Identification of ubiquitin Ser57 kinases regulating the oxidative stress response in yeast
Figures
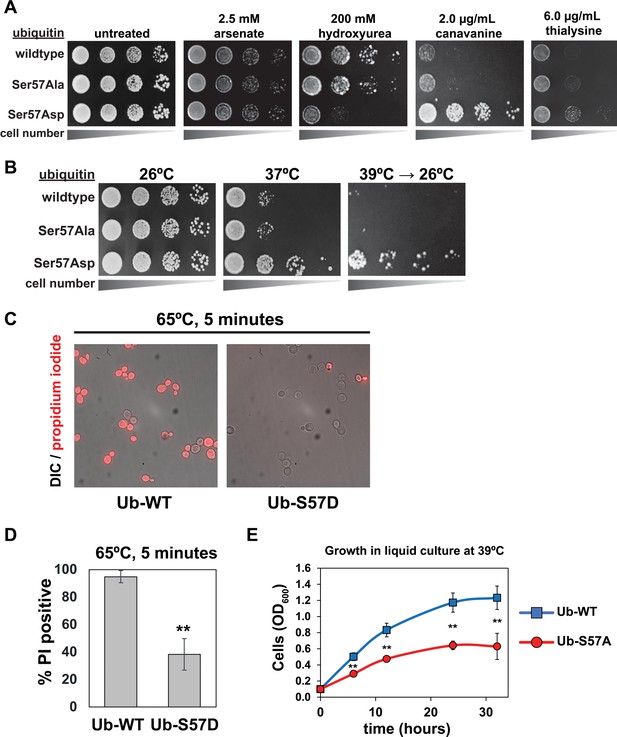
Genetic evidence of a role for Ser57 phosphorylation in the cellular response to stress.
Ubiquitin variants were shuffled into the SUB280 yeast background (as described in the Materials and Methods) to generate strains that express exclusively wildtype, S57A, or S57D ubiquitin. (A) Yeast strains expressing the indicated ubiquitin variants (wildtype, S57A or S57D) were plated in serial dilutions onto the indicated synthetic dextrose medium (SDM) agar plates. (B) Serial dilutions of yeast cells were plated onto YPD agar plates and incubated at 26°C, 37°C, or 39°C for 18 hr and shifted back to 26°C for recovery. (C) Fluorescence microscopy and (D) quantification of cells stained with propidium iodide (PI) after incubation at 65°C for 5 min. Dead cells are stained with PI. Results are means (n = 3) of % PI-positive cells ± SD (error bars). Double asterisk (**) indicates p value < 0.005 using a Student’s t-test. (E) With a starting OD600 of 0.1, cells were grown in liquid culture (-URA synthetic medium) at 39°C and OD600 was measured at different time points. Data was collected from four biological replicate experiments (n=4) and the mean OD600 ± SD (error bars) was calculated for each time point. Double asterisk (**) indicates p value < 0.005 using a Student’s t-test. For all statistical analysis associated with Figure 1, p values are reported in the Figure 1—source data 1.
-
Figure 1—source data 1
Quantification and statistical analysis for viability stain measurements (Figure 1D) and cell growth measurements (Figure 1E and Figure 1—figure supplement 1).
- https://cdn.elifesciences.org/articles/58155/elife-58155-fig1-data1-v2.xlsx
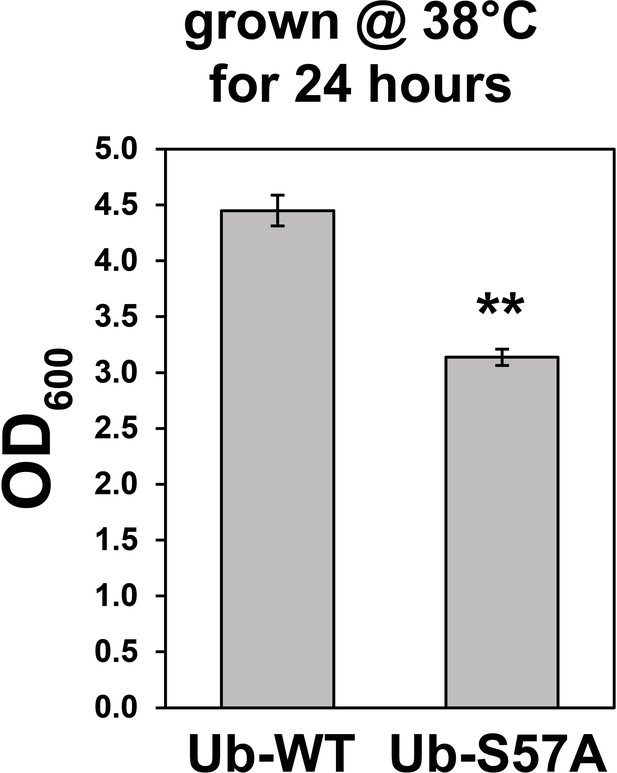
Yeast cells expressing S57A ubiquitin are sensitive to thermal stress.
Cells in liquid culture (-URA synthetic medium) were grown at 38°C with a starting OD600 of 0.1 and growth was measured after 24 hr of incubation. Values are means of OD600 from three replicates (n = 3) ± SD (error bars). Double asterisk (**) indicates p value < 0.005 using a Student’s t-test. For all statistical analysis associated with this figure supplement, p values are reported in the Figure 1—source data 1.
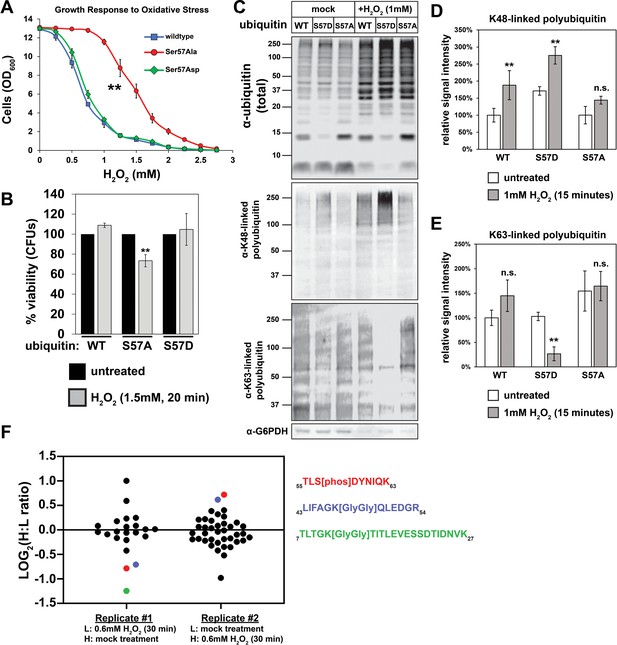
Ser57 phosphorylation of ubiquitin is important for the yeast oxidative stress response.
