Cannabidiol interactions with voltage-gated sodium channels
Figures
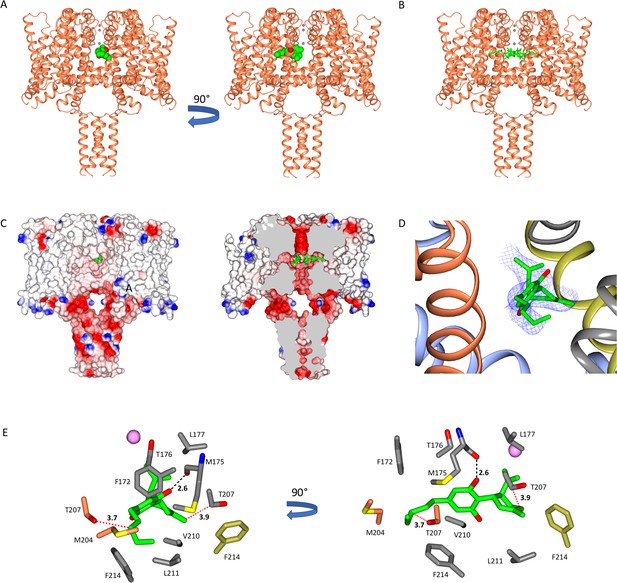
The NavMs sodium channel-cannabidiol (CBD) crystal structure.
(A) The crystal structure (2.3 Å resolution) of the NavMs sodium channel (in coral coloured ribbon depiction) with one CBD molecule (in green space-filing depiction), showing its location within the hydrophobic cavity of the channel, located adjacent to the fenestration. Three sodium ions are shown as grey spheres in the selectivity filter, for visual reference. The view on the right is rotated 90 degrees from the view on the left. (B) As in (A) but showing 4 CBD molecules in stick depiction. (C) (left) Surface view of space filling structure of NavMs coloured by electrostatics with CBD (in green) present. The orientation is the same as in the left panel of part A. The CBD (in green) is only just visible through the exterior end of one of the fenestration holes. (right) As in panel B, but with a slice through the space filling model (and through the middle of the fenestrations). For clarity only 2 CBD molecules are included, showing where the drug lies along the fenestration and blocks the ion pathway. (D) The CBD-binding site: the polypeptide backbone of the NavMs-CBD complex is depicted in ribbon motif. The ribbons are coloured by subunit (regions of the subunits that come in close contact with the one CBD molecule shown, are depicted in red, grey, blue, and yellow). The (2Fo-Fc) map (in blue mesh) was calculated at 0.7 sigma and the structure of the CBD molecule present is in stick depiction. (E) Detailed views of the residues that lie within 3.9 Å of the CBD molecule (which is depicted in green/red stick representation) are shown and coloured by domain (as in part D). The H-bond between CBD and NavMs involves the M175 backbone carbonyl group, which is shown as a dashed black line. The distance between the side chain of T207 and the CBD molecule is 3.7 Å, and is indicated by a dotted red line. This is the residue that was mutated in the electrophysiology studies. The pink sphere indicates the sodium ion site in the selectivity filter which is furthest from the extra-membranous surface (the one located farthest into the channel). On the right is the same view, rotated by 90 degrees.
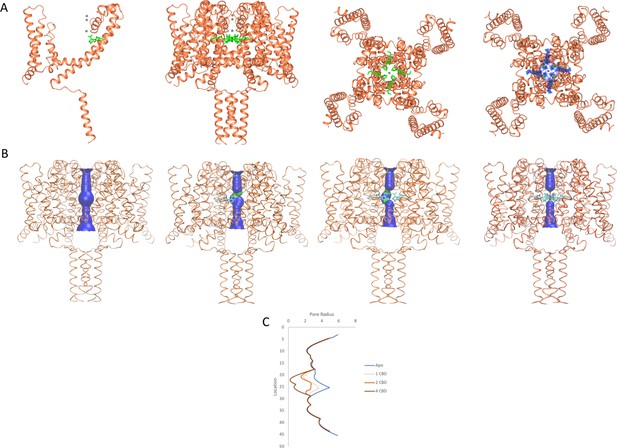
NavMs structures and pore diameters in the absence and presence of CBD.
