Cytoplasmic sharing through apical membrane remodeling
Figures
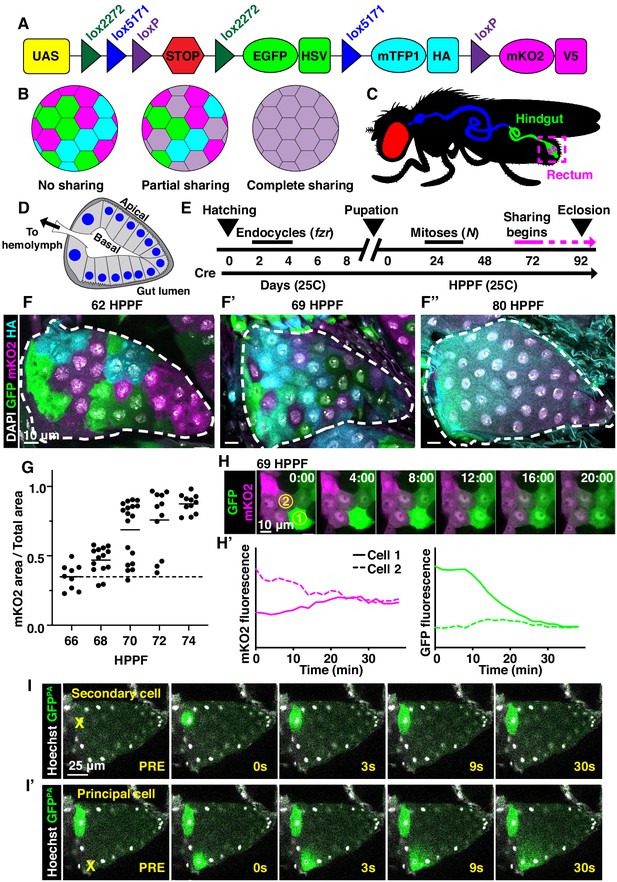
Developmentally programmed cytoplasmic sharing in Drosophila papillae.
(A) The dBrainbow construct (Hampel et al., 2011). Cre recombinase randomly excises one pair of lox sites, and approximately 1/3 of cells express either EGFP, mKO2, or mTFP1. (B) Model of dBrainbow expression with no, partial, or complete cytoplasmic sharing. (C) Drosophila digestive tract with rectum containing four papillae labeled in magenta box. (D) Cartoon of a cross-section through an adult rectal papilla. The papilla consists of an epithelial cone with the apical region facing the gut lumen and the interior basal region facing a central canal leading to the fly hemolymph. The principal papillar cells have microvilli-like projections on the apical edge. One layer of larger, secondary cells forms the base of the papilla. The papilla is covered in a cuticle layer (dark gray). Nuclei are marked in blue. (E) Approximate timeline of ubiquitous Cre induction and cytoplasm sharing onset (68–74 HPPF) within papillar development (Fox et al., 2010). Cytoplasmic sharing is temporally separate from papillar mitoses. (F–F’’) Representative dBrainbow papillae at 62 (F), 69 (F’), or 80 (F’’) hours post-puparium formation (HPPF). (G) Cytoplasmic sharing quantification during pupal development. Lines = mean at each time, which differs significantly between 66 and 74 HPPF (p<0.0001). Each point = 1 animal (N = 9–18, rep = 2). (H) Live dBrainbow-labeled papillar cells during cytoplasmic sharing (69 HPPF). (H’) Fluorescence of neighboring cells in (H). (I–I’) Representative adult papilla expressing photo-activatable GFP (GFPPA). Single cells were photo-activated (yellow X) in secondary cells (I) and principal cells (I’). Time = seconds after activation.
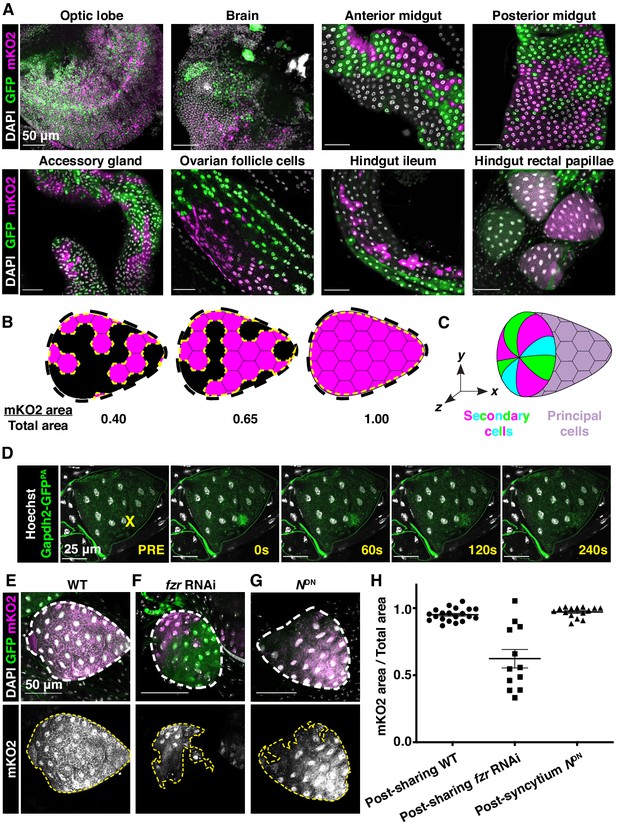
The hindgut rectal papillae share cytoplasm independent of mitosis.
(A) Representative images of dBrainbow expression in the indicated adult tissues. (B) Schematic of cytoplasmic sharing quantification. The mKO2-positive papillar area is divided by the total papillar area to give a score of cytoplasmic sharing. Numbers close to one indicate near-complete sharing. (C) Schematic of principal cells (sharing) and secondary cells (non-sharing) at the papillar base that together form each papilla. (D) Gapdh2-GFPPA activated in single cells in an adult papilla and imaged every 15 s. (E–G) Representative adults expressing dBrainbow in a (E) wild-type (WT), (F) fzr RNAi (p<0.0001), or (G) NDN background (p=0.8786). (H) Quantification of cytoplasmic sharing in adult WT, fzr RNAi, and NDN-expressing animals (N = 12–20, rep = 2).
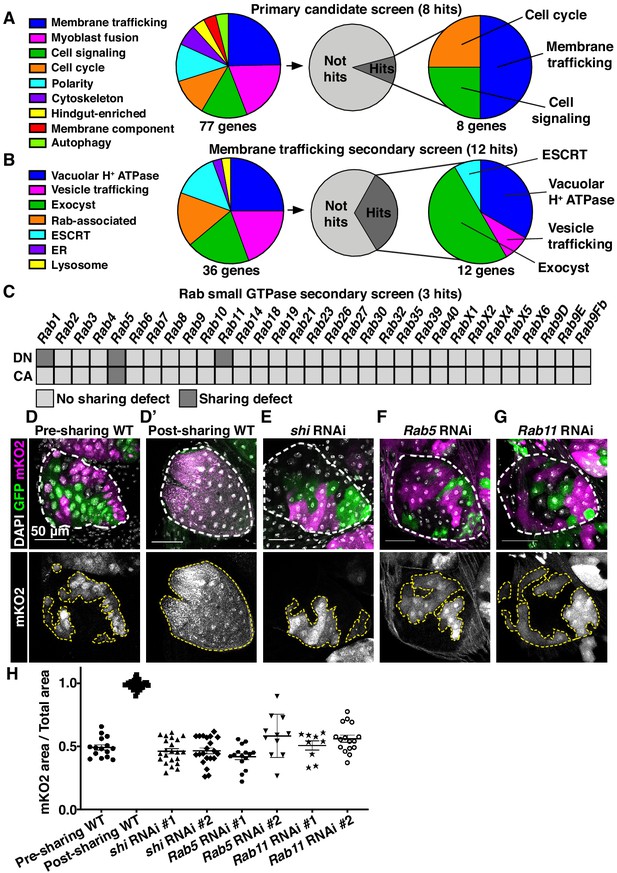
Cytoplasmic sharing requires membrane remodeling proteins.
(A) Primary dBrainbow candidate screen. RNAi and dominant-negative versions of 77 genes representing the indicated roles were screened for sharing defects, and eight genes were identified. (B) Secondary membrane trafficking screen. 36 genes were screened with 12 sharing genes identified. (C) Secondary screen of dominant-negative and constitutively-active Rab GTPases. (D–G) Representative dBrainbow in (D–D’) wild type (WT) (D) pre-sharing (48HPPF) and (D’) post-sharing (young adults), (E) adult shi RNAi, (F) adult Rab5 RNAi, (G) adult Rab11 RNAi. (H) Quantification of (D–G), including two RNAi lines for shi, Rab5, and Rab11. Pre-sharing and knock downs differ significantly from post-sharing WT (p<0.0001, N = 9–32, rep = 2–3).
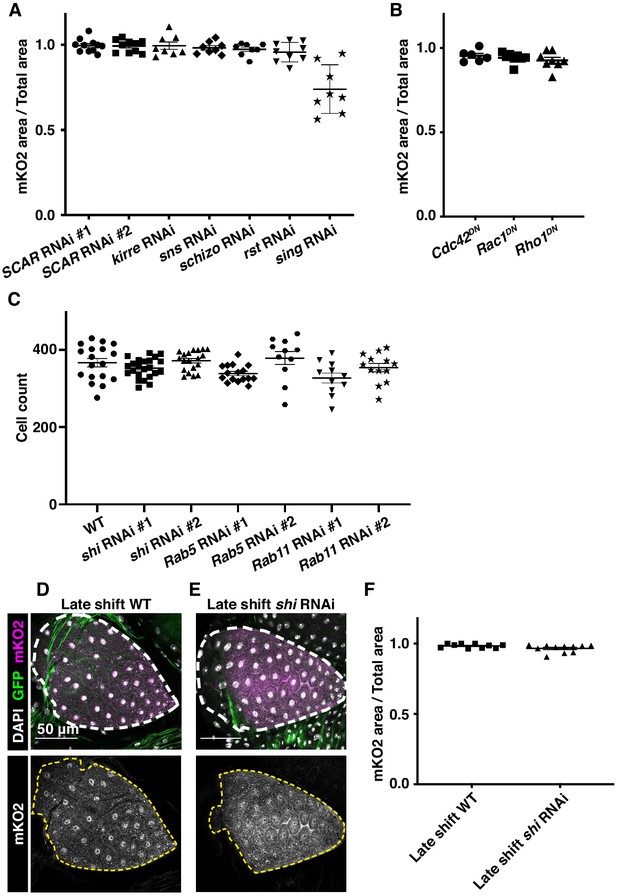
Membrane trafficking genes expressed during a developmental window regulate cytoplasm sharing.
(A) Quantification of cytoplasmic sharing in animals expressing dsRNA for myoblast fusion regulators (N = 8–11, rep = 2). All knockdown lines are previously published (Bischoff et al., 2013; Xing et al., 2018; Linneweber et al., 2015; Johnson et al., 2011; Brunetti et al., 2015). Only sing RNAi significantly differs from WT (p<0.0001). (B) Quantification of cytoplasmic sharing in animals expressing dsRNA for Rho family GTPases (N = 6–8, rep = 2). (C) Cell counts in WT and knockdown rectal papillae (N = 11–23, rep = 2). Only Rab11 #1 RNAi had a significantly different cell number than WT (p=0.0323). (D–E) Representative animals expressing dBrainbow in either a WT (D) or shi RNAi (E) genetic background were raised at 18°C until 3–4 days PPF and shifted to 29°C to induce shi knockdown at a later timepoint than in Figure 2E and H. (F) Sharing quantification in late-induced animals (N = 10–11, rep = 2).