(A) OD600 of yeast cells cultured in YPD with different concentrations of H2O2 for 48 hr (starting OD600 = 0.025). Data was collected for four biological replicate experiments (n=4) and the means ± SD (error bars) were calculated. Double asterisk (**) indicates p value < 0.005 between the range of 0.5 mM to 2.5 mM H2O2 using a Student’s t-test, which was only observed for yeast expressing S57A ubiquitin. (B) CFU count of cells treated with 1.5 mM H2O2 for 20 min. Untreated control was taken before H2O2 treatment. Data was collected for three bioloigcal replicate experiments (n = 3) and the means ± SD (error bars) were calculated. Double asterisk (**) indicates p value < 0.05 comparing untreated to H2O2 treated cells using a Student’s t-test, which revealed that only yeast expressing S57A ubiquitin experienced a significant loss of viability during H2O2 treatment in this experiment. (C) Western blot with anti-ubiquitin, anti-K48 ubiquitin, and anti-K63 ubiquitin antibodies of lysates from cells treated with 1 mM H2O2 for 30 min. (D and E) Quantification of results from the experiment shown in (C) is presented as the mean of four biological replicate experiments (n = 4) ± SD (error bars). Double asterisk (**) indicates p value < 0.005 comparing untreated to H2O2 treated cells using a Student’s t-test. Correspondingly, p values > 0.05 are labeled as not significant (n.s.). Cells used in A-E are SUB280-derived strains exclusively expressing WT, S57A, or S57D ubiquitin variants. (F) SILAC-MS of affinity purified 3XFLAG-ubiquitin from yeast cells (JMY1312) treated with 0.6 mM H2O2 for 30 minutes. The peptide corresponding to Ser57 phosphorylation is colored red, while peptides corresponding to K48- and K11-linked poly-ubiquitin are colored blue and green, respectively. All other ubiquitin-derived peptide measurements are represented with black dots. All individual measurements and p values derived from statistical tests associated with the data in this figure are reported in Figure 2—source data 1.
-
Figure 2—source data 1
Quantification and statistical analysis for the growth (Figure 2A), cell viability (Figure 2B), and anti-K48/K63 western blot analysis (Figure 2D–E) of cells in the presence or absence of H2O2.
The source data for both replicates in Figure 2F are also included.
- https://cdn.elifesciences.org/articles/58155/elife-58155-fig2-data1-v2.xlsx
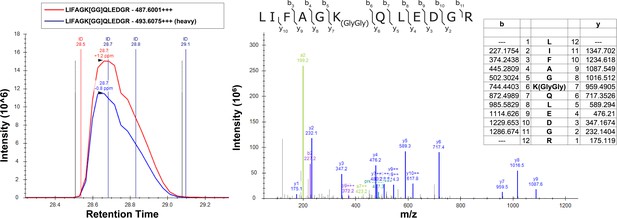
SILAC-MS fragmentation spectra of the K48-linked ubiquitin peptide isolated from yeast cells grown in the presence (light) or absence (heavy) of H2O2.
The left panel shows the chromatography of the heavy and light peptides, while the right panel shows the fragmentation pattern which confirms the identity of the peptide. Theoretical masses of b/y ions are tabulated and MS-observed masses are shown in the spectra.
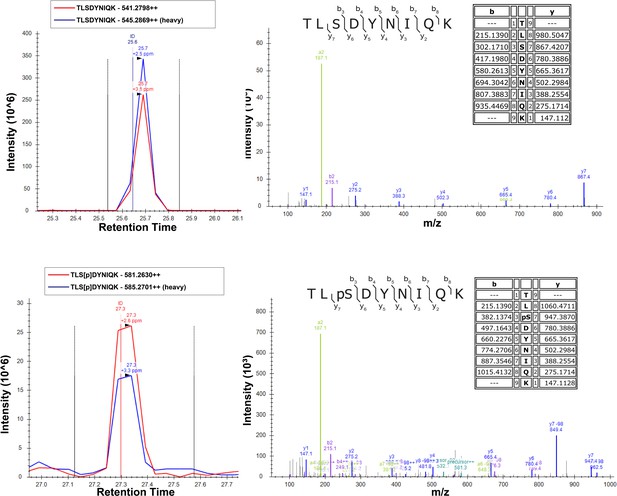
SILAC-MS fragmentation spectra of unmodified (top) and Ser57-phosphorylated (bottom) peptides of ubiquitin isolated from yeast cells grown in the presence (light) or absence (heavy) of H2O2.
Cells were grown in SILAC media (supplemented with light or heavy lysine and arginine) to the mid-log phase (OD600 of 0.6–0.7) and treated with 0.6 mM H2O2 for 30 min. Chromosomally expressed 3xFLAG-tagged ubiquitin (from the RPS31 and RPL40B loci) was isolated by affinity purification and digested with trypsin. Phosphopeptides were enriched by immobilized metal affinity chromatography (IMAC), separated by a capillary reverse-phase analytical column, and analyzed on a Q Exactive mass spectrometer. Theoretical masses of b/y ions are tabulated and MS-observed masses are shown in the spectra.
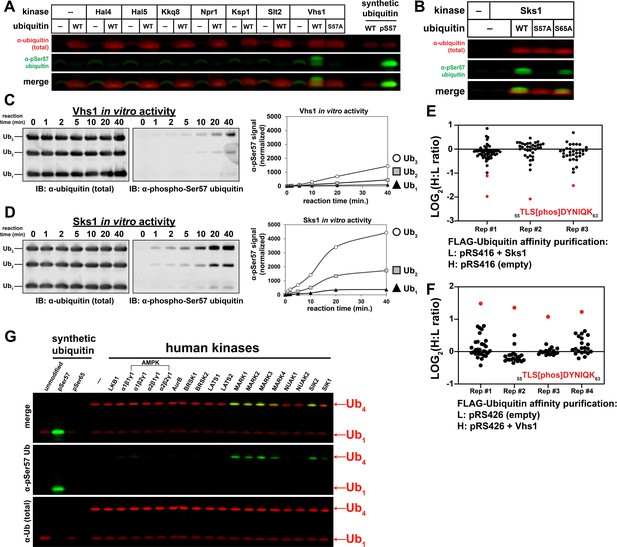
A subset of the Snf1-related family of kinases phosphorylates ubiquitin at the Ser57 position.
(A) Anti-phospho-Ser57 western blot of E. coli Rosetta 2 (DE3) lysates co-expressing ubiquitin and yeast kinases. (B) Anti-phospho-Ser57 western blot of E. coli Rosetta 2 (DE3) lysates co-expressing ubiquitin (wildtype, S57A, or S65A variants) and Sks1, a paralog of Vhs1. (C and D) In vitro reconstitution of ubiquitin Ser57 phosphorylation using purified recombinant Vhs1 (C) and Sks1 (D). Ubiquitin monomers as well as linear (M1-linked) dimers and trimers were included in equal amounts. Numerical values above each lane represent reaction time for the sample. (E and F) SILAC-MS of IP-enriched 3XFLAG ubiquitin from yeast cells (JMY1312 background) with either empty vector or with vector for overexpression of Sks1 (E) or Vhs1 (F). Black and red dots indicate resolved ubiquitin peptides and Ser57-phosphopeptides, respectively. All individual measurements are reported in Figure 3—source data 1. (G) In vitro activity assay of select human kinases of the AMPK family using equal amounts of mono-ubiquitin and M1-linked tetra-ubiquitin as substrates.
