Structural basis for capsid recruitment and coat formation during HSV-1 nuclear egress
Figures
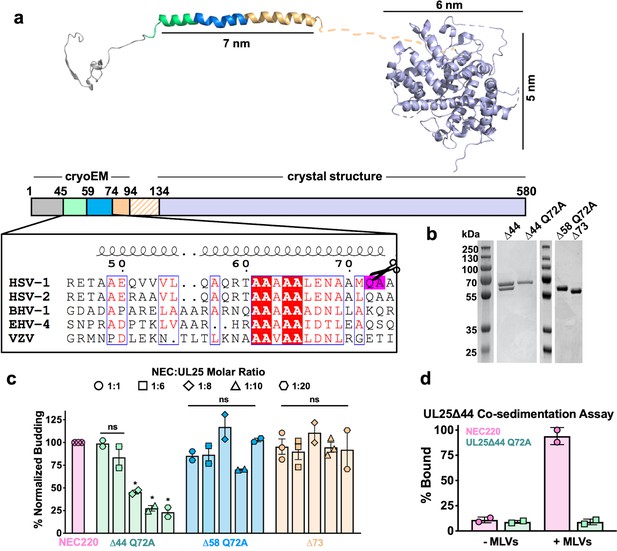
Inhibition of NEC-mediated budding by UL25 constructs.
(a) The UL25 structure and a diagram of domain organization is shown along with a multiple sequence alignment of UL25 residues 45–74 from five alphaherpesviruses. Sequence alignment was generated using Clustal Omega45 and displayed using ESPript 3.046. Identical residues are shown as white letters on a red background. Similar residues are shown as red letters in a blue box. Secondary structure derived from the cryoEM reconstruction of capsid-bound HSV-1 UL25 is shown above the alignment. The following herpesvirus sequences were used (GenBank GeneID numbers in parentheses): HSV-1, herpes simplex virus type 1, strain 17 (2703377); HSV-2, herpes simplex virus type 2, strain HG52 (1487309); BHV-1, bovine herpesvirus-1 (4783418); EHV-4, equine herpesvirus-4, strain NS80567 (1487602); and VZV, varicella-zoster virus, strain Dumas (1487687). (b) SDS-PAGE of purified UL25 constructs: UL25∆44 (cleaved product; 57 kDa), UL25∆44 Q72A (single product; 57 kDa), UL25∆58 Q72A (56 kDa) and UL25∆73 (54 kDa). (c) UL25∆44 Q72A inhibits NEC budding, whereas other UL25 constructs do not. For each condition, NEC-mediated budding was tested at 1:1, 1:6, 1:8, 1:10, and 1:20 NEC:UL25 molar ratios. Each construct was tested in at least two biological replicates, consisting of three technical replicates. Symbols show average budding efficiency of each biological replicate relative to NEC220 (100%; pink). Error bars represent the standard error of measurement for at least two individual experiments. Significance compared to NEC220 was calculated using an unpaired t-test against NEC220. *p-value<0.1. The source file with all raw data values is provided in Figure 1—source data 1. (d) UL25∆44 Q72A does not bind to acidic lipid membranes.
-
Figure 1—source data 1
Raw data and background values collected for GUV budding assays of NEC220 in the presence of either UL25∆44 Q72A, UL25∆58 Q72A or UL25∆73.
The reported biological replicate average values (%) are the points presented in Figure 1.
- https://cdn.elifesciences.org/articles/56627/elife-56627-fig1-data1-v2.xlsx
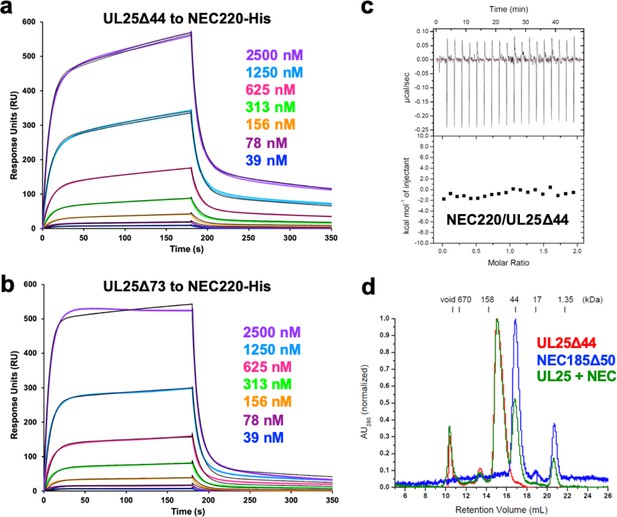
NEC-UL25 binding studies.
