Specificity in glycosylation of multiple flagellins by the modular and cell cycle regulated glycosyltransferase FlmG
Figures
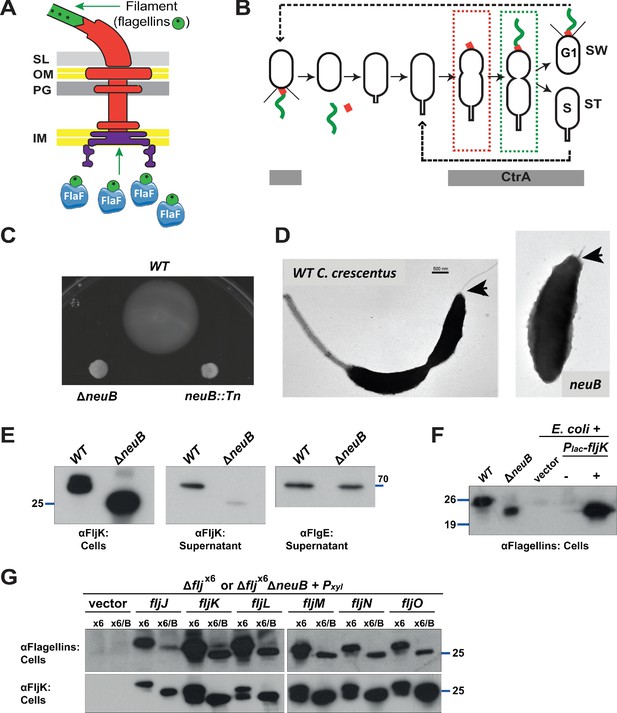
Mutation of neuB affects the assembly of the flagellar filament.
(A) Schematic of the C. crescentus flagellum with the MS- and C-ring structures in the inner membrane (IM), the hook basal body components spanning the periplasm (with the peptidoglycan – PG – layer) and outer membrane (OM), and the filament. Flagellin subunits (in green) are brought to the export machinery by the secretion chaperone FlaF. Purple star on flagellins indicates the post-translational modification by glycosylation. (B) Schematic of the C. crescentus cell cycle. The grey bar represents the time during the cell cycle when CtrA is present and activate transcription of flagellar genes. The hook structure (FlgE, in red) is synthesized in early pre-divisional cells, whereas the flagellar filament (in green) is polymerized from flagellins in late pre-divisional cells. Both,flagellar filament and hook, are shed during the swarmer (SW) to stalked (ST) cell transition. (C) Motility assay of neuB::Tn and ΔneuB mutants compared to WT strain. Overnight cultures were spotted on PYE soft agar plates and incubated for 72 hours at 30°C. Compact swarms indicate that neuB mutant cells are non-motile. (D) WT and neuB::Tn cells analyzed by transmission electron microscopy show that only a short protrusion is visible at the SW pole of neuB::Tn cells, in contrast to the WT strain (black arrow). The images suggest that in neuB mutant cells the flagellar hook is stably assembled, but not the flagellar filament. (E) Immunoblots o performed with anti-FljK (αFljK, raised against FljK produced in E. coli, see methods) and anti-FlgE (Hahnenberger and Shapiro, 1987) antibodies on cell lysates and supernatants of WT and ΔneuB cultures show that flagellins are produced in ΔneuB cells but not efficiently exported, whereas the export of the FlgE hook protein is not affected. The migration of FljK in ΔneuB cells is shifted towards lower molecular mass, suggesting that post-translational modification of flagellin is defective in the ΔneuB mutant. Molecular size standards are indicated by the blue lines with the corresponding value in kDa. (F) Immunoblot performed on extracts from E. coli cells expressing C. crescentus FljK under control of Plac from a plasmid. FljK expressed in E. coli shows the same migration profile as in ΔneuB cells, indicating that E. coli cells cannot post-translationally modify C. crescentus FljK. Molecular size standards are indicated by the blue lines with the corresponding value in kDa. Note that antibodies used in this immunoblot were raised against flagellins purified from C. crescentus (αFlagellins; Hahnenberger and Shapiro, 1987). (G) Immunoblots on extracts from Δfljx6 (x6) and Δfljx6ΔneuB (x6/B) cells expressing each flagellin from a plasmid under Pxyl control. The immunoblots were performed with antibodies raised against purified C. crescentus flagellins (upper panel) or FljK expressed and purified from E. coli (lower panel). In both cases, all six flagellins show a shift to a lower molecular mass in their migration in the absence of neuB, suggesting that all six flagellins are post-translationally modified. Molecular size standards are indicated by the blue lines with the corresponding value in kDa. Both antibodies recognize all six flagellins. Note that antibodies raised against flagellins purified from Caulobacter recognize the glycosylated form of all six flagellins better, whereas the antibodies raised against FljK expressed and purified from E. coli also efficiently recognizes unglycosylated flagellins.