-
Figure 3—source data 1
Source measurements for the H:L ratio of ubiquitin peptides resolved in all replicate experiments of SUB280 cells overexpressing Sks1 (Figure 3E) or Vhs1 (Figure 3F).
- https://cdn.elifesciences.org/articles/58155/elife-58155-fig3-data1-v2.xlsx
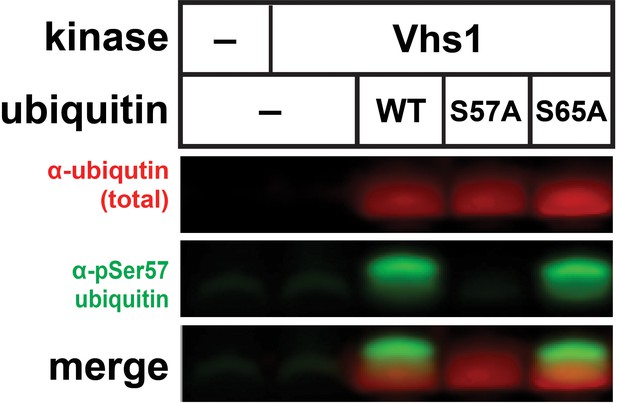
Anti-phospho-Ser57 western blot of E.coli Rosetta 2 (DE3) lysates co-expressing ubiquitin (wildtype, S57A, or S65A) and Vhs1.
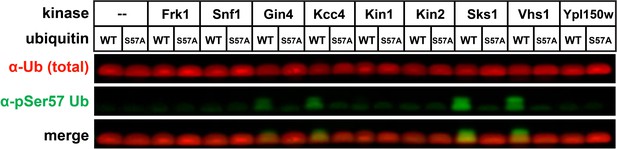
Anti-phospho-Ser57 western blot of E.coli Rosetta 2 (DE3) whole-cell lysates after heterologous co-expression of ubiquitin and yeast members of the Snf1-related family of kinases.
This is an extension of the results shown in Figure 3A.
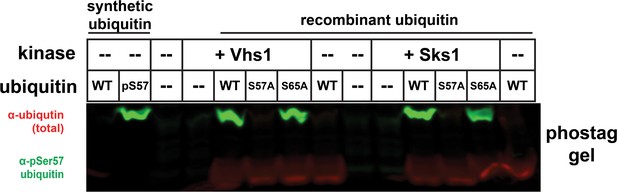
Phos-tag gel separation of ubiquitin (wildtype, S57A, or S65A variants) following co-expression with Vhs1 or Sks1.
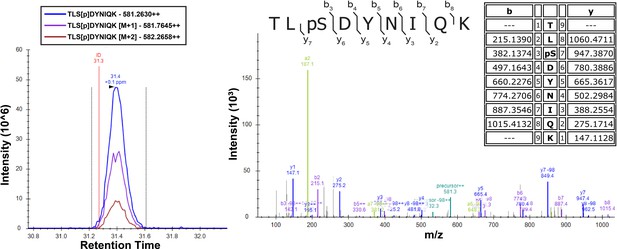
Detection of Ser57 phosphorylation of ubiquitin in the presence of Vhs1 by mass spectrometry.
Vhs1 and ubiquitin of yeast were co-expressed in E. coli Rosetta two by IPTG induction. Ubiquitin was isolated using UBD-capture pull-down, digested with trypsin, and analyzed on a Q Exactive mass spectrometer. Theoretical masses of b/y ions are tabulated and MS-observed masses are shown in the spectra.
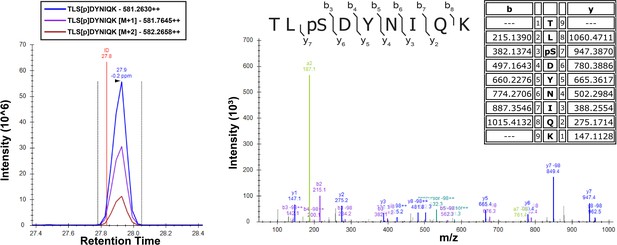
Detection of Ser57 phosphorylation of ubiquitin in the presence of Sks1 by mass spectrometry.
In vitro kinase activity assay was carried out using purified recombinant Sks1 and tetra-ubiquitin as described in the Materials and methods. Total peptide digests of the reaction mixture were separated by reverse-phase capillary column and analyzed on a Q Exactive mass spectrometer. Theoretical masses of b/y ions are tabulated and MS-observed masses are shown in the spectra.
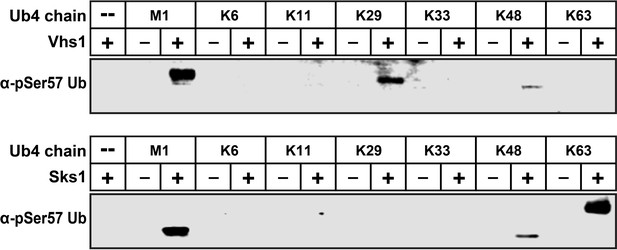
Vhs1 and Sks1 have distinct linkage-type preferences.
In vitro reconstitution assay was carried out by mixing recombinant Vhs1 (top) or Sks1 (bottom) and tetra-ubiquitin with different linkage types (M1, K6, K11, K29, K33, K48, K63) as described in the Materials and methods. Ser57 phosphorylation was detected by western blot using an anti-phospho-Ser57 ubiquitin antibody.
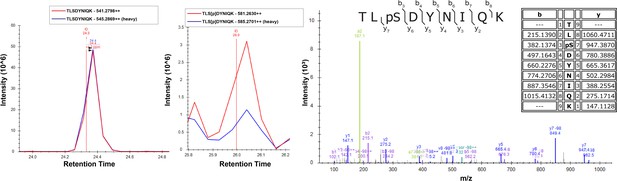
Overexpression of Sks1 in yeast cells increases Ser57 phosphorylation of ubiquitin.
Cells were grown in SILAC media supplemented with light or heavy lysine and arginine. Chromosomally expressed 3xFLAG-tagged ubiquitin (from the RPS31 and RPL40B loci) was isolated by affinity purification and digested with trypsin. Phosphopeptides were enriched by immobilized metal affinity chromatography (IMAC), separated by a capillary reverse-phase analytical column, and analyzed on a Q Exactive mass spectrometer. Theoretical masses of b/y ions are tabulated and MS-observed masses are shown in the spectra.
