Interaction mapping of endoplasmic reticulum ubiquitin ligases identifies modulators of innate immune signalling
Figures
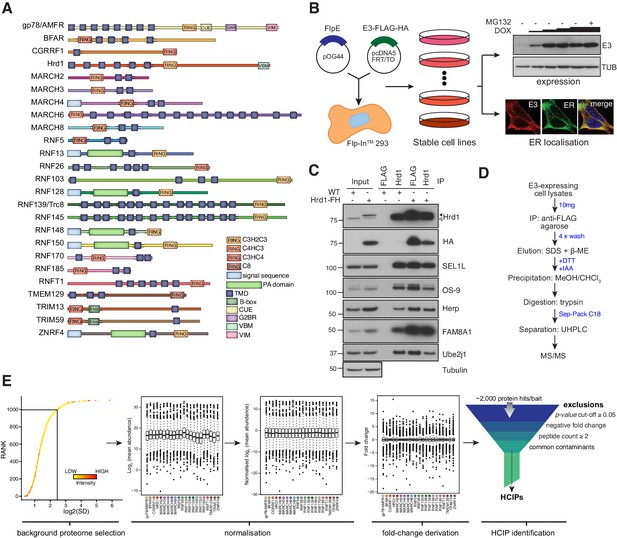
Proteomic analysis of ER-resident ubiquitin ligases.
(A) ER-resident E3s and their predicted domains. (B) Workflow to generate and validate Flp-In293 cell lines stably expressing FLAG-HA-tagged E3s (FH-E3 or E3-FH). Each Flp-In293 cell line stably integrating a tagged E3 was screened for induction and expression over increasing concentrations of DOX and MG132 treatment by western blot (anti-FLAG) as well residency in the ER by immunofluorescence, evaluating colocalisation with markers of the ER, calnexin or KDEL. (C) Co-immunoprecipitation profiles of endogenous Hrd1 and DOX-induced Hrd1-FH prepared in 1% LMNG and isolated by anti-Hrd1 or anti-FLAG, as indicated. Input (20% of total IP) is also shown. (D) Workflow of sample preparation for LC-MS/MS analysis. (E) Bioinformatic processing pipeline for identification of high-confidence candidate interacting proteins (HCIPs) for the E3 baits.
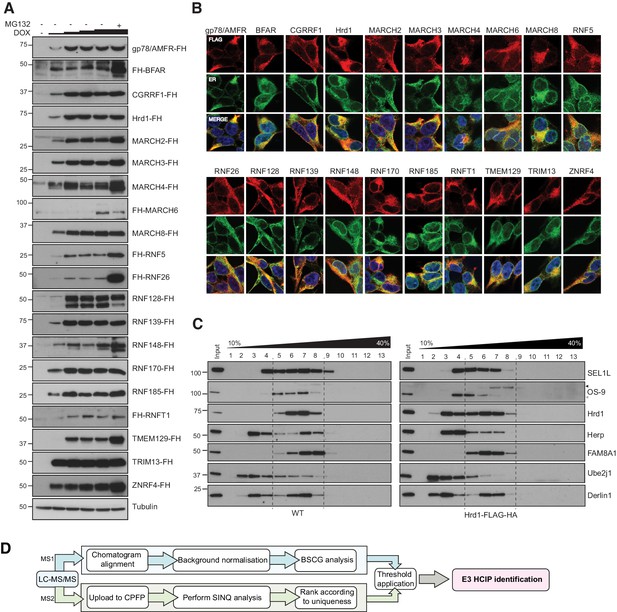
Expression and localisation of FLAG-HA-E3s.
