Cohesion is established during DNA replication utilising chromosome associated cohesin rings as well as those loaded de novo onto nascent DNAs
Figures
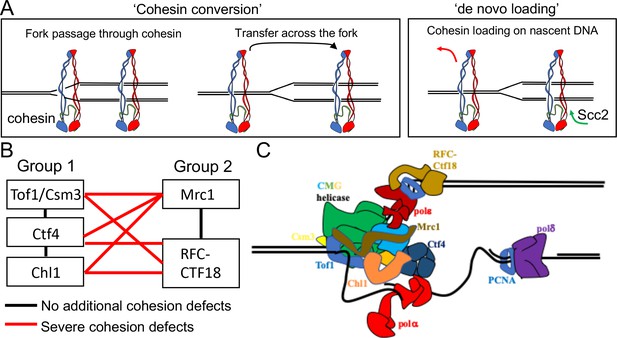
Possible mechanisms of cohesion establishment.
(A) Cohesin complexes previously associated with un-replicated DNA converted into cohesive ones behind the replication fork (Cohesin conversion) or cohesion built by de novo loading of cohesin molecules onto nascent DNAs at the replication forks (de novo loading).(B) The replisome-associated proteins that affect cohesion establishment belong to two epistasis groups, one containing Chl1, Ctf4, Csm3 and Tof1 and a second containing Mrc1 as well as the Ctf18-RFC. Combining two mutants from the same group has little or no additional effect on viability and causes no greater cohesion defect than the single mutants, while combining mutants from different groups either causes lethality or synthetic sickness and, when measured using conditional alleles, greatly increased defects in sister chromatid cohesion. Data from Xu et al., 2007. (C) The approximate positions of the Tof1/Csm3, Ctf4, Chl1, Mrc1 and the Ctf18-RFC within the replisome is shown (Baretić et al., 2020; Grabarczyk et al., 2018; Simon et al., 2014; Villa et al., 2016).
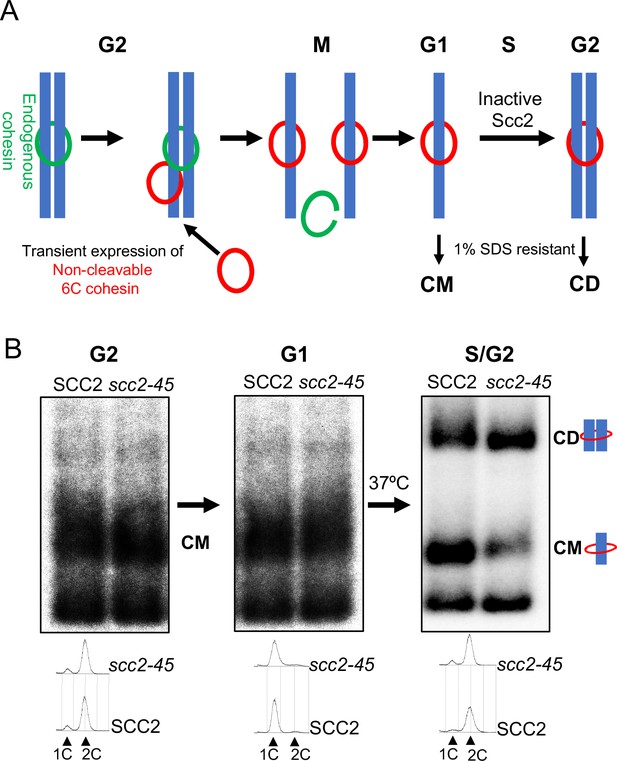
Chromosomal cohesin is converted into cohesive structures in the absence of Scc2 activity.
(A) A schematic depiction of the cohesin conversion assay, see the results and Materials and methods sections for detailed description of the assay. Green rings denote endogenous cohesin and red rings denote non-cleavable 6C cohesin. Only the latter molecules are capable of generating SDS-resistant catenated monomers (CMs) and catenated dimers (CDs). (B) Wild type (K24697) and scc2-45 (temperature sensitive mutant of Scc2) (K24738) strains that contain genes coding for 6C non cleavable cohesin (2C SMC1 2C SMC3 and GALp-2C SCC1NC) were arrested in G2 phase, the expression of 2C Scc1NC was induced by addition of galactose for 45 min. Mini-chromosome IP of the cultures at this stage shows CMs formed in both strains. The cultures were released from the G2 arrest and arrested in the subsequent the G1 phase, mini-chromosome IP of the cultures at this stage shows CMs formed in the previous G2 phase being retained in the subsequent G1 in both strains. The cultures were released from the G1 arrest and allowed to undergo replication at 37°C (in order to inactivate Scc2 in the scc2-45 strain), mini-chromosome IP shows formation of CDs in both the wild type and scc2-45 strains, in the scc2-45 mutant strain this is accompanied by a reduction in the amount of CMs. The FACS profiles of the two cultures at different stages of the experiment is shown below the respective southern blots. The data shown is representative of three independent biological repeats.