The NavMs structure (coral) is depicted in ribbon motif. (A) (from the left): For clarity, only a single NavMs monomer with one CDB (green sticks) is shown; the NavMs tetrameric structure with all 4 CBDs bound; top view of NavMs tetramer with all 4 CDBs bound; top view of NavMs tetramer showing the electron density map (blue) demonstrating all 4 CBDs are bound in the tetrameric structure. (B) Pore interior dimensions calculated using the HOLE algorithm (Smart et al., 1993). Progressively from left to right: The apo structure and structures with 1, 2, and 4 CBD molecules present. In this figure the HOLE surface is depicted in blue for pore radii greater than 2.3 Å, and green for radii less than 2.3 Å. There is no occlusion in the absence of CBD, so hydrated sodium ions could freely pass through the pore. The NavMs structure with 4 CBD molecules present shows a full occlusion in the middle of the channel transmembrane pathway (near the centre of the hydrophobic cavity, so ion transport would be prevented [Naylor et al., 2016]). (C) Accessibility plots of pore radii versus position within the pore, in the absence and presence of different numbers of CBD molecules. The plot for the apo structure is in blue, and the plots for the CBD-containing structures with 1, 2, or 4 CBD molecules are in coral, red and black, respectively. The plots for 2 and 3 CBD molecules were the same, so the latter is not shown. In cases where at least some region of the radius is <2.0 Å, sodium ions will not be able to be translocated across the channel (Naylor et al., 2016). This plot thus shows that regardless of whether one or more CBD molecules are present, ion passage will not occur. These figures were produced using VMD software (Humphrey et al., 1996).
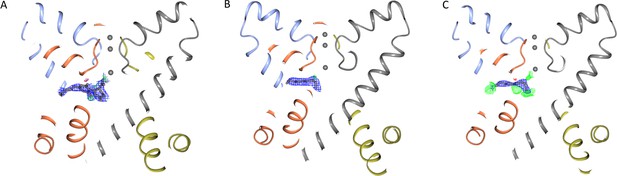
Comparisons of the electron density map of CBD in the NavMs-CBD complex with the electron density maps of Apo-NavMs crystals.
(A) CBD molecule fit into the NavMs-CBD crystal structure (overlaid by the 2Fo-Fc map contoured at 0.75 sigma), indicating the fit of the CBD to the density. (B) Aliphatic (C9H20) part of a lipid (or detergent) molecule (overlaid by the (2Fo-Fc) density (also contoured in blue at 0.75 sigma) in the crystal structure of Apo-NavMs. (C) The C9H20 structure refined into the NavMs-CBD (2Fo-Fc) (blue) map also contoured at 0.75 sigma [but without CBD structure present during refinement]. The green and red meshes (Fo-Fc maps contoured at sigma = 3) correspond to unaccounted for positive and negative densities, respectively). This clearly indicates that the density seen in the NavMs-CBD map is not due to bound lipid or detergent. The extra densities seen in the apo-NavMs maps may, however, arise from transient/partial occupancy of the cavity by a lipid, as had been indicated by molecular dynamics simulations (Ulmschneider et al., 2013), but occupancy by a lipid molecule for any significant time period is unlikely as it would inhibit the entry of hydrophobic drugs into the fenestration. In all panels the protein is shown in ribbon depiction (different colours for different polypeptide chains in the tetramer) and the three sodium ions present in the channel selectivity filter are shown as small grey balls, for visual reference.
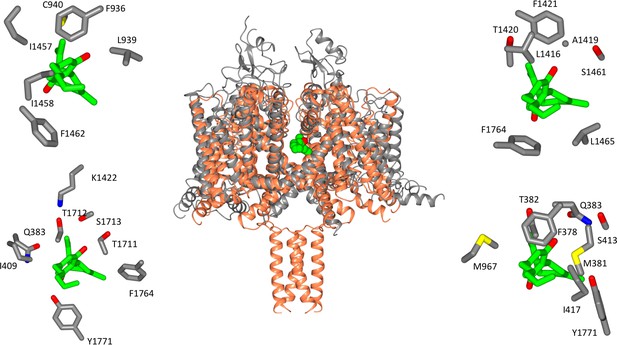
Location of CBD-binding sites in NavMs and the equivalent sites in hNav1.2.