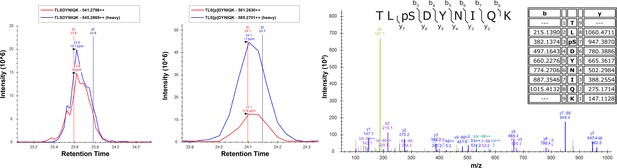
Overexpression of Vhs1 in yeast cells increases Ser57 phosphorylation of ubiquitin.
Cells were grown in SILAC media supplemented with light or heavy lysine and arginine. Chromosomally expressed 3xFLAG-tagged ubiquitin (from the RPS31 and RPL40B loci) was isolated by affinity purification and digested with trypsin. Phosphopeptides were enriched by immobilized metal affinity chromatography (IMAC), separated by a capillary reverse-phase analytical column, and analyzed on a Q Exactive mass spectrometer. Theoretical masses of b/y ions are tabulated and MS-observed masses are shown in the spectra.
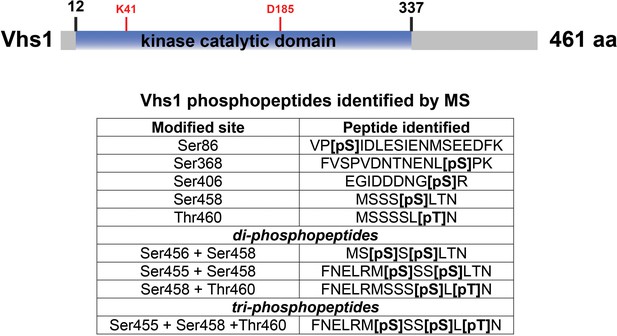

Phosphopeptides detected from Vhs1.
For the experiments shown in Figure 3F, the phosphorylation of Vhs1 was detected. A schematic of Vhs1 domain structure is shown [top] with the identified phosphopeptides listed [inset table, bottom]. Amino acids critical for kinase catalytic function – including the conserved lysine critical for ATP binding (K41) and the conserved aspartic acid (D185) critical for catalytic activity – are labeled in red.
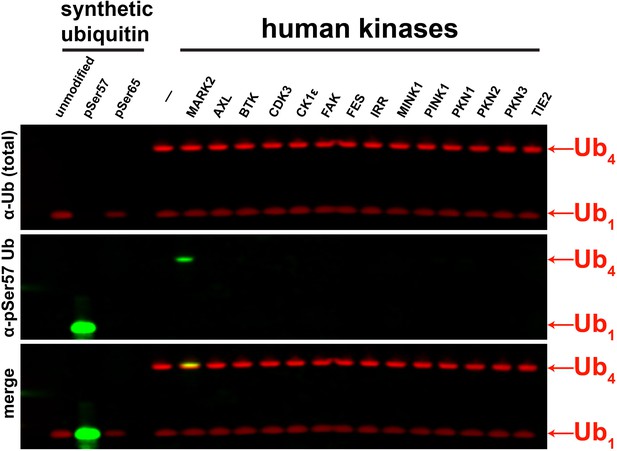
Screening of human ubiquitin Ser57 kinases by Western Blot.
Select kinases of different superfamilies were used for in vitro kinase activity assays using tetra-ubiquitin. Ubiquitin phosphorylation at the Ser57 residue was determined by Western blot using anti-phospho-Ser57 ubiquitin antibody.
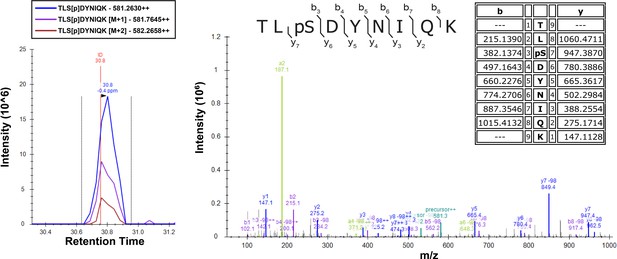
Identification and characterization of human MARK2 phosphorylation site on ubiquitin.
In vitro phosphorylation experiment was performed using recombinant GST-MARK2 and 6×His-tetra-Ub as described in the Materials and methods. Theoretical masses of b/y ions are tabulated and MS-observed masses are shown in the spectra.
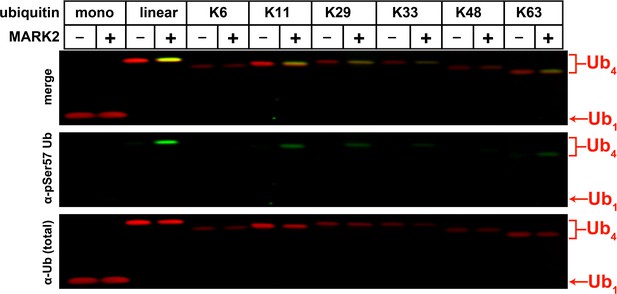
Determination of MARK2 substrate preference by analyzing activity toward tetra-ubiquitin chains with different linkages.
Ubiquitin phosphorylation at the Ser57 residue was determined by Western blot using an anti-phospho-Ser57 ubiquitin antibody.
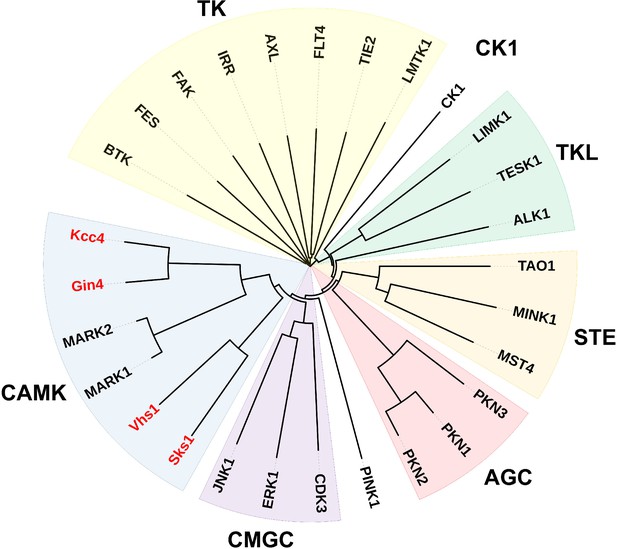
Yeast ubiquitin kinases Sks1, Vhs1, Gin4, and Kcc4 cluster with human MARKs of the CAMK superfamily of kinases.
Sequence alignment and phylogenetic neighboring of kinase domains were analyzed by Clustal Omega (EMBL-EBI) and the circular rooted phylogram was constructed using iTOL. Yeast Ub kinases (Sks1, Vhs1, Gin4, and Kcc4) are shown in red.