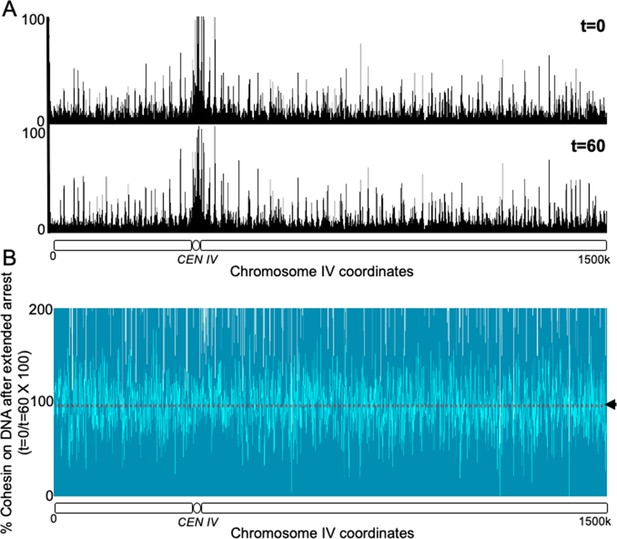
Non cleavable cohesin expressed in the G2 phase survives mitosis and remains stably associated with the chromosomes in the subsequent G1 phase.
(A) 6C non cleavable cohesin was expressed for 45 min in Wild type (K24697) strain arrested in G2 phase. The culture was released from the G2 arrest and arrested in the subsequent the G1 phase, a sample of the culture was drawn at this stage and fixed with formaldehyde (time 0). The rest of the culture was maintained in a prolonged G1 arrest by regular addition of the mating pheromone α-factor for a further 60 min. A sample was drawn at 60 min (time 60) and fixed with formaldehyde. The 0 and 60 min samples were subjected to calibrated ChIP sequencing with anti-PK antibody. The occupancy of Scc1NC along the entire chromosome IV is shown for both t = 0 and t = 60 min samples. (B) The difference in the level of Scc1NC-PK (time 60/time0) between the two conditions is shown as % of cohesin that remains on DNA upon prolonged G1 arrest. The median cohesin levels across the entire chromosome IV (dotted line) is marked with an arrowhead.
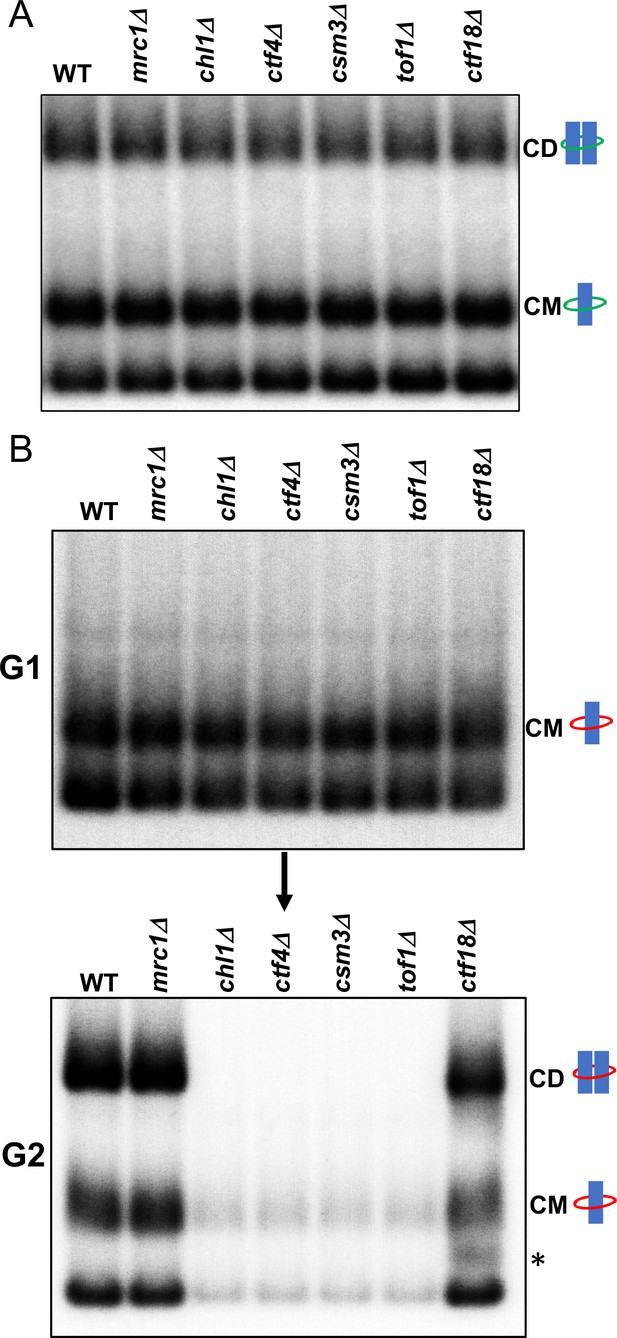
Tof1/Csm3, Ctf4 and Chl1 are essential for cohesin conversion.
