RETRACTED: Arginine methylation of SHANK2 by PRMT7 promotes human breast cancer metastasis through activating endosomal FAK signalling
Figures
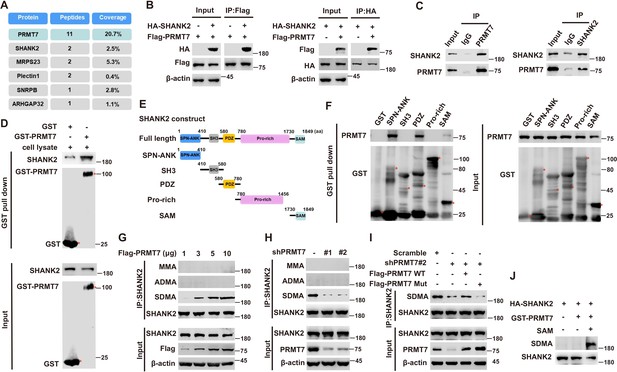
PRMT7 methylated SHANK2.
Immunoprecipitation and immunoblotting analyzes were performed with the indicated antibodies. (A) PRMT7 associated proteins from MCF7 cells expressing stable FLAG-PRMT7 were immunopurified with anti-FLAG (α-FLAG) affinity resins. The protein bands were analyzed by mass spectrometry. Representative peptide fragments of PRMT7 associated proteins and peptide coverage of the indicated proteins are shown. (B) HEK293T cells were co-transfected with Flag-PRMT7 and/or HA-SHANK2 plasmids. (C) Endogenous SHANK2 or PRMT7 was IP from MDA-MB-231 cells lysate, with anti-SHANK2 or anti-PRMT7 antibody, and the binding of PRMT7 and SHANK2 was examined by western blot. (D) Purified bacterially expressed GST or GST-PRMT7 was incubated with MDA-MB-231 cells lysate. GST-pulldown assay showed direct interaction between SHANK2 and recombinant PRMT7. (E) Diagram of the domains in SHANK2 protein. The schematics of the GST-SHANK2 expression plasmid, as well as domains and truncated mutants. (F) SHANK2 truncated mutants were incubated with MDA-MB-231 cells lysate. GST-pulldown assay showed direct interaction between PRMT7 and recombinant SHANK2. (G) HEK293T cells were transfected with increasing amounts of Flag-PRMT7 expression plasmids. Immunoprecipitation of SHANK2 with anti-SHANK2 antibody was performed. (H) MDA-MB-231 with or without stable expression of the indicated PRMT7 shRNA or a control shRNA. Immunoprecipitation of SHANK2 with anti-SHANK2 antibody was performed. (I) MDA-MB-231 cells with or without expressing PRMT7 shRNA and with or without reconstituted expression of WT PRMT7 or PRMT7 enzymatic activity mutant. Immunoprecipitation of SHANK2 with anti-SHANK2 antibody was performed. (J) HA-tagged SHANK2 was expressed in HEK293T cells. HA-purified SHANK2 protein was incubated with SAM and with or without purified bacterially expressed PRMT7. Methylation of recombinant SHANK2 protein was determined by western blot.
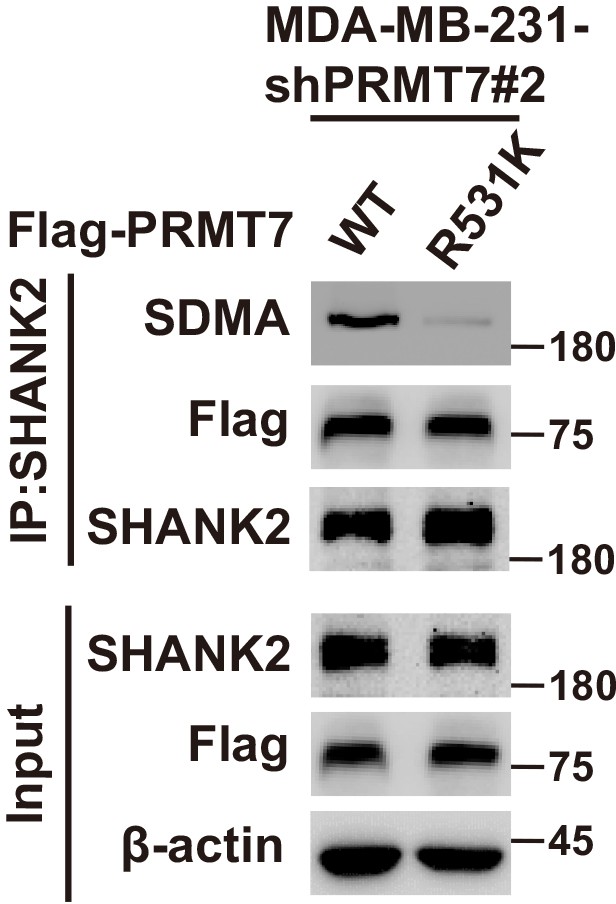
SHANK2 di-methylation was dependent on PRMT7.
Immunoprecipitation and immunoblotting analyses were performed with the indicated antibodies. MDA-MB-231 cells expressing PRMT7 shRNA and with or without reconstituted expression of PRMT7 WT or PRMT7 R531K mutant. Immunoprecipitation of SHANK2 with anti-SHANK2 antibody was performed.
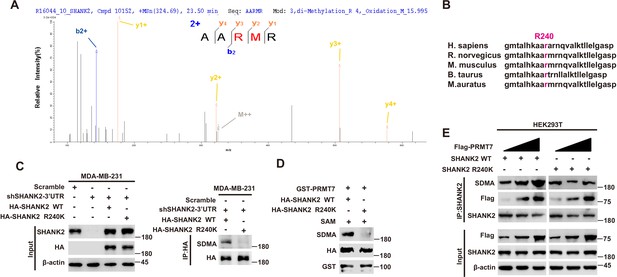
PRMT7 methylated SHANK2 at R240.