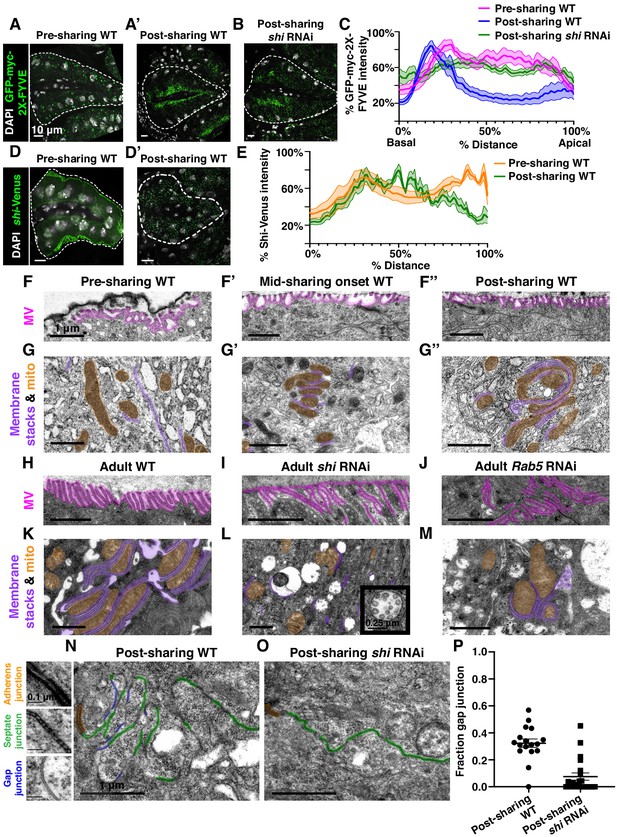
Gap junction establishment, but no membrane breaches, accompany cytoplasm sharing.
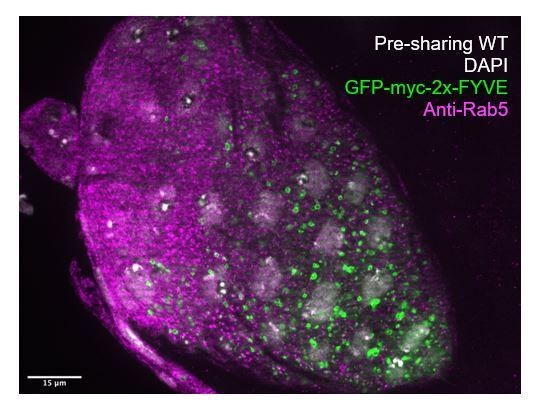
(A–A’) Endosome localization (GFP-myc-2x-FYVE), representative of (A) pre- and (A’) post-sharing onset. (B) Endosomes in shi RNAi post-sharing, see Methods. (C) Aggregated endosome line profiles for WT pre-sharing (N = 6, rep = 3), WT post-sharing (N = 7, rep = 2), and shi RNAi post-sharing (N = 10, rep = 2). Shaded area represents standard error. (D–D’) Shi-Venus localization pre- and post-sharing onset. (E) Line profiles as in (D–D’) (N = 4–5, rep = 3). (F–O) Representative Transmission Electron Micrographs (TEMs). (F–F’’) Microvillar-like structures (MV) pre- (F), mid- (F’), and post- (F’’) sharing onset. (G–G’’) Mitochondria and surrounding membrane pre- (G), mid- (G’), and post- (G’’) sharing onset. (H–J) Microvillar-like structures (MV) of adult papillae in WT (H), shi RNAi (I), and Rab5 RNAi (J). (K–M) Mitochondria and surrounding membranes of adult papillae in WT (K), shi RNAi (L), and Rab5 RNAi (M). Inset in (L) shows trapped vesicles. (N–O) WT and shi RNAi post-sharing. Adherens (orange), septate (green), and gap (blue) junctions are highlighted. (P) Quantification of the ratio of gap junction length to septate plus gap junction length (Fraction gap junction) (N = 3–4, rep = 2). p<0.0001 for the difference in gap junction ratio between WT and shi RNAi.
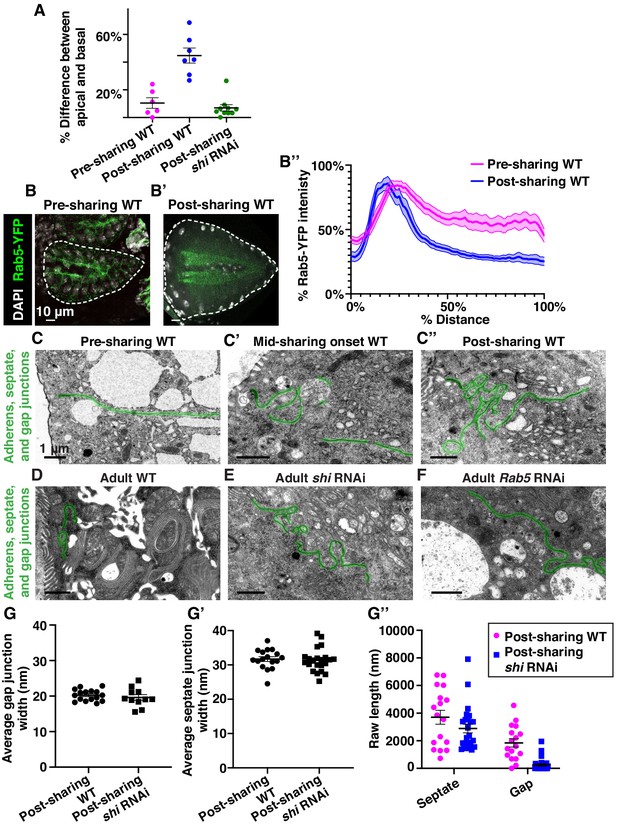
Changes in endosome polarity and apical junction shape accompany the onset of cytoplasm sharing.
(A) Quantification of the average endosome intensity difference between representative basal and apical areas across papillae in Figure 3A–C (N = 6–10, rep = 2). (B–B’) Representative localization of Rab5-YFP, green, before sharing onset (B) and after sharing onset (B’). (B’’) Aggregated line profiles of Rab5-YFP intensity before and after the beginning of sharing (N = 10, rep = 2). (C–C’’) Representative TEMs of apical (adherens, septate, and gap) junctions pre (C), mid (C’), and post (C’’) sharing onset. (D–F) Representative TEMs of apical junctions of post-sharing adult WT (D), shi RNAi (E), and Rab5 RNAi (F) papillar cells. (G–G’’) Apical junction electron micrograph measurements of post-sharing WT and shi RNAi pupal papillar cells (N = 3–4, rep = 2). Average gap junction (G) and septate junction (G’) widths were measured alongside gap and septate junction length. Width measurements were taken along the length of each cell–cell junction and averaged to give one point per cell–cell junction. (G’’) Raw septate and gap junction lengths (nm) that were used to calculate gap junction ratio in Figure 3P.
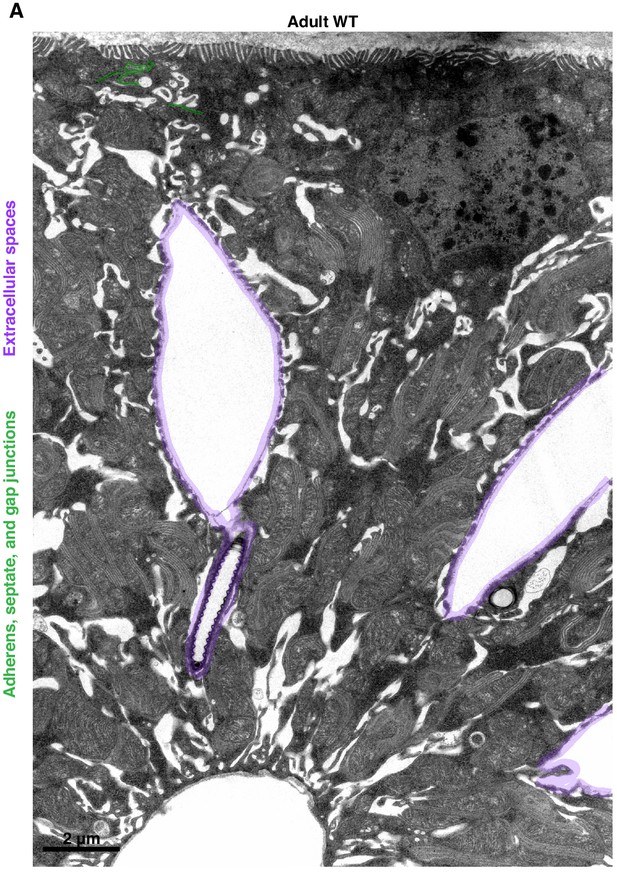
Extracellular spaces separate nuclei throughout much of the papillar lateral membrane.
(A) Representative TEM cross-section of an adult WT papilla. The apical edge facing the gut lumen is at the top; the basal edge facing the papillar central canal is at the bottom of the image.
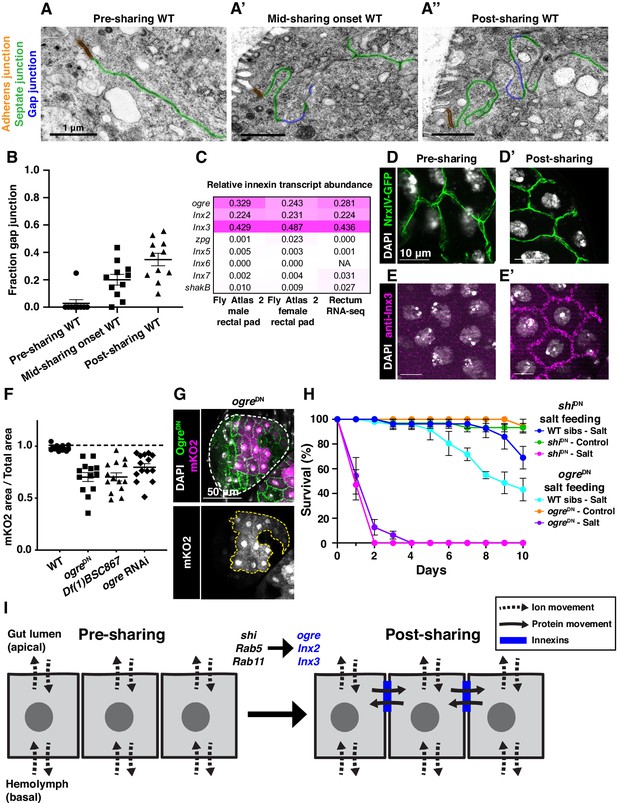
Gap junction proteins are required for cytoplasmic sharing.
(A–A’’) Representative apical junctions highlighted by junctional type in pre (A), mid (A’), and post (A’’) sharing onset. (B) Quantification of fraction gap junction (gap junction length / (gap + septate junction length)) in pre-, mid-, and post-sharing onset pupae (N = 3–4, rep = 2). (C) Drosophila innexin expression in the adult rectum (Methods). (D–D’) Adherens junctions in pre- (D) and post- (D’) sharing pupae visualized by NrxIV-GFP. (E–E’) WT pupae pre- and post-sharing onset stained with anti-Inx3. (F) Quantification of cytoplasm sharing in WT, ogreDN, Df(1)BSC867/+ (a 10-gene-deficiency covering ogre, Inx2, and Inx7), and ogre RNAi adult papillae (N = 13–14, rep = 2). (G) Representative adult rectal papilla expressing GFP-ogre and dBrainbow. (H) Survival of WT, shiDN, and ogreDN animals on a high-salt diet (N = 27–37, rep = 3). (I) Proposed model for cytoplasmic sharing in an intact papillar epithelium.
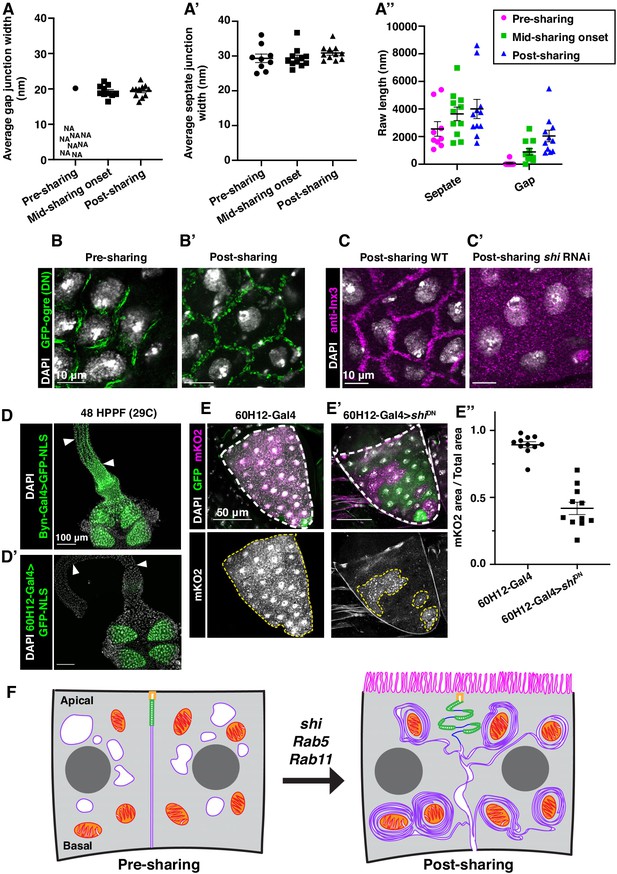
Gap junction formation coincides with cytoplasm sharing onset.
