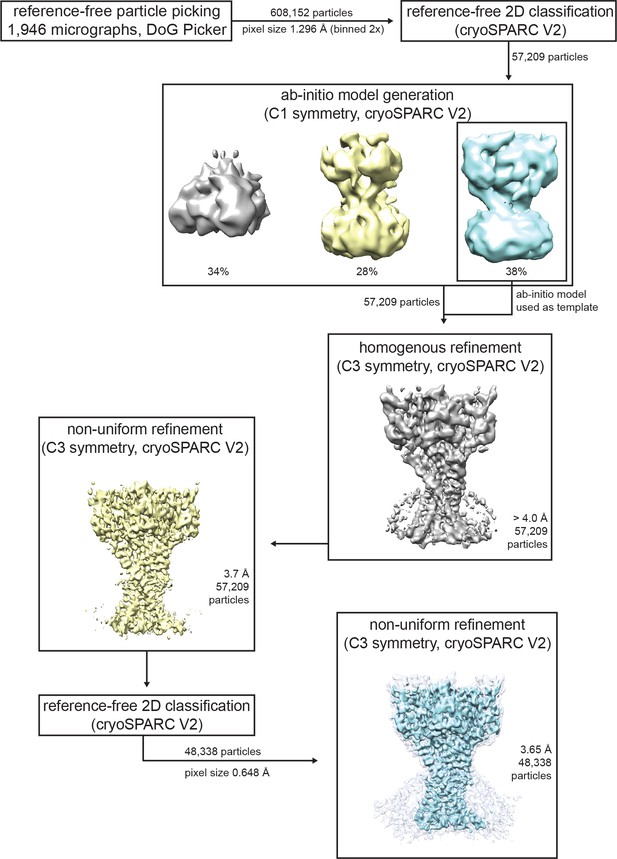
Cryo-EM data processing for cASIC1-SMA at pH 8.0.
Data processing strategy for cASIC1-SMA pH 8.0 dataset.
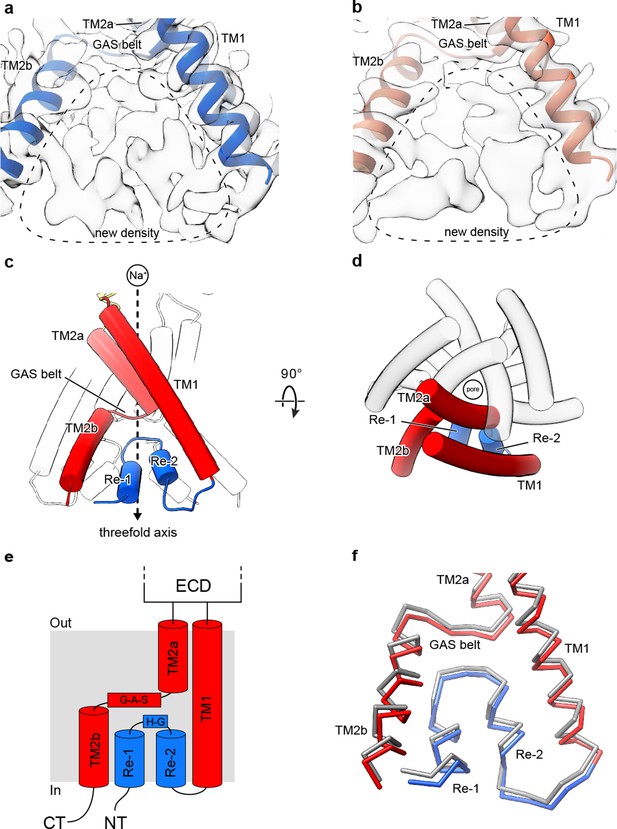
Cryo-EM density and structure of the reentrant, pre-TM1 domain.
(a-b) Cryo-EM density corresponding to amino terminal residues of cASIC1-SMA at pH 7.0 (a) and pH 8.0 (b). (c–d) Side (c) and top-down (d) views of the TMD from cASIC1-SMA in a desensitized state at low pH. A single subunit is shown, colored by domain. (e) Schematic depicting the TMD topology of cASIC1 channels. (f) Backbone superposition of cASIC1-SMA in the desensitized (colored by domain) and resting (grey) states.
