Bardet–Biedl syndrome 3 protein promotes ciliary exit of the signaling protein phospholipase D via the BBSome
Figures
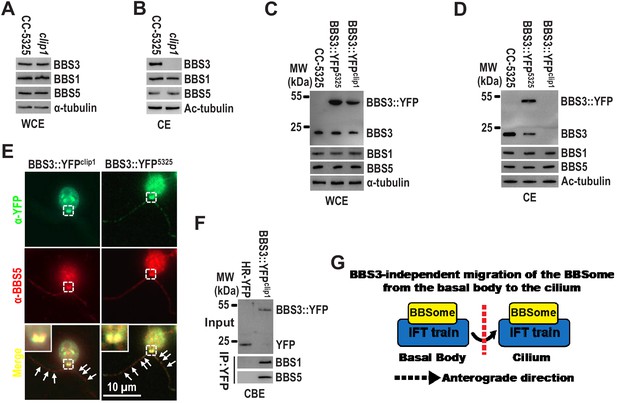
BBS3 is not required for the BBSome to enter cilia from the basal body.
(A and B) Immunoblots of whole-cell extracts (WCE) (A) and ciliary extracts (CE) (B) of CC-5325 and clip1 cells probed for BBS3, BBS1, and BBS5. (C and D) Immunoblots of WCE (C) and CE (D) of CC-5325, BBS3::YFP5325, and BBS3::YFPclip1 cells probed with α-BBS3, α-BBS1, and α-BBS5. MW: molecular weight. (E) BBS3::YFP5325 and BBS3::YFPclip1 cells stained with α-YFP (green) and α-BBS5 (red). Inset shows the basal bodies. White arrows show ciliary staining. Scale bar 10 µm. (F) Immunoblots of α-YFP captured proteins from cell body extracts (CBE) of HR-YFP (YFP-expressing CC-125 cells) and BBS3::YFPclip1 cells probed with α-BBS1 and α-BBS5. Input was quantified with α-YFP by immunoblotting. (G) Schematic presentation showing that the BBSome migrates from the basal body into cilia via IFT in a BBS3-independent manner. For immunoblotting, α-tubulin and acetylated-tubulin (Ac-tubulin) were used to adjust the loading of WCE and CE, respectively.
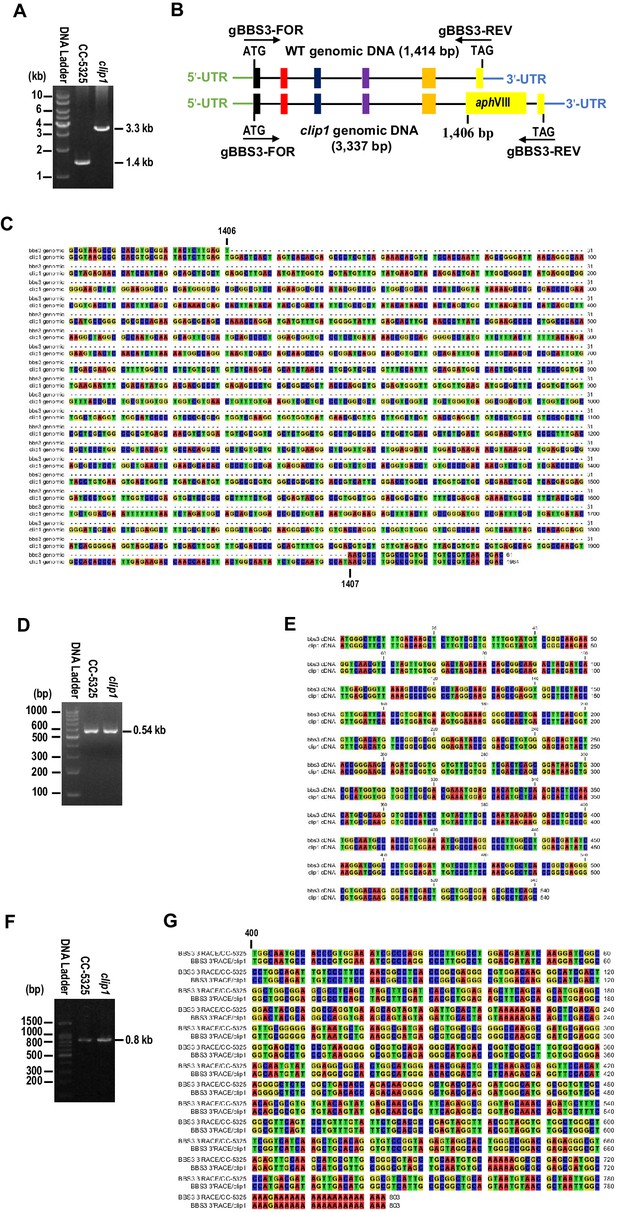
Characterization of the CLiP mutant clip1 (LMJ.RY0402.149010) at genomic and cDNA levels.