(A–A’’) Apical junction TEM measurements of pre-, mid-, and post-sharing onset pupal papillar cells (N = 3–4, rep = 2). Average gap junction (A) and septate junction (A’) widths were measured alongside gap and septate junction length. (A’’) Raw septate and gap junction lengths (nm) used to calculate gap junction ratio in Figure 4B. (B–B’) Gap junction localization visualized by UAS-GFP-ogre in pre (B) and post (B’) sharing onset pupae. (C–C’) Representative images of post-sharing WT and shi RNAi animals stained for anti-Inx3. (D) Representative image of byn-Gal4 driving GFPNLS expression throughout the pre-sharing hindgut. Arrows indicate the ileum. (D’) 60H12-Gal4 driving GFPNLS expression in pre-sharing papillae but not in the hindgut ileum or pylorus. Arrows indicate the ileum. (E) Representative image of 60H12-Gal4 driving dBrainbow in adult papillae. (E’) Representative image of 60H12-Gal4 driving shiDN expression in a dBrainbow background in adult papillae. (E’’) Quantification of cytoplasm sharing in 60H12-Gal4 and 60H12-Gal4 > shiDN animals (N = 11, rep = 2). (F) Model of membrane and junctional changes requiring membrane trafficking genes that coincide with the onset of cytoplasm sharing.
Tables
Cytoplasm sharing primary candidate screen gene results.
| Gene category | Gene | Annotation symbol | Gene ID | Sharing disrupted? |
|---|---|---|---|---|
| Autophagy | Atg1 | CG10967 | FBgn0260945 | No |
| Autophagy | Atg7 | CG5489 | FBgn0034366 | No |
| Autophagy | Atg8a | CG32672 | FBgn0052672 | No |
| Cell cycle/Chromosomes | blue | NA | FBgn0283709 | No |
| Cell cycle/Chromosomes | CapD2 | CG1911 | FBgn0039680 | No |
| Cell cycle/Chromosomes | Cdc2 | CG5363 | FBgn0004106 | Yes |
| Cell cycle/Chromosomes | Clamp | CG1832 | FBgn0032979 | No |
| Cell cycle/Chromosomes | endos | CG6513 | FBgn0061515 | No |
| Cell cycle/Chromosomes | fzr | CG3000 | FBgn0262699 | Yes |
| Cell cycle/Chromosomes | Mi-2 | CG8103 | FBgn0262519 | No |
| Cell cycle/Chromosomes | Rbp9 | CG3151 | FBgn0010263 | No |
| Cell cycle/Chromosomes | SA-2 | CG13916 | FBgn0043865 | No |
| Cell signaling | Chico | CG5686 | FBgn0024248 | No |
| Cell signaling | Egfr | CG10079 | FBgn0003731 | Yes |
| Cell signaling | grk | CG17610 | FBgn0001137 | No |
| Cell signaling | N | CG3936 | FBgn0004647 | No |
| Cell signaling | Ptp61F | CG9181 | FBgn0267487 | No |
| Cell signaling | rho | CG1004 | FBgn0004635 | Yes |
| Cell signaling | ru | CG1214 | FBgn0003295 | No |
| Cell signaling | spi | CG10334 | FBgn0005672 | No |
| Cell signaling | stet | CG33166 | FBgn0020248 | No |
| Cell signaling | wts | CG12072 | FBgn0011739 | No |
| Cell signaling | βggt-II | CG18627 | FBgn0028970 | No |
| Cytoskeleton | ALiX | CG12876 | FBgn0086346 | No |
| Cytoskeleton | Cdc42 | CG12530 | FBgn0010341 | No |
| Cytoskeleton | DCTN1-p150 | CG9206 | FBgn0001108 | No |
| Cytoskeleton | pav | CG1258 | FBgn0011692 | No |
| Cytoskeleton | wash | CG13176 | FBgn0033692 | No |
| Hindgut-enriched | dac | CG4952 | FBgn0005677 | No |
| Hindgut-enriched | Dr | CG1897 | FBgn0000492 | No |
| Hindgut-enriched | nrv3 | CG8663 | FBgn0032946 | No |
| Membrane component | Flo1 | CG8200 | FBgn0024754 | No |
| Membrane component | Flo2 | CG32593 | FBgn0264078 | No |
| Membrane component | Iris | CG4715 | FBgn0031305 | No |
| Myoblast fusion | Arf51F | CG8156 | FBgn0013750 | No |
| Myoblast fusion | Arp2 | CG9901 | FBgn0011742 | No |
| Myoblast fusion | Arp3 | CG7558 | FBgn0262716 | No |
| Myoblast fusion | Ced-12 | CG5336 | FBgn0032409 | No |
| Myoblast fusion | dock | CG3727 | FBgn0010583 | No |
| Myoblast fusion | hbs | CG7449 | FBgn0029082 | No |
| Myoblast fusion | Hem | CG5837 | FBgn0011771 | No |
| Myoblast fusion | mbc | CG10379 | FBgn0015513 | No |
| Myoblast fusion | Rac1 | CG2248 | FBgn0010333 | No |
| Myoblast fusion | Rho1 | CG8416 | FBgn0014020 | No |
| Myoblast fusion | rols | CG32096 | FBgn0041096 | No |
| Myoblast fusion | rst | CG4125 | FBgn0003285 | No |
| Myoblast fusion | SCAR | CG4636 | FBgn0041781 | No |
| Myoblast fusion | siz | CG32434 | FBgn0026179 | No |
| Myoblast fusion | WASp | CG1520 | FBgn0024273 | No |
| Polarity | Abi | CG9749 | FBgn0020510 | No |
| Polarity | CadN | CG7100 | FBgn0015609 | No |
| Polarity | cindr | CG31012 | FBgn0027598 | No |
| Polarity | cno | CG42312 | FBgn0259212 | No |
| Polarity | Gli | CG3903 | FBgn0001987 | No |
| Polarity | l(2)gl | CG2671 | FBgn0002121 | No |
| Polarity | Nrg | CG1634 | FBgn0264975 | No |
| Polarity | sdt | CG32717 | FBgn0261873 | No |
| Polarity | shg | CG3722 | FBgn0003391 | No |
| Vesicle trafficking | Atl | CG6668 | FBgn0039213 | No |
| Vesicle trafficking | Bet1 | CG14084 | FBgn0260857 | No |
| Vesicle trafficking | Chmp1 | CG4108 | FBgn0036805 | No |
| Vesicle trafficking | CHMP2B | CG4618 | FBgn0035589 | No |
| Vesicle trafficking | dnd | CG6560 | FBgn0038916 | No |
| Vesicle trafficking | Exo84 | CG6095 | FBgn0266668 | Yes |
| Vesicle trafficking | lerp | CG31072 | FBgn0051072 | No |
| Vesicle trafficking | Rab11 | CG5771 | FBgn0015790 | Yes |
| Vesicle trafficking | Rab23 | CG2108 | FBgn0037364 | No |
| Vesicle trafficking | Rab4 | CG4921 | FBgn0016701 | No |
| Vesicle trafficking | Rab7 | CG5915 | FBgn0015795 | No |
| Vesicle trafficking | Rab8 | CG8287 | FBgn0262518 | No |
| Vesicle trafficking | RabX4 | CG31118 | FBgn0051118 | No |
| Vesicle trafficking | Vha16-1 | CG3161 | FBgn0262736 | Yes |
| Vesicle trafficking | Vha55 | CG17369 | FBgn0005671 | No |
| Vesicle trafficking | VhaAC39-1 | CG2934 | FBgn0285910 | No |
| Vesicle trafficking | VhaAC39-2 | CG4624 | FBgn0039058 | No |
| Vesicle trafficking | Vps2 | CG14542 | FBgn0039402 | Yes |
| Vesicle trafficking | Vps33b | CG5127 | FBgn0039335 | No |
| Total screen results | ||||
| Sharing disrupted | 8 | |||
| No sharing phenotype | 69 | |||
| Total | 77 | |||
| Screen results by category | ||||
| Polarity | 9 | |||
| Vesicle trafficking | 19 | |||
| Myoblast fusion | 15 | |||
| Cell cycle/Chromosomes | 9 | |||
| Cell signaling | 11 | |||
| Autophagy | 3 | |||
| Cytoskeleton | 5 | |||
| Hindgut-enriched | 3 | |||
| Membrane component | 3 | |||
| Total | 77 |
Membrane trafficking primary and secondary candidate screen gene results.