(A) Doxycycline (DOX) titrations (1, 10, 100, 1000 ng/mL; 18 hr) to optimise expression conditions of individual FH-E3s in stable Flp-In293 cell lines. MG132 (10 μM, final 4 hr) was also included in one sample (DOX, 1000 ng/mL) to evaluate proteasomal degradation. Cells were lysed in RIPA buffer (50 mM Tris-HCl pH7.4, 0.1% SDS, 1% sodium deoxycholate, 1% NP-40, 2 mM EDTA, 150 mM NaCl) supplemented with cOmplete protease inhibitor cocktail (Roche), lysates separated by SDS-PAGE and western blots probed for each E3 (anti-FLAG/anti-HA). (B) Representative immunofluorescence images for each of the generated Flp-In293 cell lines. DOX-induced cell lines (18 hr) were co-stained with primary antibodies to the FLAG epitope (red) and the ER-resident protein calnexin or KDEL (ER, green). Nuclei (blue) are shown in the merged image (C) Velocity sedimentation of endogenous Hrd1 and DOX-induced Hrd1-FH complexes on 10–40% sucrose gradients. Both samples and gradients were prepared in 1% LMNG. (D) Schematic of proteomic pipeline used to determine HCIPs for ER-resident E3s.
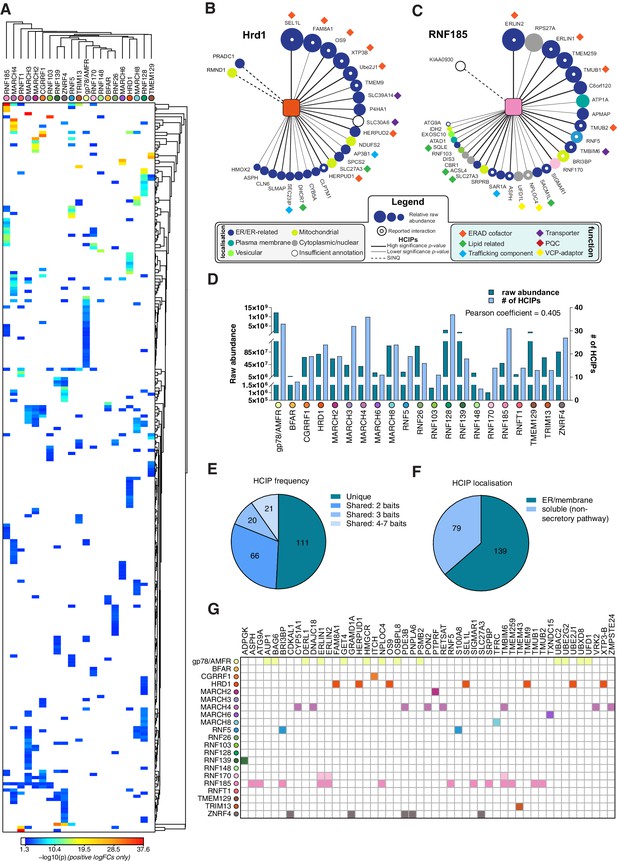
Interaction landscape of 21 ER-resident E3s.
(A) Hierarchical clustering of the 21 E3s and their associated HCIPs represented as a heat map, where the colours of individual interactors correspond to their calculated p-values. Representative HCIP interaction wheels for (B) Hrd1 and (C) RNF185. Parameters represented are described in the adjoining legend. (D) Plot showing raw abundance (RA) and number of HCIPs determined for each E3 with the Pearson coefficient indicated. (E) Distribution of HCIP interactions with E3s as unique or shared. (F) Classification of HCIPs as ER/membrane or soluble, non-secretory pathway proteins as defined by presence of validated and predicted signal peptides, glycosylation sites, disulphide bonds, and transmembrane domains (UniProt). (G) E3-HCIP interactions identified previously in BioGRID 3.5 and BioPlex 3.0.