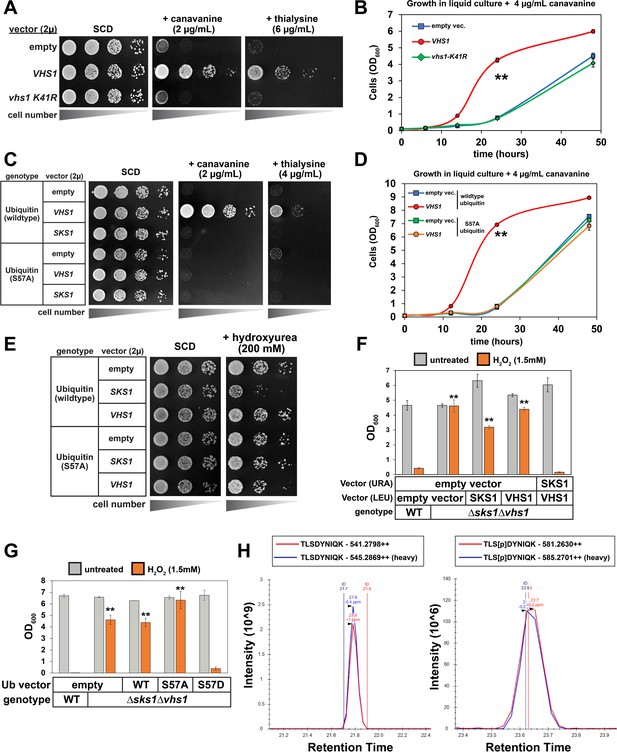
Regulation of proteotoxic stress responses by Ser57 ubiquitin kinases.
(A) Analysis of yeast strains containing high copy plasmids (empty vector, VHS1, or a kinase dead (vhs1 K41R) variant) grown on the indicated synthetic media plates. (B) Analysis of the same yeast strains in (A) grown in liquid culture containing canavanine (4 µg/mL). Values in each time point are means of three biological replicates (n = 3) and ± SD (error bars). Double asterisk (**) indicates p value < 0.005 at the 14 and 24 hr time points using a Student’s t-test, which was only observed for yeast overexpressing wildtype VHS1. (C) Analysis of yeast strains (SUB280 background, expressing either WT or S57A ubiquitin) containing the indicated high copy plasmids (empty vector or harboring VHS1 or SKS1) grown on the indicated synthetic media plates. (D) Analysis of the same yeast strains in (C) grown in liquid culture containing canavanine (4 µg/mL). Values in each time point are means of three biological replicates (n = 3) and ± SD (error bars). Double asterisk (**) indicates p value < 0.005 at the 14 and 24 hr time points using a Student’s t-test, which was only observed for yeast overexpressing wildtype VHS1 and expressing wildtype ubiquitin. (E) Analysis of yeast strains (SUB280 background, expressing either WT or S57A ubiquitin) containing the indicated high copy plasmids (empty vector or harboring VHS1 or SKS1) grown on the indicated synthetic media plates. (F) Analysis of the growth response to oxidative stress in WT and Δsks1Δvhs1 double mutant cells, with indicated complementation expression vectors. OD600 of cells (n = 4; ± SD error bars) in dropout SM media in the absence or presence of 1.5 mM H2O2 after incubation at 24 hr or 48 hr, respectively. Starting OD600 of 0.025 and data was collected for four biological replicate experiments (n=4). Double asterisk (**) indicates p value < 0.001 compared to wildtype yeast treated with H2O2 using a Student’s t-test, which was observed for yeast expressing empty vector, VHS1 individually, or SKS1 individually but not for yeast expressing both VHS1 and SKS1. (G) Analysis of the growth response to oxidative stress in WT and Δsks1Δvhs1 double mutant cells, with indicated ubiquitin expression vectors. OD600 of cells (n = 4; ± SD error bars) in dropout SM media in the absence or presence of 1.5 mM H2O2 after incubation at 24 hr or 48 hr, respectively. Starting OD600 of 0.025 and data was collected for four biological replicate experiments (n=4). Double asterisk (**) indicates p value < 0.001 compared to wildtype yeast treated with H2O2 using a Student’s t-test, which was observed for yeast expressing empty vector, wildtype ubiquitin, or S57A ubiquitin but not S57D ubiquitin. (H) Chromatography data serving as the basis for quantification of Ser57 phosphorylation corresponding to Table 1. Yeast cells expressing endogenous FLAG-ubiquitin (JMY1312 [SEY6210 background] parent cells) were cultured in heavy (H; wildtype cells [blue trace]) or light (L, isogenic Δvhs1Δsks1 deletion [red trace]) SILAC media to the mid-log phase and cells were treated with 1 mM H2O2 for 30 min before harvesting. Following cell lysis and overnight tryptic digestion of affinity-purified FLAG-ubiquitin, phospho-peptides were enriched using IMAC chromatography (see Materials and Methods) and analyzed by mass spectrometry. The Ser57 phosphopeptide is represented in the right panel, while the corresponding unmodified peptide is depicted in the left panel. All individual measurements and p values derived from statistical tests associated with the data in this figure are reported in Figure 4—source data 1.
-
Figure 4—source data 1
Quantification and statistical analysis for growth tolerance in canavanine (Figure 4B and D), thialysine (Figure 4—figure supplements 3 and 4), and hydrogen peroxide (Figure 4E and F).
- https://cdn.elifesciences.org/articles/58155/elife-58155-fig4-data1-v2.xlsx
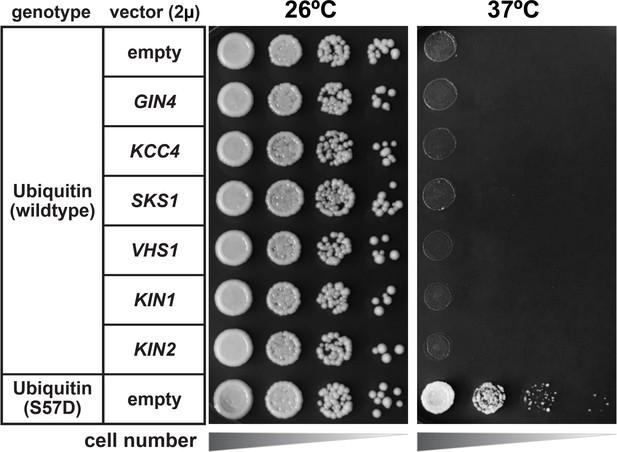
Heat stress growth phenotype of yeast strains overexpressing Ser57 ubiquitin kinases.
SUB280 cells harboring the indicated kinases cloned into a 2µ multicopy plasmid pRS327 were serially diluted onto lysine-dropout synthetic media and grown at 26°C or 37°C for 3 days. KIN1 and KIN2 were included as control kinases in the Snf1-related family that did not exhibit Ser57 ubiquitin kinase activity in our screening (Figure 3—figure supplement 2).