(Centre) Structural alignment of the NavMs-CBD crystal structure (coral) and the hNav1.2 cryo-EM structure (grey). The RMSD of the aligned structures is 3.2 Å. (Top left): equivalent binding residues between domain I and domain II of hNav1.2 found within 4 Å of the CBD site. (Top right): equivalent binding residues in hNav1.2 between domains II and III located within 4 Å of the CBD-binding site. (Bottom left): Residues in hNav1.2 between domains III and IV within 4 Å of the CBD-binding site. (Bottom right): Residues in hNAv1.2 between domains IV and I within 4 Å of the CBD-binding site. In the surrounding panels the atoms in the protein are coloured by atom type, with carbons represented in grey, oxygen in red, nitrogen in blue, and sulphur in yellow, whilst the carbon atoms of the drug are depicted in green and the oxygens in red.
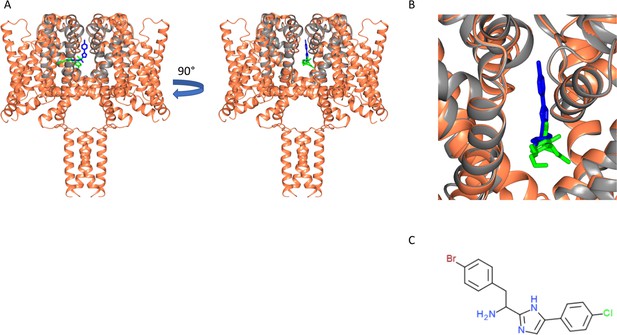
Similarity of CBD and analgesic compound binding sites.
(A) Structural alignment of the NavMs-CBD complex (protein in coral ribbon depiction, CBD in green stick depiction), with the structure of the NavMs pore-PI1 complex (Bagnéris et al., 2014) [the protein is in grey ribbon depiction and the PI1 molecule is in blue stick depiction]. PI1 is a highly potent designed analgesic compound, which binds to and inhibits flux through the NavMs channel (Bagnéris et al., 2014). Two views of the aligned structural complexes, rotated by 90 degrees, are shown. They show the similarity, but not identity of the binding sites of the two ligands. (B) Detailed view showing the locations of these molecules in the pore/fenestration area. (C) Chemical structure of the PI1 compound.
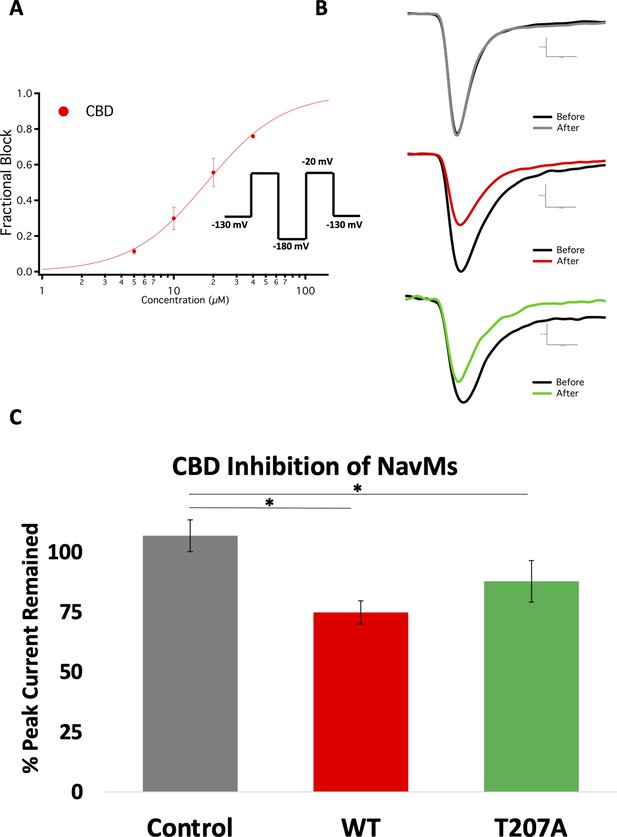
Electrophysiology studies of CBD inhibition of NavMs.