(A) Exponentially growing wild type (K23889), mrc1Δ (K28092), chl1Δ (K28082), ctf4Δ (K28084), csm3Δ (K28108), tof1Δ (K28091) and ctf18Δ (K28115) expressing endogenous 6C cohesin were synchronised in G1, released into a G2/M arrest and subjected to in vivo crosslinking and mini-chromosome IP. The CM and CD bands are marked. Data shown is representative of two independent biological repeats. (B) Wild type (K24697) mrc1Δ (K28276), chl1Δ (K28175), ctf4Δ (K28275), csm3Δ (K28282), tof1Δ (K28280) and ctf18Δ (K28285) strains that contain genes coding for 6C non cleavable cohesin (2C SMC1 2C SMC3 and GALp-2C SCC1NC) were arrested in G2 phase, the expression of 2C Scc1NC was induced by addition of galactose for 45 min. The cultures were released from the G2 arrest and arrested in the subsequent the G1 phase, mini-chromosome IP of the cultures at this stage shows CMs formed in the previous G2 phase being retained in the subsequent G1 in all the strains. The cultures were released from the G1 arrest and allowed to undergo replication and subjected to mini-chromosome IP, the CM and CD bands are marked. A species migrating between the CMs and supercoiled mini-chromosome DNA routinely detected in the ctf18Δ strain is marked with *. We currently do not know the identity of this band. Data shown is representative of two independent biological repeats.
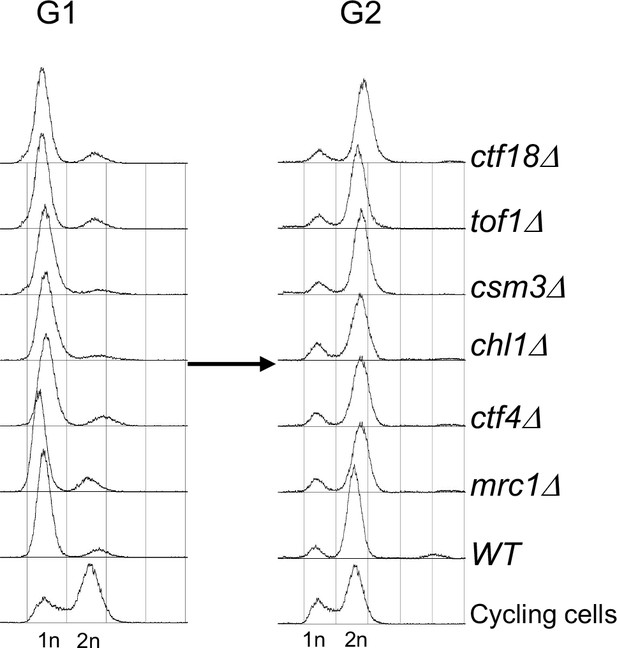
FACS profiles of the cultures described in Figure 3B.