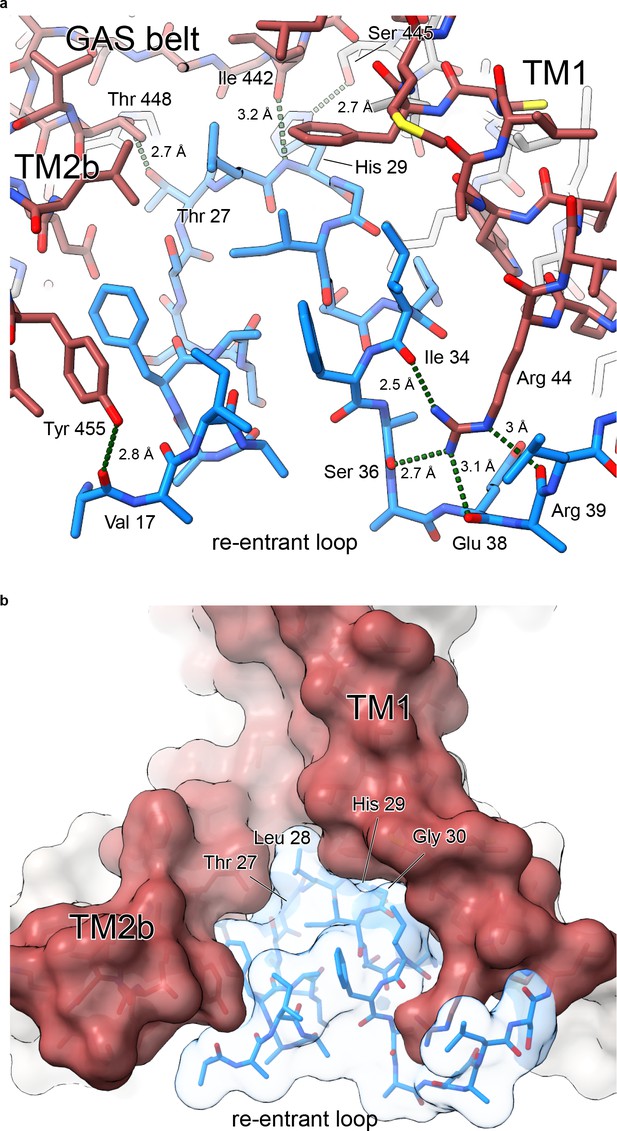
Polar contacts at the reentrant loop.
(a) Polar interactions between the reentrant loop and neighboring residues. (b) Surface representation of lower TMD showing interfaces between the reentrant loop residues and TM1 and TM2b helices.
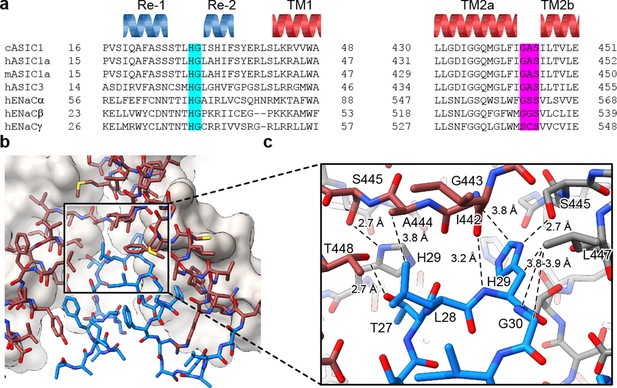
The HG motif resides at a subunit interface ‘below’ the GAS belt.
(a) Sequence alignment of selected ASIC1, ASIC3 and ENaC channels covering the pre-TM1 and TM2 domains with GAS domain and HG motif residues highlighted in pink and light blue, respectively, and with secondary structure for cASIC1 shown above the sequences. (b–c) View of the chemical environment around the reentrant loop (b) with a detailed view of the GAS domain and HG motif interface (c).