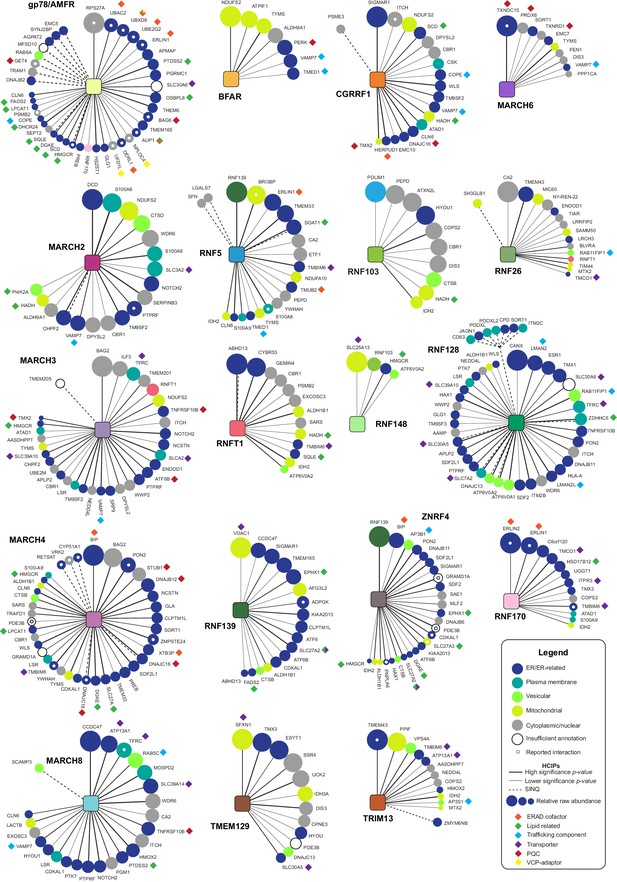
HCIP interaction network wheels for individual ER-E3s.
Protein-protein interaction wheels representing individual ER-resident E3 networks. Each HCIP is represented by a circle whose diameter reflects its raw abundance relative to the E3’s most abundant HCIP. Spoke/line thickness represents the p-value determined for each E3-HCIP interaction. Solid and dashed lines represent interactors identified by BCSG and SINQ analysis, respectively. Classifications of HCIP subcellular localisation are indicated by circle colour (described in legend), which have been manually curated using protein databases (e.g. UniProt) or when unavailable, assessed based on the predicted presence of organelle targeting features (e.g. signal sequence). HCIP function or related process, when known, is denoted by a proximal diamond (described in legend). HCIPs reported previously in BioGRID 3.5 and BioPlex 3.0 are indicated by a central white dot.
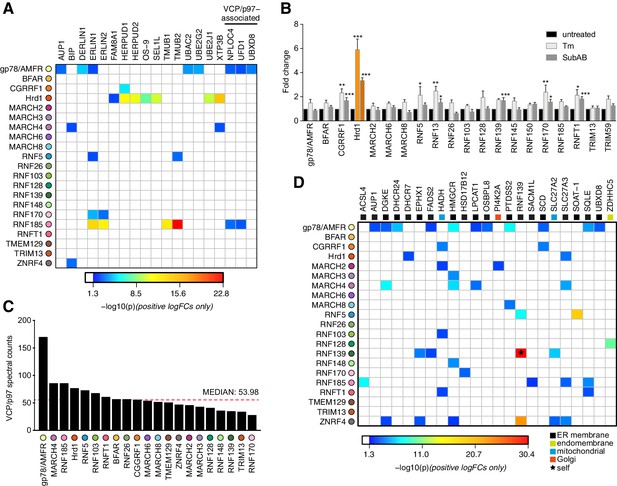
Functional associations of E3 and HCIPs.
(A) Heat map depicting established ERAD components found as HCIPs with the panel of ER-resident E3s, with colours of individual interactors corresponding to their calculated p-values. (B) Transcriptional analysis of parental Flp-In293 cells determined by NanoString. Data depict fold change of E3 transcripts measured from tunicamycin-treated (Tm, 500 ng/mL, 8 hr) and SubAB-treated (SubAB, 10 ng/mL, 8 hr) cells when compared to untreated. Mean and S.E.M. are shown from three biological repeats (n = 3). *p<0.05, **p<0.01, ***p<0.001. Detailed statistical analysis can be found in Supplementary file 1, Table 12. Hrd1 is highlighted in orange for reference. (C) Absolute number of spectral counts detected for VCP/p97 for each ER-resident E3 determined by SINQ analysis. The dotted red line shows the median spectral counts for reference (D) Heat map representing the association between proteins involved in lipid regulation (synthesis; metabolism and transport) and E3 baits. Colours associated with individual interactors correspond to their calculated p-values.
-
Figure 3—source data 1
Nanostring counts obtained for E3 transcripts in response to ER stressors used to generate Figure 3B.