SPR binding of UL25Δ44 Q72A (a) and UL25Δ73 (b) to NEC220-His indicating UL25 can bind the NEC if the NEC is able to form a type of scaffold. (c) ITC of NEC220 and UL25Δ44 showing these two proteins do not bind in solution. (d) Size-exclusion chromatography of NEC185Δ50 (crystallization construct) and UL25Δ44 shows these proteins also do not bind in solution.
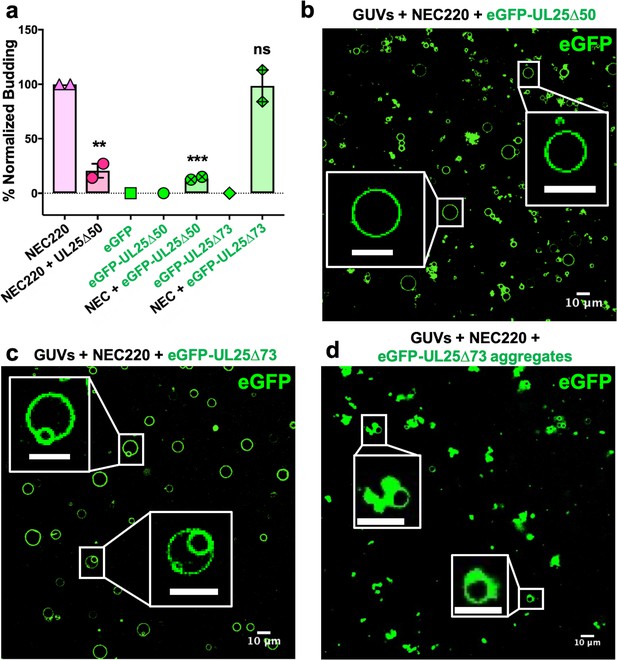
eGFP-UL25∆50 inhibits NEC budding while eGFP-UL25∆73 does not.
(a) Quantification of NEC budding in the presence of either eGFP-UL25∆50 or eGFP-UL25∆73. Each construct (except in the absence of NEC220) was tested in at least two biological replicates, each consisting of three technical replicates. Symbols show the average budding efficiency of each biological replicate relative to NEC220 (100%). Error bars represent the standard error of measurement for at least two individual experiments. Significance compared to NEC220 was calculated using an unpaired t-test against NEC220. **p-value<0.01 and ***p-value<0.001. The source file with all raw data values is provided in Figure 2—source data 1. (b) Confocal image of eGFP-UL25∆50 bound to NEC-coated vesicles. No budding is observed. (c) Confocal image of eGFP-UL25∆73 either bound to or budded into vesicles with the NEC. (d) Confocal image of eGFP-UL25∆73 aggregating on the surface of NEC-coated vesicles. All scale bars = 10 μm.
-
Figure 2—source data 1
Raw data and background values collected for GUV budding assays of NEC220 in the presence of either UL25∆50 Q72A, eGFP-UL25∆50 Q72A or eGFP-UL25∆73.
The reported biological replicate average values (%) are the points presented in Figure 2.
- https://cdn.elifesciences.org/articles/56627/elife-56627-fig2-data1-v2.xlsx
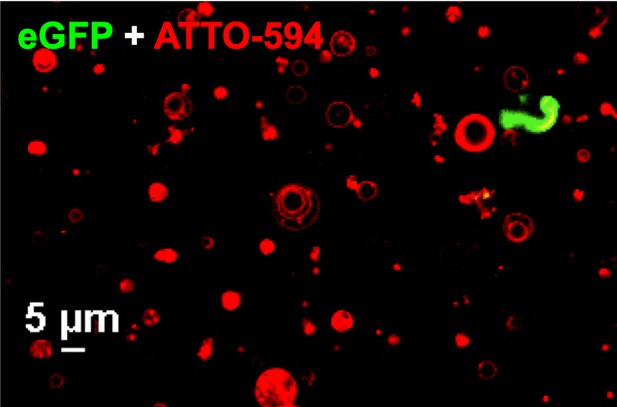
Confocal image of GUVs (red) and eGFP-UL25∆50 Q72A (green) showing that eGFP-UL25∆50 Q72A does not bind GUV membranes.