| Gene category | Gene subcategory | Gene | Annotation symbol | Gene ID | Sharing disrupted? | Screen |
|---|---|---|---|---|---|---|
| Membrane trafficking | ER | Atl | CG6668 | FBgn0039213 | No | Primary |
| Membrane trafficking | ESCRT | Chmp1 | CG4108 | FBgn0036805 | No | Primary |
| Membrane trafficking | ESCRT | CHMP2B | CG4618 | FBgn0035589 | No | Primary |
| Membrane trafficking | ESCRT | lsn | CG6637 | FBgn0260940 | No | Secondary |
| Membrane trafficking | ESCRT | Vps2 | CG14542 | FBgn0039402 | Yes | Primary |
| Membrane trafficking | ESCRT | Vps4 | CG6842 | FBgn0283469 | No | Secondary |
| Membrane trafficking | Exocyst | Exo70 | CG7127 | FBgn0266667 | No | Secondary |
| Membrane trafficking | Exocyst | Exo84 | CG6095 | FBgn0266668 | Yes | Primary |
| Membrane trafficking | Exocyst | Sec10 | CG6159 | FBgn0266673 | Yes | Secondary |
| Membrane trafficking | Exocyst | Sec15 | CG7034 | FBgn0266674 | Yes | Secondary |
| Membrane trafficking | Exocyst | Sec5 | CG8843 | FBgn0266670 | Yes | Secondary |
| Membrane trafficking | Exocyst | Sec6 | CG5341 | FBgn0266671 | Yes | Secondary |
| Membrane trafficking | Exocyst | Sec8 | CG2095 | FBgn0266672 | Yes | Secondary |
| Membrane trafficking | Lysosome | lerp | CG31072 | FBgn0051072 | No | Primary |
| Membrane trafficking | Rab-associated | CG41099 | CG41099 | FBgn0039955 | No | Secondary |
| Membrane trafficking | Rab-associated | mtm | CG9115 | FBgn0025742 | No | Secondary |
| Membrane trafficking | Rab-associated | nuf | CG33991 | FBgn0013718 | No | Secondary |
| Membrane trafficking | Rab-associated | Rala | CG2849 | FBgn0015286 | No | Secondary |
| Membrane trafficking | Rab-associated | Rep | CG8432 | FBgn0026378 | No | Secondary |
| Membrane trafficking | Rab-associated | Rip11 | CG6606 | FBgn0027335 | No | Secondary |
| Membrane trafficking | Vacuolar H+ ATPase | Vha16-1 | CG3161 | FBgn0262736 | Yes | Primary |
| Membrane trafficking | Vacuolar H+ ATPase | Vha16-2 | CG32089 | FBgn0028668 | No | Secondary |
| Membrane trafficking | Vacuolar H+ ATPase | Vha16-3 | CG32090 | FBgn0028667 | No | Secondary |
| Membrane trafficking | Vacuolar H+ ATPase | Vha16-5 | CG6737 | FBgn0032294 | Yes | Secondary |
| Membrane trafficking | Vacuolar H+ ATPase | Vha55 | CG17369 | FBgn0005671 | No | Primary |
| Membrane trafficking | Vacuolar H+ ATPase | VhaAC39-1 | CG2934 | FBgn0285910 | No | Primary |
| Membrane trafficking | Vacuolar H+ ATPase | VhaAC39-2 | CG4624 | FBgn0039058 | No | Primary |
| Membrane trafficking | Vacuolar H+ ATPase | VhaPPA1-1 | CG7007 | FBgn0028662 | Yes | Secondary |
| Membrane trafficking | Vacuolar H+ ATPase | VhaPPA1-2 | CG7026 | FBgn0262514 | Yes | Secondary |
| Membrane trafficking | Vesicle trafficking | Bet1 | CG14084 | FBgn0260857 | No | Primary |
| Membrane trafficking | Vesicle trafficking | Chc | CG9012 | FBgn0000319 | No | Secondary |
| Membrane trafficking | Vesicle trafficking | dnd | CG6560 | FBgn0038916 | No | Primary |
| Membrane trafficking | Vesicle trafficking | shi | CG18102 | FBgn0003392 | Yes | Secondary |
| Membrane trafficking | Vesicle trafficking | Vps29 | CG4764 | FBgn0031310 | No | Secondary |
| Membrane trafficking | Vesicle trafficking | Vps33b | CG5127 | FBgn0039335 | No | Primary |
| Membrane trafficking | Vesicle trafficking | Vps35 | CG5625 | FBgn0034708 | No | Secondary |
| Total screen results | ||||||
| Sharing disrupted | 12 | |||||
| No sharing phenotype | 24 | |||||
| Total | 36 | |||||
| Screen results by category | Total | Hits | ||||
| ER | 1 | 0 | ||||
| ESCRT | 5 | 1 | ||||
| Exocyst | 7 | 6 | ||||
| Lysosome | 1 | 0 | ||||
| Rab-associated | 6 | 0 | ||||
| Vacuolar H+ ATPase | 9 | 4 | ||||
| Vesicle trafficking | 7 | 1 | ||||
| Total | 36 |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (D. melanogaster) | w1118 | Bloomington Drosophila Stock Center | BDSC:3605; FLYB:FBst0003605; RRID:BDSC_3605 | w1118 |
| Genetic reagent (D. melanogaster) | tub-Gal4 | Bloomington Drosophila Stock Center | BDSC:5138; FLYB:FBst0005138; RRID:BDSC_5138 | y1 w*; P{tubP-GAL4}LL7/TM3, Sb1 Ser1 |
| Genetic reagent (D. melanogaster) | tub-Gal80ts | NA | NA | NA |
| Genetic reagent (D. melanogaster) | UAS-dBrainbow | Bloomington Drosophila Stock Center; (Hampel et al., 2011) | BDSC:34513; FLYB:FBst0034513; RRID:BDSC_34513 | w1118; P{UAS-Brainbow}attP2 |
| Genetic reagent (D. melanogaster) | UAS-dBrainbow | Bloomington Drosophila Stock Center; (Hampel et al., 2011) | BDSC:34514; FLYB:FBst0034514; RRID:BDSC_34514 | w1118; P{UAS-Brainbow}attP40 |
| Genetic reagent (D. melanogaster) | Hsp70>cre | Bloomington Drosophila Stock Center | BDSC:851; FLYB:FBst0000851; RRID:BDSC_851 | y1 w67c23 P{Crey}1b; D*/TM3, Sb1 |
| Genetic reagent (D. melanogaster) | UAS-fzr RNAi | Vienna Drosophila Resource Center | VDRC:25550; FLYB:FBst0455950 | w1118; P{GD9960}v25550 |
| Genetic reagent (D. melanogaster) | UAS-shi RNAi #1 | Bloomington Drosophila Stock Center | BDSC:28513; FLYB:FBst0028513; RRID:BDSC_28513 | y1 v1; P{TRiP.JF03133}attP2 |
| Genetic reagent (D. melanogaster) | UAS-shi RNAi #2 | Bloomington Drosophila Stock Center | BDSC:36921; FLYB:FBst0036921; RRID:BDSC_36921 | y1 sc* v1 sev21; P{TRiP.HMS00154}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Rab5 RNAi #1 | Bloomington Drosophila Stock Center | BDSC:30518; FLYB:FBst0030518; RRID:BDSC_30518 | y1 v1; P{TRiP.JF03335}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Rab5 RNAi #2 | Bloomington Drosophila Stock Center | BDSC:67877; FLYB:FBst0067877; RRID:BDSC_67877 | y1 sc* v1 sev21; P{TRiP.GL01872}attP40 |
| Genetic reagent (D. melanogaster) | UAS-Rab11 RNAi #1 | Bloomington Drosophila Stock Center | BDSC:27730; FLYB:FBst0027730; RRID:BDSC_27730 | y1 v1; P{TRiP.JF02812}attP2 |
| Genetic reagent (D. melanogaster) | UAS-Rab11 RNAi #2 | Vienna Drosophila Resource Center | VDRC:22198; FLYB:FBst0454467 | w1118; P{GD11761}v22198 |
| Genetic reagent (D. melanogaster) | UAS-SCAR RNAi #1 | Bloomington Drosophila Stock Center | BDSC:36121; FLYB:FBst0036121; RRID:BDSC_36121 | y1 sc* v1 sev21; P{TRiP.HMS01536}attP40 |
| Genetic reagent (D. melanogaster) | UAS-SCAR RNAi #2 | Bloomington Drosophila Stock Center | BDSC:51803; FLYB:FBst0051803; RRID:BDSC_51803 | y1 v1; P{TRiP.HMC03361}attP40 |
| Genetic reagent (D. melanogaster) | UAS-kirre RNAi | Vienna Drosophila Resource Center | VDRC:27227; FLYB:FBst0456824 | w1118; P{GD14476}v27227 |
| Genetic reagent (D. melanogaster) | UAS-sns RNAi | Vienna Drosophila Resource Center | VDRC:877; FLYB:FBst0471238 | w1118; P{GD65}v877/TM3 |
| Genetic reagent (D. melanogaster) | UAS-schizo RNAi | Vienna Drosophila Resource Center | VDRC:36625; FLYB:FBst0461775 | w1118; P{GD14895}v36625 |
| Genetic reagent (D. melanogaster) | UAS-sing RNAi | Vienna Drosophila Resource Center | VDRC:12202; FLYB:FBst0450437 | w1118; P{GD3396}v12202/TM3 |
| Genetic reagent (D. melanogaster) | UAS-Cdc42DN | Bloomington Drosophila Stock Center | BDSC:6288; FLYB:FBst0006288; RRID:BDSC_6288 | w*; P{UAS-Cdc42.N17}3 |
| Genetic reagent (D. melanogaster) | UAS-Rac1DN | Bloomington Drosophila Stock Center | BDSC:6292; FLYB:FBst0006292; RRID:BDSC_6292 | y1 w*; P{UAS-Rac1.N17}1 |
| Genetic reagent (D. melanogaster) | UAS-Rho1DN | Bloomington Drosophila Stock Center | BDSC:7328; FLYB:FBst0007328; RRID:BDSC_7328 | w*; P{UAS-Rho1.N19}2.1 |
| Genetic reagent (D. melanogaster) | UAS-GFPNLS | Bloomington Drosophila Stock Center | BDSC:4776; FLYB:FBst0004776; RRID:BDSC_4776 | w1118; P{UAS-GFP.nls}8 |
| Genetic reagent (D. melanogaster) | UAS-GFP-Myc-2x-FYVE | Bloomington Drosophila Stock Center | BDSC:42712; FLYB:FBst0042712; RRID:BDSC_42712 | w*; P{UAS-GFP-myc-2xFYVE}2 |
| Genetic reagent (D. melanogaster) | UAS-YFP-Rab5 | Bloomington Drosophila Stock Center | BDSC:9775; FLYB:FBst0009775; RRID:BDSC_9775 | y1 w*; P{UASp-YFP.Rab5}Pde808b |
| Genetic reagent (D. melanogaster) | 60H12-Gal4 | Bloomington Drosophila Stock Center | BDSC:39268; FLYB:FBst0039268; RRID:BDSC_39268 | w1118; P{GMR60H12-GAL4}attP2 |
| Genetic reagent (D. melanogaster) | UAS-shiDN | Bloomington Drosophila Stock Center | BDSC:5822; FLYB:FBst0005822; RRID:BDSC_5822 | w*; TM3, P{UAS-shi.K44A}3-10/TM6B, Tb1 |
| Genetic reagent (D. melanogaster) | NrxIV-GFP | Bloomington Drosophila Stock Center | BDSC:50798; FLYB:FBst0050798; RRID:BDSC_50798 | y1 w*; P{PTT-GA}Nrx-IVCA06597 |
| Genetic reagent (D. melanogaster) | Df(1)BSC867 | Bloomington Drosophila Stock Center | BDSC:29990; FLYB:FBst0029990; RRID:BDSC_29990 | Df(1)BSC867, w1118/Binsinscy |
| Genetic reagent (D. melanogaster) | UAS-ogre RNAi | Vienna Drosophila Resource Center | VDRC:7136; FLYB:FBst0470569 | w1118; P{GD3264}v7136 |
| Genetic reagent (D. melanogaster) | byn-Gal4 | Singer et al., 1996 | FLYB:FBal0137290 | P{GawB}bynGal4 |
| Genetic reagent (D. melanogaster) | UAS-GFPPA | Lynn Cooley; McLean and Cooley, 2013 | FLYB:FBti0148163 | P{20XUAS-IVS-Syn21-mC3PA-GFP-p10} |
| Genetic reagent (D. melanogaster) | UAS-NDN | Rebay et al., 1993 | NA | NA |
| Genetic reagent (D. melanogaster) | UAS-shi-Venus | Stefano De Renzis; Fabrowski et al., 2013 | NA | NA |
| Genetic reagent (D. melanogaster) | UAS-GFP-ogre | Andrea Brand; Spéder and Brand, 2014 | FLYB:FBtp0127574 | ogreUAS.N.GFP |
| Genetic reagent (D. melanogaster) | UAS-Gapdh2-GFPPA | This paper | NA | Transgenic line created through gene synthesis and embryo injection. Codon-optimized D. melanogaster Gapdh2 fused to GFPPAunder UAS control. |
| Antibody | anti-GFP(Rabbit polyclonal) | Thermo Fisher Scientific | Cat# A11122; RRID:AB_221569 | IF (1:1000) |
| Antibody | anti-HA (Rat monoclonal) | Roche | Cat# 11867423001; RRID:AB_390918 | IF (1:100) |
| Antibody | anti-Inx3(Rabbit polyclonal) | Reinhard Bauer; Lehmann et al., 2006 | RRID:AB_2568555 | IF (1:75) |
| Antibody | Anti-Rabbit Alexa Fluor 488 (Goat) | Thermo Fisher Scientific | Cat# A32731; RRID:AB_2633280 | IF (1:2000) |
| Antibody | Anti-Rabbit Alexa Fluor 568 (Goat) | Thermo Fisher Scientific | Cat# A-11011; RRID:AB_143157 | IF (1:2000) |
| Antibody | Anti-Rat Alexa Fluor 633 (Goat) | Thermo Fisher Scientific | Cat# A-21094; RRID:AB_2535749 | IF (1:2000) |
| Other | DAPI stain | Sigma-Aldrich | Cat# D9542 | (1:5000) |
Primary and secondary candidate screen stock numbers used and results.