- https://cdn.elifesciences.org/articles/57306/elife-57306-fig3-data1-v1.xlsx
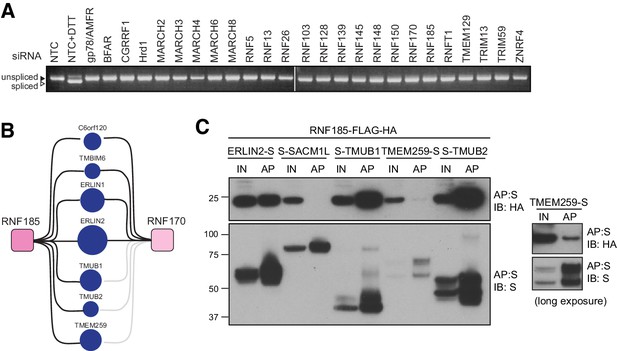
Validation of ER-resident E3 interactions.
(A) Induction of ER stress as determined by splicing of XBP1 in Flp-In293 cells knocked down for individual ER-resident E3s by siRNAs. Splicing was validated by treatment of siNTC (non-targeting control)-transfected cells with DTT (5 mM, 2 hr). (B) Diagram representing shared HCIPs of RNF185 and RNF170. (C) Validation of RNF185 HCIPs. Transient expression of S-tagged HCIPs in Flp-In293 cells stably expressing FH-RNF185. Complexes were affinity purified from 1% LMNG-solubilised lysates by S-protein agarose, separated by SDS-PAGE and the resulting western blots probed for RNF185 (anti-HA) and the HCIP (anti-S-tag). Because of weaker expression, TMEM259-S western blots are also presented in a longer exposure. Input (IN, 20%) and affinity purified (AP) material are shown.
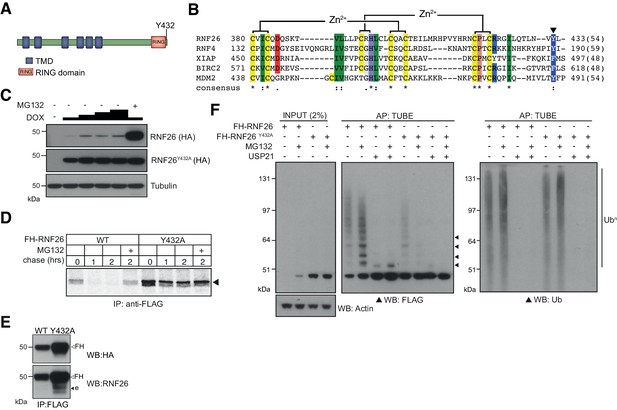
Characterisation of the RNF26 ubiquitin ligase complex.
(A) Domain organisation of the human RNF26 protein. (B) Protein sequence alignment of the RING domain and C-terminus of RNF26 with those of human RNF4 (P78317), XIAP (P98170), BIRC2 (Q13490) and MDM2 (Q00987). Conserved residues are demarcated according to the Rasmol colour scheme. (C) DOX titration of Flp-In293 cells stably expressing FH-RNF26WT or FH-RNF26Y432A ± MG132 with lysates separated by SDS-PAGE and resulting western blots probed for RNF26 (anti-HA) and tubulin. (D) 35S-Met/Cys pulse-chase assay (0, 1, 2 hr) of DOX-induced Flp-In293 cells stably expressing FH-RNF26WT or FH-RNF26Y432A,±MG132 and immunoprecipitated by anti-FLAG agarose. (E) Co-immunoprecipitation of endogenous RNF26 from DOX-induced Flp-In293 cells stably expressing FH-RNF26WT or FH-RNF26Y432A by anti-FLAG beads. Detection of FLAG-HA (FH) and endogenous (e) RNF26 by western blot using the indicated antibodies. (F) TUBE pulldowns from FH-RNF26WT or FH-RNF26Y432A Flp-In293 cell lysates,±MG132 and Usp21.
-
Figure 4—figure supplement 1—source data 1
Nanostring data of E3 transcript abundance relative to CNX used to generate Figure 4—figure supplement 1B.