Both the ATTO-594 and eGFP channels are shown. The scale bar represents 5 μm.
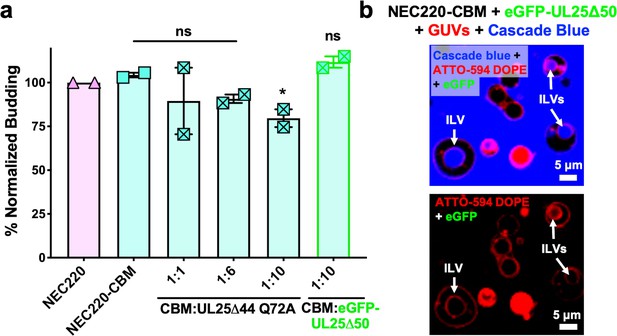
UL25 inhibits NEC220-CBM budding to a lesser extent.
(a) NEC220-CBM budding is not inhibited to the same extent as NEC220 budding by either UL25Δ44 Q72A or eGFP-UL25Δ50 Q72A. Budding was tested at 1:1, 1:6 and 1:10 NEC220-CBM:UL25 molar ratios for UL25Δ44 Q72A and at a 1:10 NEC-CBM:UL25 molar ratio for eGFP-UL25Δ50 Q72A. Each condition was tested in at least two biological replicates, each consisting of three technical replicates. Symbols represent average budding efficiency of each biological replicate relative to NEC220 (100%). Error bars represent the standard error of measurement for at least two individual experiments. Significance compared to NEC220 was calculated using an unpaired t-test against NEC220. *p-value<0.1. The source file with all raw data values is provided in Figure 3—source data 1. (b) Confocal microscopy image showing eGFP-UL25Δ50 Q72A does not bind to NEC220-CBM coated GUVs as indicated by the lack of green signal on the membranes of intraluminal vesicles (ILVs) formed by NEC220-CBM budding (indicated by white arrows). Top panel shows red (ATTO-594 DOPE), green (eGFP), and blue (Cascade Blue) channels. Bottom panel shows red (ATTO-594 DOPE) and green (eGFP) channels only. Scale bars = 5 μm.
-
Figure 3—source data 1
Raw data and background values collected for GUV budding assays of NEC220-CBM in the presence of either UL25∆44 Q72A or GFP-UL25∆50 Q72A.
The reported biological replicate average values (%) are the points presented in Figure 3.
- https://cdn.elifesciences.org/articles/56627/elife-56627-fig3-data1-v2.xlsx
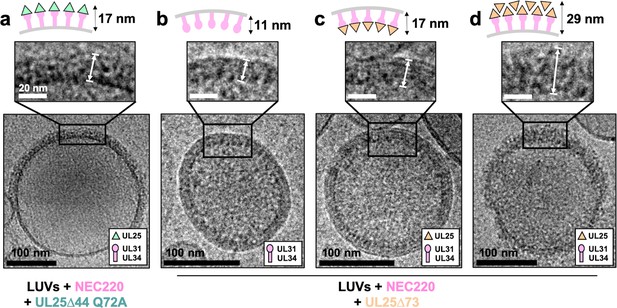
CryoEM shows UL25∆44 Q72A inhibits NEC220 budding while UL25∆73 does not.
(a) UL25∆44 Q72A bound to the NEC220 on the outside of the unbudded lipid vesicles, forming a fence-like array (~17 nm). In the presence of UL25∆73, three scenarios have been observed: (b) NEC220 alone bound to the inner surface of the budded lipid vesicles (~11 nm); (c) UL25∆73 bound to the NEC220, which is itself bound to the inner surface of the budded lipid vesicles (~17 nm), and (d) UL25∆73 aggregates bound to the NEC on the outside of the unbudded lipid vesicles (>29 nm). Budded lipid vesicles in panels b and c are no longer contained within a ‘mother’ lipid vesicle and represent the end-product of budding. Scale bars = 100 nm. Inset scale bars = 20 nm. All inset panels are shown on the same scale. White arrows in insets define measurement boundaries of vesicle-bound proteins displayed in the corresponding cartoon models.