| Gene | Annotation symbol | Gene ID | Mutant or UAS transgene | Stock center | Stock number | Chr | Sharing disrupted? | Notes |
|---|---|---|---|---|---|---|---|---|
| Abi | CG9749 | FBgn0020510 | RNAi | BDSC | 51455 | 2 | No | |
| ALiX | CG12876 | FBgn0086346 | RNAi | BDSC | 33417 | 3 | No | |
| ALiX | CG12876 | FBgn0086346 | RNAi | BDSC | 50904 | 2 | No | |
| Arf51F | CG8156 | FBgn0013750 | RNAi | BDSC | 51417 | 3 | No | |
| Arf51F | CG8156 | FBgn0013750 | Mutant | BDSC | 17076 | 2 | No | |
| Arf51F | CG8156 | FBgn0013750 | RNAi | BDSC | 27261 | 3 | No | |
| Arp2 | CG9901 | FBgn0011742 | RNAi | BDSC | 27705 | 3 | No | |
| Arp3 | CG7558 | FBgn0262716 | RNAi | BDSC | 32921 | 3 | No | |
| Atg1 | CG10967 | FBgn0260945 | RNAi | BDSC | 44034 | 2 | No | |
| Atg1 | CG10967 | FBgn0260945 | RNAi | BDSC | 26731 | 3 | No | |
| Atg7 | CG5489 | FBgn0034366 | RNAi | BDSC | 34369 | 3 | No | |
| Atg7 | CG5489 | FBgn0034366 | RNAi | BDSC | 27707 | 3 | No | |
| Atg8a | CG32672 | FBgn0052672 | RNAi | BDSC | 28989 | 3 | No | |
| Atg8a | CG32672 | FBgn0052672 | RNAi | BDSC | 58309 | 2 | No | |
| Atg8a | CG32672 | FBgn0052672 | RNAi | BDSC | 34340 | 3 | No | |
| Atl | CG6668 | FBgn0039213 | RNAi | BDSC | 36736 | 2 | No | |
| Bet1 | CG14084 | FBgn0260857 | RNAi | BDSC | 41927 | 2 | No | |
| blue | NA | FBgn0283709 | RNAi | BDSC | 44094 | 3 | No | |
| blue | NA | FBgn0283709 | RNAi | BDSC | 41637 | 2 | No | |
| CadN | CG7100 | FBgn0015609 | RNAi | BDSC | 27503 | 3 | No | |
| CadN | CG7100 | FBgn0015609 | RNAi | BDSC | 41982 | 3 | No | |
| CapD2 | CG1911 | FBgn0039680 | Mutant | BDSC | 59393 | 3 | No | |
| Cdc2 | CG5363 | FBgn0004106 | RNAi | VDRC | 41838 | 3 | Yes | |
| Cdc2 | CG5363 | FBgn0004106 | RNAi | BDSC | NA | 3 | No | |
| Cdc42 | CG12530 | FBgn0010341 | RNAi | BDSC | 42861 | 2 | No | |
| Cdc42 | CG12530 | FBgn0010341 | DN | BDSC | 6288 | 2 | No | |
| Ced-12 | CG5336 | FBgn0032409 | RNAi | BDSC | 28556 | 3 | No | |
| Ced-12 | CG5336 | FBgn0032409 | RNAi | BDSC | 58153 | 2 | No | |
| Chc | CG9012 | FBgn0000319 | DN | BDSC | 26821 | 2 | No | |
| Chc | CG9012 | FBgn0000319 | RNAi | BDSC | 27350 | 3 | No | |
| Chc | CG9012 | FBgn0000319 | RNAi | BDSC | 34742 | 3 | No | |
| Chico | CG5686 | FBgn0024248 | RNAi | BDSC | 36788 | 2 | No | |
| Chmp1 | CG4108 | FBgn0036805 | RNAi | BDSC | 33928 | 3 | No | |
| CHMP2B | CG4618 | FBgn0035589 | RNAi | BDSC | 28531 | 3 | No | |
| CHMP2B | CG4618 | FBgn0035589 | RNAi | BDSC | 38375 | 2 | No | |
| cindr | CG31012 | FBgn0027598 | RNAi | BDSC | 35670 | 3 | No | |
| cindr | CG31012 | FBgn0027598 | RNAi | BDSC | 38976 | 2 | No | |
| Clamp | CG1832 | FBgn0032979 | RNAi | BDSC | 27080 | 3 | No | |
| cno | CG42312 | FBgn0259212 | RNAi | BDSC | 33367 | 3 | No | |
| cno | CG42312 | FBgn0259212 | RNAi | BDSC | 38194 | 2 | No | |
| dac | CG4952 | FBgn0005677 | RNAi | BDSC | 26758 | 3 | No | |
| dac | CG4952 | FBgn0005677 | RNAi | BDSC | 35022 | 3 | No | |
| DCTN1-p150 | CG9206 | FBgn0001108 | DN | BDSC | 51645 | 2 | No | |
| dnd | CG6560 | FBgn0038916 | RNAi | BDSC | 27488 | 3 | No | |
| dnd | CG6560 | FBgn0038916 | RNAi | BDSC | 34383 | 3 | No | |
| dock | CG3727 | FBgn0010583 | RNAi | BDSC | 27728 | 3 | No | |
| dock | CG3727 | FBgn0010583 | RNAi | BDSC | 43176 | 3 | No | |
| dock | CG3727 | FBgn0010583 | Mutant | BDSC | 11385 | 2 | No | |
| Dr | CG1897 | FBgn0000492 | RNAi | BDSC | 26224 | 3 | No | |
| Dr | CG1897 | FBgn0000492 | RNAi | BDSC | 42891 | 2 | No | |
| Egfr | CG10079 | FBgn0003731 | DN | BDSC | 5364 | 2 | Yes | |
| Egfr | CG10079 | FBgn0003731 | RNAi | VDRC | 43267 | 3 | Yes | |
| endos | CG6513 | FBgn0061515 | RNAi | BDSC | 53250 | 3 | No | |
| endos | CG6513 | FBgn0061515 | RNAi | BDSC | 65996 | 3 | No | |
| Exo70 | CG7127 | FBgn0266667 | RNAi | BDSC | 28041 | 3 | No | |
| Exo70 | CG7127 | FBgn0266667 | RNAi | BDSC | 55234 | 3 | No | |
| Exo84 | CG6095 | FBgn0266668 | RNAi | BDSC | 28712 | 3 | Yes | |
| Flo1 | CG8200 | FBgn0024754 | RNAi | BDSC | 36700 | 3 | No | |
| Flo1 | CG8200 | FBgn0024754 | RNAi | BDSC | 36649 | 2 | No | |
| Flo2 | CG32593 | FBgn0264078 | RNAi | BDSC | 55212 | 3 | No | |
| Flo2 | CG32593 | FBgn0264078 | RNAi | BDSC | 40833 | 2 | No | |
| fzr | CG3000 | FBgn0262699 | RNAi | VDRC | 25550 | 2 | Yes | |
| Gli | CG3903 | FBgn0001987 | RNAi | BDSC | 31869 | 3 | No | |
| Gli | CG3903 | FBgn0001987 | RNAi | BDSC | 58115 | 2 | No | |
| grk | CG17610 | FBgn0001137 | RNAi | BDSC | 38913 | 3 | No | |
| hbs | CG7449 | FBgn0029082 | RNAi | BDSC | 57003 | 2 | No | |
| Hem | CG5837 | FBgn0011771 | Mutant | BDSC | 8752 | 3 | No | |
| Hem | CG5837 | FBgn0011771 | Mutant | BDSC | 8753 | 3 | No | |
| Hem | CG5837 | FBgn0011771 | RNAi | BDSC | 29406 | 3 | No | |
| Hem | CG5837 | FBgn0011771 | RNAi | BDSC | 41688 | 3 | No | |
| Hsc70Cb | CG6603 | FBgn0026418 | RNAi | BDSC | 33742 | 3 | No | |
| Hsc70Cb | CG6603 | FBgn0026418 | DN | BDSC | 56497 | 2 | No | |
| Iris | CG4715 | FBgn0031305 | RNAi | BDSC | 50587 | 2 | No | |
| Iris | CG4715 | FBgn0031305 | RNAi | BDSC | 63582 | 2 | No | |
| l(2)gl | CG2671 | FBgn0002121 | RNAi | BDSC | 31517 | 3 | No | |
| lerp | CG31072 | FBgn0051072 | RNAi | BDSC | 57436 | 2 | No | |
| lilli | CG8817 | FBgn0041111 | RNAi | BDSC | 26314 | 3 | No | |
| lilli | CG8817 | FBgn0041111 | RNAi | BDSC | 34592 | 3 | No | |
| mbc | CG10379 | FBgn0015513 | RNAi | BDSC | 32355 | 3 | No | |
| mbc | CG10379 | FBgn0015513 | RNAi | BDSC | 33722 | 3 | No | |
| Mi-2 | CG8103 | FBgn0262519 | RNAi | BDSC | 16876 | 3 | No | |
| mtm | CG9115 | FBgn0025742 | RNAi | BDSC | 38339 | 3 | No | |
| N | CG3936 | FBgn0004647 | DN | Rebay Lab | NA | 2 | No | |
| N | CG3936 | FBgn0004647 | RNAi | Sara Bray | NA | 1 | No | |
| Nrg | CG1634 | FBgn0264975 | RNAi | BDSC | 28724 | 3 | No | |
| Nrg | CG1634 | FBgn0264975 | RNAi | BDSC | 38215 | 2 | No | |
| Nrg | CG1634 | FBgn0264975 | RNAi | BDSC | 37496 | 2 | No | |
| nrv3 | CG8663 | FBgn0032946 | RNAi | BDSC | 29431 | 3 | No | |
| nrv3 | CG8663 | FBgn0032946 | RNAi | BDSC | 50725 | 3 | No | |
| nuf | CG33991 | FBgn0013718 | RNAi | BDSC | 31493 | 3 | No | |
| pav | CG1258 | FBgn0011692 | RNAi | BDSC | 35649 | 3 | No | |
| pav | CG1258 | FBgn0011692 | RNAi | BDSC | 43963 | 2 | No | |
| Ptp61F | CG9181 | FBgn0267487 | RNAi | BDSC | 32426 | 3 | No | |
| Ptp61F | CG9181 | FBgn0267487 | RNAi | BDSC | 56036 | 2 | No | |
| Rab1 | CG3320 | FBgn0285937 | CA | BDSC | 9758 | 3 | No | |
| Rab1 | CG3320 | FBgn0285937 | DN | BDSC | 9757 | 3 | Yes | Requires 60H12-Gal4 |
| Rab1 | CG3320 | FBgn0285937 | RNAi | BDSC | 27299 | 3 | Yes | |
| Rab1 | CG3320 | FBgn0285937 | RNAi | BDSC | 34670 | 3 | No | |
| Rab2 | CG3269 | FBgn0014009 | CA | BDSC | 9761 | 2 | No | |
| Rab2 | CG3269 | FBgn0014009 | DN | BDSC | 9759 | 2 | No | |
| Rab3 | CG7576 | FBgn0005586 | CA | BDSC | 9764 | 3 | No | |
| Rab3 | CG7576 | FBgn0005586 | DN | BDSC | 9766 | 2 | No | |
| Rab4 | CG4921 | FBgn0016701 | CA | BDSC | 9770 | 3 | No | |
| Rab4 | CG4921 | FBgn0016701 | DN | BDSC | 9768 | 2 | No | |
| Rab4 | CG4921 | FBgn0016701 | DN | BDSC | 9769 | 3 | No | |
| Rab5 | CG3664 | FBgn0014010 | CA | BDSC | 9773 | 3 | Yes | |
| Rab5 | CG3664 | FBgn0014010 | DN | BDSC | 42704 | 3 | Yes | Requires 60H12-Gal4 |
| Rab5 | CG3664 | FBgn0014010 | RNAi | BDSC | 67877 | 2 | Yes | |
| Rab5 | CG3664 | FBgn0014010 | RNAi | BDSC | 30518 | 3 | Yes | |
| Rab5 | CG3664 | FBgn0014010 | RNAi | BDSC | 51847 | 2 | No | |
| Rab6 | CG6601 | FBgn0015797 | CA | BDSC | 9776 | 3 | No | |
| Rab6 | CG6601 | FBgn0015797 | DN | BDSC | 23250 | 3 | No | |
| Rab7 | CG5915 | FBgn0015795 | CA | BDSC | 9779 | 3 | No | |
| Rab7 | CG5915 | FBgn0015795 | DN | BDSC | 9778 | 3 | No | |
| Rab7 | CG5915 | FBgn0015795 | DN | BDSC | 9778 | 3 | No | |
| Rab8 | CG8287 | FBgn0262518 | DN | BDSC | 9780 | 3 | No | |
| Rab8 | CG8287 | FBgn0262518 | CA | BDSC | 9781 | 2 | No | |
| Rab8 | CG8287 | FBgn0262518 | DN | BDSC | 9780 | 3 | No | |