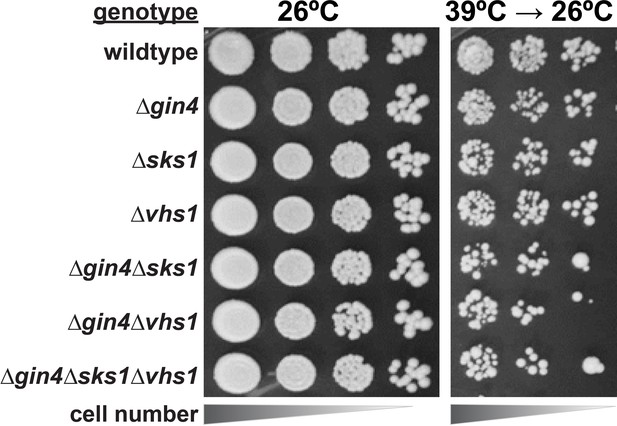
Analysis of cell tolerance to heat stress in yeast kinase deletion cells.
Null-mutants derived from SEY6210 yeast strain were serially diluted onto YPD plates and incubated at 26°C or 39°C for 18 hr and shifted to 26°C for growth recovery.
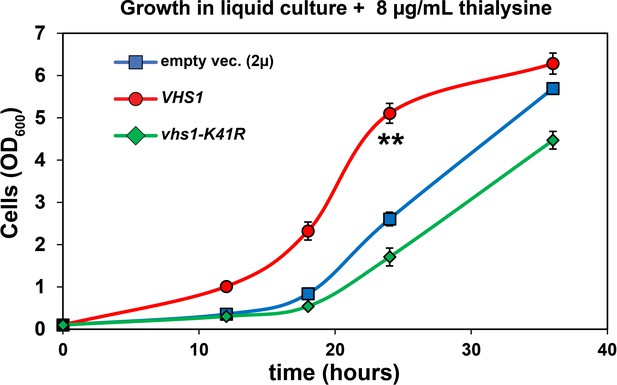
Analysis of cell tolerance to thialysine of yeast cells overexpressing VHS1 or a kinase dead (vhs-K41R) variant (kinase dead).
SUB280 cells expressing wildtype ubiquitin were transformed with VHS1 or vhs-K41R (in 2µ multicopy plasmid pRS327) and grown in lysine-dropout synthetic liquid media supplemented with 8 µg/mL thialysine. OD600 of cells were measured in different time points and the results were plotted as means of four biological replicates (n=4) with ± SD expressed as error bars. Double asterisk (**) indicates p value < 0.001 at the 12, 18, and 24 hr time points using a Student’s t-test, which was only observed for yeast overexpressing wildtype VHS1. For all statistical analysis associated with this figure supplement, p values are reported in the Figure 4—source data 1.
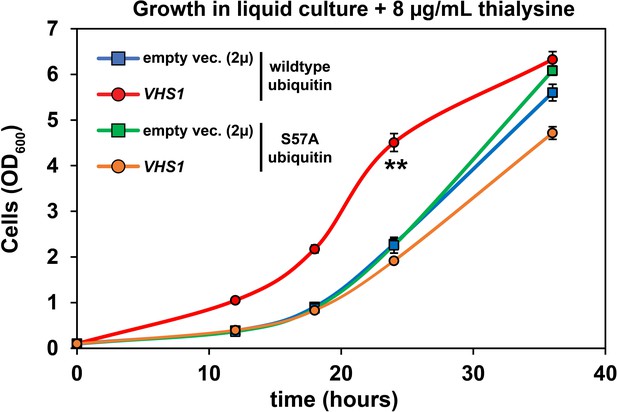
Analysis of cell tolerance to thialysine of yeast cells overexpressing VHS1.
Vhs1 (cloned into a 2µ multicopy plasmid pRS327) was overexpressed in SUB280 cells expressing ubiquitin (exclusively wildtype or S57A). Cells were grown in lysine dropout synthetic liquid media supplemented with 8 µg/mL thialysine. Values at each time point are means of four biological replicates with ± SD expressed as error bars. Double asterisk (**) indicates p value < 0.001 at the 12, 18, and 24 hr time points using a Student’s t-test, which was only observed for yeast overexpressing wildtype VHS1 in the context of wildtype ubiquitin expression. For all statistical analysis associated with this figure supplement, p values are reported in the Figure 4—source data 1.
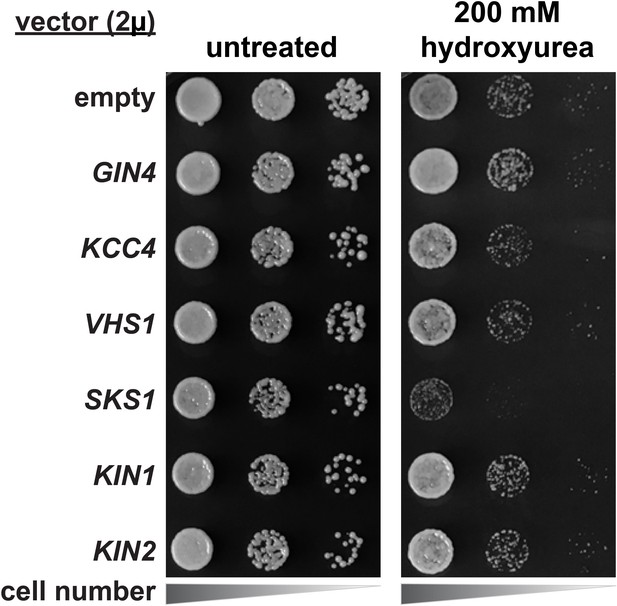
Hydroxyurea growth phenotype of yeast strains overexpressing Ser57 ubiquitin kinases.
SUB280 cells harboring the indicated kinases (expressed from the 2µ multicopy plasmid pRS327) were serially diluted onto lysine-dropout synthetic media or equivalent media supplemented with 200 mM hydroxyurea and grown at 26°C. KIN1 and KIN2 were included as control kinases in the Snf1-related family that did not exhibit Ser57 ubiquitin kinase activity in our screening (Figure 3—figure supplement 2).
Tables
Corresponds to Figure 4H.
Analysis of ubiquitin phosphorylation in Δvhs1Δsks1 mutants. Yeast cells expressing endogenous FLAG-ubiquitin (JMY1312 [SEY6210 background] parent cells) were cultured in heavy (H; wildtype cells) or light (L, isogenic Δvhs1Δsks1 deletion) SILAC media to the mid-log phase. In Experiment #1, cells were untreated, and in Experiment #2 cells were treated with 1 mM H2O2 for 30 min before harvesting cells. Following cell lysis and overnight digestion of lysates with trypsin, phospho-peptides were enriched using IMAC chromatography (see Materials and methods) and analyzed by mass spectrometry. Phosphorylated ubiquitin peptides resolved in these experiments are indicated and the values presented in the table represent the H:L SILAC ratio, which has been normalized to the total ubiquitin H:L ratio for each experiment and log-transformed (log2). ‘n.d.’ for Ser19 in Experiment #1 indicates that this peptide was identified but could not be quantified in this experiment due to poor quality of chromatographic data.