(A) Agarose gel electrophoresis showing the PCR products amplified from the BBS3 genomic DNA of CC-5325 and clip1 cells. The primer pair cBBS3-FOR and cBBS3-REV that targets the nucleotide sequence between the initial codon (ATG) and the stop codon (TAG) of BBS3 genomic DNA was used to amplify the BBS3 genomic DNA of 1414 bp from CC-5325 cells. A single DNA fragment of ~3.3 kb (3337 bp) was amplified from the clip1 cells using the same primer pair. (B) A diagram showing an insertion in the clip1 genomic DNA of clip1 cells. A paromomycin-resistant gene sequence of 1923 bp was inserted in the 3' distal region (insertion site is at 1406 bp) of the fifth exons of the clip1 genomic DNA in clip1 cells. Black lines and colorful boxes represent five introns and six exons, respectively, in the clip1 genomic DNA. The paromomycin-resistant gene (aphVIII) insertion was marked as a yellow box. The 5'- and 3'-untranslated regions (UTRs) are also shown. The primer pair used for PCR in A is shown. (C) Sequence alignment of the BBS3 genomic DNA between CC-5325 and clip1 cells. The clip1 cells contained an aphVIII insertion of 1923 bp between 1406 and 1407 bp of the BBS3 genomic DNA. (D) Agarose gel electrophoresis showing the BBS3 cDNA amplified from CC-5325 and clip1 cells using the primer pair cBBS3-FOR and cBBS3-REV. A single DNA band of 540 bp was amplified from both cells. (E) Sequence alignment of the BBS3 cDNA between CC-5325 and clip1 cells. Both cells contained intact wild-type BBS3 cDNA. (F) Agarose gel electrophoresis showing the BBS3 cDNA amplified from CC-5325 and clip1 cells by 3'-RACE assay. A single DNA band of 800 bp was amplified from both cells. (G) Sequence alignment of the BBS3 3'-partial cDNA between CC-5325 and clip1 cells. Both cells contained intact 3'-end of wild-type BBS3 cDNA. The number (400) stands for the nucleotides of the BBS3 cDNA starting from the initial codon ATG.
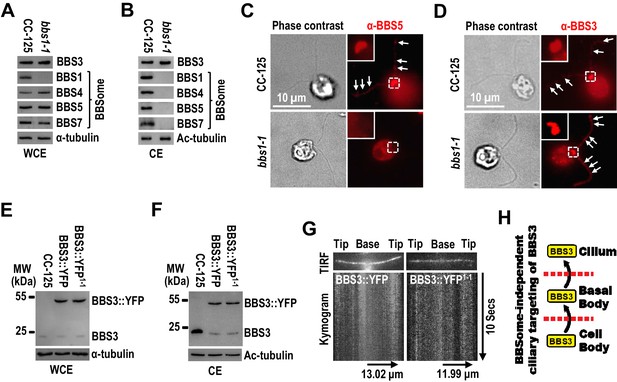
BBS3 enters cilia without relying on the BBSome.
(A and B) Immunoblots of whole-cell extracts (WCE) (A) and ciliary extracts (CE) (B) of CC-125 and bbs1-1 cells probed for BBS3 and the BBSome subunits BBS1, BBS4, BBS5, and BBS7. (C and D) CC-125 and bbs1-1 cells stained with α-BBS5 (red) (C) and α-BBS3 (red) (D). The corresponding phase contrast images are present. Inset shows the basal bodies. White arrows show ciliary staining. Scale bar 10 µm. (E and F) Immunoblots of WCE (E) and CE (F) of CC-125, BBS3::YFP (BBS3::YFP-expressing CC-125 cells), and BBS3::YFP1-1 (BBS3::YFP-expressing bbs1-1 cells) cells probed with α-BBS3. (G) Total internal reflection fluorescence images and corresponding kymograms of BBS3::YFP and BBS3::YFP1-1 cells (Figure 2—videos 1 and 2, 15 fps). The time and transport lengths are indicated on the right and on the bottom, respectively. (H) Schematic presentation showing that BBS3 can traffic from the cell body to the basal body for entering cilia in a BBSome-independent manner. For panels A, B, E, and F, α-tubulin and Ac-tubulin were used to adjust the loading of WCE and CE, respectively.
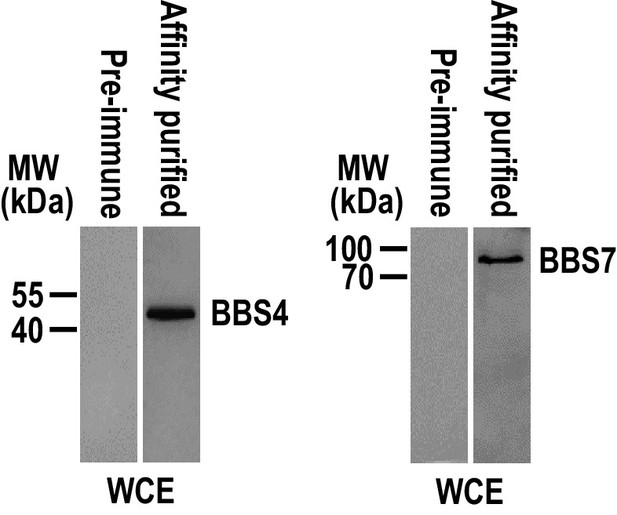
Affinity-purified polyclonal antisera against BBS4 and BBS7 can recognize their target proteins specifically.
