The role of extracellular matrix phosphorylation on energy dissipation in bone
Figures
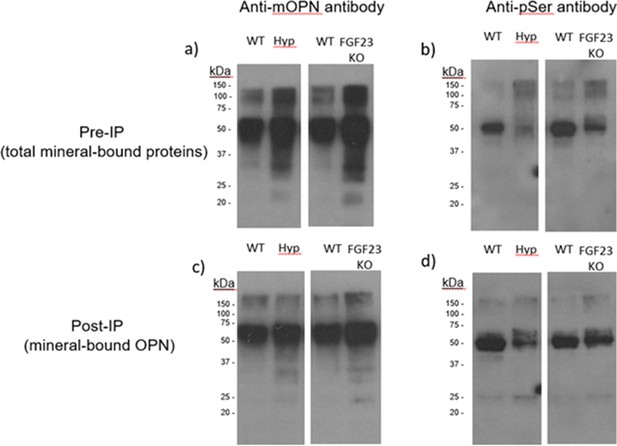
Pre-immunoprecipation (Pre-IP) of mineral-bound OPN.
(a) and global phosphorylation (b) in protein extracts of long bones from WT, Hyp and Fgf23-/- mice. Post-immunoprecipation (Post-IP) indicates that despite similar levels of OPN (c), phosphorylation of OPN is reduced in these disease models (d).
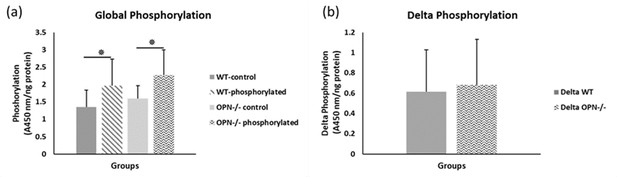
Mean global protein phosphorylation.
(a) and change in phosphorylation (b) for WT and Opn KO groups. * indicates significance at p<0.05 and error bars represent standard deviation.
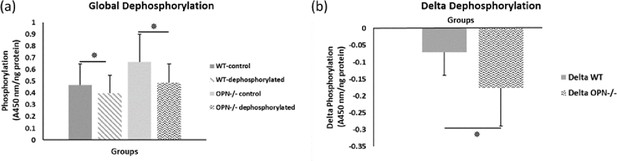
Mean global protein phosphorylation.
(a) and change in phosphorylation (b) after removal of phosphate groups (dephosphorylation) for WT and Opn KO groups. * indicates significance at p<0.05 and error bars represent standard deviation.
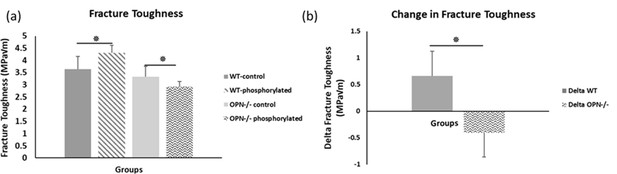
Mean fracture toughness (a) and change in fracture toughness (b) due to ex-vivo phosphorylation for WT and Opn KO groups.
* Indicates significance at p<0.05 and error bars represent standard deviation.
-
Figure 4—source data 1
Fracture toughness of phosphoryled WT and Opn KO mice.
- https://cdn.elifesciences.org/articles/58184/elife-58184-fig4-data1-v3.xlsx
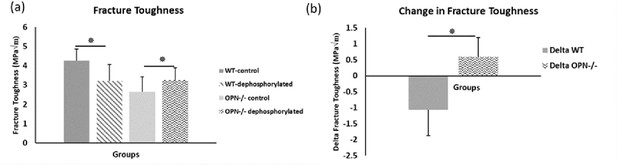
Mean fracture toughness (a) and change in fracture toughness (b) attributable to ex-vivo dephosphorylation for WT and Opn KO groups.
* Indicates significance at p<0.05 and error bars represent standard deviation.
-
Figure 5—source data 1
Fracture toughness of dephosphoryled WT and Opn KO mice.