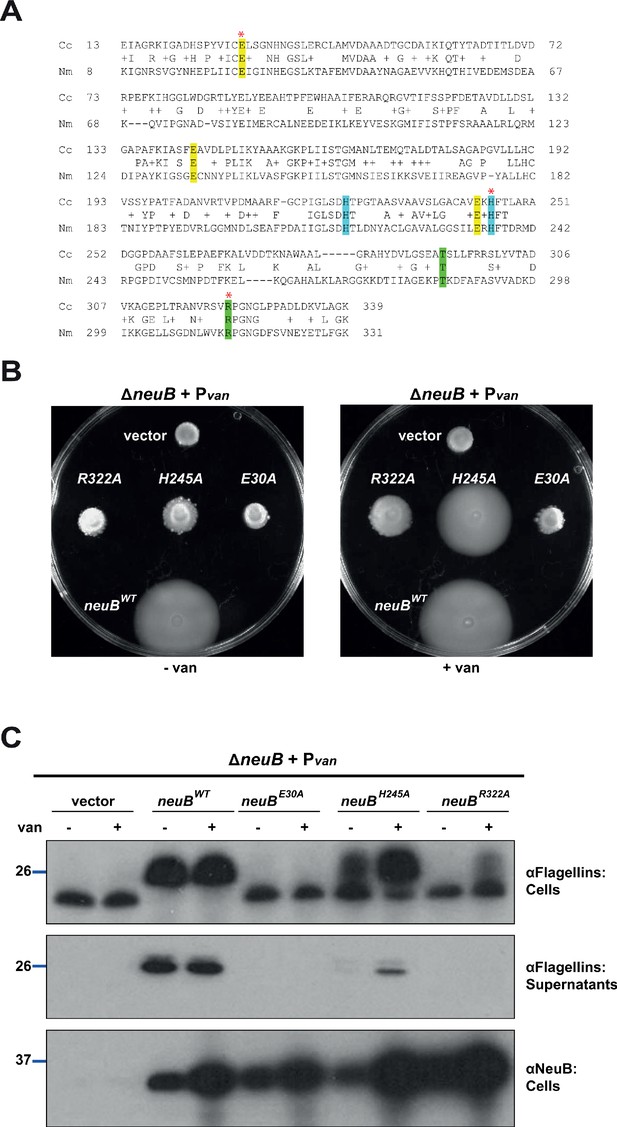
NeuB putative catalytic activity is required for motility and flagellin modification.
(A) Sequence alignment of C. crescentus NeuB (Cc) to N. meningitidis sialic acid synthase (Nm). The three glutamate residues that have been proposed to stabilize the reaction intermediate are highlighted in yellow; the two histidine residues that coordinate the Mn2+ cofactor are highlighted in blue; the threonine and arginine residues highlighted in green are located in the antifreeze-like C-terminal domain and protrude into the active site of the other subunit in the N. meningitidis sialic acid synthase dimer. The residues selected for site-directed mutagenesis in C. crescentus are indicated by a red asterisk. (B) Motility assay of ΔneuB cells complemented with different neuB alleles expressed from Pvan on a plasmid. Only the WT neuB allele can fully complement the motility defect of the ΔneuB strain, whereas the allele encoding NeuB(H245A) complements partially and the NeuB(E30A) and (R322A) versions do not restore motility. (C) Immunoblots showing the levels of flagellins and NeuB in ΔneuB cells complemented with different NeuB versions expressed from Pvan on a plasmid. All the NeuB variants were expressed (lower panel). Immunoblotting for flagellins in whole cell lysates (cells, upper panel) indicates that only the WT NeuB version can restore the migration profile of flagellins, whereas the E30A and R322A variants are inactive and the H245A variant shows an intermediate phenotype. The middle panel shows that only upon induction of the H245A version can the flagellins be detected in the culture supernatant, in agreement with the motility assay shown in panel B. Molecular size standards are indicated by the blue lines, with the corresponding value in kDa. Note that antibodies used in this immunoblot wereraised against flagellins purified from C. crescentus (αFlagellins; Hahnenberger and Shapiro, 1987) and they detect the glycosylated version of the flagellin better than the other flagellin antiserum (αFljK, see Figure 1).