(A) The purified SHANK2 from MDA-MB-231 cells transfected with HA-SHANK2 was analyzed for methylation by mass spectrometry. The SHANK2 R240 residue in fragmentation of KAARMRN was methylated. The Mascot score was 27.28, and the expectation value was 3.74E-04. (B) Alignment of the consensus SHANK2 sequences between different species near the R240 methylation site. (C) MDA-MB-231 cells with or without expressing SHANK2 shRNA and with or without reconstituted expression of WT SHANK2 or SHANK2 R240K mutant. Immunoprecipitation of HA with anti-HA antibody was performed. (D) Flag-tagged SHANK2 WT or SHANK2 R240K was transfected into MDA-MB-231 cells. HA-purified SHANK2 WT or SHANK2 R240K proteins were incubated with purified bacterially expressed PRMT7 and SAM. Methylation of SHANK2 protein was determined by western blot. (E) HEK293T cells were co-transfected with increasing amounts of Flag-PRMT7 and SHANK2 WT or SHANK2 R240K expression plasmids. Immunoprecipitation of SHANK2 with anti-SHANK2 antibody was performed.
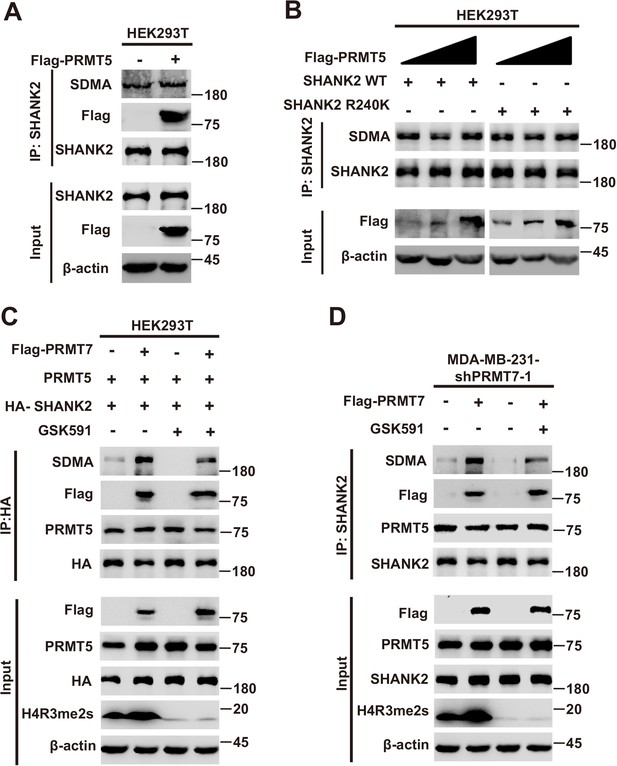
The role of PRMT5 in PRMT7 mediated SHANK2 methylation.
Immunoprecipitation and immunoblotting analyses were performed with the indicated antibodies. (A) HEK293T cells were co-transfected with control pcDNA3.1 or PRMT5 plasmids. Immunoprecipitation of SHANK2 with anti-SHANK2 antibody was performed. (B) HEK293T cells were co-transfected with increasing amounts of Flag-PRMT5 and SHANK2 WT or SHANK2 R240K expression plasmids. Immunoprecipitation of SHANK2 with anti-SHANK2 antibody was performed. (C) HEK293T cells with increasing amounts of Flag-PRMT7, PRMT5 and SHANK2 were treated with GSK591 (5 µM) or DMSO as control. Immunoprecipitation of HA with anti-HA antibody was performed. Western blot was performed with the indicated antibodies. (D) MDA-MB-231 cells with increasing amounts of Flag-PRMT7 were treated with GSK591 (5 µM) or DMSO as control. Immunoprecipitation of SHANK2 with anti-SHANK2 antibody was performed. Western blot was performed with the indicated antibodies.
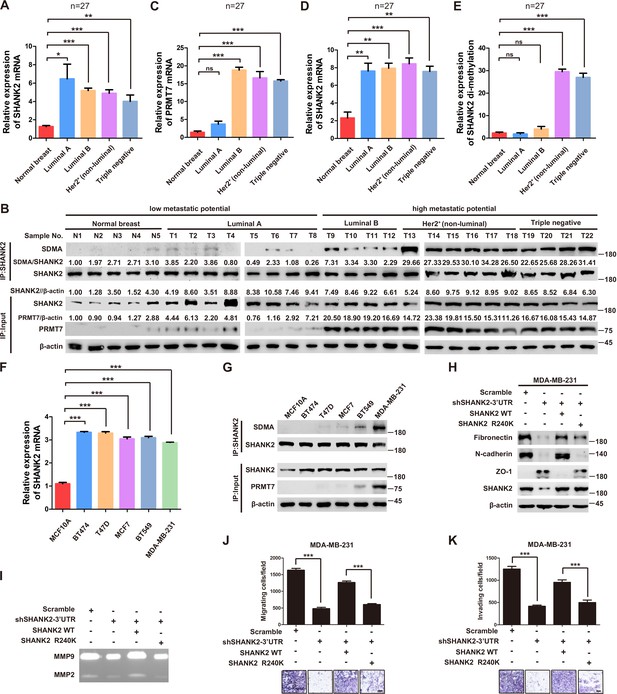
SHANK2 R240 methylation promoted migration and invasion of breast cancer cells.
(A) Comparison of SHANK2 expression level in breast cancer tissues of different breast cancer subtypes with that in normal tissues by RT-PCR. The logarithmic scale of 2-ΔΔCt was used to measure the fold-change. β-actin was used as an internal reference. n = 27, *p<0.05, **p<0.01, ***p<0.001, one-way ANOVA test. (B–E) Comparison of PRMT7, SHANK2 and di-methylated SHANK2 levels in breast cancer tissues using western blot analysis. Data are based on the analysis of independent samples (n = 27). Immunoprecipitation of SHANK2 with anti-SHANK2 antibody was performed. *p<0.05, **p<0.01, ***p<0.001, ns = not significant, one-way ANOVA test. (F) SHANK2 mRNA expression in indicated breast cancer cell lines by RT-PCR. The logarithmic scale of 2-ΔΔCt was used to measure the fold-change. β-actin was used as an internal reference. ***p<0.001, Student’s t test. (G) SHANK2 was IP from indicated breast cancer cell lines lysate, and input lysates and IP samples were analyzed using anti-methylation, anti-SHANK2 or anti-PRMT7 antibodies, as indicated. (H) MDA-MB-231 cells expressing SHANK2 shRNA and reconstituted expression of SHANK2 WT or SHANK2 R240K mutant. Western blot was performed with the indicated EMT marker antibodies. (I) MDA-MB-231 cells with or without stable expression of the indicated SHANK2 shRNA were transfected with SHANK2 WT or SHANK2 R240K mutant. Assessment of MMP2 and MMP9 enzymatic activities by gelatin zymography. (J and K) MDA-MB-231 cells expressing SHANK2 shRNA and reconstituted expression of SHANK2 WT or SHANK2 R240K mutant. Cell migration (J) and Invasion (K) were determined by transwell assays. (n = 3, independent experiments). Scale bars: 100 µm. Data represent the mean ± SD of three independent experiments. ***p<0.001, Student’s t test.