Immunoblots of whole-cell extracts isolated from CC-125 cells with the affinity-purified BBS4 or BBS7 antisera identified one band with a size of approximately 46 or 80 kDa as expected, respectively. In contrast, the pre-immune serum did not recognize any bands. MW: molecular weight.
Total internal reflection fluorescence imaging of BBS3::YFP movement in cilia of CC-125 cells (BBS3::YFP cells).
A frame from this movie and kymogram are shown in Figure 2G. Play speed is real time (15 fps).
Total internal reflection fluorescence imaging of BBS3::YFP movement in cilia of bbs1-1 cells (BBS3::YFP1-1 cells).
A frame from this movie and kymograph are shown in Figure 2G. Play speed is real time (15 fps).
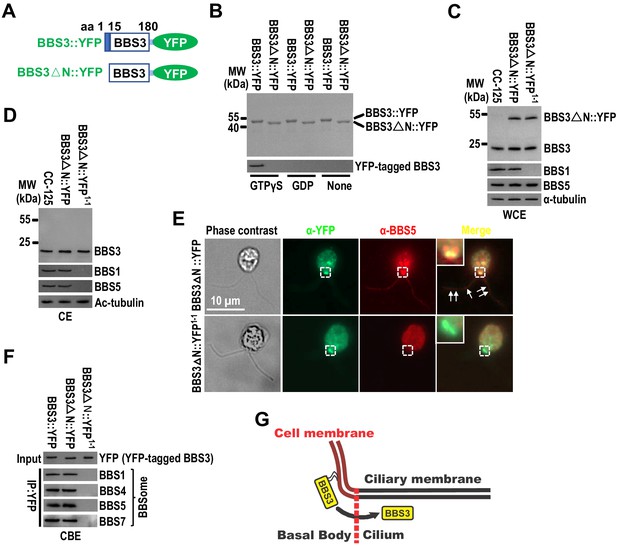
Membrane association is a prerequisite for BBS3 to enter cilia.
(A) Schematic presentation of BBS3::YFP and BBS3△N::YFP. △N: N-terminal 15 amino acids of BBS3 deleted. (B) SDS-PAGE visualization of bacterially expressed BBS3::YFP and BBS3△N::YFP and immunoblots of liposome-incubated BBS3::YFP and BBS3△N::YFP in the presence of GTPγS, GDP, or neither probed with α-BBS3. (C and D) Immunoblots of whole-cell extracts (WCE) (C) and ciliary extracts (CE) (D) of CC-125, BBS3△N::YFP (BBS3△N::YFP-expressing CC-125 cells), and BBS3△N::YFP1-1 (BBS3△N::YFP-expressing bbs1-1 cells) cells probed with α-BBS3, α-BBS1, and α-BBS5. α-tubulin and Ac-tubulin were used to adjust the loading of WCE and CE, respectively. (E) BBS3△N::YFP and BBS3△N::YFP1-1 cells stained with α-YFP (green) and α-BBS5 (red). The corresponding phase contrast images are present. Inset shows the basal bodies. White arrows show ciliary staining. Scale bar 10 µm. (F) Immunoblots of α-YFP-captured proteins from cell body extracts (CBE) of BBS3::YFP, BBS3△N::YFP, and BBS3△N::YFP1-1 cells probed with α-BBS1, α-BBS4, α-BBS5, and α-BBS7. Input was quantified with α-YFP by immunoblotting. (G) Schematic presentation of how BBS3 translocates from the basal bodies to cilia on the membrane.
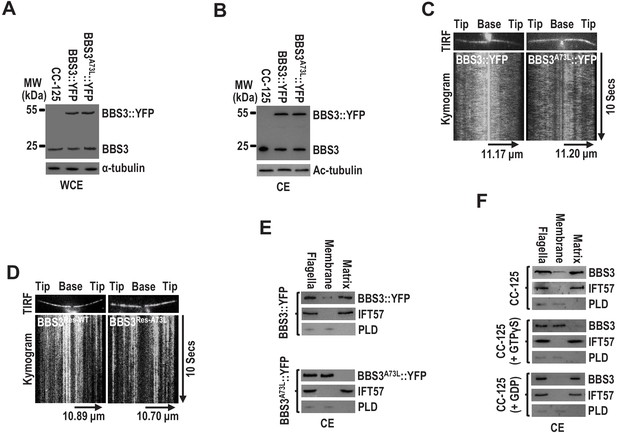
BBS3 associates with the ciliary membrane in a GTP-dependent manner.