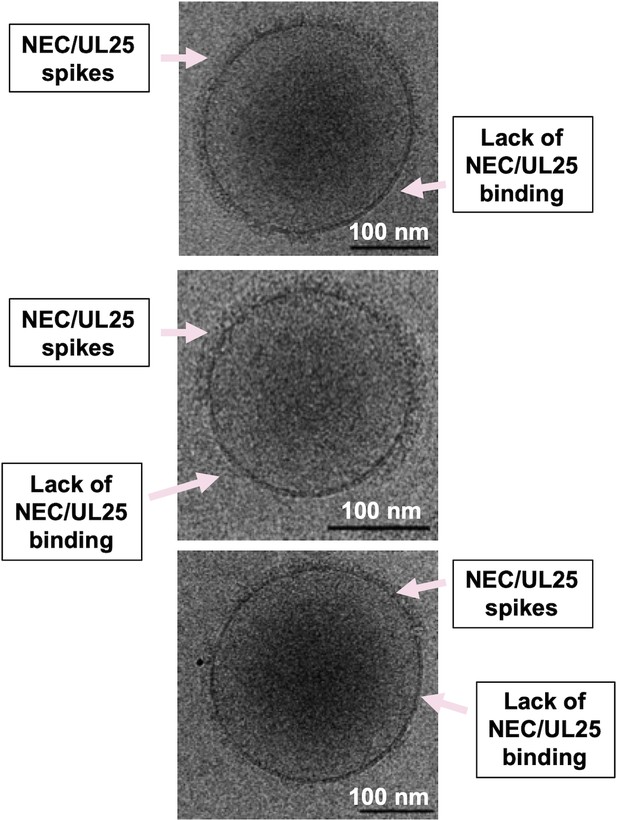
Incomplete distribution of NEC-UL25 around vesicles.
Slices of selected tomograms of NEC220/UL25∆44 Q72A-bound vesicles used to generate cryoET averages in Figure 5. Regions of either protein binding or lack of protein binding are indicated by pink arrows. Scale bar represents 100 nm.
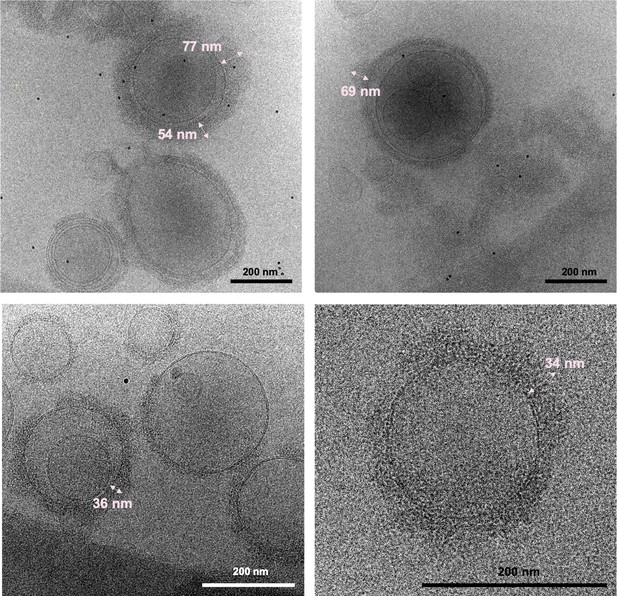
Slices of selected tomograms of NEC220/UL25∆73-bound vesicles.
Selected regions of UL25∆73 aggregation are indicated by brackets along with the corresponding measurements of each selection. Scale bars represent 200 nm.