- https://cdn.elifesciences.org/articles/58184/elife-58184-fig5-data1-v3.xlsx
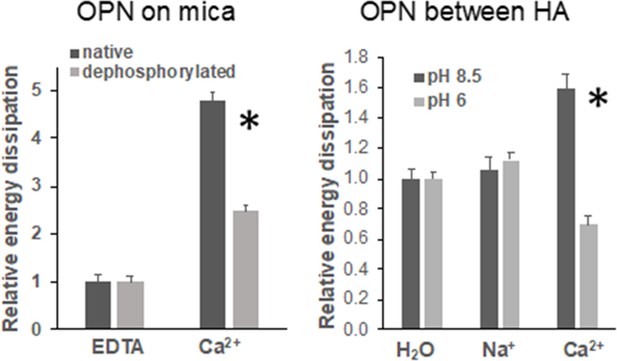
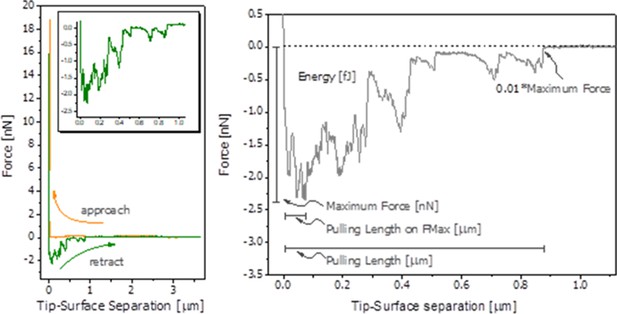
Energy dissipation of OPN networks during AFM-FS experiments.
Energies are normalized to dissipation levels in EDTA for OPN deposited on mica and pulled with a pristine AFM tip (pH 7.4) and to dissipation levels in H2O for OPN deposited on HA and pulled with a HA-functionalized tip. All values are significantly different except OPN between HA, pH 8.5 H2O vs. Na+. It should be noted that the relative differences are similar to what is seen for quantitative values, except for EDTA and H2O levels due to normalization. These values are provided in Supplementary files 1 and 2. * indicates significance at p<0.05 and error bars represent standard error (SE) of the mean.
-
Figure 6—source data 1
Energy dissipation of native (phosphorylated) and dephosphorylated OPN film on mica in EDTA and calcium solution.
- https://cdn.elifesciences.org/articles/58184/elife-58184-fig6-data1-v3.xlsx
-
Figure 6—source data 2
Energy dissipation of native (phosphorylated) OPN film on HA under various pH and ionic conditions.
- https://cdn.elifesciences.org/articles/58184/elife-58184-fig6-data2-v3.xlsx
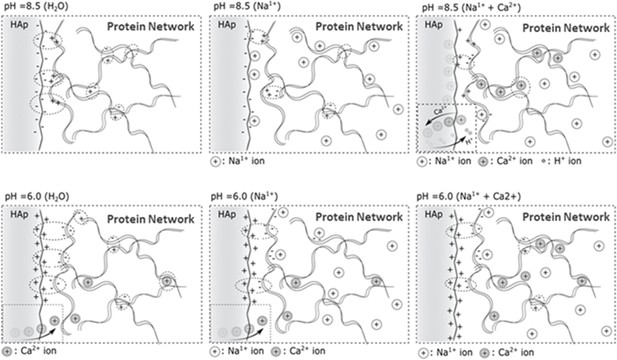
Proposed model for the OPN-HA interaction in different ionic- and pH environments.