(A and B) Immunoblots of whole-cell extracts (WCE) (A) and ciliary extracts (CE) (B) of CC-125, BBS3::YFP, and BBS3A73L::YFP cells probed with α-BBS3. α-tubulin and Ac-tubulin were used to adjust the loading of WCE and CE, respectively. (C and D) Total internal reflection fluorescence images and corresponding kymograms of BBS3::YFP and BBS3A73L::YFP cells (Figure 4—videos 1 and 2, 15 fps) (C), and BBS3Res-WT and BBS3Res-A73L cells (Figure 4—videos 3 and 4, 15 fps) (D). The time and transport lengths are indicated on the right and on the bottom, respectively. (E) Immunoblots of flagella and ciliary membrane and matrix fractions of BBS3::YFP and BBS3A73L::YFP cells probed with α-BBS3, α-IFT57, and α-PLD. (F) Immunoblots of flagella and ciliary membrane and matrix fractions of CC-125 cells, in the presence of GTPγS, GDP, or neither, probed for BBS3, IFT57, and PLD. For panels E and F, IFT57 is solely present in the ciliary matrix fraction, while PLD is only detected in the ciliary membrane fraction.
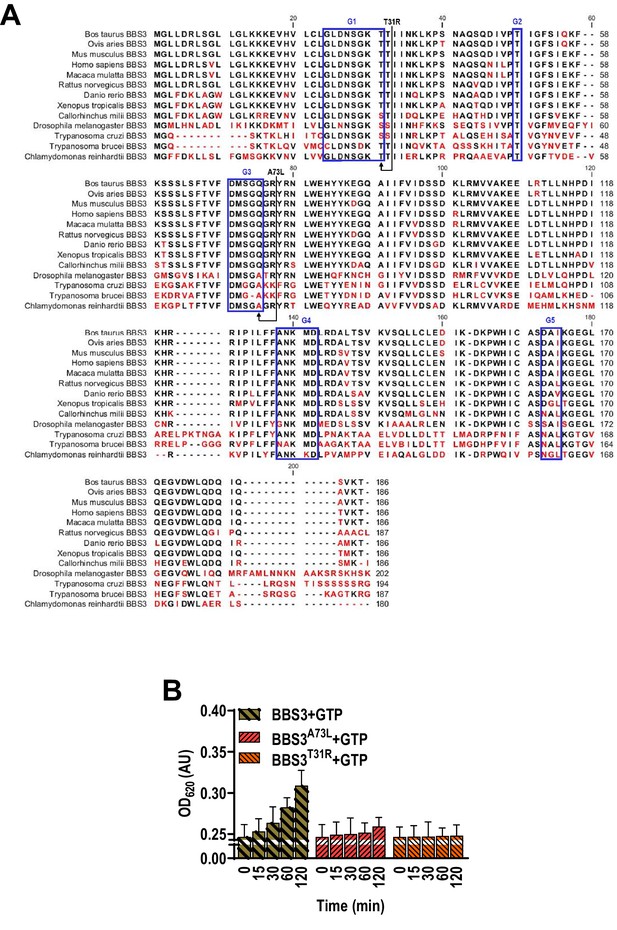
BBS3 is a highly conserved protein across ciliated species.
(A) Sequence alignment of deduced amino acid sequences from 13 invertebrate and vertebrate BBS3 orthologues and modified BBS3 sequences. Alignment was generated using CLC main workbench (version 6.8); the most conserved residues are shown in blue, the less conserved are in red. BBS3 contains five conserved domains including G1, G2, G3, G4, and G5, as labeled in the boxes. The P-loop threonine (T31) and alanine or glutamine (A73 or Q73) were marked. Dashes indicate gaps introduced to optimize the alignment. Arrowheads indicate missense mutations created in dominant-negative and constitutive-active BBS3 mutants. The BBS3 accession numbers are Bos Taurus, NP_001069250.1, Ovis aries, XP_004002902.1, Homo sapiens, AAH24239.1, Macaca mulatta, NP_001244689.1, Mus musculus, NP_062639.3, Rattus norvegicus, NP_001102312.1, Callorhinchus milii, XP_007899465.1, Xenopus tropicalis, XP_012813411.1, Danio rerio, NP_001008733.1, Drosophila melanogaster, NP_611421.1, Trypanosoma brucei, 228 .m00796, Trypanosoma cruzi, RNC37227.1, and Chlamydomonas reinhardtii, XP_001691012.1. (B) GTP hydrolysis of BBS3, BBS3A73L, and BBS3T31R. Introduction of a leucine at the position of A73 of BBS3 dramatically reduces the intrinsic GTPase activity, indicating that BBS3A73L is a GTP-locked, constitutive-active variant. No GTPase activity was observed for BBS3T31R, indicating that BBS3T31R appears to be an inactive GDP-locked or simply a dominant-negative variant.
-
Figure 4—figure supplement 1—source data 1
Source data for the GTP hydrolysis experiment shown in Figure 4—figure supplement 1B.