(A) Block was measured after ~6 min wash and incubation in CBD. The IC50 measurement was from CBD inhibition data obtained from whole-cell voltage-clamp recordings and fit with the Hill-Langmuir equation. The IC50 for CBD inhibition of wild-type NavMs is 17.8 ± 0.5 µM with a Hill slope of 1.5 ± 0.1 (the S.E. values quoted are errors of the fit, n = 8 panel-wide). (B) Sample traces of (top) wild-type NavMs without CBD added, (middle) wild-type NavMs before (black) and after (red)10 µM CBD perfusion (compound effect was measured after ~6 min of wash/incubation, 1 Hz), and (bottom), as in the middle panel, but using the T207A mutant (in green). (C) Bar graphs showing percentage of peak sodium current remaining over time after control (no CBD) (grey) and with CBD (red, green, as above) perfusions at 10 µM (n = 4–6). Statistical comparison against control: WT (p<0.0027) and T207A (p=0.0269).
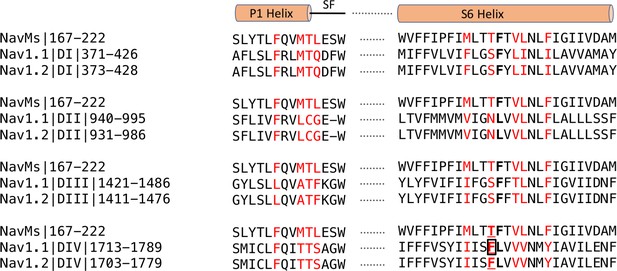

Sequence alignments of CBD-binding site regions of NavMs With corresponding regions in hNav1.1 and hNav1.2.
In red are the CBD-binding residues (within 3.9 Å of the compound) in NavMs and the equivalent residues in hNav1.1 and hNav1.2. The bold black F indicates the site of the NavMs F208L mutant used in these studies. It was changed from F to L in NavMsL because in half of the human Nav domains it is an F and in the other half it is an L (both indicated in bold black in the other sequences). However, as shown in Figure 6—figure supplement 2, the residue type present at this site makes essentially no difference in the structure. Residues located in the binding site are found within the P1 pore helix, the selectivity filter loop, and the S6 helix. The residue, which when mutated to alanine in hNav1.1 reduces the binding affinity of CBD (Ghovanloo et al., 2018), is boxed. This corresponds to T207 in NavMs, which is the residue that was mutated to alanine in the electrophysiology characterisations in the present study. The sequence alignment was carried out using Clustal Omega (Sievers et al., 2011) and annotated manually.
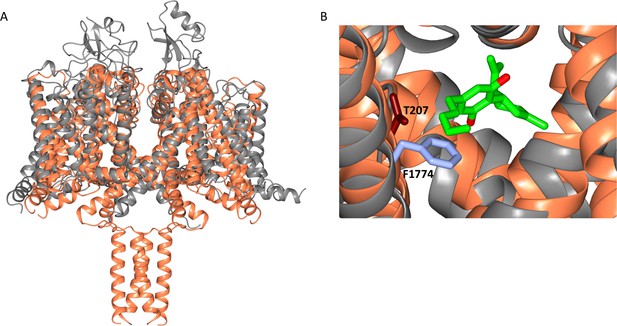
Alignment of the NavMs-CBD structure (coral) with a homology model of hNav1.1 (grey).
The homology model was created in SWISS-MODEL (Waterhouse et al., 2009) using hNav1.2 (PDB code 6J8E) as a template. (A) Overall structural alignment. (RMSD = 3.34 Å for the transmembrane regions). (B) Detail of this overlay showing the location of CBD (green and red stick depiction) in the region of NavMs residue T207 (light blue stick representation); this residue corresponds with residue F1774 in hNav1.1, which is a residue that has been identified as being in the local anaesthetic binding site, and was the residue that was mutated in electrophysiology experiments (Ghovanloo et al., 2018), which showed the effects of CBD inhibition on hNav1.2.