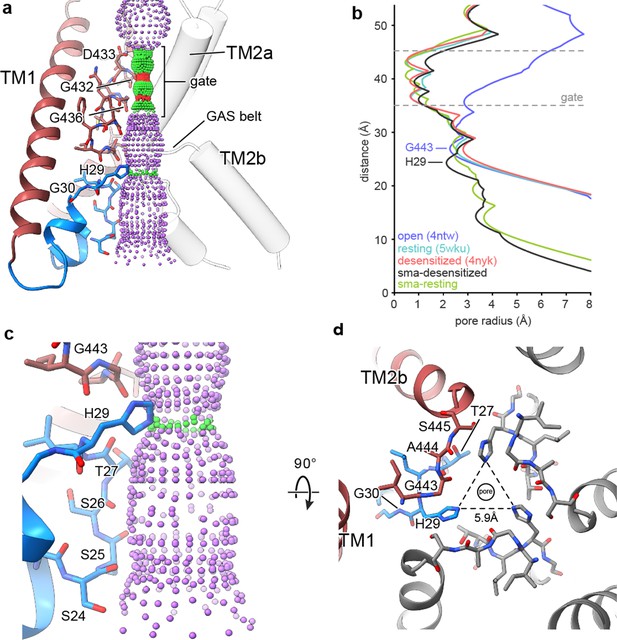
The reentrant loop forms the lower ion permeation pathway.
(a) Overview of pore-forming residues of desensitized cASIC1-SMA channels beginning at the ion channel gate. Pore profile calculated with HOLE software (pore radius: red <1.15 Å<green < 2.3 Å<purple) is shown. (b) Plot of pore radius corresponding to the view in (a) for resting (Yoder et al., 2018a) (PDB 5WKU), open (Baconguis et al., 2014) (PDB 4NTW) and desensitized (Baconguis et al., 2014; Gonzales et al., 2009) (PDB 4NYK) X-ray and cASIC1-SMA cryo-EM structures. (c) Detailed view of the ‘lower’ ion permeation pathway formed by pre-TM1 residues. (d) Top-down view of the constriction ‘below’ the ion channel gate formed by His 29, as visualized in the desensitized cASIC1-SMA state. The GAS belt residues are in the foreground.
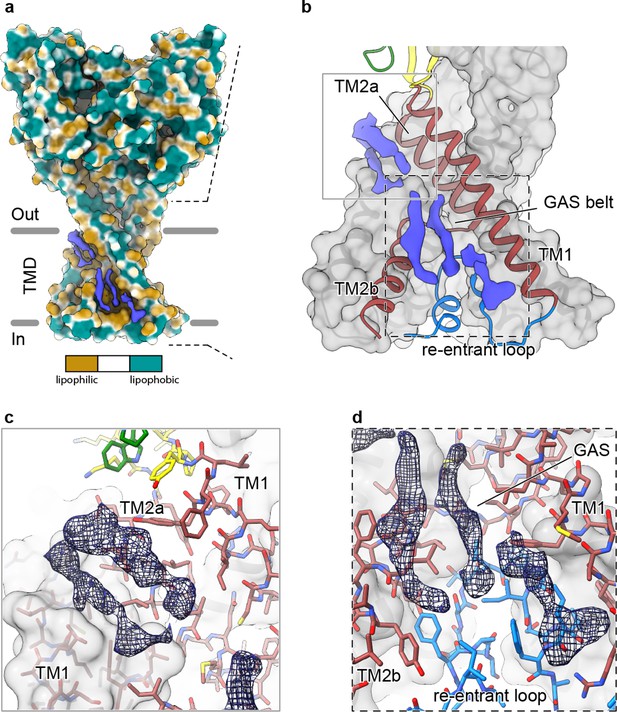
Elongated density within lipophilic channels at the TMD of cASIC1-SMA channels.
(a) Surface representation of cASIC1-SMA in the desensitized state colored by lipophilicity potential calculated with pyMLP (Laguerre et al., 1997) in ChimeraX (Goddard et al., 2018). (b) Hybrid cartoon and surface representation of putative lipid sites at the TMD. (c–d) Putative lipid densities between TM1 and TM2a (c) and between TM1 and TM2b adjacent to the reentrant loop (d) of cASIC1-SMA.
Tables
Additional files
-
Supplementary file 1
Cryo-EM data collection, processing and validation statistics.
- https://cdn.elifesciences.org/articles/56527/elife-56527-supp1-v2.doc
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/56527/elife-56527-transrepform-v2.pdf