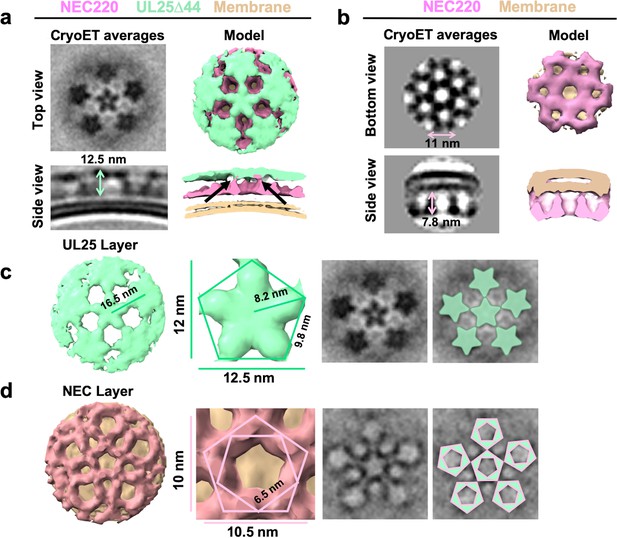
CryoET of UL25-mediated inhibition of NEC budding.
(a) CryoET averages of NEC220 in the presence of UL25Δ44 Q72A (top and side views). Corresponding 3D models are shown with NEC220 (pink) and UL25Δ44 Q72A (green). The vesicle bilayer is shown in beige. The models show the UL25 layer coating the NEC layer in five-pointed stars on the outside of the vesicles. The length of the NEC-UL25 spikes is 12.5 nm. Black arrows indicate the point of tilt within the NEC layer. (b) CryoET averages of NEC220 forming hexameric lattices in the presence of membranes (bottom and side views). Corresponding 3D models are shown with NEC (pink) and the vesicle bilayer (beige). The diameter of the hexameric rings is ~11 nm, while the length of the spikes is 7.8 nm. (c) CryoET model and averages of the UL25 layer (green) highlighting the five-pointed star formation of UL25 (represented here as a pentamer of dimers) in the presence of NEC. (d) CryoET model and averages of the NEC layer showing NEC220 forming a pentagonal lattice (pink pentagons), rather than hexagonal (as seen for wild-type in panel b). Green triangles indicate location of UL25 binding to the NEC.
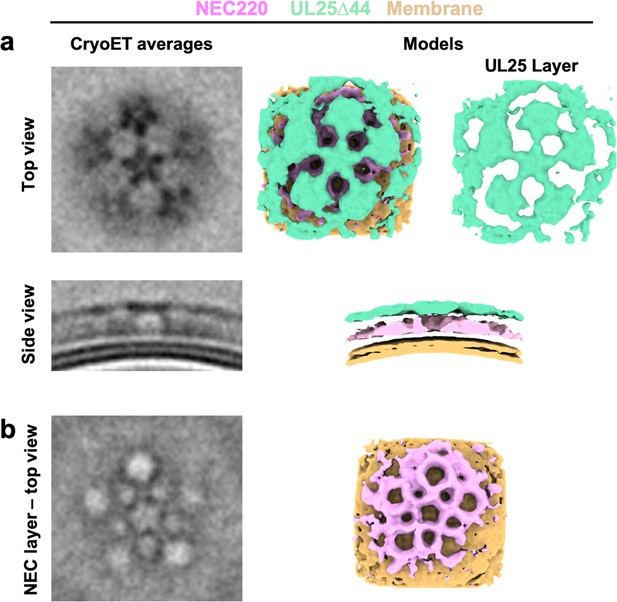
CryoET averages of NEC220 in the presence of UL25Δ44 Q72A (top and side views) prior to applying five-fold symmetry.
Corresponding 3D models are shown with NEC220 (pink) and UL25Δ44 Q72A (green). The vesicle bilayer is shown in beige. The models show the UL25 layer coating the NEC layer in five-pointed stars on the outside of the vesicles (a) and that the NEC layer forms a pentagonal lattice (b), similar to what is observed after symmetry is applied.
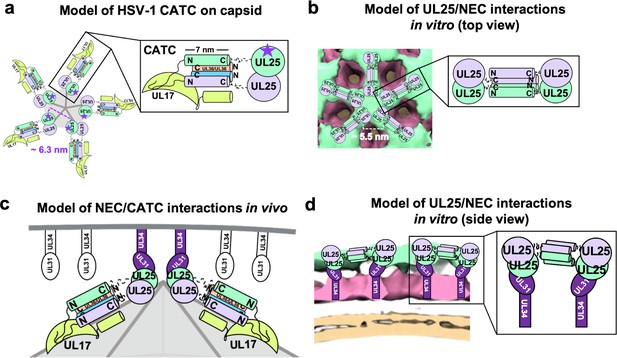
Models of UL25/UL25 and UL25/NEC interactions in vitro and in vivo.