| Rab9 | CG9994 | FBgn0032782 | CA | BDSC | 9785 | 3 | No | |
| Rab9 | CG9994 | FBgn0032782 | DN | BDSC | 23642 | 3 | No | |
| Rab10 | CG17060 | FBgn0015789 | CA | BDSC | 9787 | 3 | No | |
| Rab10 | CG17060 | FBgn0015789 | DN | BDSC | 9786 | 3 | No | |
| Rab11 | CG5771 | FBgn0015790 | CA | BDSC | 9791 | 3 | No | |
| Rab11 | CG5771 | FBgn0015790 | DN | BDSC | 23261 | 3 | Yes | |
| Rab11 | CG5771 | FBgn0015790 | RNAi | BDSC | 27730 | 3 | Yes | |
| Rab11 | CG5771 | FBgn0015790 | RNAi | VDRC | 108382 | 2 | Yes | |
| Rab11 | CG5771 | FBgn0015790 | RNAi | VDRC | 22198 | 3 | Yes | |
| Rab11 | CG5771 | FBgn0015790 | Mutant | BDSC | 42708 | 3 | Yes | |
| Rab14 | CG4212 | FBgn0015791 | CA | BDSC | 9795 | 2 | No | |
| Rab14 | CG4212 | FBgn0015791 | DN | BDSC | 23264 | 3 | No | |
| Rab18 | CG3129 | FBgn0015794 | CA | BDSC | 9797 | 3 | No | |
| Rab18 | CG3129 | FBgn0015794 | DN | BDSC | 23238 | 3 | No | |
| Rab19 | CG7062 | FBgn0015793 | CA | BDSC | 9800 | 3 | No | |
| Rab19 | CG7062 | FBgn0015793 | DN | BDSC | 9799 | 3 | No | |
| Rab21 | CG17515 | FBgn0039966 | CA | BDSC | 23864 | 2 | No | |
| Rab21 | CG17515 | FBgn0039966 | DN | BDSC | 23240 | 3 | No | |
| Rab23 | CG2108 | FBgn0037364 | RNAi | BDSC | 36091 | 3 | No | |
| Rab23 | CG2108 | FBgn0037364 | RNAi | BDSC | 55352 | 2 | No | |
| Rab23 | CG2108 | FBgn0037364 | CA | BDSC | 9806 | 3 | No | |
| Rab23 | CG2108 | FBgn0037364 | DN | BDSC | 9804 | 3 | No | |
| Rab26 | CG34410 | FBgn0086913 | CA | BDSC | 23243 | 3 | No | |
| Rab26 | CG34410 | FBgn0086913 | DN | BDSC | 9808 | 3 | No | |
| Rab27 | CG14791 | FBgn0025382 | CA | BDSC | 9811 | 2 | No | |
| Rab27 | CG14791 | FBgn0025382 | DN | BDSC | 23267 | 2 | No | |
| Rab30 | CG9100 | FBgn0031882 | CA | BDSC | 9814 | 2 | No | |
| Rab30 | CG9100 | FBgn0031882 | DN | BDSC | 9813 | 3 | No | |
| Rab32 | CG8024 | FBgn0002567 | CA | BDSC | 23280 | 3 | No | |
| Rab32 | CG8024 | FBgn0002567 | DN | BDSC | 23281 | 2 | No | |
| Rab35 | CG9575 | FBgn0031090 | CA | BDSC | 9817 | 3 | No | |
| Rab35 | CG9575 | FBgn0031090 | DN | BDSC | 9820 | 3 | No | |
| Rab39 | CG12156 | FBgn0029959 | CA | BDSC | 9823 | 3 | No | |
| Rab39 | CG12156 | FBgn0029959 | DN | BDSC | 23247 | 3 | No | |
| Rab40 | CG1900 | FBgn0030391 | CA | BDSC | 9827 | 3 | No | |
| Rab40 | CG1900 | FBgn0030391 | DN | BDSC | 9829 | 2 | No | |
| Rab9D | CG32678 | FBgn0067052 | CA | BDSC | 9835 | 3 | No | |
| Rab9D | CG32678 | FBgn0067052 | DN | BDSC | 23257 | 2 | No | |
| Rab9E | CG32673 | FBgn0052673 | CA | BDSC | 9832 | 2 | No | |
| Rab9E | CG32673 | FBgn0052673 | DN | BDSC | 23255 | 3 | No | |
| Rab9Fb | CG32670 | FBgn0052670 | CA | BDSC | 9844 | 3 | No | |
| Rab9Fb | CG32670 | FBgn0052670 | DN | BDSC | 9845 | 2 | No | |
| RabX1 | CG3870 | FBgn0015372 | CA | BDSC | 9839 | 2 | No | |
| RabX1 | CG3870 | FBgn0015372 | DN | BDSC | 23252 | 3 | No | |
| RabX2 | CG2885 | FBgn0030200 | CA | BDSC | 9842 | 3 | No | |
| RabX2 | CG2885 | FBgn0030200 | DN | BDSC | 9843 | 2 | No | |
| RabX4 | CG31118 | FBgn0051118 | RNAi | BDSC | 28704 | 3 | No | |
| RabX4 | CG31118 | FBgn0051118 | RNAi | BDSC | 44070 | 2 | No | |
| RabX4 | CG31118 | FBgn0051118 | CA | BDSC | 23277 | 2 | No | |
| RabX4 | CG31118 | FBgn0051118 | DN | BDSC | 9849 | 3 | No | |
| RabX5 | CG7980 | FBgn0035255 | CA | BDSC | 9852 | X | No | |
| RabX5 | CG7980 | FBgn0035255 | DN | BDSC | 9853 | 2 | No | |
| RabX6 | CG12015 | FBgn0035155 | CA | BDSC | 9855 | 2 | No | |
| RabX6 | CG12015 | FBgn0035155 | DN | BDSC | 9856 | 3 | No | |
| CG41099 | CG41099 | FBgn0039955 | RNAi | BDSC | 34883 | 3 | No | |
| Rac1 | CG2248 | FBgn0010333 | RNAi | BDSC | 28985 | 3 | No | |
| Rac1 | CG2248 | FBgn0010333 | DN | BDSC | 6292 | 3 | No | |
| Rala | CG2849 | FBgn0015286 | DN | BDSC | 32094 | 2 | No | |
| Rala | CG2849 | FBgn0015286 | RNAi | BDSC | 34375 | 3 | No | |
| Rbp9 | CG3151 | FBgn0010263 | RNAi | BDSC | 42796 | 3 | No | |
| Rep | CG8432 | FBgn0026378 | RNAi | BDSC | 28047 | 3 | No | |
| rho | CG1004 | FBgn0004635 | Mutant | BDSC | 1471 | 3 | Yes | |
| rho | CG1004 | FBgn0004635 | RNAi | BDSC | 38920 | 3 | Yes | |
| rho | CG1004 | FBgn0004635 | RNAi | BDSC | 41699 | 2 | Yes | |
| Rho1 | CG8416 | FBgn0014020 | DN | BDSC | 7328 | 3 | No | |
| Rho1 | CG8416 | FBgn0014020 | DN | BDSC | 58818 | 2 | No | |
| Rho1 | CG8416 | FBgn0014020 | RNAi | BDSC | 32383 | 3 | No | |
| Rip11 | CG6606 | FBgn0027335 | RNAi | BDSC | 38325 | 3 | No | |
| rols | CG32096 | FBgn0041096 | RNAi | BDSC | 56986 | 2 | No | |
| rols | CG32096 | FBgn0041096 | RNAi | BDSC | 58262 | 2 | No | |
| rst | CG4125 | FBgn0003285 | RNAi | BDSC | 28672 | 3 | No | |
| ru | CG1214 | FBgn0003295 | RNAi | BDSC | 41593 | 3 | No | |
| ru | CG1214 | FBgn0003295 | RNAi | BDSC | 58065 | 2 | No | |
| SA-2 | CG13916 | FBgn0043865 | RNAi | VDRC | 108267 | 2 | No | |
| SCAR | CG4636 | FBgn0041781 | RNAi | BDSC | 31126 | 3 | No | |
| SCAR | CG4636 | FBgn0041781 | RNAi | BDSC | 51803 | 2 | No | |
| SCAR | CG4636 | FBgn0041781 | Mutant | BDSC | 8754 | 2 | No | |
| sdt | CG32717 | FBgn0261873 | RNAi | BDSC | 33909 | 3 | No | |
| sdt | CG32717 | FBgn0261873 | RNAi | BDSC | 35291 | 3 | No | |
| Sec10 | CG6159 | FBgn0266673 | RNAi | BDSC | 27483 | 3 | Yes | |
| Sec15 | CG7034 | FBgn0266674 | RNAi | BDSC | 27499 | 3 | Yes | |
| Sec5 | CG8843 | FBgn0266670 | RNAi | VDRC | 28873 | 3 | Yes | |
| Sec5 | CG8843 | FBgn0266670 | RNAi | BDSC | 50556 | 3 | No | |
| Sec6 | CG5341 | FBgn0266671 | RNAi | VDRC | 105836 | 2 | Yes | |
| Sec6 | CG5341 | FBgn0266671 | RNAi | BDSC | 27314 | 3 | Yes | |
| Sec8 | CG2095 | FBgn0266672 | RNAi | BDSC | 57441 | 2 | Yes | |
| shg | CG3722 | FBgn0003391 | RNAi | BDSC | 27689 | 3 | No | |
| shi | CG18102 | FBgn0003392 | DN | BDSC | 5822 | 3 | Yes | Requires 60H12-Gal4 |
| shi | CG18102 | FBgn0003392 | RNAi | BDSC | 28513 | 3 | Yes | |
| shi | CG18102 | FBgn0003392 | RNAi | BDSC | 36921 | 3 | Yes | |
| siz | CG32434 | FBgn0026179 | RNAi | BDSC | 39060 | 2 | No | |
| spi | CG10334 | FBgn0005672 | RNAi | BDSC | 28387 | 3 | No | |
| spi | CG10334 | FBgn0005672 | RNAi | BDSC | 34645 | 3 | No | |
| stet | CG33166 | FBgn0020248 | RNAi | BDSC | 57698 | 3 | No | |
| Vha16-1 | CG3161 | FBgn0262736 | RNAi | BDSC | 40923 | 2 | Yes | |
| Vha16-1 | CG3161 | FBgn0262736 | RNAi | VDRC | 104490 | 2 | Yes | |
| Vha16-1 | CG3161 | FBgn0262736 | RNAi | VDRC | 49291 | 2 | Yes | |
| Vha16-2 | CG32089 | FBgn0028668 | RNAi | BDSC | 65167 | 2 | No | |
| Vha16-3 | CG32090 | FBgn0028667 | RNAi | BDSC | 57474 | 2 | No | |
| Vha16-5 | CG6737 | FBgn0032294 | RNAi | BDSC | 25803 | 3 | Yes | |
| Vha55 | CG17369 | FBgn0005671 | RNAi | BDSC | 40884 | 2 | No | |
| VhaAC39-1 | CG2934 | FBgn0285910 | RNAi | BDSC | 35029 | 3 | No | |
| VhaAC39-2 | CG4624 | FBgn0039058 | Mutant | BDSC | 62725 | 3 | No | |
| VhaAC39-2 | CG4624 | FBgn0039058 | RNAi | VDRC | 34303 | 2 | No | |
| VhaPPA1-1 | CG7007 | FBgn0028662 | RNAi | BDSC | 57729 | 2 | Yes | |
| VhaPPA1-2 | CG7026 | FBgn0262514 | RNAi | BDSC | 65217 | 2 | Yes | |
| Vps2 | CG14542 | FBgn0039402 | RNAi | VDRC | 24869 | 3 | Yes | |
| Vps2 | CG14542 | FBgn0039402 | RNAi | BDSC | 38995 | 2 | Yes | |
| lsn | CG6637 | FBgn0260940 | RNAi | BDSC | 38289 | 2 | No | |
| Vps29 | CG4764 | FBgn0031310 | RNAi | BDSC | 53951 | 2 | No | |
| Vps33b | CG5127 | FBgn0039335 | RNAi | BDSC | 44006 | 2 | No | |
| Vps35 | CG5625 | FBgn0034708 | RNAi | BDSC | 38944 | 2 | No | |
| Vps4 | CG6842 | FBgn0283469 | RNAi | BDSC | 31751 | 3 | No | |
| wts | CG12072 | FBgn0011739 | RNAi | BDSC | 41899 | 3 | No | |
| wash | CG13176 | FBgn0033692 | RNAi | BDSC | 62866 | 2 | No | |
| WASp | CG1520 | FBgn0024273 | RNAi | BDSC | 25955 | 3 | No | |
| WASp | CG1520 | FBgn0024273 | RNAi | BDSC | 51802 | 2 | No | |
| βggt-II | CG18627 | FBgn0028970 | RNAi | BDSC | 50516 | 2 | No | |
| βggt-II | CG18627 | FBgn0028970 | RNAi | BDSC | 34902 | 3 | No |
Fly stocks used in addition to the screens.
