No relationship between frontal alpha asymmetry and depressive disorders in a multiverse analysis of five studies
Figures
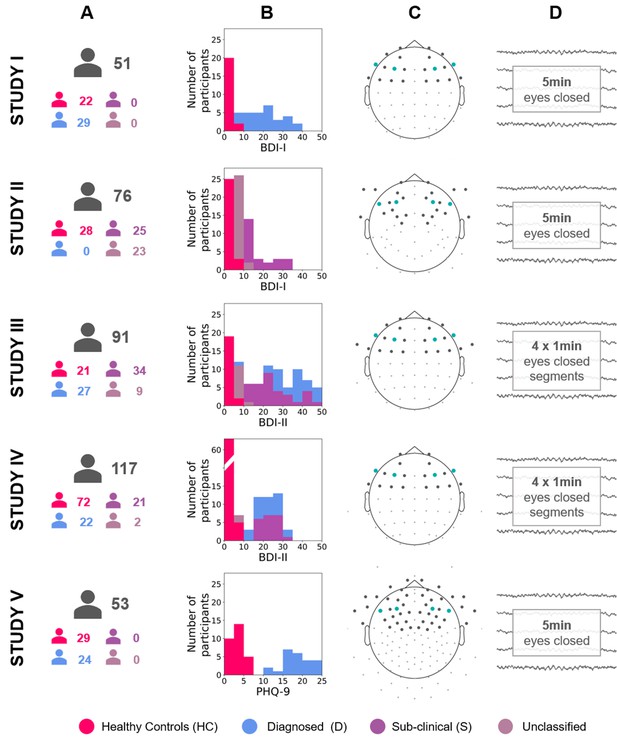
Diagram describing the five studies included in this article (Studies I, II, III, IV, and V).
(A) Number of participants for each study and group (see Table 12 for details). (B) Stacked histograms showing the distribution of BDI or PHQ-9 scores in each study and each group. (C) Channel montage. Frontal channels used in cluster-based analyses are marked with gray dots. Channels used in channel-pairs analysis are marked with teal dots (F3–F4, F7–F8, and corresponding channels in the EGI montage). (D) Rest period length and scheme.
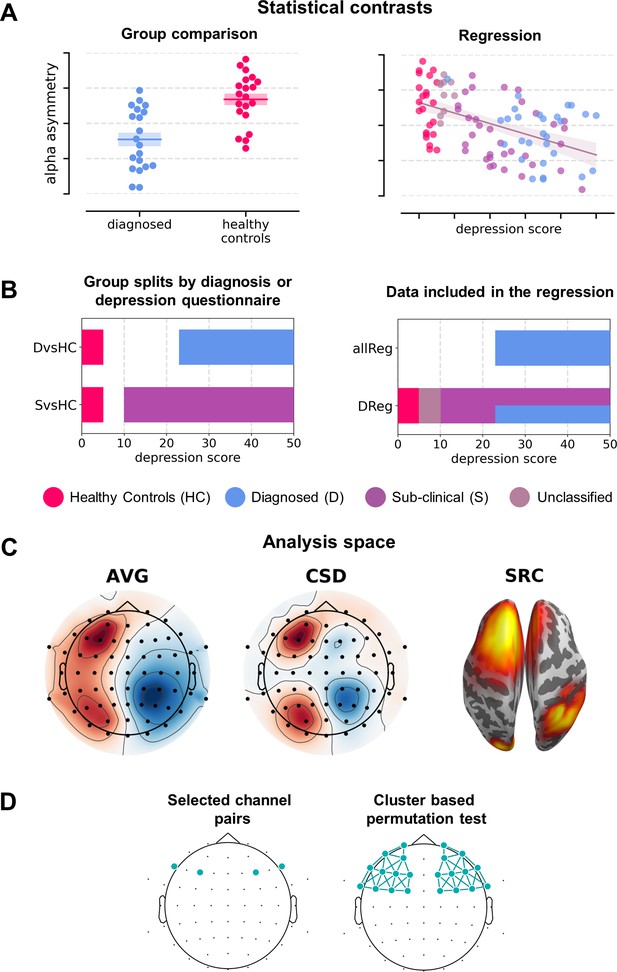
Analysis variants used (described in detail in Variants of statistical analysis section).
(A) Schematic depiction of given statistical contrast: group comparisons (left) vs regression (right). (B) Specification of each contrast against depression scores. Left panel shows a schematic range of depression scores for each contrast: diagnosed vs healthy controls (DvsHC) and sub-clinical vs healthy controls (SvsHC). Right panel shows the range of depression scores for data included in each linear contrast: regression on diagnosed subjects (DReg) uses only subjects with clinical diagnosis, while regression on all subjects (allReg) uses all subject groups. The color legend for the subject groups is presented below these figures. (C) Analysis space: AVG – channel level, average reference; CSD – channel level, current source density; SRC – source level, DICS beamforming. (D) Schematic depiction of analysis method: selected channel pairs versus all frontal channels with cluster-based correction for multiple comparisons.
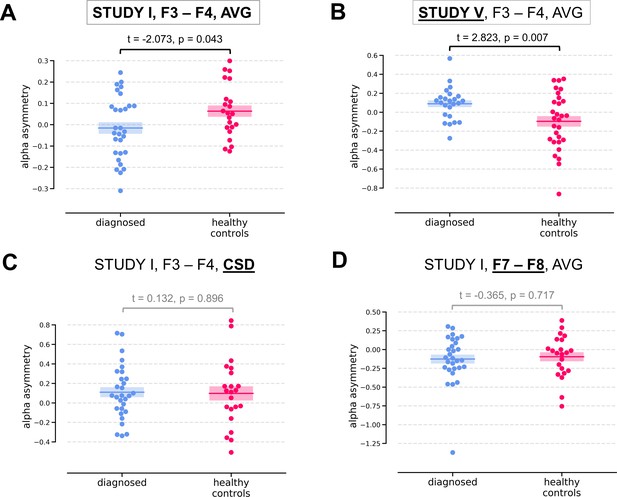
Selected results for channel-pairs level analyses, depressed vs healthy controls (DvsHC) contrast.