(a) A schematic representation of the pentagonal HSV-1 CATC [two copies of UL25 (green and purple), two copies of C-terminal UL36 (peach and blue) and one copy of UL17 (lime green)] arrangement at the capsid vertex. Inset shows a close-up view of the characteristic antiparallel four-helix bundle composed of two UL25 helices and two UL36 helices. Purple stars indicate the proposed UL25 copies that bind to the NEC upon capsid docking. The distance between the centers of two adjacent inner UL25 cores (green) in the capsid (Dai and Zhou, 2018) is ~6.3 nm. (b) Proposed model of the UL25 stars formed in vitro. The distance between the centers of two adjacent UL25 dimers is ~5.5 nm. Inset shows a close-up view of the proposed antiparallel four-helix bundle composed of two pairs of UL25 helices from adjacent stars. We hypothesize that four-helix bundles link the neighboring UL25 stars into a net. (c) Proposed side-view model of the NEC (purple) interacting with the most surface exposed capsid-bound UL25 (green), resulting in a pentameric NEC (indicated by dark purple coloring). NEC molecules prior to capsid binding are shown in an unknown oligomeric state (white). (d) Side view of the proposed NEC/UL25 interactions in vitro.
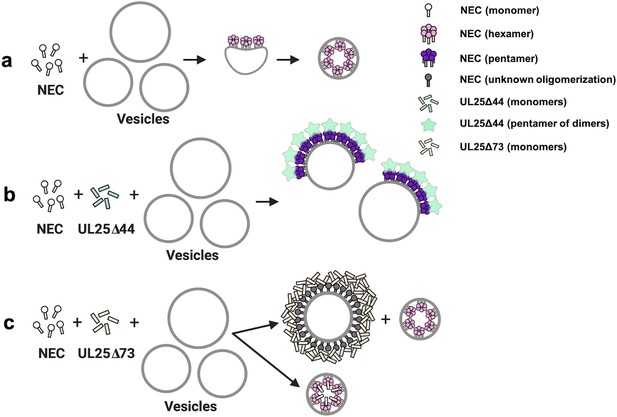
A model of NEC-mediated budding in the absence and presence of UL25, in vitro.
(a) NEC-mediated budding requires only the NEC, which vesiculates membranes by forming hexagonal coats (pink) that, potentially, contain irregular defects to achieve curvature. (b) UL25∆44 Q72A (green) inhibits NEC-mediated budding by inducing the formation of a pentagonal NEC coat (purple) suboptimal for budding. (c) UL25∆73 (peach) aggregates around some NEC-coated vesicles, which blocks budding. Sequestration of UL25∆73 at a few locations reduces its concentration elsewhere and enables budding. Binding of UL25∆73 to NEC in the absence of aggregation does not interfere with budding, and bound UL25∆73 buds into vesicles with the NEC. This figure was created with Biorender.com.
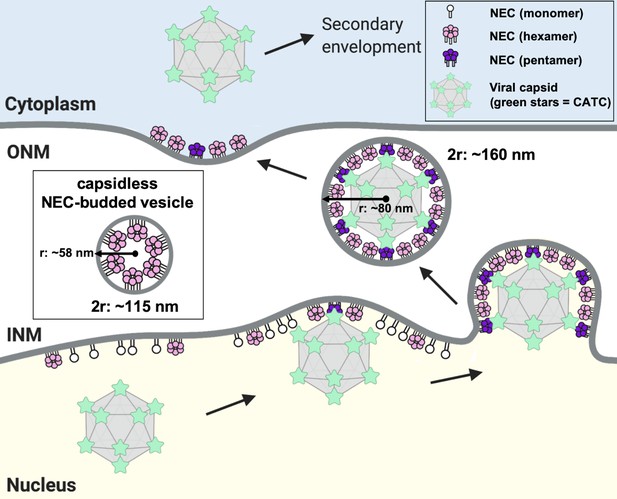
A model of NEC-mediated budding in HSV-1 infected cells.