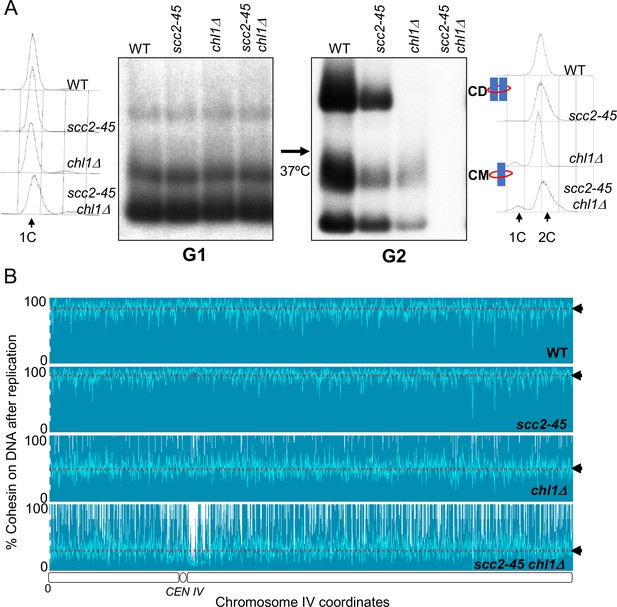
Chromosome associated cohesin is evicted during S phase in the absence of the conversion pathway.
(A) Wild type (K24697), scc2-45 (K24738), chl1Δ (K28175) and chlΔ scc2-45 (K28061) strains that contain genes coding for 6C non cleavable cohesin (2C SMC1 2C SMC3 and GALp-2C SCC1NC) were arrested in G2 phase, the expression of 2C Scc1NC was induced by addition of galactose for 45 min. The cultures were released from the G2 arrest and arrested in the subsequent the G1 phase, mini-chromosome IP of the cultures at this stage shows CMs formed in the previous G2 phase being retained in the subsequent G1 in all the strains. The cultures were released from the G1 arrest and allowed to undergo replication at 37°C (in order to inactivate Scc2 in the scc2-45 strain). Mini-chromosome IP shows formation of CDs in both the wild type, and scc2-45 strains, the chl1Δ strain did not form CD and showed a reduction in CM level and the chlΔ scc2-45 strain failed to IP any mini-chromosome DNA. The FACS profiles of the cultures at different stages of the experiment are shown next to the respective southern blots. Data shown is representative of two independent biological repeats. (B) Samples drawn from the cultures described above during the G1 phase and after replication at 37°C were subjected to calibrated ChIP sequencing with anti-PK antibody to measure Scc1NC levels (See Figure 4—figure supplement 2 for the ChIP profiles). The change in Scc1NC levels on chromosome IV after replication is shown, median level of Scc1NC across the entire chromosome IV (dotted line) is marked with arrowheads.
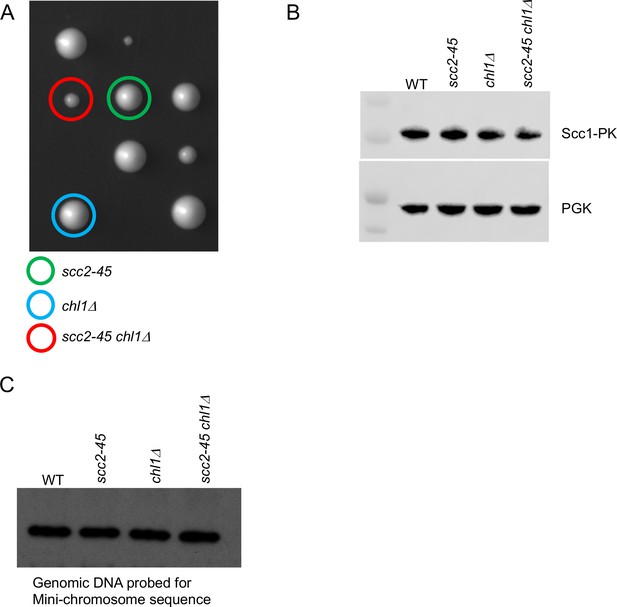
Chromosome associated cohesin is evicted during S phase in the absence of the conversion pathway.
(A) The haploid progeny from crossing a chl1Δ and scc2-45 strain are shown, the genotype of selected haploids is shown. (B) The cultures that described in Figure 4A, after undergoing replication at 37°C were analysed by western blotting against the indicated antibodies. (C) 10 μg of the genomic DNA extracted from the aforementioned cultures were subjected to southern blotting and hybridization with TRP probe to detect mini-chromosome levels.