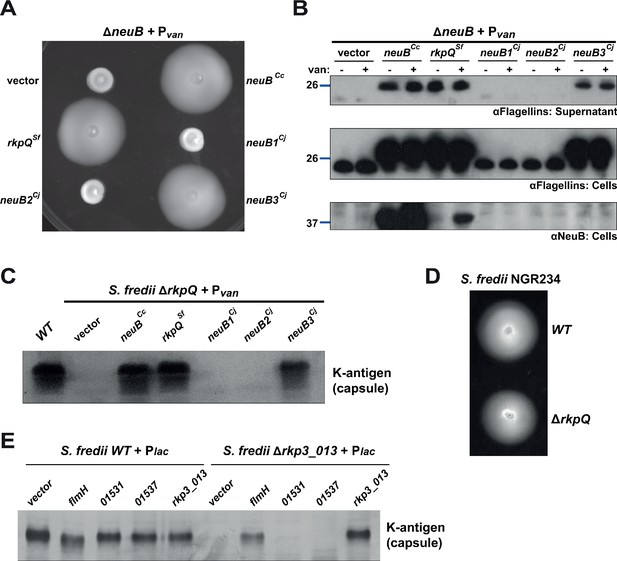
Heterologous complementation of the ΔneuB mutant with pseudaminic acid synthases.
(A) Motility assay of ΔneuB cells complemented with different neuB homologs expressed from Pvan on plasmid. Only NeuB from C. crescentus and the homologs known to be pseudaminic acid synthases (RkpQ from S. fredii and NeuB3 from C. jejuni) can fully complement the motility defect of the ΔneuB strain, whereas NeuB1 and NeuB2 from C. jejuni do not restore motility. (B) Immunoblots showing the intracellular levels of flagellins and NeuB in ΔneuB cells complemented with different NeuB homologs expressed from Pvan on a plasmid. Consistent with the motility assay shown in panel A, only NeuBCc, RkpQSf, and NeuB3Cj can restore the flagellin migration profile (in whole cell lysates, middle panel) and the secretion of flagellin in the supernatant (upper panel) in ΔneuB cells. RkpQSf is the protein that shows the highest similarity to NeuBCc, as shown by the fact that RkpQSf is detected by the antibodies against NeuBCc (lower panel). The blue lines on the left indicate the migration of the molecular size standards, with the corresponding value in kDa. Note that antibodies used in this blot were raised against flagellins purified from C. crescentus (αFlagellins; Hahnenberger and Shapiro, 1987). (C) Capsular polysaccharide profile of S. fredii NGR234 ΔrkpQ mutant cells expressing different NeuB homologs from Pvan on a plasmid. Cells with a ΔrkpQ mutation do not produce capsular polysaccharide, but production of capsular polysaccharide in ΔrkpQ cells can be restored by NeuBCc or NeuB3Cj (but not by NeuB1Cj or NeuB2Cj), which indicates that NeuBCc is a pseudaminic acid synthase. (D) Motility assay of S. fredii NGR234 WT and ΔrkpQ mutant showing that mutation of the pseudaminic acid synthase does not affect motility in S. fredii. (E) Capsular polysaccharide profile of S. fredii WT and Δrkp3_013 expressing putative C. crescentus acetyltransferases from Plac on a plasmid. Production of capsular polysaccharide in Δrkp3_013 cells can be restored only by expression of flmH, which suggests that FlmH can participate in the pseudaminic acid biosynthetic pathway.