- https://cdn.elifesciences.org/articles/59119/elife-59119-fig4-figsupp1-data1-v2.xlsx
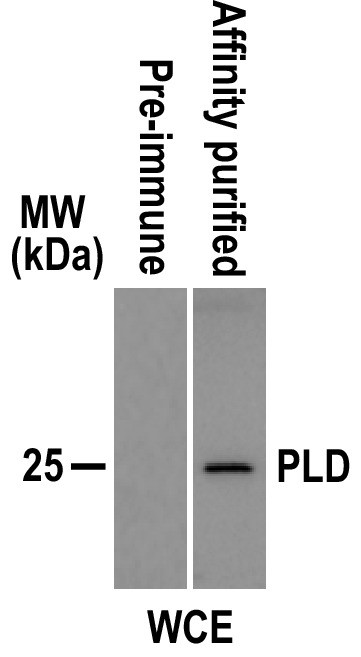
Affinity-purified polyclonal antisera against PLD can recognize its target protein specifically.
Immunoblots of whole-cell extracts (WCE) isolated from CC-125 cells with the affinity-purified PLD antisera identified one band with a size of approximately 25 kDa as expected. In contrast, the pre-immune serum did not recognize any bands. MW: molecular weight.
Total internal reflection fluorescence imaging of BBS3::YFP movement in cilia of CC-125 cells (BBS3::YFP cells).
A frame from this movie and kymograph are shown in Figure 4C. Play speed is real time (15 fps).
Total internal reflection fluorescence imaging of BBS3A73L::YFP movement in cilia of CC-125 cells (BBS3A73L::YFP cells).
A frame from this movie and kymograph are shown in Figure 4C. Play speed is real time (15 fps).
Total internal reflection fluorescence imaging of BBS3::GFP movement in cilia of BBS3miRNA cells (BBS3Res-WT cells).
A frame from this movie and kymograph are shown in Figure 4D. Play speed is real time (15 fps).
Total internal reflection fluorescence imaging of BBS3A73L::GFP movement in cilia of BBS3miRNA cells (BBS3Res-A73L cells).
A frame from this movie and kymograph are shown in Figure 4D. Play speed is real-time (15 fps).
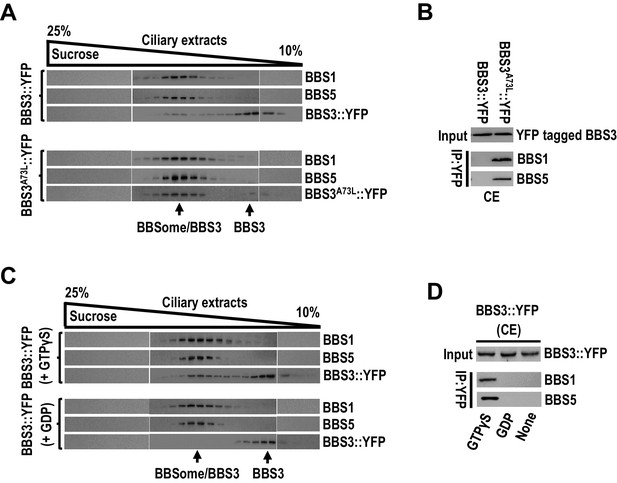
BBS3 interacts with the BBSome in cilia in a GTP-dependent manner.
(A) Immunoblots of sucrose density gradient of ciliary extracts (CE) of BBS3::YFP and BBS3A73L::YFP cells probed with α-BBS1, α-BBS5, and α-YFP. (B) Immunoblots of α-YFP-captured proteins from CE of BBS3A73L::YFP and BBS3::YFP cells probed with α-BBS1 and α-BBS5. (C) Immunoblots of sucrose density gradient of CE of BBS3::YFP cells, in the presence of GTPγS or GDP, probed with α-BBS1, α-BBS5, and α-GFP. (D) Immunoblots of α-YFP-captured proteins from CE of BBS3::YFP cells, in the presence of GTPγS, GDP, or neither, probed with α-BBS1 and α-BBS5. For panels B and D, input was quantified with α-YFP by immunoblotting.
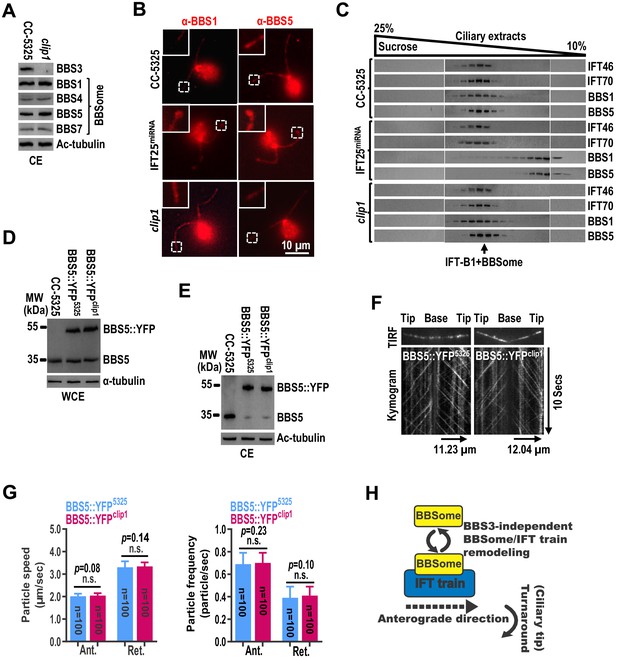
The BBSome cycles through cilia normally in the absence of BBS3 in cilia.