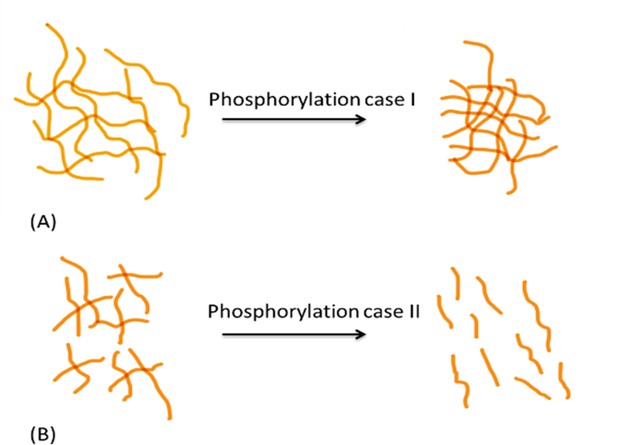
Schematic diagram showing differential effects of phosphorylation on conformation of protein systems.
In protein system (A), phosphorylation tends to increase inter- and intrafilament interactions, hence the interfilament distance is reduced. In protein system (B), phosphorylation tends to create interfilament repellant, hence increasing the protein system alignment and inter- filament distance.
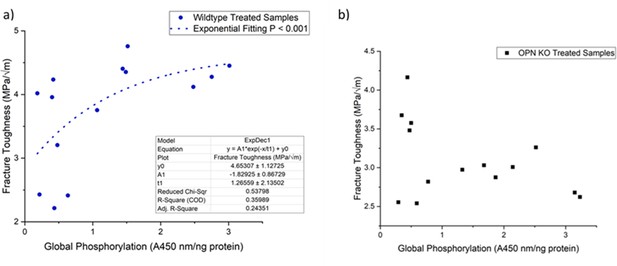
Schematic of the relationship between global protein phosphorylation and fracture toughness of wild-type (a) and Opn KO (b) mice.
By continuing the increase in phosphorylation of WT bone, fracture toughness improves exponentially. There is no significant relationship between global phosphorylation and fracture toughness in Opn KO mice following ex-vivo phosphorylation and dephosphorylation.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (M. musculus) | C57BL/6NCrl | Charles River | RRID:IMSR_CRL:27 | |
| Genetic reagent (M. musculus) | B6.Cg-PhexHyp/J | Jackson Laboratory | Cat#: 000528 RRID:IMSR_JAX:000528 | Animals maintained in Dr M Mckee lab. |
| Genetic reagent (M. musculus) | Fgf23-/- | PMID:15579309 | Animals were a gift from Dr. B. Lanske | |
| Genetic reagent (M. musculus) | B6.129S6(Cg)-Spp1tm1Blh/J | PMID:9661074 | Animals were a gift from Dr S. Rittling. | |
| Genetic Reagent (B. taurus) | Milk protein (Mammary gland) | PMID:8320368 | Provided by Dr ES Sorensen | |
| Chemical compound, drug | Synthetic hydroxyapatite | Andriotis et al., 2010. Crystal Research and Technology | Produced by Dr N. Bouropoulos | |
| Commercial assay or kit | pIMAGO-biotin HRP Detection | Tymora Analytical | Cat# 900–100 | |
| Antibody | anti-OPN (goat polyclonal) | R and D Systems | Cat# AF808, RRID:AB_2194992 | (1:100,000 µL) |
| Antibody | anti-phosphoserine (rabbit polyclonal) | Thermo Fisher Scientific | Cat# 61–8100, RRID:AB_2533940 | (1:2500 µL) |
Additional files
-
Supplementary file 1
Adhesive properties of native (phosphorylated) and dephosphorylated OPN film on mica.
- https://cdn.elifesciences.org/articles/58184/elife-58184-supp1-v3.docx
-
Supplementary file 2
Adhesive properties of native (phosphorylated) OPN film on HA.
- https://cdn.elifesciences.org/articles/58184/elife-58184-supp2-v3.docx
-
Supplementary file 3
Mean maximum force of native (phosphorylated) OPN film on HA.
- https://cdn.elifesciences.org/articles/58184/elife-58184-supp3-v3.docx
-
Supplementary file 4
Mean maximum force of native (phosphorylated) and dephosphorylated OPN film on mica.
- https://cdn.elifesciences.org/articles/58184/elife-58184-supp4-v3.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/58184/elife-58184-transrepform-v3.docx