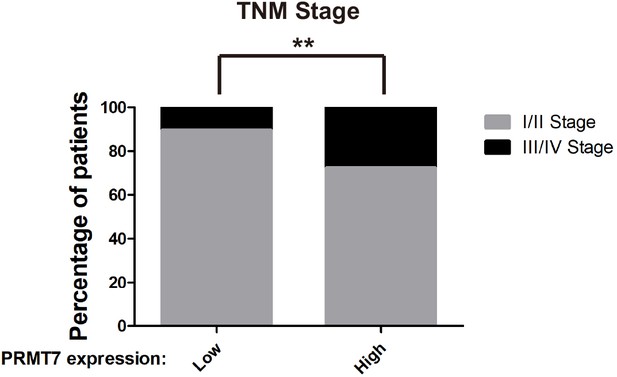
PRMT7 was associated with clinical outcomes.
Correlation between PRMT7 expression and clinical outcomes through analysis of stage of breast cancer patients in TCGA Breast (BRCA) database(n = 1075 patients). **p<0.01, Chi2 test.
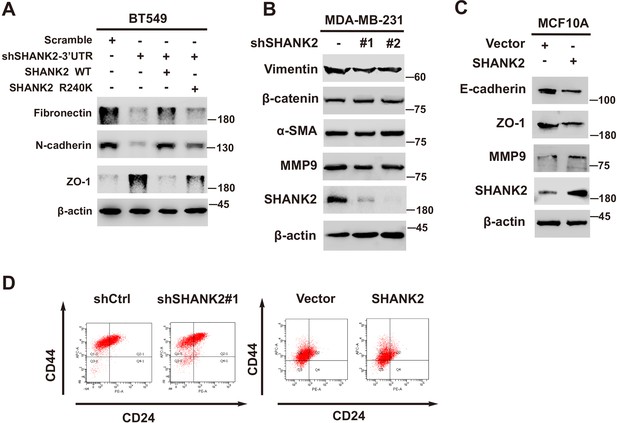
SHANK2 was involved in breast cancer cells EMT.
(A) BT549 cells with or without expressing SHANK2 shRNA and with or without reconstituted expression of SHANK2 WT or SHANK2 R240K mutant. Western blot was performed with the indicated EMT marker antibodies. (B) MDA-MB-231 with or without stable expression of the indicated SHANK2 shRNA or a control shRNA. Western blot was performed with the indicated EMT marker antibodies. (C) MCF10A cells expressing the indicated HA-tagged SHANK2. Western blot was performed with the indicated EMT marker antibodies. (D) Stem-like CD44high/CD24low profile in MDA-MB-231 cells transfected with stable expression of the indicated SHANK2 shRNA or a control shRNA, and MCF10A cells expressing the indicated HA-tagged SHANK2.
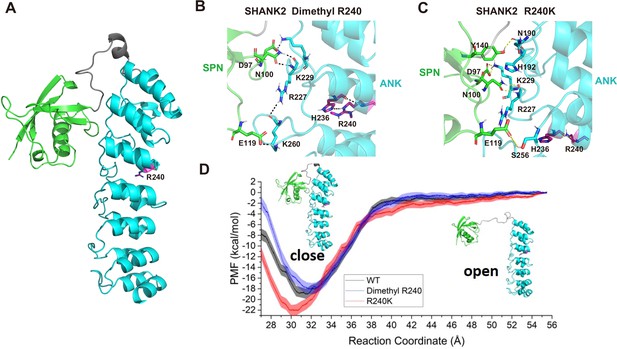
R240 methylation impaired SPN-ANK domain interaction of SHANK2.
(A) Definition of the SPN domain (coloured in green) and the ANK domain (coloured in cyan) in SHANK2. The location of R240 was represented with sticks. (B) Key residues forming hydrogen bonds between SPN and ANK in SHANK2 WT. R240 can form hydrogen bonds with H236 in ANK (coloured in magenta). (C) Key residues forming hydrogen bonds between SPN and ANK in SHANK2 R240K. K240 can still form hydrogen bonds with H236 in ANK (coloured in magenta). (D) Potential of mean forces for the conformational changes (from closed state to the open state) of SPN and ANK in the wild type (WT, black line), methylated (Dimethyl R240, blue line) and mutant (R240K, red line) SHANK2.
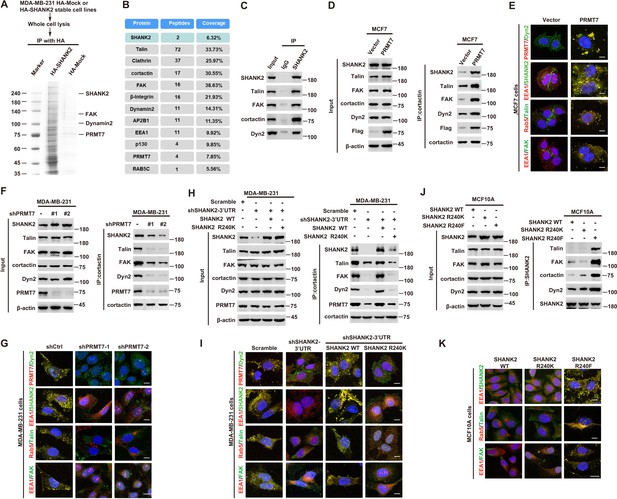
SHANK2 R240 methylation promotes FAK/dynamin2/talin complex co-localized with endosome.