(A) Immunoblots of ciliary extracts (CE) of CC-5325 and clip1 cells probed for BBS3 and the BBSome subunits BBS1, BBS4, BBS5, and BBS7. Ac-tubulin was used to adjust the loading. (B) CC-5325, IFT25miRNA, and clip1 cells stained with α-BBS1 (red) and α-BBS5 (red). Inset shows ciliary tips. Scale bar 10 µm. (C) Immunoblots of sucrose density gradient of CE of CC-5325, IFT25miRNA, and clip1 probed with α-IFT46, α-IFT70, α-BBS1, and α-BBS5. (D and E) Immunoblots of whole-cell extracts (WCE) (D) and CE (E) of CC-5325, BBS5::YFP5325, and BBS5::YFPclip1 cells probed with α-BBS5. α-tubulin and Ac-tubulin were used to adjust the loading of WCE and CE, respectively. (F) Total internal reflection fluorescence images and corresponding kymograms of BBS5::YFP5325 and BBS5::YFPclip1 cells (Figure 6—videos 1 and 2, 15 fps). The time and transport lengths are indicated on the right and on the bottom, respectively. (G) Speeds and frequencies of BBS5::YFP inside cilia of BBS5::YFP5325 and BBS5::YFPclip1 cells are similar. The speeds of BBS5::YFP were 2.01 ± 0.12 (anterograde) and 3.31 ± 0.27 μm/sec (retrograde), and the frequencies were 0.69 ± 0.10 (anterograde) and 0.39 ± 0.10 particles/sec (retrograde) for BBS5::YFP5325 cells. In cilia of BBS5::YFPclip1 cells, the speeds of BBS5::YFP were 2.04 ± 0.11 (anterograde) and 3.34 ± 0.19 μm/sec (retrograde), and the frequencies were 0.70 ± 0.09 (anterograde) and 0.41 ± 0.08 particles/sec (retrograde). Error bar indicates SD. n: number of cilia analyzed; n.s.: non-significance. (H) Schematic presentation showing that the BBSome and IFT trains remodel at the ciliary tip in a BBS3-independent manner.
-
Figure 6—source data 1
Source data for the speed and frequency experiment shown in Figure 6G.
- https://cdn.elifesciences.org/articles/59119/elife-59119-fig6-data1-v2.xlsx
Total internal reflection fluorescence imaging of BBS5::YFP movement in cilia of CC-5325 cells (BBS5::YFP5325 cells).
A frame from this movie and kymograph are shown in Figure 6F. Play speed is real time (15 fps).
Total internal reflection fluorescence imaging of BBS5::YFP movement in cilia of clip1 cells (BBS5::YFPclip1 cells).
A frame from this movie and kymograph are shown in Figure 6F. Play speed is real time (15 fps).
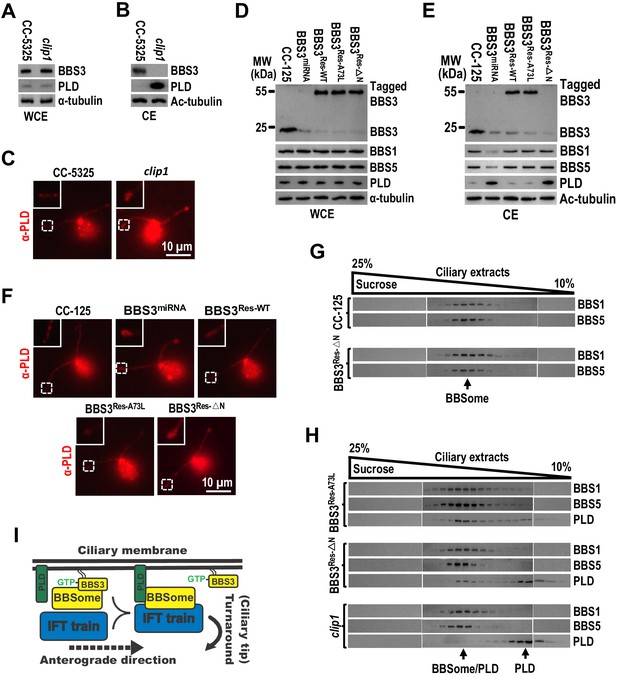
BBS3 is essential for PLD to associate with the BBSome for ciliary exit.