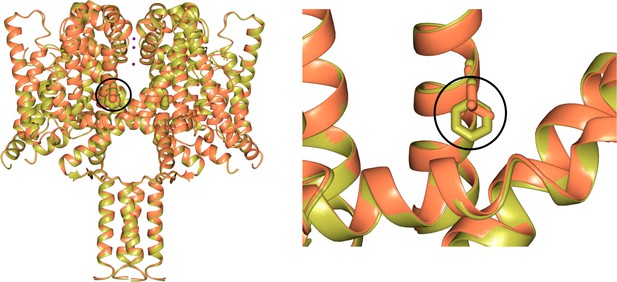
Comparisons of wild type NavMs (gold) [PDB ID 5HVX] and the NavMsL mutant (coral) [PDB ID 6YZ2] structures.
These figures show (left) the close overall similarities of the structures, and (right) the detailed region surrounding mutated residue F208L (which is circled in both panels, and is indicated in bold in the sequence alignment shown in Figure 6).
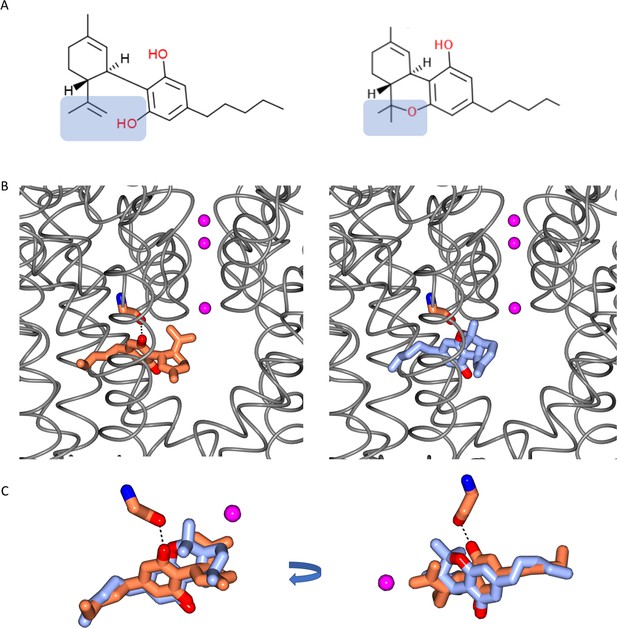
Comparisons of CBD and THC.
(A) Chemical structures of cannabidiol (CBD) (left) and Δ-9-tetrahydrocannabinol (THC) (right). The difference between the two structures is highlighted in blue background. The formation of a pyran ring in THC removes the hydrogen from the hydroxyl group which is present in CBD. (B) The locations of CBD in the CBD-NavMs crystal structure (left), and THC (right) modelled into this site. The NavMs structure is depicted in grey ribbons, and the three sodium ions sites in the selectively filter are indicated by the pink balls, as a reference point. (C) (left) A detailed overlay of CBD (coral) and THC (blue) shows the additional hydrogen bond between the protein and drug for CBD, by comparison to THC, which, without the corresponding hydroxyl group does not have the potential to form such a hydrogen bond. The view on the right is rotated from the view on the left to clearly visualise the alignment.
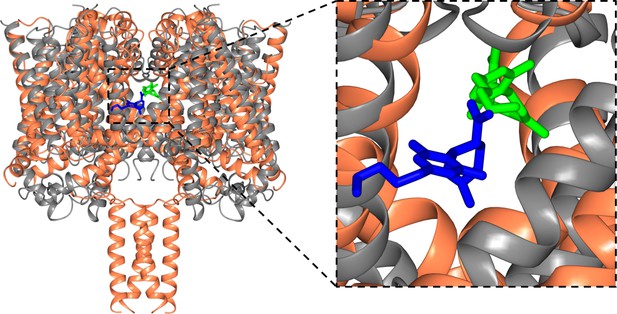
Structural alignments of the NavMs-CBD (coral ribbons) crystal structure and the TRPV2-CBD cryo-EM structure [PDB ID 6U88] (grey ribbons).