- https://cdn.elifesciences.org/articles/57306/elife-57306-fig2-v1.xlsx
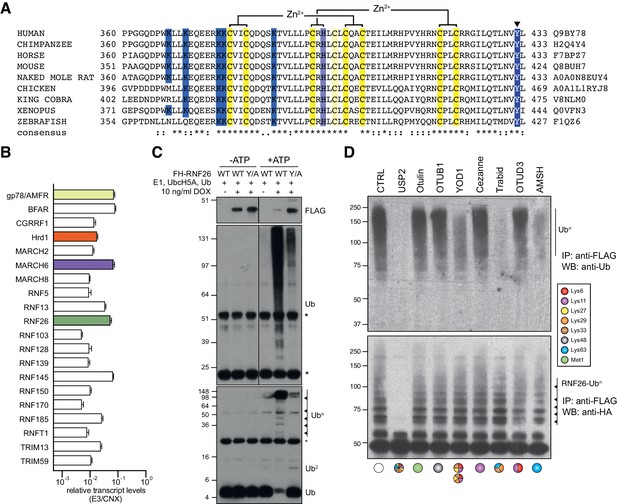
RNF26 ubiquitination.
(A) Protein sequence alignment of the RING domain and C-terminus of RNF26 form different species. (B) E3 transcript abundance in Flp-In293 cells determined by NanoString and normalised to calnexin levels. (C) FH-RNF26WT or FH-RNF26Y432A were isolated from their respective control or DOX-induced Flp-In293 cell lines with anti-FLAG and combined with recombinant E1, UbcH5a, Ub, ±ATP to perform in vitro ubiquitination reactions. Resulting western blots were probed with antibodies against FLAG (RNF26) and Ub. Low (top panel) and high (bottom panel) percentage SDS-PAGE gels are presented to better resolve both poly- and mono-/di-ubiquitin forms, respectively. (D) Ub linkages present on FH-RNF26WT as determined by UbiCREST analysis.
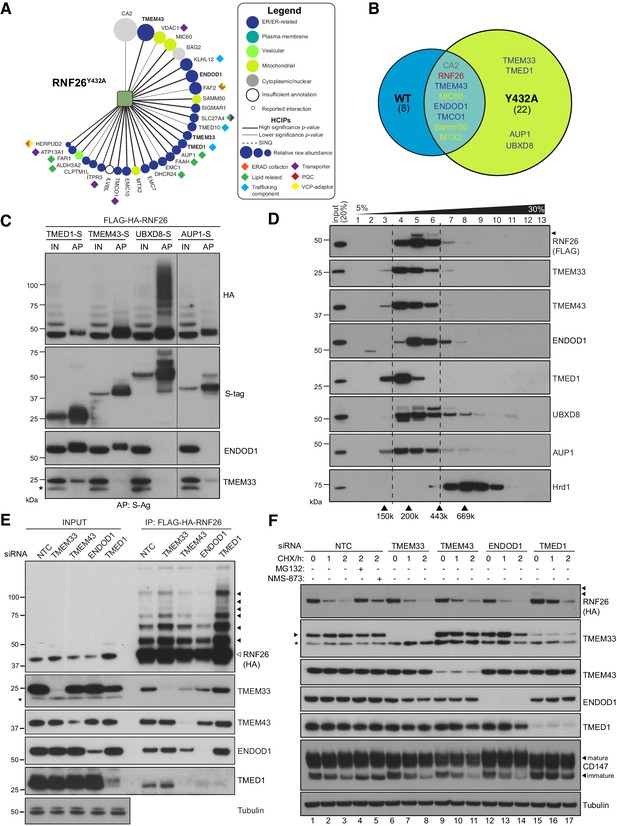
RNF26 assembles with HCIPs in the ER.