(A and B) Immunoblots of whole-cell extracts (WCE) (A) and ciliary extracts (CE) (B) of CC-5325 and clip1 cells probed with α-BBS3 and α-PLD. (C) CC-5325 and clip1 cells stained with α-PLD (red). (D and E) Immunoblots of WCE (D) and CE (E) of CC-125, BBS3miRNA, BBS3Res-WT, BBS3Res-A73L, and BBS3Res-△N cells probed with α-BBS3, α-BBS1, α-BBS5, and α-PLD. (F) CC-125, BBS3miRNA, BBS3Res-WT, BBS3Res-A73L, and BBS3Res-△N cells stained with α-PLD (red). (G) Immunoblots of sucrose density gradient of CE of CC-125 and BBS3Res-△N cells probed with α-BBS1 and α-BBS5. (H) Immunoblots of sucrose density gradient of CE of BBS3Res-A73L, BBS3Res-△N, and clip1 cells probed with α-BBS1, α-BBS5, and α-PLD. (I) Schematic presentation of how BBS3 binds and recruits the BBSome to the ciliary membrane for interacting with PLD and how PLD-associated BBSome loads onto IFT trains for ciliary exit. For panels A, B, D, and E, α-tubulin and Ac-tubulin were used to adjust the loading for WCE and CE, respectively. For panels C and F, the inset shows ciliary tips. Scale bar 10 µm.
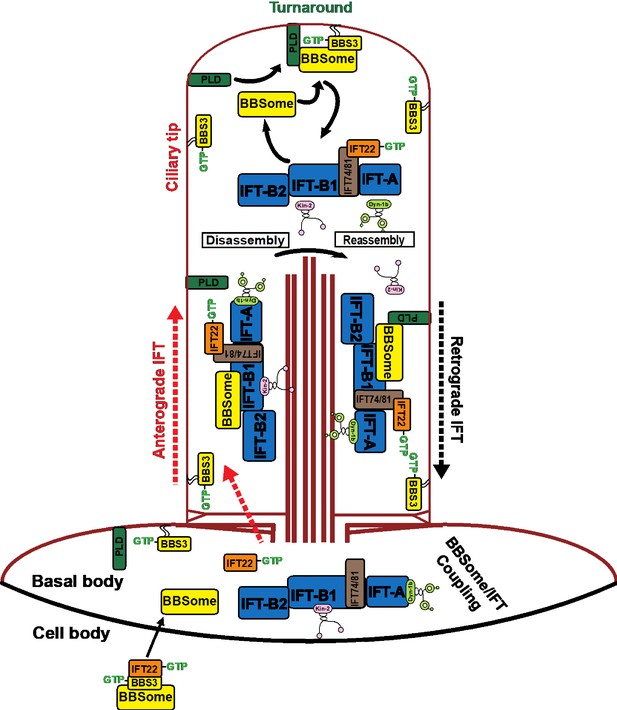
Hypothetical model of how BBS3 enters cilia to promote ciliary exit of PLD via the BBSome in C. reinhardtii.
IFT22 and BBS3 both in a GTP-bound configuration recruit the BBSome to the basal body, where the BBSome separates from BBS3 and loads onto the anterograde IFT trains by coupling with IFT-B1 and enters cilia via IFT (Xue et al., 2020). GTP-bound IFT22 is released from BBS3 and binds IFT-B1 through a direct interaction between IFT22 and the IFT74/81 sub-complex for ciliary entry (Bhogaraju et al., 2013; Lucker et al., 2010; Taschner et al., 2014). GTP-bound BBS3 then associates with the cell membrane to diffuse into cilia. PLD enters cilia by diffusion on membrane (Liu and Lechtreck, 2018). At the ciliary tip, the BBSome is released from anterograde IFT trains through BBSome/IFT train remodeling (Xue et al., 2020). GTP-bound BBS3 binds and recruits the BBSome to the ciliary membrane for interacting with PLD (Liu and Lechtreck, 2018). PLD-laden BBSomes then load onto retrograde IFT trains by coupling with IFT-B1 for ciliary exit (Xue et al., 2020). The kinesin-II anterograde motor and the cytoplasmic dynein-1b retrograde motor are also shown (Cole et al., 1998; Pazour et al., 1998). After turnaround at the ciliary tip, kinesin-II diffuses back to the ciliary base (Chien et al., 2017; Hendel et al., 2018).