(A) SHANK2 associated proteins from HA-SHANK2 expressing stable MDA-MB-231 cells were immunopurified with anti-HA (α-HA) affinity resins. The protein bands were analyzed by mass spectrometry. (B) Representative peptide fragments of SHANK2 associated proteins and peptide coverage of the indicated proteins are shown. (C) Endogenous talin, FAK, dynamin2, EEA1 and SHANK2 were IP from MDA-MB-231 cells, with indicated antibodies, and the binding of talin, FAK, dynamin2 and SHANK2 was examined by western blot. (D) MCF7 cells with expression of the indicated Flag-Vector or Flag-tagged PRMT7. Immunoprecipitation of cortactin with anti-cortactin antibody was performed. (E) Confocal images of PRMT7, dynamin2, SHANK2, EEA1, talin, Rab5, FAK, cortactin and DAPI staining in MCF7 cells with expression of the indicated Flag-Vector or Flag-tagged PRMT7. Scale bars, 10 µm. (F) MDA-MB-231cells with or without stable expression of the indicated PRMT7 shRNA or a control shRNA. Immunoprecipitation of cortactin with anti-cortactin antibody was performed. (G) confocal images of PRMT7, dynamin2, SHANK2, EEA1, talin, Rab5, FAK, cortactin and DAPI staining in MDA-MB-231cells with or without stable expression of the indicated PRMT7 shRNA or a control shRNA. Scale bars, 10 µm. (H) MDA-MB-231cells expressing SHANK2 shRNA with or without reconstituted expression of WT SHANK2 or SHANK2 R240K mutant. Immunoprecipitation of cortactin with anti-cortactin antibody was performed. (I) Confocal images of PRMT7, dynamin2, SHANK2, EEA1, talin, Rab5, FAK, cortactin and DAPI staining in MDA-MB-231cells expressing SHANK2 shRNA with or without reconstituted expression of WT SHANK2 or SHANK2 R240K mutant. Scale bars, 10 µm. (J) MCF10A cells with reconstituted expression of SHANK2 WT, SHANK2 R240K or SHANK2 R240F (arginine to phenylalanine) mutant. Immunoprecipitation of SHANK2 with anti-SHANK2 antibody was performed. (K) Confocal images of PRMT7, dynamin2, SHANK2, EEA1, talin, Rab5, FAK, cortactin in MCF10A cells with reconstituted expression of SHANK2 WT, SHANK2 R240K or SHANK2 R240F mutant. Scale bars, 10 µm.
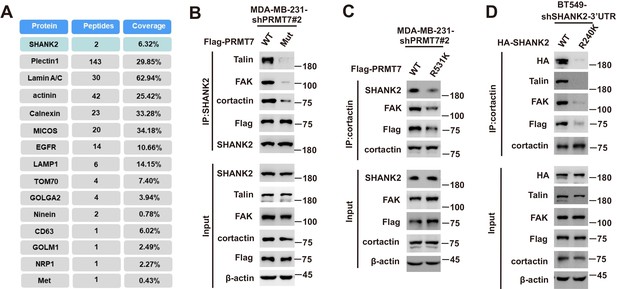
SHANK2 was essential for the interaction and activity between talin and FAK.
Immunoprecipitation and immunoblotting analyses were performed with the indicated antibodies. (A) Representative peptide fragments of SHANK2 associated proteins and peptide coverage of the indicated proteins are shown. (B) MDA-MB-231 cells expressing PRMT7 shRNA and with or without reconstituted expression of WT PRMT7 or PRMT7 enzymatic activity mutant. Immunoprecipitation of SHANK2 with anti-SHANK2 antibody was performed. (C) MDA-MB-231 cells expressing PRMT7 shRNA and with or without reconstituted expression of PRMT7 WT or PRMT7 R531K mutant. Immunoprecipitation of cortactin with anti-SHANK2 antibody was performed. (D) BT549 cells with or without expressing SHANK2 shRNA and with or without reconstituted expression of SHANK2 WT or SHANK2 R240K mutant. Immunoprecipitation of cortactin with anti-SHANK2 antibody was performed.
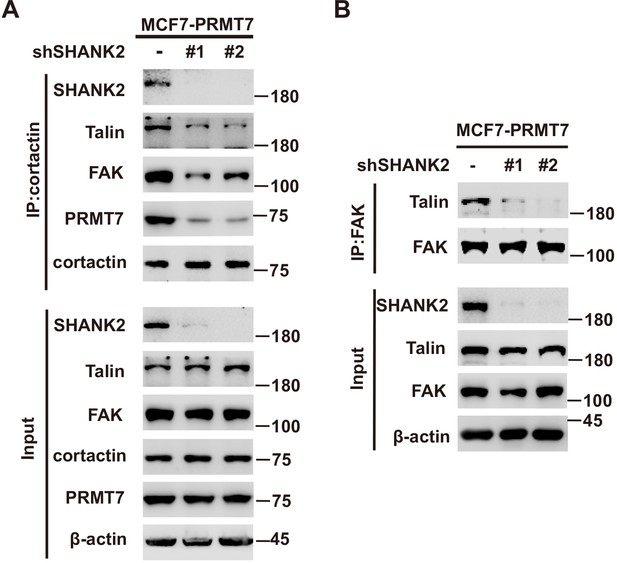
SHANK2 was necessary for the interaction between talin and FAK.
(A) MCF7 cells with reconstituted expression of PRMT7 and with or without stable expression of the indicated SHANK2 shRNA. Immunoprecipitation of cortactin with anti-cortactin antibody was performed. (B) MCF7 cells with reconstituted expression of PRMT7 and with or without stable expression of the indicated SHANK2 shRNA. Immunoprecipitation of FAK with anti-FAK antibody was performed.
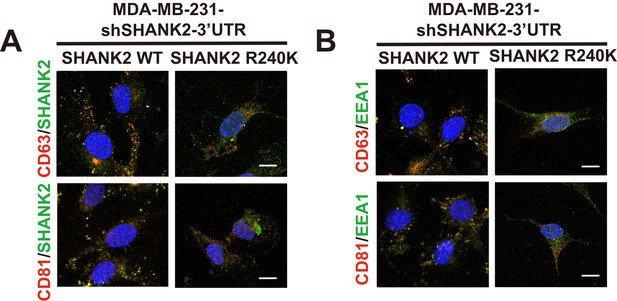
SHANK2 was co-localized with exosome markers.