(A) HCIP interaction network wheel for FH-RNF26Y432A. Legend described in Figure 2—figure supplement 1. (B) Venn diagram of HCIPs identified by LC-MS/MS for FH-RNF26WT and FH-RNF26Y432A. ER-resident HCIPs are indicated in blue. (C) Co-precipitation of FH-RNF26WT from stable Flp-In293 cells by transiently expressed S-tagged HCIPs (TMED1, TMEM43, UBXD8 and AUP1). Cells were solubilised in 1% LMNG and protein complexes affinity purified from the resulting lysates by S-protein agarose. Western blots of affinity purified material (AP) and input lysate (IN, 20%) were probed with antibodies recognising RNF26 (HA), HCIPs (S-tag), TMEM33 and ENDOD1. (D) Velocity sedimentation of FH-RNF26 complexes from 1% LMNG-solubilised lysates on a sucrose gradient (5–30%), with individual TCA-precipitated fractions (1-13) subsequently separated by SDS-PAGE and the resulting western blots probed for the indicated proteins. Ubiquitinated forms of RNF26 are indicated by black arrowheads. Molecular weight of gel filtration standards solubilised and sedimented in equivalent buffer conditions are shown for comparison beneath the Hrd1 blot, which in turn serves to highlight a complex of a different mass. (E) siRNA-mediated knockdown of HCIPs in FH-RNF26WT Flp-In293 cells alters interaction profiles. RNF26 complexes immunoprecipitated by anti-FLAG agarose were separated by SDS-PAGE with resulting western blots probed by antibodies for RNF26 and the indicated HCIPs. Tubulin was used as a loading control. Ubiquitinated forms of RNF26 are indicated by black arrowheads. (F) Cycloheximide (CHX) chase assays (100 μg/ml; 0, 1, 2 hr) of FH-RNF26WT Flp-In293 cells (DOX, 1 µg/mL, 18 hr) knocked down for individual HCIPs by siRNAs with the resulting western blots probed for the indicated antibodies. MG132 (10 µM, 2 hr) and NMS-873 (10 µM, 2 hr) were included with NTC samples where indicated. Ubiquitinated forms of RNF26 are denoted by black arrowheads. Mature and immature forms of CD147 are indicated by black arrowheads.
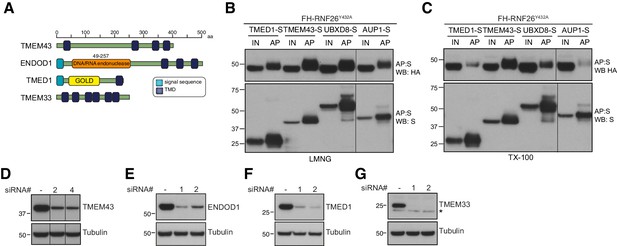
Validation of RNF26 interactions with HCIPs.
(A) Domain organisation of human TMEM43, TMEM33, ENDOD1 and TMED1 proteins. Transient expression of S-tagged HCIPs in the FH-RNF26Y432A Flp-In293 cell line to validate robustness of RNF26-HCIP interactions. Cells solubilised in either 1% LMNG (B) or 1% TX-100 (C) yielded protein complexes affinity purified by S-protein agarose that were probed on western blots by antibodies against the S- and HA-tags. Input (IN, 20%) and affinity purified (AP) material are shown. (D–G) Validation of knockdown of TMEM43, TMEM33, ENDOD1 and TMED1 by two independent siRNAs. Western blots are probed using HCIP-specific antibodies and tubulin as a loading control.
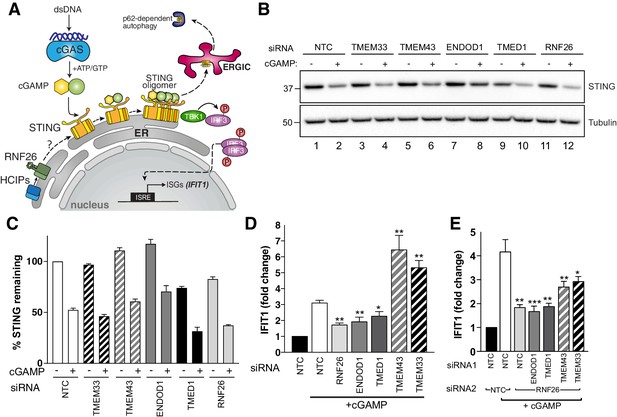
RNF26 and its interactors modulate STING-dependent innate immune signalling.