Capsid-bound UL25 induces the formation of pentagonal insertions (purple pentamers) within the NEC coat (pink hexamers and white monomers) as it is forming, which enables the formation of an NEC coat of appropriate size and curvature around the capsid. Inset shows a capsidless perinuclear vesicle formed in NEC-expressing uninfected cells that forms a hexagonal coat with presumably irregular defects, similar to the NEC coat formed in vitro. This figure was created with Biorender.com.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (HSV-1 KOS) | UL25 | Geneart | JQ673480.1 | |
| Strain, strain background (Escherichia coli) | BL21(DE3) | Kerafast | LoBSTr | Chemically competent cells |
| Recombinant DNA reagent | eGFP-N2 (plasmid) | Clontech | eGFP-N2 | |
| Recombinant DNA reagent | pKH90 (plasmid) | PMID:24916797 | UL31 1–306 | |
| Recombinant DNA reagent | pJB02 (plasmid) | PMID:24916797 | UL34 1–220 | |
| Recombinant DNA reagent | pJB104 (plasmid) | This paper | UL25∆44 | See Materials and methods, Cloning |
| Recombinant DNA reagent | pJB118 (plasmid) | This paper | NEC-CBM | See Materials and methods, Cloning |
| Recombinant DNA reagent | pJB123 (plasmid) | This paper | UL25∆73 | See Materials and methods, Cloning |
| Recombinant DNA reagent | pED03 (plasmid) | This paper | UL25∆44 Q72A | See Materials and methods, Cloning |
| Recombinant DNA reagent | pED05 (plasmid) | This paper | eGFP-UL25∆73 | See Materials and methods, Cloning |
| Recombinant DNA reagent | pED13 (plasmid) | This paper | UL25∆50 | See Materials and methods, Cloning |
| Recombinant DNA reagent | pED14 (plasmid) | This paper | eGFP-UL25∆50 | See Materials and methods, Cloning |
| Peptide, recombinant protein | UL25∆44 | This paper | Purified from E. coli BL21(DE3) LoBSTr cells | |
| Peptide, recombinant protein | UL25∆44 Q72A | This paper | Purified from E. coli BL21(DE3) LoBSTr cells | |
| Peptide, recombinant protein | UL25∆50 Q72A | This paper | Purified from E. coli BL21(DE3) LoBSTr cells | |
| Peptide, recombinant protein | UL25∆58 Q72A | This paper | Purified from E. coli BL21(DE3) LoBSTr cells | |
| Peptide, recombinant protein | UL25∆73 | This paper | Purified from E. coli BL21(DE3) LoBSTr cells | |
| Peptide, recombinant protein | eGFP-UL25∆50 Q72A | This paper | Purified from E. coli BL21(DE3) LoBSTr cells | |
| Peptide, recombinant protein | eGFP-UL25∆73 | This paper | Purified from E. coli BL21(DE3) LoBSTr cells | |
| Peptide, recombinant protein | NEC220 | This paper | Purified from E. coli BL21(DE3) LoBSTr cells | |
| Peptide, recombinant protein | NEC-CBM | This paper | Purified from E. coli BL21(DE3) LoBSTr cells | |
| Chemical compound, drug | Cascade Blue hydrazide | Thermo Fisher Scientific | Thermo Fisher Scientific: C687 | |
| Software, algorithm | ImageJ | ImageJ | RRID:SCR_003070 | |
| Software, algorithm | IMOD | IMOD | RRID:SCR_003297 | |
| Other | 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphate | Avanti Polar Lipids | Avanti Polar Lipids:850857 | POPA |
| Other | 1-palmitoyl-2-oleoyl-glycero-3-phosphocholine | Avanti Polar Lipids | Avanti Polar Lipids:850457 | POPC |
| Other | 1-palmitoyl-2-oleoyl-sn-glycero-3-phospho-L-serine | Avanti Polar Lipids | Avanti Polar Lipids:840034 | POPS |
Additional files
-
Supplementary file 1
Supplementary Tables 1 and 2.
UL25∆44 Q72A/NEC particles used for cryoET averaging. List of primers used for cloning procedures described in Materials and methods.
- https://cdn.elifesciences.org/articles/56627/elife-56627-supp1-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/56627/elife-56627-transrepform-v2.docx