(A) confocal images of SHANK2, CD63, CD81 and DAPI staining in MDA-MB-231cells expressing SHANK2 shRNA with or without reconstituted expression of WT SHANK2 or SHANK2 R240K mutant. Scale bars, 10 µm. (B) confocal images of EEA1, CD63, CD81 and DAPI staining in MDA-MB-231cells expressing SHANK2 shRNA with or without reconstituted expression of WT SHANK2 or SHANK2 R240K mutant. Scale bars, 10 µm. .
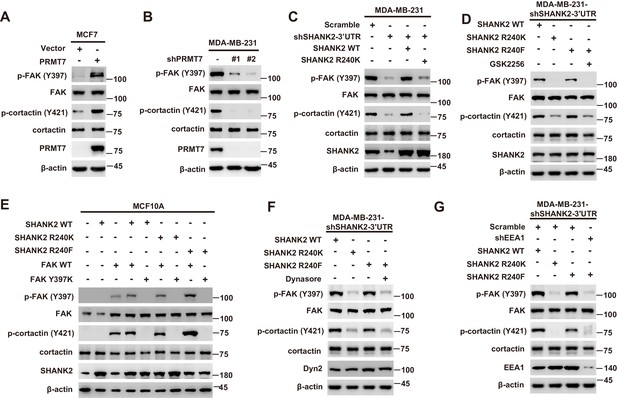
SHANK2 R240 methylation activated endosomal FAK signals.
(A) Phosphorylated/total FAK and cortactin were determined in MCF-7 cells expressing the indicated Flag-tagged PRMT7. Western blot was performed with the indicated antibodies. (B) Phosphorylated/total FAK and cortactin were determined in MDA-MB-231 cells with or without stable expression of the indicated SHANK2 shRNA or a control shRNA. Western blot was performed with the indicated antibodies. (C) Phosphorylated/total FAK and cortactin were determined in MDA-MB-231 cells with or without expressing SHANK2 shRNA and with or without reconstituted expression of WT SHANK2 or SHANK2 R240K mutant. Western blot was performed with the indicated antibodies. (D) Phosphorylated/total FAK and cortactin were determined in MDA-MB-231 cells expressing SHANK2 shRNA with or without reconstituted expression of WT SHANK2 or SHANK2 R240K or SHANK2 R240F mutant, SHANK2 R240F were treated with GSK2256098 (10 µM) or DMSO as control. Western blot was performed with the indicated antibodies. (E) Phosphorylated/total FAK and cortactin were determined in MCF10A cells co-transfected with SHANK2 and/or FAK plasmids. Western blot was performed with the indicated antibodies. (F) Phosphorylated/total FAK and cortactin were determined in MDA-MB-231 cells expressing SHANK2 shRNA with or without reconstituted expression of WT SHANK2, SHANK2 R240K or SHANK2 R240F mutant. SHANK2 R240F group was treated with dynasore (80 µM) or DMSO as control. Western blot was performed with the indicated antibodies. (G) Phosphorylated/total FAK and cortactin were determined in MDA-MB-231 cells expressing SHANK2 shRNA with or without reconstituted expression of WT SHANK2, SHANK2 R240K or SHANK2 R240F mutant. SHANK2 R240F with or without stable expression of the indicated EEA1 shRNA or a control shRNA. Western blot was performed with the indicated antibodies.
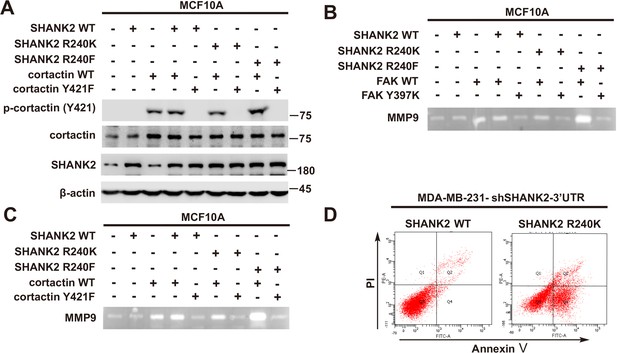
SHANK2 R240F activated FAK and cortactin.
(A) Phosphorylated/total FAK and cortactin were determined in MCF10A cells co-transfected with SHANK2 and/or cortactin plasmids. Western blot was performed with the indicated antibodies. (B) MCF10A cells co-transfected with SHANK2 and/or cortactin plasmids. Assessment of MMP9 enzymatic activities by gelatin zymography. (C) MCF10A cells co-transfected with SHANK2 and/or cortactin plasmids. Assessment of the MMP9 enzymatic activities by gelatin zymography. (D) MDA-MB-231 cells with or without expressing SHANK2 shRNA and with or without reconstituted expression of SHANK2 WT or SHANK2 R240K mutant. Flow cytometric analysis of annexin V-FITC/PI staining was conducted to examine cell death.
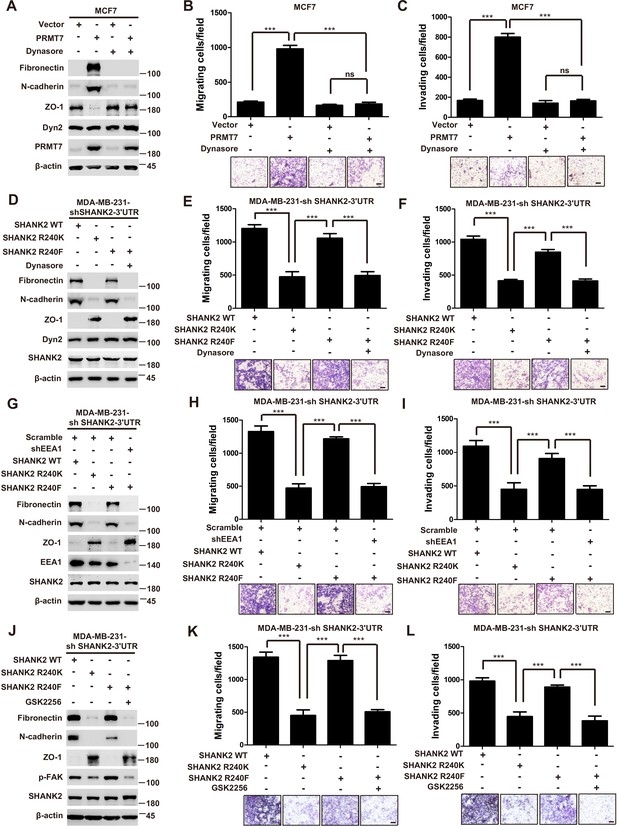
SHANK2 R240 methylation promoted migration and invasion of breast cancer cells by activating endosomal FAK signals.