(Left) Overall alignment of the structures. The TRPV2 structure was trimmed to remove the extramembranous regions for clarity. The RMSD of the alignment is 4.2 Å. The CBD in NavMs is shown in green and that in TRPV2 is in blue. The CBD site in NavMs appears to be located further into the fenestration than it is in TRPV2. (Right) Detailed view of the CBD sites, highlighting the similarity and differences in orientation and location in the two channel types.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Magnetococcus marinus MC-1) | NavMs | Uniprot | Mmc1_0789 | A0L5S6_MAGMM |
| Strain, strain background (Escherichia coli) | Over Express C41 (DE3) | Sigma-Aldrich | CMC0017 | Chemically competent E. coli for expression of toxic proteins |
| Cell line (Cricetulus griseus) | CHO-K1 | CedarLine Laboratories | RRID:CVCL_0214 | |
| Recombinant DNA reagent | pET15b plasmid encoding NavMs | PMID:28205548 | Plasmid for NavMs and NavMsL expression for structural study | |
| Recombinant DNA reagent | His-NavMs | addgene.org | 100004 | pTracer-CMV2, IRES GFP plasmid encoding NavMs |
| Sequence-based reagent | F208L_F | This Paper | PCR Primer for NavMsL (forward) | 5'-CTCACCACCCTGA CCGTGCTCAACCTGT TTATTGG-3′ |
| Squence-based reagent | F208L_R | This Paper | PCR Primer for NavMsL (reverse) | 5′-GAGCACGGTCAGG GTGGTGAGCATGATG +AACGGGATG-3′ |
| Chemical compound, drug | Cannabidiol (CBD) | Sigma-Aldrich | C7515 | |
| Chemical compound, drug | Cannabidiol (CBD) | Toronto Research Chemicals | F175300 | |
| Software, algorithm | XDS | PMID:20124692 | RRID:SCR_015652 | Data Processing |
| Software, algorithm | Aimless | doi:10.1107/S0907444913000061 | RRID:SCR_015747 | Data Processing |
| Software, algorithm | CCP4 | PMID:15299374 | RRID:SCR_007255 | Structure Determination/Refinement |
| Software, algorithm | Phaser | PMID:19461840 | RRID:SCR_014219 | Structure Determination/Refinement |
| Software, algorithm | Coot | PMID:20383002 | RRID:SCR_014222 | Structure Determination/Refinement |
| Software, algorithm | REFMAC | PMID:21460454 | RRID:SCR_014225 | Structure Determination/Refinement |
| Software, algorithm | PROCHECK | doi:10.1107/s0021889892009944 | RRID:SCR_019043 | Structure Determination/Refinement |
| Software, algorithm | Molprobity | PMID:29067766 | RRID:SCR_014226 | Structure Determination/Refinement |
| Software, algorithm | BUSTER | PMID:22505257 | RRID:SCR_015653 | Structure Determination/Refinement |
| Software, algorithm | CCP4mg | PMID:21460457 | RRID:SCR_019041 | Structure Presentation |
| Software, algorithm | PatchMaster | HEKA Elektronik | RRID:SCR_000034 | Data Acquisition |
| Software, algorithm | FitMaster | HEKA Elektronik | RRID:SCR_016233 | Data Analysis |
| Software, algorithm | IGOR Pro | Wavemetrics, Lake Oswego, OR | RRID:SCR_000325 | Data Analysis |
| Software, algorithm | Clustal Omega | PMID:24170397 | RRID:SCR_001591 | Sequence Alignment |
| Software, algorithm | HOLE | PMID:9195488 | www.holeprogram.org | Channel Pore Dimension Analysis |
Additional files
-
Supplementary file 1
Crystal structure parameters.
- https://cdn.elifesciences.org/articles/58593/elife-58593-supp1-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/58593/elife-58593-transrepform-v2.docx