(A) Diagram of the cGAS-STING signalling pathway. (B) Representative western blot of cGAMP-treated Flp-In293 cells transfected with siRNA targeting RNF26 and HCIPs (TMEM33, TMEM43, ENDOD1, TMED1) and probed for STING and tubulin. (C) Quantification of 3 biological replicates for (b) with mean and S.E.M. shown (n = 3). (D) qRT-PCR for IFIT1 and GAPDH from cGAMP-treated Flp-In293 cells (5 μg/ml, 6 hr) transfected with siRNAs targeting RNF26, ENDOD1, TMED1, TMEM43 and TMEM33, along with a non-targeted control (NTC). Normalised IFIT1 levels in cGAMP-treated cells are shown relative to their untreated counterpart for each siRNA. Mean and S.E.M are shown for at least four biological replicates. (E) Same as (D) but including siRNA targeting RNF26 along with HCIPs. Mean and S.E.M are shown for four biological replicates. For all statistical analysis, *p<0.05, **p<0.01, ***p<0.001. Details of statistical analysis are in Supplementary file 1, Table 12.
-
Figure 6—source data 1
STING protein quantification and IFIT1 fold change data used to generate Figure 6.
- https://cdn.elifesciences.org/articles/57306/elife-57306-fig6-data1-v1.xlsx
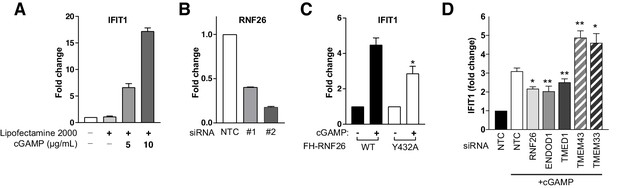
Modulation of the interferon response by RNF26 and its HCIPs.
(A) Fold change in IFIT1 transcription in response to cGAMP. Flp-In293 cells were transfected with cGAMP (0, 5, 10 μg/ml) using Lipefectamine2000 (6 hr) with IFIT1 and GAPDH transcript levels measured by qRT-PCR (Taqman). Mean and S.E.M. from three biological replicates are presented. (B) Validation of RNF26 knockdown with two independent siRNAs by qPCR. Mean and S.E.M. from three biological repeats are presented. (C) Fold change in IFIT1 transcription in response to cGAMP in FH-RNF26WT and FH-RNF26Y432A Flp-In293 cell lines. Mean and S.E.M. from four biological replicates are presented. (D) As described in Figure 6D for an independent set of siRNAs. For all statistical analysis, *p<0.05, **p<0.01,***p<0.001. Details of statistical analysis can be found in Supplementary file 1, Table 12.
-
Figure 6—figure supplement 1—source data 1
IFIT1 fold change and RNF26 transcript abundance data used to generate Figure 6—figure supplement 1.
- https://cdn.elifesciences.org/articles/57306/elife-57306-fig6-figsupp1-data1-v1.xlsx
Additional files
-
Supplementary file 1
Data Tables 1-12: ER-E3 LC-MS/MS (1-8) and cDNA/siRNA sequence (9-12) supporting information.
Supplemental Table 1- ER-resident E3 features and positioning of FLAG-HA epitopes. Supplemental Table 2- Unique protein signatures of ER-resident E3 pulldowns identified by LC-MS/MS and BSCG analysis. Supplemental Table 3- Common contaminants removed from BSCG and SINQ analyses. Supplemental Table 4- Unique ER-resident E3 interactors identified by SINQ analysis of LC-MS/MS dataset. Supplemental Table 5- High-confidence candidate interacting proteins (HCIPs) of ER-resident E3s. Supplemental Table 6- NanoString quantitative transcriptomic codeset of ER-resident E3s. Supplemental Table 7- Targets of UPR transcription factors among ER-resident E3 HCIPs. Supplemental Table 8- BSCG analysis of RNF26Y432A compared to RNF26WT. Supplemental Table 9- ORF sources for ER-resident E3. Supplemental Table 10- ORF sources for HCIPs Supplemental Table 11- siRNA sequences Supplemental Table 12. Statistical analyses.
- https://cdn.elifesciences.org/articles/57306/elife-57306-supp1-v1.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/57306/elife-57306-transrepform-v1.docx