(A) MCF7cells expressing the indicated Flag-tagged PRMT7. MCF7-Vector and MCF7-PRMT7 groups were treated with dynasore (80 µM) or DMSO as control. Western blot was performed with the indicated EMT marker antibodies. (B and C) MCF7 cells expressing the indicated Flag-tagged PRMT7. MCF7-Vector and MCF7-PRMT7 groups were treated with dynasore (80 µM) or DMSO as control. Cell migration (B) and Invasion (C) were determined by transwell assays. (n = 3, independent experiments). Scale bars: 50 µm. Data represent the mean ± SD of three independent experiments. Scale bars,100 µm. ***p<0.001, Student’s t test. (D) MDA-MB-231 cells expressing SHANK2 shRNA with or without reconstituted expression of WT SHANK2, SHANK2 R240K or SHANK2 R240F mutant. SHANK2 R240F group was treated with dynasore (80 µM) or DMSO as control. Western blot was performed with the indicated EMT marker antibodies. (E and F) MDA-MB-231 cells expressing SHANK2 shRNA with or without reconstituted expression of WT SHANK2, SHANK2 R240K or SHANK2 R240F mutant. SHANK2 R240F group was treated with dynasore (80 µM) or DMSO as control. Cell migration (E) and Invasion (F) were determined by transwell assays. (n = 3, independent experiments). Scale bars: 100 µm. ***p<0.001, Student’s t test. (G) MDA-MB-231 cells expressing SHANK2 shRNA with or without reconstituted expression of WT SHANK2, SHANK2 R240K or SHANK2 R240F mutant. SHANK2 R240F with or without stable expression of the indicated EEA1 shRNA or a control shRNA. Western blot was performed with the indicated EMT marker antibodies. (H and I) MDA-MB-231 cells expressing SHANK2 shRNA with or without reconstituted expression of WT SHANK2, SHANK2 R240K or SHANK2 R240F mutant. SHANK2 R240F with or without stable expression of the indicated EEA1 shRNA or a control shRNA. Cell migration (H) and Invasion (I) were determined by transwell assays. (n = 3, independent experiments). Scale bars: 100 µm. ***p<0.001, Student’s t test. (J) MDA-MB-231 cells expressing SHANK2 shRNA with or without reconstituted expression of WT SHANK2, SHANK2 R240K or SHANK2 R240F mutant. SHANK2 R240F group was treated with GSK2256098 (10 µM) or DMSO as control. Western blot was performed with the indicated EMT marker antibodies.
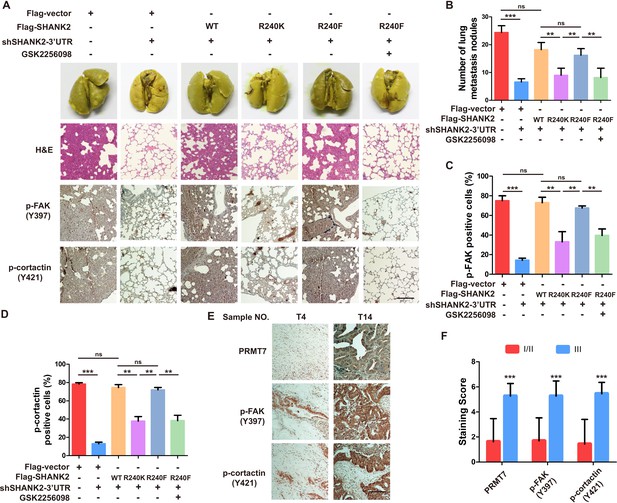
Arginine methylation at SHANK2 R240 promoted metastasis through activates FAK signalling.
(A) Equal numbers of MDA-MB-231 cells expressing SHANK2 shRNA with or without reconstituted expression of WT SHANK2, SHANK2 R240K or SHANK2 R240F mutant. SHANK2 R240F group was treated with GSK2256098 or DMSO. Cells were tail-vein injected into BALB/c nude mice, and lung metastasis was determined. The gross appearance of the lungs and the tumour nodules on the lungs were examined. Tumours were paraffin embedded and stained with H and E at Day12. n = 5 tumours in each group. IHC analyses of tumour tissues were performed with anti-p-FAK or anti-p-cortactin antibody. Scale bars,100 µm. (B) The number of visible surface metastatic lesions in lungs was counted. (n = 5 mice for each group, **p<0.01, ***p<0.001, ns = not significant, one-way ANOVA test). (C, D) p-FAK and p-cortactin -positive cells were quantified in 10 microscope fields. **p<0.01, ***p<0.001, ns = not significant, one-way ANOVA test. (E, F) Typical pictures of the immunohistochemical staining of PRMT7, p-FAK and p-cortactin in breast cancer samples. Scale bars,100 µm. n = 27, **p<0.01, Student’s t test.