| Stock name | Stock number | Origin | References |
|---|---|---|---|
| w1118 | 3605 | BDSC | |
| tub-Gal4 | 5138 | BDSC | |
| tub-Gal80ts | NA | NA | |
| UAS-dBrainbow | 34513 | BDSC | Hampel et al., 2011 |
| UAS-dBrainbow | 34514 | BDSC | Hampel et al., 2011 |
| Hsp70 > cre | 851 | BDSC | |
| UAS-fzr RNAi | 25550 | VDRC | Fox et al., 2010; Schoenfelder et al., 2014 |
| UAS-shi RNAi #1 | 28513 | BDSC | |
| UAS-shi RNAi #2 | 36921 | BDSC | |
| UAS-Rab5 RNAi #1 | 30518 | BDSC | |
| UAS-Rab5 RNAi #2 | 67877 | BDSC | |
| UAS-Rab11 RNAi #1 | 27730 | BDSC | |
| UAS-Rab11 RNAi #2 | 22198 | VDRC | |
| UAS-SCAR RNAi #1 | 36121 | BDSC | Bischoff et al., 2013 |
| UAS-SCAR RNAi #2 | 51803 | BDSC | Xing et al., 2018 |
| UAS-kirre RNAi | 27227 | VDRC | Linneweber et al., 2015 |
| UAS-sns RNAi | 877 | VDRC | Linneweber et al., 2015 |
| UAS-schizo RNAi | 36625 | VDRC | Johnson et al., 2011 |
| UAS-sing RNAi | 12202 | VDRC | Brunetti et al., 2015 |
| UAS-Cdc42DN | 6288 | BDSC | |
| UAS-Rac1DN | 6292 | BDSC | |
| UAS-Rho1DN | 7328 | BDSC | |
| UAS-GFPNLS | 4776 | BDSC | |
| UAS-GFP-Myc-2x-FYVE | 42712 | BDSC | Gillooly et al., 2000; Wucherpfennig et al., 2003 |
| UAS-YFP-Rab5 | 9775 | BDSC | |
| 60H12-Gal4 | 39268 | BDSC | |
| UAS-shiDN | 5822 | BDSC | |
| NrxIV-GFP | 50798 | BDSC | |
| Df(1)BSC867 | 29990 | BDSC | |
| UAS-ogre RNAi | 7136 | VDRC | Holcroft et al., 2013; Spéder and Brand, 2014 |
| byn-Gal4 | - | NA | Singer et al., 1996 |
| UAS-GFPPA | - | Lynn Cooley | Datta et al., 2008 |
| UAS-NDN | - | NA | Rebay et al., 1993 |
| UAS-shi-Venus | - | Stefano De Renzis | Fabrowski et al., 2013 |
| UAS-GFP-ogre | - | Andrea Brand | Spéder and Brand, 2014 |
| UAS-Gapdh2-GFPPA | - | - | This paper |
Additional Methods.
| Panel | Additional methods |
|---|---|
| Figure 1—figure supplement 1F-F'' | Hsp70 > cre; UAS-dBrainbow; byn-Gal4 papillae dissected at 62 (D), 69 (D’), or 80 (D’’) hours post-puparium formation (HPPF) at 25°C. Hindguts were stained with Rabbit anti-GFP (Thermo-Fisher, A11122, 1:1000), Rat anti-HA (Sigma, 3F10, 1:100), and DAPI at 5 μg/ml. |
| Figure 1G | Hsp70 > cre; UAS-dBrainbow; byn-Gal4 papillae dissected at various HPPF at 25°C. The area labeled by mKO2 was divided by total papillar area. |
| Figure 1H | Hsp70 > cre; UAS-dBrainbow; byn-Gal4 papillae live-imaged at 69HPPF at 25°C. |
| Figure 1H' | Fluorescence intensity measured in neighboring cells during sharing onset (1H). |
| Figure 1I-I' | byn-Gal4/UAS-GFPPA, live-imaged during adulthood. Single secondary and principal cells were photoactivated and imaged every 3 s. |
| Figure 2A | UAS-RNAis and dominant-negative versions of 77 genes representing a wide range of cellular roles were screened (Hsp70 > cre; UAS-dBrainbow; byn-Gal4) for sharing defects. Animals expressing both UAS-dBrainbow and an UAS-driven RNAi or mutant gene were raised at 25°C and shifted to 29°C at L3. If a given RNAi or DN line was lethal when expressed with the byn-Gal4 driver, a Gal80ts was crossed in and the animals raised at 18°C with a shift to 29°C at pupation. Given the robustness of cytoplasmic sharing in WT animals, gene knockdowns or mutants with even single cell defects in sharing were considered ‘hits’. |
| Figure 2B | Secondary screen of 36 genes representing various categories of membrane trafficking (Hsp70 > cre; UAS-dBrainbow; byn-Gal4) for sharing defects. Animals expressing both UAS-dBrainbow and an UAS-driven RNAi were raised at 25°C and shifted to 29°C at L3. If a given RNAi line was lethal when expressed with the byn-Gal4 driver, a Gal80ts was crossed in and the animals raised at 18°C with a shift to 29°C at pupation. Given the robustness of cytoplasmic sharing in WT animals, gene knockdowns with even single cell defects in sharing were considered ‘hits’. |
| Figure 2C | Secondary screen (Hsp70 > cre; UAS-dBrainbow; byn-Gal4) of dominant-negative and constitutively-active variants of the Drosophila Rab GTPases. UAS-Rab11DN and UAS-Rab14DN required a Gal80ts repressor and temperature shifts from 18 to 29°C at pupation. UAS-Rab1DN and UAS-Rab5DN required papillar-specific expression using an alternative Gal4 driver (60 H12-Gal4), Gal80ts repressor, and temperature shifts from 18 to 29°C at pupation. |
| Figure 2D | Hsp70 > cre; UAS-dBrainbow; byn-Gal4, Gal80ts animals dissected pre-sharing (48 HPPF at 29°C). |
| Figure 2D' | Hsp70 > cre; UAS-dBrainbow; byn-Gal4, Gal80ts animals raised at 18°C and shifted to 29°C at pupation and dissected post-sharing (young adult). |
| Figure 2E | Young adult animals expressing UAS-shi RNAi #1 in a Hsp70 > cre; UAS-dBrainbow; byn-Gal4, Gal80ts background. Animals were shifted from 18 to 29°C at pupation to maximize RNAi and minimize animal lethality. |
| Figure 2F | Young adult animals expressing UAS-Rab5 RNAi #1 in a Hsp70 > cre; UAS-dBrainbow; byn-Gal4, Gal80ts background. Animals were shifted from 18 to 29°C at 1–2 days PPF to maximize RNAi and minimize animal lethality. |
| Figure 2G | Young adult animals expressing UAS-Rab11 RNAi #2 in a Hsp70 > cre; UAS-dBrainbow; byn-Gal4, Gal80ts background. Animals were shifted from 18 to 29°C at 1–2 days PPF to maximize RNAi and minimize animal lethality. |
| Figure 2H | Animals were shifted and dissected as in 2D-G. Additionally, Hsp70 > cre; UAS-dBrainbow; byn-Gal4, Gal80ts animals expressing UAS-shi RNAi #2 were raised at 18°C and shifted to 29°C at pupation, animals expressing UAS-Rab5 RNAi #2 were raised at 18°C and shifted to 29°C at L3, and animals expressing UAS-Rab11 RNAi #1 were raised at 18°C and shifted to 29°C at 1–2 days PPF. |
| Figure 3A-A' | Pupae expressing the early and late endosome marker UAS-GFP-myc-2x-FYVE were dissected pre (A, 48HPPF at 29°C) and post (A’, 72HPPF at 29°C) sharing onset. |
| Figure 3B | Pupae expressing UAS-GFP-myc-2x-FYVE in a UAS-shi RNAi #1 background at a post-sharing time point (24HPPF at 18°C + 72 hr at 29°C). |
| Figure 3C | Aggregated line profiles of UAS-GFP-myc-2x-FYVE intensity across papilla. |
| Figure 3D-D' | Pupae expressing UAS-shi-Venus were dissected pre (D, 48HPPF at 29°C) and post (D’, 72HPPF at 29°C) sharing onset. |
| Figure 3E | Aggregated line profiles of Shi-Venus intensity from the basal (0% distance) to the apical (100% distance) edges of the papilla. See 3C. |
| Figure 3F-F'' | Transmission electron micrographs of the microvillar-like structures of pupal papillae pre (F, 60HPPF at 25°C), mid (F’, 66HPPF at 25°C), and post (F’’, 69HPPF at 25°C) cytoplasm sharing onset. |
| Figure 3G-G'' | Electron micrographs of mitochondria and surrounding membrane material pre (G, 60HPPF at 25°C), mid (G’, 66HPPF at 25°C), and post (G’’, 69HPPF at 25°C) |
| Figure 3H | Electron micrograph of microvillar-like structures of WT (w1118) young adult papillar cells. |
| Figure 3I | Electron micrograph of microvillar-like structures of young adult byn-Gal4, Gal80ts, UAS-shi RNAi #2 (raised at 18°C, shifted at pupation to 29°C). |
| Figure 3J | Electron micrograph of microvillar-like structures of young adult byn-Gal4, Gal80ts, UAS-Rab5 RNAi #1 animals (raised at 18°C, shifted at 1–2 days PPF to 29°C). |
| Figure 3K | Electron micrograph of mitochondria and surrounding membrane material of WT (w1118) young adult papillar cells. |
| Figure 3L | Electron micrograph of mitochondria and surrounding membrane material of young adult byn-Gal4, Gal80ts, UAS-shi RNAi #2 (raised at 18°C, shifted at pupation to 29°C). |
| Figure 3M | Electron micrograph of mitochondria and surrounding membrane material of young adult byn-Gal4, Gal80ts, UAS-Rab5 RNAi #1 animals (raised at 18°C, shifted at 1–2 days PPF to 29°C). |
| Figure 3N | Electron micrograph of post-sharing WT (TM3/UAS-shi RNAi #1) pupa (24HPPF at 18°C, shifted to 29°C for 50 hr, then dissected) |
| Figure 3O | Electron micrograph of post-sharing byn-Gal4, Gal80ts,UAS-shi RNAi #1 pupa (24HPPF at 18°C, shifted to 29°C for 50 hr, then dissected) |
| Figure 3P | Gap junction length / (gap junction length + septate junction length) measured in WT and UAS-shi RNAi #1 pupae (see 3N-3O). Each point represents an image of a junction. |
| Figure 4A-A'' | Electron micrographs of apical junctions (adherens, septate, and gap) pre (A, 60HPPF at 25°C), mid (A’, 66HPPF at 25°C), and post (A’’, 69HPPF at 25°C) |
| Figure 4B | Gap junction length / (gap junction length + septate junction length) measured in pupae pre (60HPPF at 25°C), mid (66HPPF at 25°C), and post (69HPPF at 25°C) sharing onset. Each point represents an image of a junction. |
| Figure 4C | Relative innexin transcript abundance (innexin X transcripts/total innexin transcripts) using data from Fly Atlas 2 (Leader et al., 2018) and RNA-seq of adult w1118 rectums performed in the Fox Lab. |
| Figure 4D-D' | Pupae with endogenously GFP-tagged NrxIV (NrxIV-GFP) dissected pre (D, 48HPPF) and post (D', 72HPPF) sharing onset. |
| Figure 4E-E' | Pupae stained with Inx3 antibody (gift from Reinhard Bauer, rabbit, 1:75) pre (E, 48HPPF) and post (E', 58HPPF, papillae do not stain well at later timepoints) sharing onset. |
| Figure 4F | Young adult animals expressing no transgene (WT), UAS-ogreDN, UAS-ogre RNAi, or containing a deficiency covering ogre, Inx2, and Inx7 in a Hsp70 > cre; UAS-dBrainbow; byn-Gal4, Gal80ts background. Animals were raised at 25°C until L3 and then shifted to 29°C until dissection at young adulthood. |