| Peptide | Phospho- site | log2 (H:L Ratio) | |
|---|---|---|---|
| Expt #1 | Expt #2 | ||
| TITLEVE[pS]SDTIDNVK | Ser19 | n.d. | 0.18 |
| TL[pS]DYNIQK | Ser57 | 0.06 | −0.06 |
| ES[pT]LHLVLR | Thr66 | −0.22 | 0.11 |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (S. cerevisiae) | SUB280 | D. Finley Lab; Finley et al., 1994 PMID:8035826 PMCID:PMC359070 | MATa, lys2-801, leu2-3, 112, ura3-52, his3-D200, trp 1–1, ubi1-D1::TRP1, ubi2-D2::ura3, ubi3-Dub-2, ubi4-D2::LEU2 [pUB39 Ub, LYS2] [pUB100, HIS3] | |
| Strain, strain background (S. cerevisiae) | SUB280 Ub (WT, S57A, S57D) | Lee et al., 2017 PMID:29130884 PMCID:PMC5706963 | MATa, SUB280 [RPS31 (WT, S57A, S57D; URA3)] | |
| Strain, strain background (S. cerevisiae) | NHY318 | This paper | MATa, SUB280 arg4::KANMX | |
| Strain, strain background (S. cerevisiae) | SEY6210 | S. Emr Lab; Robinson et al., 1988 PMID:3062374 PMCID:PMC365587 | RRID:ATCC: 96099 | WT: MATα leu2-3,112 ura4-52 his3-D200 trp1-D901 lys2-801 suc2-D9 |
| Strain, strain background (S. cerevisiae) | SEY6210.1 | S. Emr Lab; Robinson et al., 1988 PMID:3062374 PMCID:PMC365587 | WT: MATa leu2-3,112 ura4-52 his3-D200 trp1-D901 lys2-801 suc2-D9 | |
| Strain, strain background (S. cerevisiae) | NHY175 | This paper | MATα, SEY6210 gin4::NATMX | |
| Strain, strain background (S. cerevisiae) | NHY117 | This paper | MATα, SEY6210 sks1::TRP1 | |
| Strain, strain background (S. cerevisiae) | NHY118 | This paper | MATα, SEY6210 vhs1::TRP1 | |
| Strain, strain background (S. cerevisiae) | NHY196 | This paper | MATa, SEY6210.1 sks1::TRP1 vhs1::TRP1 | |
| Strain, strain background (S. cerevisiae) | NHY177 | This paper | MATα, SEY6210 gin4::NATMX sks1::TRP1 | |
| Strain, strain background (S. cerevisiae) | NHY179 | This paper | MATα, SEY6210 gin4::NATMX vhs1::TRP1 | |
| Strain, strain background (S. cerevisiae) | NHY181 | This paper | MATα, SEY6210 gin4::NATMX sks1::TRP1 vhs1::TRP1 | |
| Strain, strain background (S. cerevisiae) | JMY1312 | This paper | MATα, SEY6210 arg4::KANMX FLAG-RPS3::TRP1, FLAG-RPL40B::TRP1 | |
| Strain, strain background (S. cerevisiae) | NHY326 | This paper | MATa, JMY1312 sks1::TRP1 vhs1::TRP1 | |
| Strain, strain background (E. coli) | Rosetta 2 (DE3) | Millipore (Burlington, MA) | Cat# 71400 | F- ompT hsdSB(rB- mB-) gal dcm (DE3) pRARE2 (CamR) Competent Cells - Novagen |
| Strain, strain background (E. coli) | C41 (DE3) | Lucigen Corporation (Middleton, WI) | F– ompT hsdSB (rB - mB -) gal dcm (DE3) Competent Cells | |
| Recombinant DNA | pRS415 | Stratagene | (Empty Backbone) Yeast Centromere Plasmid (LEU2) GenBank: U03449 | |
| Recombinant DNA | pRS416 | Sikorski and Hieter, 1989 | (Empty Backbone) Yeast Centromere Plasmid (URA3) GenBank: U03450 | |
| Recombinant DNA | pJAM1303 | This paper | pNative-Sks1 in pRS416 (URA3); yeast expression | |
| Recombinant DNA | pJAM1304 | This paper | pNative-Sks1 in pRS415 (LEU2); yeast expression | |
| Recombinant DNA | pJAM1280 | This paper | pTDH3-Sks1 in pRS415 (LEU2); yeast expression | |
| Recombinant DNA | pJAM1253 | This paper | pNative_Vhs1 in pRS415 (LEU2); yeast expression | |
| Recombinant DNA | pRS327 | PMID:14989082 | RRID:Addgene_51787 | (Empty Backbone) multicopy (YEp) vector with a 2 μm origin of replication (LYS2); yeast expression |
| Recombinant DNA | pJAM1746 | This paper | pTDH3-Gin4 in pRS327; yeast expression | |
| Recombinant DNA | pJAM1749 | This paper | pTDH3-Kcc4 in pRS327; yeast expression | |
| Recombinant DNA | pJAM1743 | This paper | pTDH3-Sks1 in pRS327; yeast expression | |
| Recombinant DNA | pJAM1740 | This paper | pTDH3-Vhs1 in pRS327; yeast expression | |
| Recombinant DNA | pJAM1771 | This paper | pTDH3-Vhs1 K41R (kinase dead) in pRS327; yeast expression | |
| Recombinant DNA | pJAM1240 | This paper | 6XHis-Sks1 in pET15b, AmpR; E. coli expression | |
| Recombinant DNA | pJAM1169 | This paper | Vhs1-6XHis in pET23d, AmpR; E. coli expression | |
| Recombinant DNA | pJAM983 | This paper | Gin4-6XHis in pET23d, AmpR; E. coli expression | |
| Recombinant DNA | pJAM1116 | This paper | Hal4-6XHis in pET23d, AmpR; E. coli expression | |
| Recombinant DNA | pJAM1118 | This paper | Hal5-6XHis in pET23d, AmpR; E. coli expression | |
| Recombinant DNA | pJAM1120 | This paper | Kkq8-6XHis in pET23d, AmpR; E. coli expression | |
| Recombinant DNA | pJAM1336 | This paper | Snf1-6XHis in pET23d, AmpR; E. coli expression | |
| Recombinant DNA | pJAM585 | This paper | 6XHis-Npr1 in pCOLADuet, KmR; E. coli expression | |
| Recombinant DNA | pJAM1270 | This paper | Ksp1-6XHis in pET23d, AmpR; E. coli expression | |
| Recombinant DNA | pJAM1186 | This paper | GST-Slt2 in pGEX6p1; E. coli expression | |
| Recombinant DNA | pJAM1335 | This paper | Frk1-6XHis in pET23d, AmpR; E. coli expression | |
| Recombinant DNA | pJAM987 | This paper | Kcc4-6XHis in pET23d, AmpR; E. coli expression | |
| Recombinant DNA | pJAM1306 | This paper | Kin1-6XHis in pET23d, AmpR; E. coli expression | |
| Recombinant DNA | pJAM1307 | This paper | Kin2-6XHis in pET23d, AmpR; E. coli expression | |
| Recombinant DNA | pJAM1272 | This paper | Ypl150w-6XHis in pET23d, AmpR; E. coli expression | |
| Recombinant DNA | pJAM1167 | This paper | 6XHis-GST-UBD, AmpR;E. coli expression | |