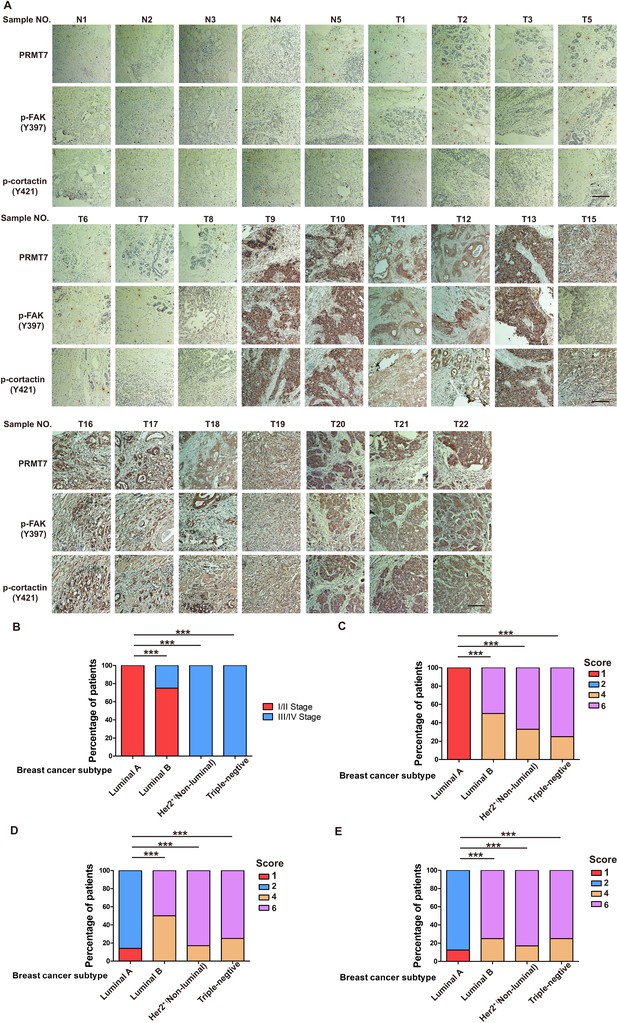
PRMT7 was associated with III stage breast cancer.
(A) Typical pictures of the immunohistochemical staining of PRMT7, p-FAK and p-cortactin in breast cancer samples. Scale bars,100 µm. (B–E) Statistic of stage and staining correlates with subtype. n = 27, ***p<0.001, Chi2 test.
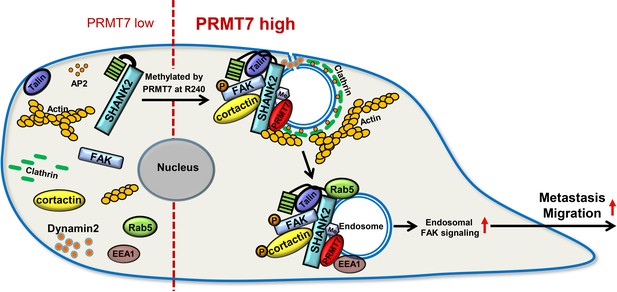
A model for PRMT7-mediated SHANK2 methylation that promotes breast cancer migration through activating endosomal FAK signalling In the absence of PRMT7, SHANK2 cannot interact with dynamin2/talin/FAK/cortactin complex to endosome.
In the presence of PRMT7, R240-methylated SHANK2 disturbs SPN-ANK domain blockade of SHANK2, thereby promoting the co-localization of dynamin2/talin/FAK/cortactin complex on endosome, further activating FAK signalling to promote tumour cells migration.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Antibody | Anti- N-cadherin (Mouse monoclonal) | BD Biosciences | Cat# 561553, RRID:AB_10713831 | WB (1:1000) |
| Antibody | Anti- fibronectin (Mouse monoclonal) | BD Biosciences | Cat# 610070, RRID:AB_2105706 | WB (1:1000) |
| Antibody | Anti-β-catenin (Mouse monoclonal) | BD Biosciences | Cat# 610154 | WB (1:1000) |
| Antibody | Anti- EEA1 (Mouse monoclonal) | BD Biosciences | Cat# 610456, RRID:AB_397829 | IF (1:100) |
| Antibody | Anti- Rab5 (Mouse polyclonal) | BD Biosciences | Cat# 610724, RRID:AB_398047 | IF (1:100) |
| Antibody | Anti-β-actin (Mouse monoclonal) | Sigma-Aldrich | Cat# A5228, RRID:AB_262054 | WB (1:5000) |
| Antibody | Anti- HA (Mouse monoclonal) | Sigma-Aldrich | Cat# H9658, RRID:AB_260092 | WB (1:1000) IP (4 μg) |
| Antibody | Anti- MMP2 (Rabbit polyclonal) | GeneTex | Cat# GTX-104577 RRID:AB_1950932 | WB (1:1000) |
| Antibody | Anti-MMP9 (Rabbit polyclonal) | GeneTex | Cat# 40086 | WB (1:1000) |
| Antibody | Anti- p-cortactin (Rabbit polyclonal) | Bioworld | Cat# BS4778, RRID:AB_1663129 | IHC (1:100) |
| Antibody | Anti- p-cortactin (Rabbit polyclonal) | Abcam | Cat# ab47768, RRID:AB_869231 | WB (1:1000) |
| Antibody | mono methyl Arginine (Mouse monoclonal) | Abcam | Cat# ab415, RRID:AB_304323 | WB (1:500) |
| Antibody | asymmetric dimethyl Arginine(Mouse monoclonal) | Abcam | Cat# ab412, RRID:AB_304292 | WB (1:500) |
| Antibody | Dyn2 (Rabbit polyclonal) | Abcam | Cat# ab3457, RRID:AB_2093679 | WB (1:1000) IF (1:100) |
| Antibody | Flag (Mouse monoclonal) | Abmart | Cat# M20008, RRID:AB_2713960 | WB (1:1000) |
| Antibody | p-FAK (Rabbit polyclonal) | Invitrogen | Cat# 700255, RRID:AB_2532307 | IHC (1:100) IP (4 μg) |
| Antibody | PRMT7(Mouse monoclonal) | santa cruz | Cat# sc-376077, RRID:AB_10990266 | IHC (1:200) |
| Antibody | PRMT7 (Rabbit monoclonal) | Cell Signaling Technology | Cat# 14762, RRID:AB_2798599 | WB (1:1000) IP (4 μg) IF (1:100) |
| Antibody | SHANK2 (Rabbit polyclonal) | Cell Signaling Technology | Cat# 12218, RRID:AB_2797848 | WB (1:500) IP (4 μg) IF (1:100) |
| Antibody | p-FAK(Rabbit monoclonal) | Cell Signaling Technology | Cat# 8556, RRID:AB_10891442 | WB (1:1000) |
| Antibody | FAK (Rabbit polyclonal) | Cell Signaling Technology | Cat# 3285, RRID:AB_2269034 | WB (1:1000) IF (1:100) |
| Antibody | p-talin(Rabbit monoclonal) | Cell Signaling Technology | Cat# 13589, RRID:AB_2798267 | WB (1:1000) |
| Antibody | talin(Rabbit monoclonal) | Cell Signaling Technology | Cat# 4021, RRID:AB_2204018 | WB (1:1000) IF (1:100) |
| Antibody | ZO-1(Rabbit polyclonal) | Cell Signaling Technology | Cat# 5406, RRID:AB_1904187 | WB (1:1000) |
| Antibody | symmetric dimethyl Arginine(Rabbit polyclonal) | Millipore | Cat# 07–412, RRID:AB_11212396 | WB (1:500) |
| Antibody | PRMT5(Rabbit polyclonal) | Millipore | Cat# 07–405, RRID:AB_310589 | WB (1:1000) |