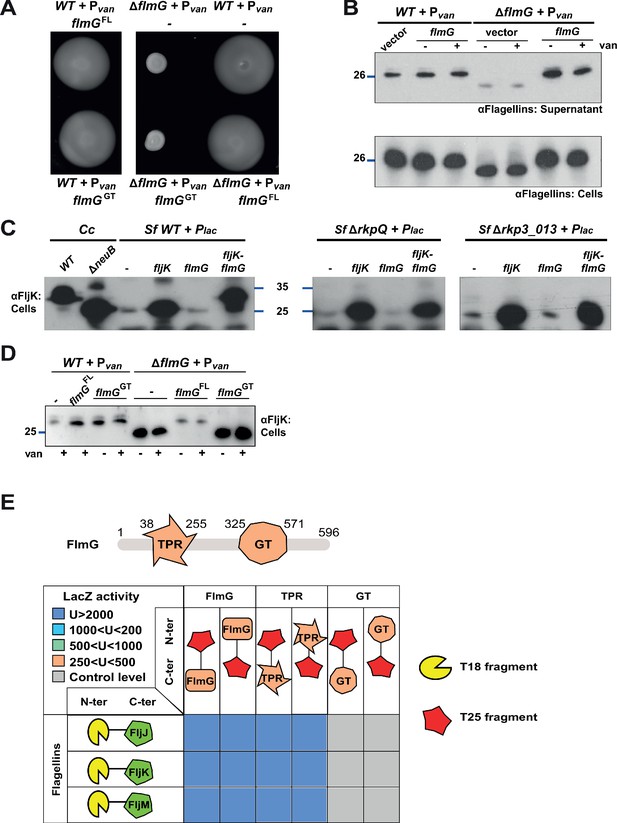
FlmG is the putative glycosyltransferase required to post-translationally modify flagellins.
(A) Motility assay of WT and ΔflmG cells complemented with Pvan-flmG full-length (FL) or glycosyltransferase CTD (GT) on plasmid. ΔflmG cells are non-motile, as indicated by the compact swarming. The expression of FlmG full-length from Pvan restores motility, in contrast to the GT domain alone. (B) Immunoblot showing the levels of flagellins in supernatants and whole cell lysates from WT and ΔflmG cells complemented with Pvan-flmG on a plasmid. Flagellins produced by ΔflmG cells show the same migration profile as those produced by ΔneuB cells, with faster migration and reduced abundance in the supernatant. The blue lines on the left indicate the molecular size standard, with the corresponding value in kDa. Note that antibodies used in this blot were raised against flagellins purified from C. crescentus (αFlagellins; Hahnenberger and Shapiro, 1987). (C) Immunoblot on C. crescentus FljK expressed in S. fredii NGR234 strains. The left panel shows that when expressed alone in WT S. fredii FljK migrates faster, like in the C. crescentus ΔneuB mutant. By contrast, co-expression of FlmG and FljK in S. fredii NGR234 results in a shift in the migration profile of FljK, similar to that observed in C. crescentus WT cells. When FlmG and FljK are co-expressed in S. fredii strains unable to synthesize pseudaminic acid (ΔrkpQ, middle panel; Δrkp3_013, right panel) FljK shows only the fast migrating band, independently from the presence of FlmG. The blue lines indicate the molecular size standards, with the corresponding value in kDa. (D) Immunoblot with anti-FljK antibodies on whole cell lysates from WT and ΔflmG cells expressing FlmG full-length (FL) or the glycosyltransferase CTD (GT) from Pvan on a plasmid. The expression of the GT domain alone does not restore the migration profile of flagellins in ΔflmG cells, in agreement with the motility defect shown in panel A. (E) BACTH assay showing the interaction of FlmG with flagellins. The cartoon represents FlmG, with the N-terminal TPR domain and the C-terminal glycosyltransferase domain. The scheme below shows the β-galactosidase activity, expressed in Miller units (U) of E. coli BTH101 cells containing the pair-wise combinations of the different constructs used for the BACTH assay. T18 and T25 correspond to the two fragments of the adenylate cyclase and are represented as yellow shape and red star, respectively. Hybrids with the proteins of interest (FlmG, FljJ, FljK, and FljM) were created as N-ter or C-ter fusions, as mentioned. In the case of FlmG, fusions were created with the full-length protein, the TPR domain or the glycosyltransferase (GT) domain only. Gray squares correspond to β-galactosidase activity similar to the control (empty plasmid, U < 250), orange squares indicate an activity between 250 and 500 U, light green squares between 500 and 1000 U, light blue between 1000 and 2000 U, and dark blue above 2000 U. The values of β-galactosidase correspond to the mean and standard deviation of three independent experiments and are listed in Supplementary file 1 Table S1.