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (C. reinhardtii) | CC-125 | Chlamydomonas Genetic Center (CGC) | CC-125 | Wild-type strain |
| Genetic reagent (C. reinhardtii) | CC-5325 | CGC | CC-5325 | Wild-type strain |
| Genetic reagent (C. reinhardtii) | clip1 | CGC | LMJ.RY0402.149010 | CLiP mutant/CC-5323 |
| Genetic reagent (C. reinhardtii) | bbs1-1 | Lechtreck et al., 2009 | BBS1-null mutant/CC-125 | |
| Genetic reagent (C. reinhardtii) | BBS3::YFP | This study | pBKS-gBBS3::YFP-Paro/CC-125 | |
| Genetic reagent (C. reinhardtii) | BBS3A73L::YFP | This study | pBKS-gBBS3A73L::YFP-Paro/CC-125 | |
| Genetic reagent (C. reinhardtii) | BBS5::YFP | Xue et al., 2020 | pBKS-gBBS5::YFP-Paro/CC-125 | |
| Genetic reagent (C. reinhardtii) | BBS3::YFP5325 | This study | pBKS-gBBS3::YFP-Ble/CC-5325 | |
| Genetic reagent (C. reinhardtii) | BBS3::YFPclip1 | This study | pBKS-gBBS3::YFP-Ble/clip1 | |
| Genetic reagent (C. reinhardtii) | BBS5::YFP5325 | Xue et al., 2020 | pBKS-gBBS5::YFP-Ble/CC-5325 | |
| Genetic reagent (C. reinhardtii) | BBS5::YFPclip1 | This study | pBKS-gBBS5::YFP-Ble/clip1 | |
| Genetic reagent (C. reinhardtii) | BBS3::YFP1-1 | This study | pBKS-gBBS3::YFP-Ble/bbs1-1 | |
| Genetic reagent (C. reinhardtii) | BBS3△N::YFP1-1 | This study | pBKS-gBBS3△N::YFP-Ble/bbs1-1 | |
| Genetic reagent (C. reinhardtii) | BBS3△N::YFP | This study | pBKS-gBBS3△N::YFP-Ble/CC-125 | |
| Genetic reagent (C. reinhardtii) | BBS3miRNA | Xue et al., 2020 | pMI-BBS3-Paro/CC-125 | |
| Genetic reagent (C. reinhardtii) | BBS3Res-WT | Xue et al., 2020 | pBKS-gBBS3::GFP-Ble/BBS3miRNA | |
| Genetic reagent (C. reinhardtii) | BBS3Res-A73L | Xue et al., 2020 | pBKS-gBBS3A73L::GFP-Ble/BBS3miRNA | |
| Genetic reagent (C. reinhardtii) | IFT25miRNA | Dong et al., 2017b | pMI-IFT25-Paro/CC-125 | |
| Genetic reagent (C. reinhardtii) | BBS3Res-△N | This study | pBKS-gBBS3△N::YFP-Ble/BBS3miNRA | |
| Genetic reagent (C. reinhardtii) | HR-YFP | This study | pBKS-HSP 70A-RBCS 2-YFP-Ble/CC-125 | |
| Antibody | Anti-IFT46 (rabbit polyclonal) | Dong et al., 2017b | immunoblotting (1:1,000) | |
| Antibody | Anti-IFT57 (rabbit polyclonal) | Dong et al., 2017b | immunoblotting (1:1,000) | |
| Antibody | Anti-IFT70 (rabbit polyclonal) | Dong et al., 2017b | immunoblotting (1:1,000) | |
| Antibody | Anti-BBS1 (rabbit polyclonal) | Xue et al., 2020 | immunoblotting (1:1,000) immunofluorescence (1:200) | |
| Antibody | Anti-BBS3 (rabbit polyclonal) | Xue et al., 2020 | immunoblotting (1:500) immunofluorescence (1:200) | |
| Antibody | Anti-BBS4 (rabbit polyclonal) | This study | immunoblotting (1:500) | |
| Antibody | Anti-BBS5 (rabbit polyclonal) | Xue et al., 2020 | immunoblotting (1:1000) immunofluorescence (1:200) | |
| Antibody | Anti-BBS7 (rabbit polyclonal) | This study | immunoblotting (1:1000) | |
| Antibody | Anti-PLD (rabbit polyclonal) | This study | immunoblotting (1:1000) | |
| Antibody | Anti-α-tubulin (mouse monoclonal) | Sigma-Aldrich | B512 | immunoblotting (1:25,000) |
| Antibody | Anti-Ac-tubulin (mouse monoclonal) | Sigma-Aldrich | 6-11B-1 | immunoblotting (1:50,000) |
| Antibody | Anti-GFP (YFP) (mouse monoclonal) | Roche | 7.1 and 13.1 | immunoblotting (1:1,000) |
| Antibody | HRP-conjugated goat anti-mouse IgG | Jackson Lab. | RRID:AB_10015289 | immunoblotting (1:10,000) |
| Antibody | HRP-conjugated goat anti-rabbit IgG | Jackson Lab. | RRID:AB_2313567 | immunoblotting (1:10,000) |
| Antibody | Alexa-Fluor 594-conjugated goat anti-rabbit IgG | Molecular Probes | RRID:AB_142057 | immunofluorescence (1:400) |
| Antibody | Alexa-Flour 488-conjugated goat anti-mouse IgG | Molecular Probes | RRID:AB_2536097 | immunofluorescence (1:400) |