| Antibody | cortactin(Mouse monoclonal) | Millipore | Cat# 16–228, RRID:AB_441969 | WB (1:1000) IF (1:100) |
| Sequenced-based reagent | Interference in the primer:PRMT7#1 | Addgene | GGAACAAGCTATTTCCCATCC | |
| Sequenced-based reagent | Interference in the primer:PRMT7#2 | Addgene | GGATGCAGTGTGTGTACTTCC | |
| Sequenced-based reagent | Interference in the primer:SHANK2#1 | Addgene | GGAGTTAGTCAAAGCACAAAG | |
| Sequenced-based reagent | Interference in the primer:SHANK2#2 | Addgene | GCTTGGAGCAAGAGAGAATTT | |
| Sequenced-based reagent | Interference in the primer:shEEA1#1 | Addgene | GCGGAGTTTAAGCAGCTACAA | |
| Sequenced-based reagent | PCR Primer: β-actin forward: | Addgene | PCR Primers | GAGCACAGAGCCTCGCCTTT |
| Sequenced-based reagent | PCR Primer: β-actin reverse: | Addgene | PCR Primers | ATCCTTCTGACCCATGCCCA |
| Sequenced-based reagent | PCR Primer: SHANK2 forward: | Addgene | PCR Primers | CGGGTAATCCTCCCAAA |
| Sequenced-based reagent | PCR Primer: SHANK2 reverse: | Addgene | PCR Primers | CTTTATCCCGGCGTTTCATC |
| Recombinant DNA reagent | Flag-cortactin (plasmid) | The lab of Dr. Alpha S. Yap.This paper | ||
| Recombinant DNA reagent | HA-SHANK2 (plasmid) | The lab of Dr. Min Goo Lee.This paper | ||
| Recombinant DNA reagent | pcDNA3-HA-SHANK2 (plasmid) | Addgene | ||
| Recombinant DNA reagent | pWPXLD-SHANK2 (plasmid) | Addgene | ||
| Recombinant DNA reagent | pWPXLD-Flag-PRMT7 (plasmid) | Addgene | ||
| Recombinant DNA reagent | pWPXLD-PRMT7 (plasmid) | Addgene | ||
| Recombinant DNA reagent | pWPXLD-Flag-PRMT5 (plasmid) | Addgene | ||
| Recombinant DNA reagent | pWPXLD-PRMT5 (plasmid) | Addgene | ||
| Chemical compound, drug | Dynasore | Selleck | Cat# S8047 | |
| Chemical compound, drug | GSK591 | Selleck | Cat# S8111 | |
| Chemical compound, drug | GSK2256098 | Selleck | Cat# S5823 | |
| Chemical compound, drug | EGF | Sigma-Aldrich | Cat# E9644 | |
| Chemical compound, drug | hydrocortisone | Sangon | Cat# A610506 | |
| Chemical compound, drug | cholera toxin | Sigma-Aldrich | Cat# C8180 | |
| Chemical compound, drug | insulin | Gibco, Grand Island, NY, USA | Cat# 12585–014 | |
| Chemical compound, drug | hFGF basic | R and D Systems, Minneapolis, MN, USA | Cat# P09038 | |
| Strain, strain background Mus musculus | BALB/c nude (CAnN.Cg-Foxn1nu/Crl) | Charles River Labs | Cat#CRL:194,RRID:IMSR_CRL:194 | |
| Cell line (Homo-sapiens) | MCF10A (human; female) | American Type Culture Collection | Cat# CRL-10317, RRID:CVCL_0598 | |
| Cell line (Homo-sapiens) | MCF7 (human; female) | American Type Culture Collection | Cat# HTB-22, RRID:CVCL_0031 | |
| Cell line (Homo-sapiens) | T47D (human; female) | American Type Culture Collection | Cat# HTB-133, RRID:CVCL_0553 | |
| Cell line (Homo-sapiens) | BT474(human; female) | American Type Culture Collection | Cat# HTB-20, RRID:CVCL_0179 | |
| Cell line (Homo-sapiens) | MDA-MB-231(human; female) | American Type Culture Collection | Cat# HTB-26, RRID:CVCL_0062 | |
| Cell line (Homo-sapiens) | BT549(human; female) | American Type Culture Collection | Cat# HTB-122, RRID:CVCL_1092 | |
| Cell line (Homo-sapiens) | HEK293T (human; fetus) | American Type Culture Collection | Cat# CRL-3216, RRID:CVCL_0063 | |
| Software, algorithm | ImageJ | Wayne Rasband, National Institutes of Health | RRID:SCR_003070 | |
| Software, algorithm | GraphPad Prism 8 | GraphPad Software | Version 8.3 RRID:SCR_002798 |
Additional files
-
Supplementary file 1
Clinical breast cancer adjacent tissues.
- https://cdn.elifesciences.org/articles/57617/elife-57617-supp1-v2.xlsx
-
Supplementary file 2
Clinical breast cancer tissues.
- https://cdn.elifesciences.org/articles/57617/elife-57617-supp2-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/57617/elife-57617-transrepform-v2.docx