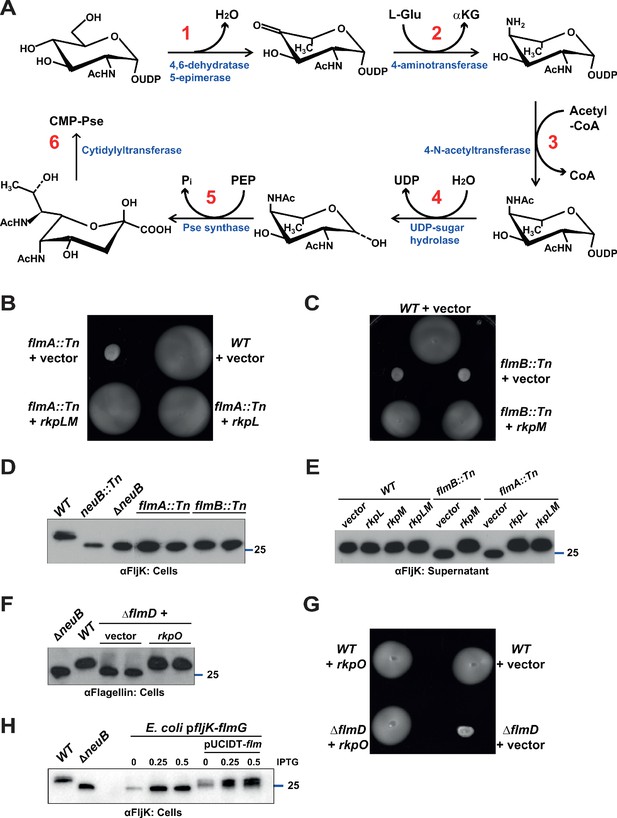
Pseudaminic acid biosynthetic pathway in C. crescentus.
(A) Schematic of the pseudaminic acid biosynthetic pathway as it has been described in C. jejuni and H. pylori (reviewed in Salah Ud-Din and Roujeinikova, 2018). The different steps are catalyzed by PseB (1), PseC (2), PseH (3), PseG (4), PseI (5), and PseF (6). (B) Motility assay of WT and flmA::Tn cells expressing S. fredii NGR234 rkpL and rkpM from Pvan ona plasmid. (C) Motility assay of WT and flmB::Tn cells expressing S. fredii NGR234 rkpM from Pvan on a plasmid. (D) Immunoblots of extracts from WT and mutant cells probed with polyclonal anti-FljK antibodies on whole cell lysates. Flagellins produced by flmA::Tn and flmB::Tn cells show the same migration profile as in ΔneuB mutant cells. The blue line indicates the migration of the molecular size standard, with the corresponding size in kDa. (E) Immunoblots of supernatants from WT, flmA::Tn and flmB::Tn cells probed with polyclonal anti-FljK antibodies. The expression of the S. fredii homologs RkpL and RkpM from Pvan on a plasmid restores the migration profile and secretion of flagellin in flmA::Tn and flmB::Tn cells, respectively. The blue line indicates the migration of the molecular size standard, with the corresponding size in kDa. (F) Immunoblot of extracts from WT and ΔflmD cells. Mutation of flmD impairs post-translational flagellin modification, and the defect is complemented by the expression of the S. fredii NGR234 homologue RkpO from a plasmid. The blue line indicates the migration of the molecular size standard, with the corresponding size in kDa. Note that antibodies used in this blot were raised against flagellins purified from C. crescentus (αFlagellins; Hahnenberger and Shapiro, 1987). (G) Motility assay of WT and ΔflmD cells expressing S. fredii NGR234 RkpO from Pvan on a plasmid. (H) Immunoblot with anti-FljK antibodies on whole cell lysates from E. coli expressing fljK and flmG from Plac on a plasmid, in the presence or absence of a compatible plasmid carrying the complete set of Caulobacter genes for the pseudaminic acid biosynthetic pathway (pUCIDT-flm, see Materials and methods and Supplementary file 1 Table S3). In the absence of pUCIDT-flm, FljK shows the same migration profile as in Caulobacter ΔneuB cells, whereas in the presence of pUCITD-flm FljK migration is shifted toward higher molecular mass, as in Caulobacter WT cells. The values above the panel indicate the concentration of the inducer for Plac-fljK-flmG (mM IPTG). The blue line indicates the migration of the molecular size standard, with the corresponding size in kDa.