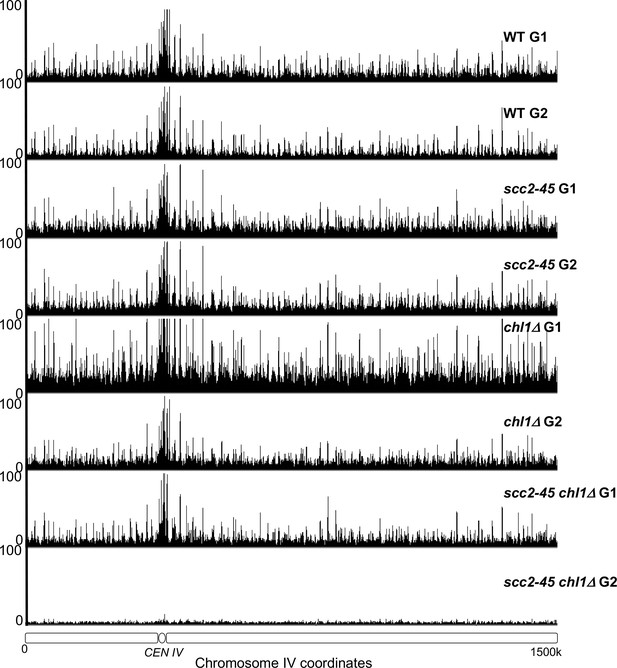
The occupancy of Scc1NC along chromosome IV in the experiment described in Figure 4B.
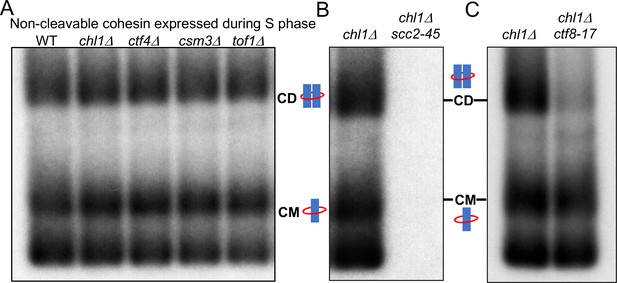
Cohesion can also be established by CTF18-RFC and Scc2 dependent de novo loading of nucleoplasmic cohesin.
(A) Wild type (K24697), chl1Δ (K28175), ctf4Δ (K28275), csm3Δ (K28282) and tof1Δ (K28280) strains that contain genes coding for 6C non cleavable cohesin (2C SMC1 2C SMC3 and GALp-2C SCC1NC) were arrested in G1 phase, the expression of 2C Scc1NC was induced by addition of galactose for 15 min. The cultures were released from the G1 arrest into medium containing galactose in order to ensure constant expression of Scc1NC during DNA replication, mini-chromosome IP of the cultures after replication shows CM and CD formation in all the strains. Data shown is representative of two independent biological repeats. (B) chl1Δ (K28175) and chl1Δ scc2-45 (K28061) strains that contain genes coding for 6C non-cleavable cohesin (2C SMC1 2C SMC3 and GALp-2C SCC1NC) were arrested in G1 phase, the expression of 2C Scc1NC was induced by addition of galactose for 15 min. The cultures were released from the G1 arrest into medium containing galactose in order to ensure constant expression of Scc1NC during DNA replication at 37°C (in order to inactivate Scc2 in the scc2-45 strain). Mini-chromosome IP of the cultures after replication shows CM and CD formation in the chl1Δ while in the chl1Δ scc2-45 we failed to IP any mini-chromosome DNA. Data shown is representative of two independent biological repeats. (C) chl1Δ (K28175) and chl1Δ ctf8-17 (K28295) strains that contain genes coding for 6C non cleavable cohesin (2C SMC1 2C SMC3 and GALp-2C SCC1NC) were arrested in G1 phase, the expression of 2C Scc1NC was induced by addition of galactose for 15 min. The cultures were released from the G1 arrest into medium containing galactose in order to ensure constant expression of Scc1NC during DNA replication at 37°C (in order to inactivate Ctf8 in the ctf8-17 strain). Mini-chromosome IP of the cultures after replication shows CM and CD formation in the chl1Δ. While the chl1Δ ctf8-17 strains showed the presence of CMs, the level of CDs was greatly reduced. Data shown is representative of two independent biological repeats.