| Recombinant DNA | pJAM1235 | This paper | Ubiquitin in pBG100; E. coli expression | |
| Recombinant DNA | pJAM1236 | This paper | Ubiquitin S57A in pBG100; E. coli expression | |
| Recombinant DNA | pJAM1237 | This paper | Ubiquitin S65A in pBG100; E. coli expression | |
| Recombinant DNA | pJAM995 | This paper | di-Ub in pET23d, AmpR; E. coli expression | |
| Recombinant DNA | pJAM996 | This paper | tri-Ub in pET23d, AmpR; E. coli expression | |
| Antibody | α-Flag (FLAG(R) M2) mouse monoclonal | Sigma (St. Louis, MO) | RRID:AB_262044 | WB (1:1000) |
| Antibody | α-Ubiquitin (VU101) mouse monoclonal; clone VU-1 | LifeSensors (Malvern, PA) | Cat# VU101 RRID:AB_2716558 | WB (1:1000) |
| Antibody | α-Glucose-6-Phosphate Dehydrogenase (G6PDH), yeast rabbit polyclonal | Sigma (St. Louis, MO) | Cat# A9521 RRID:AB_258454 | WB (1:10000) |
| Antibody | α-pSer57 Ubiquitin rabbit polyclonal | This paper | WB (1:1000) | |
| Antibody | K48-linkage Specific Polyubiquitin rabbit monoclonal; clone D9D5 | Cell Signaling Technology (Danvers, MA) | Cat# 8081 RRID:AB_10859893 | WB (1:1000) |
| Antibody | K63-linkage Specific Polyubiquitin rabbit monoclonal; clone Apu3 | Millipore (Burlington, MA) | Cat# 05–1308 RRID:AB_1587580 | WB (1:1000) |
| Antibody | IRDye 680RD-Goat anti-mouse polyclonal | LI-COR Biosciences (Lincoln, NE) | Cat# 926–68070 RRID:AB_10956588 | WB (1:10000) |
| Antibody | IRDye 800CW-Goat anti-rabbit polyclonal | LI-COR Biosciences (Lincoln, NE) | Cat# 926–32211 RRID:AB_621843 | WB (1:10000) |
| Commercial assay, kit | PTMScan Ubiquitin Remnant Motif (α-K-ε-GG) | Cell Signaling Technology (Danvers, MA) | RRID:Cat# 5562 | bead-conjugated for immunoprecipitation of –GG-εK-conjugated peptides |
| Chemical compound, drug | EZ view Red Anti-FLAG M2 Affinity Gel | Sigma (St. Louis, MO) | Cat# F2426-1ML RRID:AB_2616449 | mouse-monoclonal; clone M2 bead-conjugated for immunoprecipitation of FLAG-conjugated proteins |
| Chemical compound, drug | L-Lysine-13C6,13N2hydrochloride | Sigma (St. Louis, MO) | Cat# 608041–1G | |
| Chemical compound, drug | L-Arginine-13C6,13N4hydrochloride | Sigma (St. Louis, MO) | Cat# 608033–1G | |
| Chemical compound, drug | Sodium arsenate dibasic heptahydrate | Sigma (St. Louis, MO) | Cat# A6756 | Synonym: arsenate |
| Chemical compound, drug | Hydrogen peroxide | Fisher (Hampton, NH) | Cat# H325-500 | Synonym: H2O2 |
| Chemical compound, drug | Hydroxyurea | INDOFINE Chemical Company, Inc (Hillsborough, NJ) | Cat# BIO-216 | |
| Chemical compound, drug | S-(2-Aminoethyl)-L-cysteine hydrochloride | Sigma (St. Louis, MO) | Cat# A2636-1G | Synonym: L-4-Thialysine hydrochloride, thialysine |
| Chemical compound, drug | L-Canavanine | Sigma (St. Louis, MO) | Cat. # C1625 | Synonym: canavanine |
| Chemical compound, drug | DL-2-Aminoadipic acid | Sigma (St. Louis, MO) | Cat. # A0637 | |
| Chemical compound, drug | MG132 | Apexbio (Houston, TX) | Cat. # A2585 | |
| Chemical compound, drug | Protease inhibitor cocktail (cOmplete Tablets, Mini) | Roche (Basel, Switzerland) | Cat# 04693159001 | |
| Chemical compound, drug | PhosSTOP | Roche (Basel, Switzerland) | Cat# 04906837001 | |
| Chemical compound, drug | Phos-tag Acrylamide | NARD | Cat# AAL-107 | |
| Software, algorithm | MaxQuant Version 1.6.5.0 | Max Planck Institute of Biochemistry | RRID:SCR_014485 | |
| Software, algorithm | Skyline Version 20.1.0.155 | MacCoss Lab Software | RRID:SCR_014080 | |
| Software, algorithm | Adobe Illustrator CS5.1 Version 15.1.0 | Adobe (San Jose, CA) | RRID:SCR_010279 | |
| Software, algorithm | Image Studio Lite Version 5.2 | LI-COR Biosciences (Lincoln, NE) | RRID:SCR_013715 |
Additional files
-
Supplementary file 1
Corresponds to Figure 2—figure supplement 1.
Yeast cells (NHY318 background) expressing either wildtype or S57D ubiquitin were cultured in heavy (H; expressing wildtype ubiquitin) or light (L, expressing S57D ubiquitin) SILAC media to the mid-log phase and treated with 1 mM H2O2 for 30 min before harvesting cells. Following cell lysis and digestion of lysates with trypsin for 24 hr, ubiquitin-remnant peptides were enriched (see Materials and methods) and analyzed by mass spectrometry. Three biological replicate experiments were analyzed. Since the peptide corresponding to K63-linked poly-ubiquitin also harbors the residue mutated in phosphomimetic (S57D) ubiquitin, K63 linkages are a blind spot for SILAC measurements in these experiments. ‘n.d.’ indicates not detected.
- https://cdn.elifesciences.org/articles/58155/elife-58155-supp1-v2.docx
-
Supplementary file 2
Corresponds to Figure 3A and Figure 3—figure supplement 2.
For yeast Ser57 ubiquitin kinases, we analyzed consensus phosphorylation motifs as determined from a previous study based on in vitro activity analysis on peptide libraries (Mok et al., 2010). Values in parentheses are the quantified selectivity values, based on site preference of in vitro activity. Only amino acids selected at a position with a value >2 are shown.
- https://cdn.elifesciences.org/articles/58155/elife-58155-supp2-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/58155/elife-58155-transrepform-v2.docx