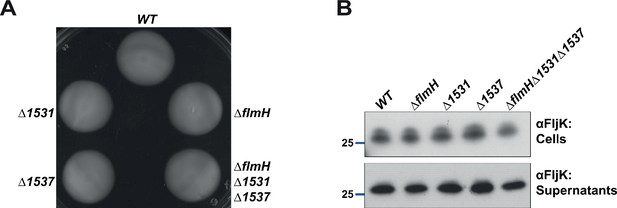
Acetyltransferase function is likely redundant in Caulobacter.
(A) Motility assay of strains mutated for the three putative acetyltransferases (ΔflmH, Δ1531, Δ1537, and the triple mutant ΔflmHΔ1531Δ1537). None of the mutants show a motility defect compared to WT cells. (B) Immunoblot with polyclonal anti-FljK antibodies on whole cell lysates and supernatants of strains mutated for the three putative acetyltransferases (ΔflmH, Δ1531, Δ1537, and the triple mutant ΔflmHΔ1531Δ1537). None of the mutants show a defect in flagellins migration or secretion. The blue line indicates the migration of the molecular size standard, with the corresponding size in kDa.
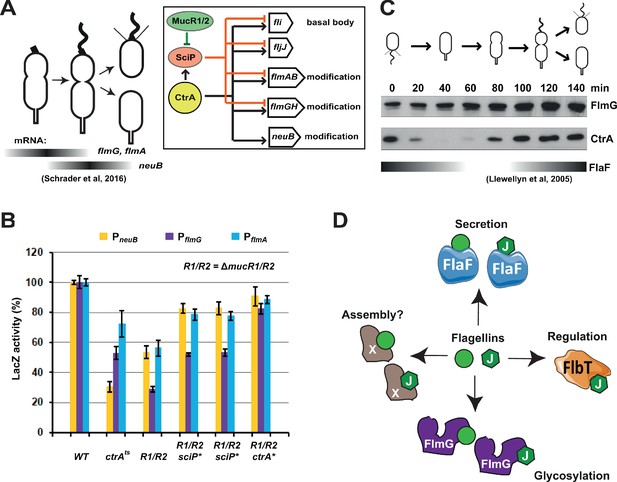
Transcription of genes encoding flagellin-modification proteins is cell cycle regulated.