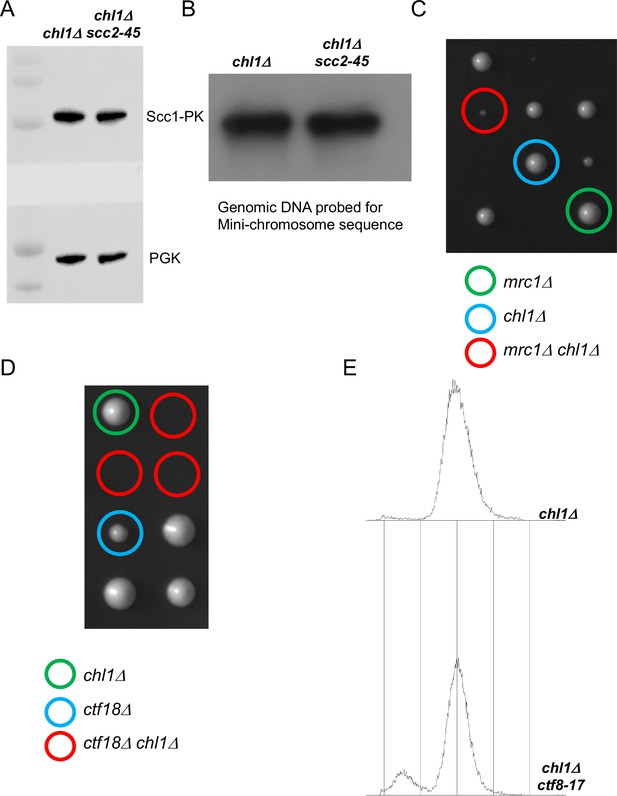
Cohesion can also be established by CTF18-RFC and Scc2 dependent de novo loading of nucleoplasmic cohesin.
(A) chl1Δ (K28175) and chl1Δ scc2-45 (K28061) cultures from the final step of the experiment described in Figure 5B were analysed by western botting against the indicated antibodies and in (B), the genomic DNAs from these cultures were analysed by southern blotting against the TRP probe to detect the mini-chromosome. (C) Tetrad analysis of the haploids resulting from a cross of mrc1Δ and chl1Δ strains, the genotypes of selected haploid spores are indicated. (D) Tetrad analysis of the haploids resulting from a cross of chl1Δ and ctf18Δ strains, the genotypes of selected haploid spores are indicated. (E) FACS profiles of the chl1Δ (K28175) and chl1Δ ctf8-17 (K28295) cultures from the end of the experiment described in Figure 5C.
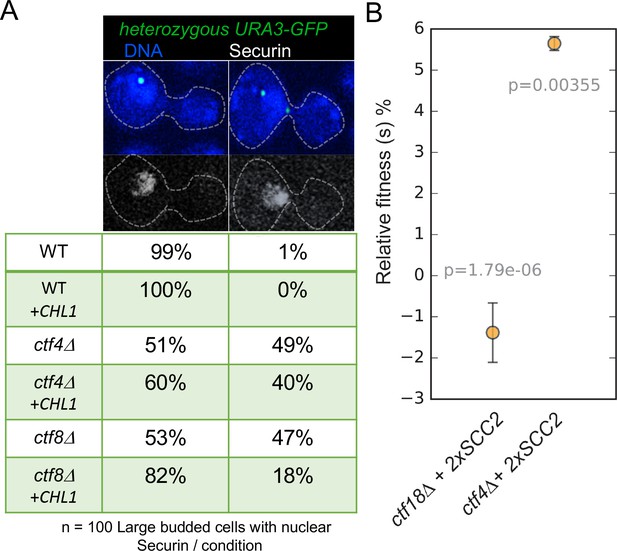
Augmentation of either cohesion establishment pathway suppresses the cohesion defects caused by abrogation of the other.
(A) Asynchronous cultures of wild-type (K10003), ctf4Δ (K11692), ctf8Δ (K10349) that carry either an empty 2μ vector or a 2μ vector containing the CHL1 gene were fixed and stained for DNA and with an antibody against the Myc epitope as detailed in Materials and methods. The strains express Pds1-Myc18 (securin) and contain chromosome V marked by GFP at the URA3 locus, 35 kb away from the centromere. Fluorescence images of representative large budded metaphase cell showing one or two URA3-GFP dots (green), nuclear securin (white) and DNA (blue) is shown. The proportion of securin-positive cells containing one or two URA3-GFP signals in the different strains is listed below the images. 100 metaphase cells were scored for each strain in this experiment. (B) The fitness of ancestral ctf4Δ and ctf18Δ strains carrying an extra copy of the SCC2 gene, relative to the ctf4Δ and ctf18Δ ancestors respectively. Error bars represent standard deviation, the p values from unequal variances t-tests are also shown. The source data is presented in Source data 1.