| Figure 4G | See Figure 4F. |
| Figure 4H | 60 H12-Gal4, Gal80ts driving UAS-shiDN and WT siblings were shifted from 18 to 29°C at pupation. byn-Gal4, Gal80ts driving UAS-ogreDN animals and WT siblings were raised at 25°C and shifted to 29°C at L3. Animals 1–3 days post-eclosion were sorted into sex-matched groups and fed a control diet or a high salt (2% NaCl) diet. Survival was assessed once per day for 10 days. |
| Figure 1—figure supplement 1A | Hsp70 > cre; UAS-dBrainbow; tubulin-Gal4 animals raised at 29°C. Tissues dissected at adulthood. |
| Figure 1—figure supplement 1D | byn-Gal4/UAS-Gapdh2-GFPPA raised at 29°C and live-imaged during adulthood. Principal cells were photoactivated and imaged every 15 s. |
| Figure 1—figure supplement 1E | Hsp70 > cre; UAS-dBrainbow; byn-Gal4 animals were shifted from 25 to 29°C during L3 and dissected at adulthood. |
| Figure 1—figure supplement 1F | Hsp70 > cre; UAS-dBrainbow/UAS-fzr RNAi; byn-Gal4 animals were shifted from 25 to 29°C during L2 to maximize fzr knock down during endocycling. Animals were dissected at adulthood. |
| Figure 1—figure supplement 1G | Hsp70 > cre; UAS-dBrainbow; byn-Gal4/UAS-NDN animals were shifted from 25 to 29°C during L3 to ensure maximum UAS-NDN expression during mitoses. Animals were dissected at adulthood. |
| Figure 2—figure supplement 1A | Hsp70 > cre; UAS-dBrainbow; byn-Gal4, Gal80ts animals expressing various previously published myoblast fusion RNAis raised at 25°C and shifted to 29°C at L3 and dissected post-sharing (young adult). |
| Figure 2—figure supplement 1B | Hsp70 > cre; UAS-dBrainbow; byn-Gal4, Gal80ts animals expressing various previously published UAS-dominant-negative active regulators raised at 18°C and shifted to 29°C at L3 and dissected post-sharing (young adult). |
| Figure 2—figure supplement 1C | Papillar cells were identified using byn-Gal4, Gal80ts, driving UAS-GFPNLS expression. Cells were counted in one, z-sectioned half of the papillae and multiplied by two to give an approximate cell count. |
| Figure 2—figure supplement 1D | Hsp70 > cre; UAS-dBrainbow; byn-Gal4, Gal80ts animals were raised at 18°C until 3–4 days PPF and shifted to 29°C and dissected at young adulthood. |
| Figure 2—figure supplement 1E | Hsp70 > cre; UAS-dBrainbow; byn-Gal4, Gal80ts animals expressing UAS-shi RNAi #1 were raised at 18°C until 3–4 days PPF and shifted to 29°C and dissected at young adulthood. |
| Figure 3—figure supplement 1A | See Figure 3A-C. Basal and apical membrane defined as 10–20% and 90–100% total distance of papillae, respectively. |
| Figure 3—figure supplement 1B-B' | byn-Gal4 > UAS-Rab5-YFP animals dissected pre (48HPPF, 29°C) and post (72HPPF, 29°C) sharing onset. |
| Figure 3—figure supplement 1B'' | See Figure 3—figure supplement 1B-B' and Figure 3C. |
| Figure 3—figure supplement 1C-C'' | Electron micrographs of apical junctions (adherens, septate, and gap) pre (D, 60HPPF at 25°C), mid (D’, 66HPPF at 25°C), and post (D’’, 69HPPF at 25°C) |
| Figure 3—figure supplement 1D | Electron micrograph of apical junctions (adherens, septate, and gap) of WT (w1118) young adult papillar cells. |
| Figure 3—figure supplement 1E | Electron micrograph of apical junctions (adherens, septate, and gap) of young adult byn-Gal4, Gal80tsts, UAS-shi RNAi #2 (raised at 18°C, shifted at pupation to 29°C). |
| Figure 3—figure supplement 1F | Electron micrograph of apical junctions (adherens, septate, and gap) of young adult byn-Gal4, Gal80tsts, UAS-Rab5 RNAi #1 animals (raised at 18°C, shifted at 1–2 days PPF to 29°C). |
| Figure 3—figure supplement 1G | See Figure 3N-O. Junction width was measured throughout and averaged per image. Each point represents one image of a junction. |
| Figure 3—figure supplement 1G' | See Figure 3N-O. Junction width was measured throughout and averaged per image. Each point represents one image of a junction. |
| Figure 3—figure supplement 1G'' | See Figure 3N-O. Raw lengths shown were used to calculate ‘fraction gap junction’ in 3P. Each point represent one image of a junction. |
| Figure 3—figure supplement 2A | TEM of young adult (w1118) papilla. |
| Figure 4—figure supplement 1A | See Figure 4A-B. Junction width was measured throughout and averaged per image. Each point represents one image of a junction. |
| Figure 4—figure supplement 1A' | See Figure 4A-B. Junction width was measured throughout and averaged per image. Each point represents one image of a junction. |
| Figure 4—figure supplement 1A'' | See Figure 4A-B. Raw lengths shown were used to calculate ‘fraction gap junction’ in 3P. Each point represent one image of a junction. |
| Figure 4—figure supplement 1B-B' | Pupae expressing byn-Gal4, Gal80tsts, UAS-ogreDN (UAS-GFP-ogre) dissected pre (B, 48HPPF, 29°C) and post (B', 72HPPF, 29°C) sharing onset. |
| Figure 4—figure supplement 1C | byn-Gal4, Gal80ts pupae raised at 18°C until 0HPPF and then shifted to 29°C until dissection at 58HPPF. Pupal rectums were stained with Inx3 antibody (gift from Reinhard Bauer, rabbit, 1:75). |
| Figure 4—figure supplement 1C' | byn-Gal4, Gal80tsts, UAS-shi RNAi #2 pupae raised at 18°C until 0HPPF and then shifted to 29°C until dissection at 58HPPF. Pupal rectums were stained with Inx3 antibody (gift from Reinhard Bauer, rabbit, 1:75). |
| Figure 4—figure supplement 1D | byn-Gal4 > UAS-GFPNLS dissected pre (48HPPF, 29°C) sharing onset. |
| Figure 4—figure supplement 1D' | 60H12-Gal4 > UAS-GFPNLS dissected pre (48HPPF, 29°C) sharing onset. The pan-hindgut driver used in previous experiments, brachyenteron (byn-Gal4), causes animal lethality with shi, Rab5, and Rab11 knockdown within a few days. We therefore screened for and identified an alternative, papillae-specific driver (60H12-Gal4), derived from regulatory sequences of the hormone receptor gene Proctolin Receptor. 60H12-Gal4 > shiDN animals are viable on a control diet allowing us to test papillar function on a high-salt diet. |
| Figure 4—figure supplement 1E | Hsp70 > cre; UAS-dBrainbow; 60H12-Gal4 animals raised at 18°C and shifted to 29°C at pupation and dissected as young adults. |
| Figure 4—figure supplement 1E' | Hsp70 > cre; UAS-dBrainbow; 60H12-Gal4 / UAS-shiDN animals raised at 18°C and shifted to 29°C at pupation and dissected as young adults. |
| Figure 4—figure supplement 1E'' | See Figure 4—figure supplement 1E-E'. |
Additional statistics.
| Panel | N (animals) per group | Bio. reps | Statistical test | P-value |
|---|---|---|---|---|
| Figure 1G | 9–18 | 2 | Unpaired t-test | 66HPPF:74HPPF < 0.0001 |
| Figure 2H | 9–32 | 2–3 | One-way ANOVA with Tukey's multiple comparisons test | ANOVA:<0.0001 Pre:WT < 0.0001 WT:shi #1 < 0.0001 WT:shi #2 < 0.0001 WT:Rab5 #1 < 0.0001 WT:Rab5 #2 < 0.0001 WT:Rab11 #1 < 0.0001 WT:Rab11 #2 < 0.0001 shi #1:Rab5 #2 0.0181 shi #1:Rab11 #2 0.0428 shi #2:Rab5 #2 0.0263 Rab5 #1:Rab5 #2 0.0009 Rab5 #1:Rab11 #2 0.0020 all others, ns |
| Figure 3C | 6–10 | 2–3 | see 3-S1A | see Figure 3—figure supplement 1A |
| Figure 3E | 4–5 | 3 | Unpaired t-test | Apical region: Pre:Post < 0.0001 |
| Figure 3P | 3–4 | 2 | Unpaired t-test | WT:shi RNAi < 0.0001 |
| Figure 4B | 3–4 | 2 | Unpaired t-test | Pre:Post < 0.0001 |
| Figure 4F | 13–14 | 2 | One-way ANOVA with Tukey's multiple comparisons test | ANOVA:<0.0001 WT:ogreDN < 0.0001 WT:Df < 0.0001 WT:ogre RNAi 0.0007 |
| Figure 4H | 27–37 | 3 | One-way ANOVA with Tukey's multiple comparisons test (mean death at 10 days in each group) | ANOVA:<0.0001 WTsalt:shiDNreg ns, 0.7173 WTsalt:shiDNsalt < 0.0001 shiDNsalt:shiDNreg < 0.0001 ANOVA:<0.0001 WTsalt:ogreDNreg < 0.0001 WTsalt:ogreDNsalt < 0.0001 ogreDNsalt:ogreDNreg < 0.0001 |
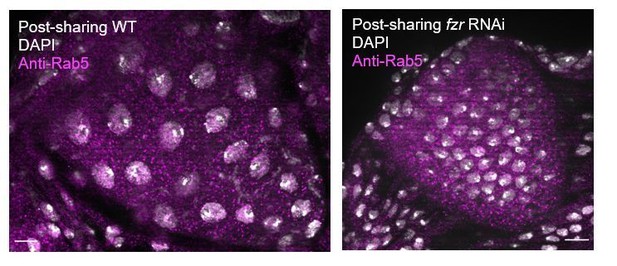
| Figure 1—figure supplement 1H | 12–20 | 2 | Unpaired t-test | WT:fzr RNAi < 0.0001 WT:NDN ns, 0.1786 |
| Figure 2—figure supplement 1A | 8–11 | 2 | One-way ANOVA with Tukey's multiple comparisons test | ANOVA:<0.0001 Sing RNAi:all others < 0.0001 All others: ns |
| Figure 2—figure supplement 1B | 6–8 | 2 | One-way ANOVA | ANOVA: ns, 0.3692 |
| Figure 2—figure supplement 1C | 11–23 | 2 | One-way ANOVA with Tukey's multiple comparisons test | ANOVA: 0.0044 shi RNAi #1:Rab11 RNAi #1 0.0244 Rab5 RNAi #2:Rab11 RNAi #1 0.0193 All others: ns |
| Figure 2—figure supplement 1F | 10–11 | 2 | Unpaired t-test | ns, 0.0782 |
| Figure 3—figure supplement 1A | 6–10 | 2 | One-way ANOVA with Tukey's multiple comparisons test | ANOVA:<0.0001 Pre:Post < 0.0001 Pre:shi RNAi ns, 0.7882 Post:shi RNAi < 0.0001 |
| Figure 3—figure supplement 1B'' | 10 | 2 | Unpaired t-test | Apical basal difference (see 1-S3A) Pre:Post 0.0007 |
| Figure 3—figure supplement 1G | 3–4 | 2 | Unpaired t-test | ns, 0.2203 |
| Figure 3—figure supplement 1G' | 3–4 | 2 | Unpaired t-test | ns, 0.4754 |
| Figure 3—figure supplement 1G'' | 3–4 | 2 | Multiple unpaired t-tests | Septate: WT:shi RNAi ns, 0.1547 Gap: WT:shi RNAi < 0.0001 |
| Figure 4—figure supplement 1A | 3–4 | 2 | One-way ANOVA | ns, 0.8973 |
| Figure 4—figure supplement 1A' | 3–4 | 2 | One-way ANOVA | ns, 0.3994 |
| Figure 4—figure supplement 1A'' | 3–4 | 2 | Multiple unpaired t-tests | Septate: all ns Gap: Pre:Post 0.0004 Gap: all others, ns |
| Figure 4—figure supplement 1E'' | 11 | 2 | Unpaired t-test | WT:shiDN < 0.0001 |