(A) Schematic of the transcriptional regulatory network controlling the expression of flagellar and pseudaminic acid biosynthetic genes in C. crescentus pre-divisional cells. The gray bars represent the time during the cell cycle when flmA, flmG, and neuB mRNA levels peak (Schrader et al., 2016). The transcriptional regulatory network is represented in the box: CtrA promotes the expression of genes encoding the basal body (fli genes), the regulatory flagellin FljJ and several components of the flagellin glycosylation pathway (flmAB and flmGH operons, neuB). The expression of these same genes is repressed by SciP (with the exception of neuB), whose expression is under control of CtrA (positively) and MucR1/2 (negatively). (B) β-galactosidase activity of PneuB, PflmG, and PflmA transcriptional fusions. Transcription of neuB, flmG and flmAB is significantly reduced in ctrA401 (T170I, temperature sensitive allele ctrAts) and double ΔmucR1ΔmucR2 (R1/R2) strains. β-galactosidase activity of PneuB, PflmG, and PflmA is partially restored in ΔmucR1ΔmucR2 cells carrying ctrA(T170A), sciP(T24I), or sciP(T65A) alleles (ctrA* and sciP*). Values are expressed as percentages (activity in WT NA1000 set at 100%). (C) Immunoblot showing FlmG and CtrA protein levels in synchronized C. crescentus cells. The gray bar represents the abundance of the flagellin chaperone FlaF along the cell cycle, as previously shown (Llewellyn et al., 2005). (D) Model depicting a flagellin-centric view of the events during flagellar assembly in C. crescentus. The FljJ flagellin is illustrated as a green hexagon, whereas the other flagellins (FljKLMNO) are depicted as green circles. The events acting on flagellins are translational regulation (binding of FljJ by FlbT), glycosylation by FlmG, secretion by FlaF and possibly assembly by an unknown factor (X).
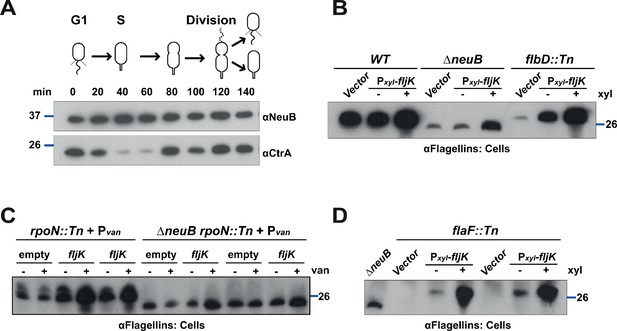
NeuB is stable along the cell cycle and flagellins can be glycosylated in HBB mutants or in the absence of the FlaF chaperone.
(A) Immunoblot showing protein levels in synchronized C. crescentus cells: NeuB levels do not change significantly over the cell cycle. The blue lines indicate the migration of the molecular size standards, with the corresponding size in kDa. (B) Immunoblots on WT, ΔneuB and flbD::Tn (lacks the hook, the rings structures in the peptidoglycan and in the outer membrane) cells expressing fljK from Pxyl on a plasmid. Upon induction with 0.3% xylose, FljK is produced in flbD::Tn cells and shows the same migration profile as in WT cells, which indicates that FljK is glycosylated in the flbD::Tn mutant. The blue line indicates the migration of the molecular size standard, with the corresponding size in kDa. Note that antibodies used in this blot were raised against flagellins purified from C. crescentus (αFlagellins; Hahnenberger and Shapiro, 1987). (C) Immunoblot on rpoN::Tn (lacks the hook, the rings structures in the peptidoglycan and in the outer membrane) and ΔneuB rpoN::Tn cells expressing fljK from Pvan on plasmid. FljK produced in rpoN::Tn cells shows a migration profile consistent with its modification by glycosylation, compared to ΔneuB rpoN::Tn cells. The blue line indicates the migration of the molecular size standard, with the corresponding size in kDa. Note that antibodies used in this blot were raised against flagellins purified from C. crescentus (αFlagellins; Hahnenberger and Shapiro, 1987). (D) Immunoblot on flaF::Tn cells expressing fljK from Pxyl on a plasmid. Upon induction with 0.3% xylose, FljK is produced in flaF::Tn cells and shows the same migration profile as in WT cells, which indicates that FljK is glycosylated in the absence of the FlaF chaperone. The blue line indicates the migration of the molecular size standard, with the corresponding size in kDa. Note that antibodies used in this blot were raised against flagellins purified from C. crescentus (αFlagellins; Hahnenberger and Shapiro, 1987).