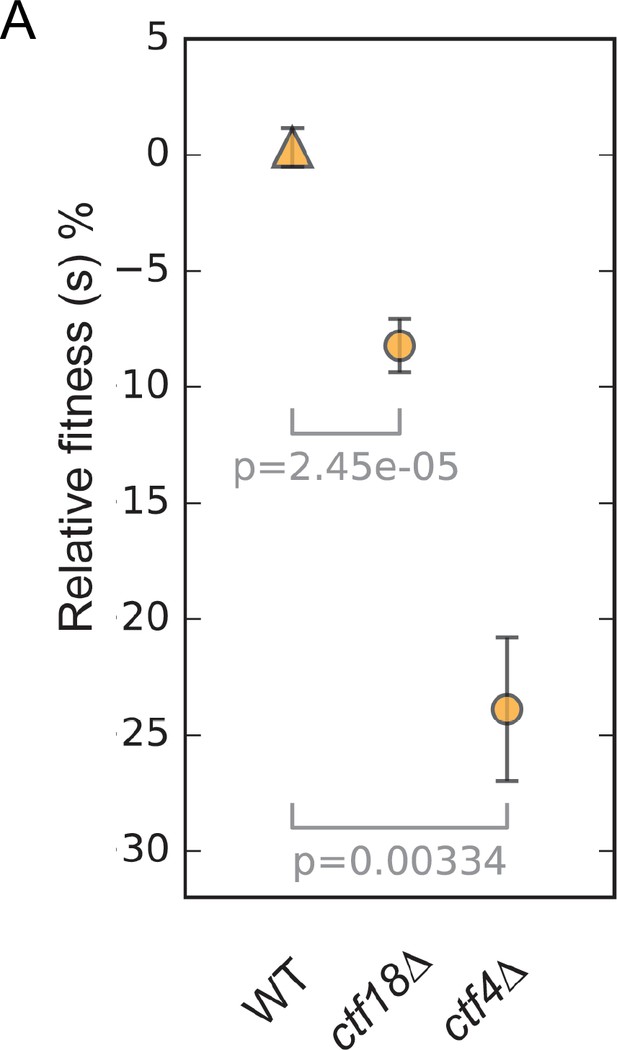
The fitness of wild type, ctf18Δ and ctf4Δ strains relative to the wild-type ancestor.
Error bars represent standard deviation, the p values from unequal variances t-tests are also shown.
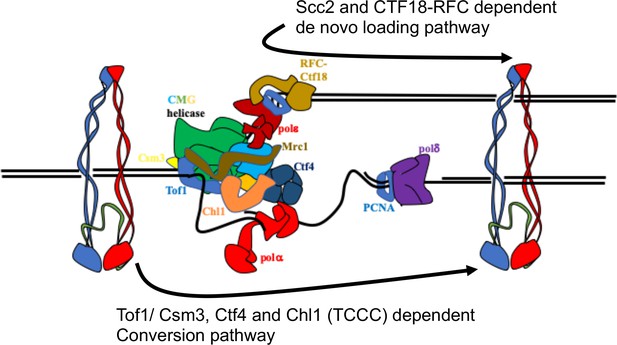
Two parallel pathways for cohesion establishment at the replication fork: chromosome associated cohesin is converted into cohesive structures during S phase.