Tables
Homology of the C. crescentus enzymes involved in the pseudaminic acid biosynthesis pathway to S. fredii NGR234 and C. jejuni proteins.
| C. crescentus | S. fredii NGR234 | C. jejuni NCTC 11168 | Enzymatic activity |
|---|---|---|---|
| FlmA (CCNA_00233) | RkpL 52% id; 68% sim | PseB 57% id; 72% sim | Dehydratase |
| FlmB (CCNA_00234) | RkpM 44% id; 58% sim | PseC 34% id; 52% sim | Aminotransferase |
| FlmH (CCNA_01523) | Rkp3_013† 26% id; 37% sim | PseH‡ 26% id; 50% sim | N-acetyltransferase |
| FlmD (CCNA02947) | RkpO 36% id; 49% sim | PseG 22% id; 41% sim | UDP-sugar hydrolase |
| NeuB (CCNA_02961) | RkpQ 56% id; 70% sim | PseI 43% id; 59% sim | Pseudaminic acid synthase |
| FlmC§ (CCNA_02946) | RkpN | PseF | Cytidyltransferase |
| FlmG (CCNA_01524) | - | - | Protein Glycosyltransferase |
-
* Although C. crescentus genomes encodes for several putative acetyltransferases, only CCNA_01531 has significant homology to FlmH (35% identity [id] and 54% similarity [sim]).
† FlmH aligns to Rkp3_013 only for about 50% of its length.
-
‡ FlmH aligns to PseH for about 75% of its length.
§ FlmC shows no homology to RkpN or PseF; RkpN and PseF share 46% identity and 68% similarity.
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, Strain Background (Caulobacter crescentus NA1000) | Caulobacter crescentus NA1000 | Evinger and Agabian, 1977; PMID:334726 | See Supplementary file 2 Table S2 | |
| Antibody | CtrA Rabbit polyclonal | Domian et al., 1997; PMID:9267022 | Home-made antibodies raised against full-length protein of C. crescentus | Immunoblot: 1/10000 dilution Figure 6, Figure 6—figure supplement 1 |
| Antibody | FlgE Rabbit polyclonal | Hahnenberger and Shapiro, 1987; PMID:3039149 | Home-made antibodies raised against FlgE protein of C. crescentus | Immunoblot: 1/50000 dilution Figure 1 |
| Antibody | Flagellins Rabbit polyclonal | Hahnenberger and Shapiro, 1987; PMID:3039149 | Home-made antibodies raised against purified flagellins from C. crescentus | Immunoblot: 1/20000 dilution Figure 1, Figure 2, Figure 3, Figure 4, Figure 5, Figure 6—figure supplement 1 |
| Antibody | FljK Rabbit polyclonal | Ardissone et al., 2020 | Home-made antibodies raised against full-length protein of C. crescentus | Immunoblot: 1/10000 dilution Figure 1, Figure 4, Figure 5, Figure 5—figure supplement 1 |
| Antibody | NeuB Rabbit polyclonal | This paper | Home-made antibodies raised against full-length protein of C. crescentus | Immunoblot: 1/10000 dilution Figure 2, Figure 3, Figure 6—figure supplement 1 |
| Antibody | FlmG Rabbit polyclonal | This paper | Home-made antibodies raised against FlmG (amino acids 301–500) of C. crescentus | Immunoblot: 1/10000 dilution Figure 6 |
| Recombinant DNA reagent | Plasmids | This paper | See Supplementary file 2 Table S2 | |
| Sequence-based reagent | PCR primers | This paper | See Supplementary file 3 Table S3 | |
| Sequence-based reagent | FljK synthetic sequence | This paper | FljK coding sequence, codon-optimized for E. coli | See Supplementary file 3 Table S3 |
| Sequence-based reagent | Pseudaminic acid biosynthesis operon synthetic sequence | This paper | FlmA, FlmB, FlmH, FlmD, NeuB and FlmC coding sequences, codon-optimized for E. coli | See Supplementary file 3 Table S3 |
Additional files
-
Supplementary file 1
Table S1.
LacZ activity of BACTH assay.
- https://cdn.elifesciences.org/articles/60488/elife-60488-supp1-v1.docx
-
Supplementary file 2
Table S2.
Strains and plasmids used in this study.
- https://cdn.elifesciences.org/articles/60488/elife-60488-supp2-v1.docx
-
Supplementary file 3
Table S3.
Oligonucleotides and synthetic genes used in this study.
- https://cdn.elifesciences.org/articles/60488/elife-60488-supp3-v1.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/60488/elife-60488-transrepform-v1.pdf