This process does not require Scc2 (except to load cohesin in the first place), but is strictly dependent on Tof1, Csm3, Chl1, and Ctf4 (the TCCC pathway). A second pathway that operates in parallel is concerned exclusively with de novo loading of cohesin onto nascent DNAs, in a Ctf18-RFC and Scc2-dependent manner (de novo pathway). See Discussion for details.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (S. cerevisiae) | NCBITaxon:4932 | This paper | Yeast strains | Supplementary file 1 |
| Antibody | Mouse monoclonal Anti-V5 | BioRad | Cat# MCA1360 | (1:1000) |
| Antibody | Mouse monoclonal Anti PGK1 | ThermoFisher Scientific | Cat#459250 | (1:5000) |
| Chemical compound | Acid-washed glass beads | Sigma | Cat# G8722 | N/A |
| Chemical compound | ATP α-32P | Hartmann Analytic | Cat# SRP-203 | N/A |
| Chemical compound | Bismaleimidoethane (BMOE) | ThermoFisher | Cat# 22323 | (5 mM) |
| Chemical compound | Complete EDTA free protease inhibitor cocktail | Roche | Cat# 4693132001 | (1:50 ml) |
| Chemical compound | Dithiothreitol | Fluka | Cat# BP172 | (5 mM) |
| Chemical compound | DMSO | Sigma | Cat# D8418 | N/A |
| Chemical compound | Immobilon Western ECL | Millipore | Cat# WBLKS0500 | N/A |
| Chemical compound | Trisodium citrate | Sigma | Cat# W302600 | N/A |
| Chemical compound | RNase A | Roche | Cat# 10109169001 | N/A |
| Chemical compound | Nocodazole | Sigma | Cat# M1404 | N/A |
| Chemical compound | PMSF | Sigma | Cat# 329-98-6 | N/A |
| Chemical compound | Potassium chloride | Sigma | Cat# P5405 | N/A |
| Chemical compound | Proteinase K | Roche | Cat# 03115836001 | N/A |
| Chemical compound | Sodium sulfite | Sigma | Cat# 71988 | N/A |
| Biological Sample | α-factor peptide | CRUK Peptide Synthesis Service | N/A | N/A |
| Commercial Assay or Kit | ChIP Clean and Concentrator Kit | Zymo Research | Cat# D5205 | N/A |
| Commercial Assay or Kit | E-Gel SizeSelect II Agarose Gels, 2% | ThermoFisher | Cat# G661012 | N/A |
| Commercial Assay or Kit | Library Quantification Kit Ion Torrent Platforms | KAPA Biosystems | Cat# 28-9537-67 | N/A |
| Commercial Assay or Kit | NEBNext Fast DNA library prep set for Ion Torrent | NEB | Cat# Z648094 | N/A |
| Commercial Assay or Kit | NuPAGE 3–8% Tris-Acetate Protein Gels, 1.5 mm, 10-well | ThermoFisher | Cat# E6270L | N/A |
| Commercial Assay or Kit | Prime-it II Random Primer Labelling Kit | Agilent | Cat# NP0321BOX | N/A |
| Commercial Assay or Kit | Protein G dynabeads | ThermoFisher | Cat# 300385 | N/A |
| Software, algorithm | Galaxy platform | Giardine et al., 2005 | ||
| Software, algorithm | FastQC | Galaxy tool version 1.0.0 | https://usegalaxy.org | N/A |
| Software, algorithm | Trim sequences | Galaxy tool version 1.0.0 | https://usegalaxy.org | N/A |
| Software, algorithm | Filter FASTQ | Galaxy tool version 1.0.0 | https://usegalaxy.org | N/A |
| Software, algorithm | Bowtie2 | Langmead and Salzberg, 2012 Galaxy tool version 0.2 | https://usegalaxy.org | N/A |
| Software, algorithm | Bam to BigWig | Galaxy tool version 0.1.0 | https://usegalaxy.org | N/A |
| Software, algorithm | Samtools | Li et al., 2009 | https://usegalaxy.org | N/A |
| Software, algorithm | IGB browser | Nicol et al., 2009 | http://samtools.sourceforge.net/ | N/A |
| Software, algorithm | Filter SAM or BAM | Li et al., 2009 Galaxy tool version 1.1.0 | http://bioviz.org/igb/ | N/A |
| Software, algorithm | chr_position.py | This study | https://usegalaxy.org | N/A |
| Software, algorithm | filter.py | This study; Petela, 2019 | https://github.com/naomipetela/nasmythlab-ngs | N/A |
| Software, algorithm | bcftools call | Li et al., 2009 | https://github.com/naomipetela/nasmythlab-ngs | N/A |
Additional files
-
Source data 1
Source data for the Relative fitness assay.
- https://cdn.elifesciences.org/articles/56611/elife-56611-data1-v1.xlsx
-
Supplementary file 1
Strain list.
- https://cdn.elifesciences.org/articles/56611/elife-56611-supp1-v1.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/56611/elife-56611-transrepform-v1.docx