Panels A and B show 2 of 3 significant channel pair results: average referenced F3–F4 channel pair for Studies I (A) and V (B). The remaining panels (C and D) show other channel pair analysis variants: specifically those that differ by exactly one parameter (underlined text) from the result presented in the panel A. Horizontal lines represent averages for each group, and shaded areas show standard error of the mean.
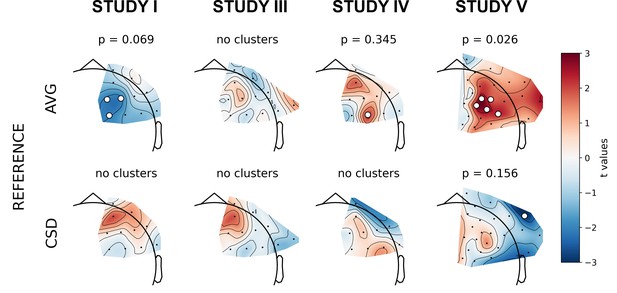
Selected results of cluster-based analyses: topographies of DvsHC contrast effects in a reference by study matrix.
More positive (red) t values indicate more right-sided (less left-sided) alpha asymmetry for diagnosed participants. More negative (blue) t values indicate more right-sided (less left-sided) FAA for healthy controls. Channels that are part of a cluster are marked with white dots.

Results of cluster-based analyses for allReg contrast (regression on all subjects).
Topographies of effects are presented in a reference by study matrix. Color bar on the right presents color coding for the t values: more positive (red)/negative (blue) relationship between FAA and BDI. Channels that are part of a cluster are marked with white dots.
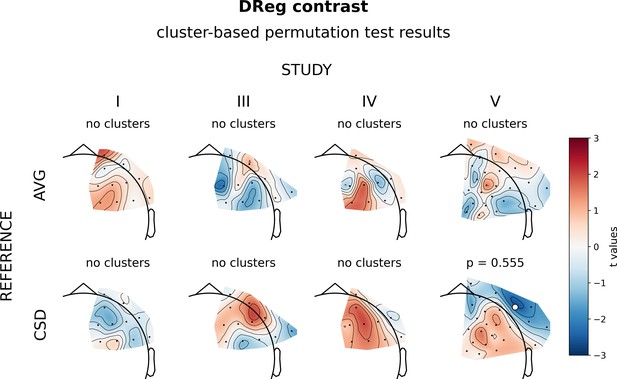
Results of cluster-based analyses for DReg contrast (regression on diagnosed subjects).
Topographies of effects are presented in a reference by study matrix. Color bar on the right presents color coding for the t values: more positive (red)/negative (blue) relationship between FAA and BDI. Channels that are part of a cluster are marked with white dots.
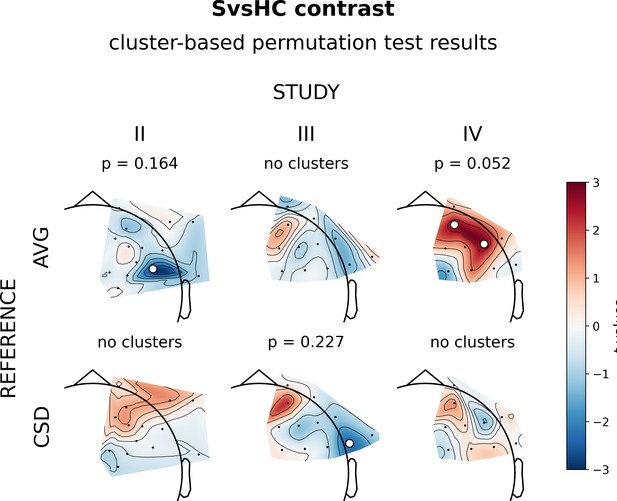
Results of cluster-based analyses for SvsHC (comparison between subclinical and healthy controls) contrast.
Topographies of different contrast effects in a reference by study matrix. More positive (red) t values indicate more right-sided (less left-sided) alpha asymmetry for subclinical participants. More negative (blue) t values indicate more right-sided (less left-sided) FAA for healthy controls. Channels that are part of a cluster are marked with white dots.
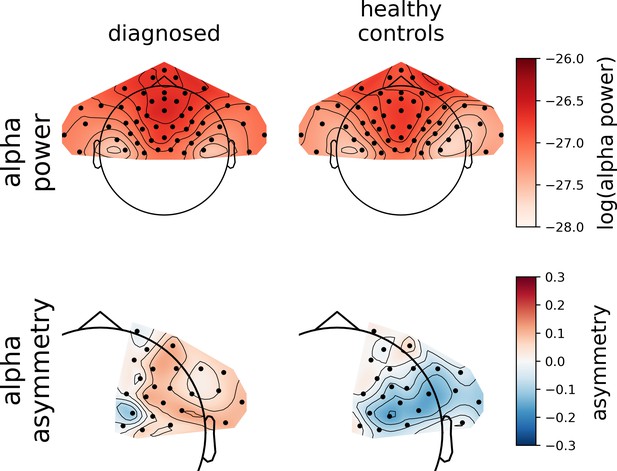
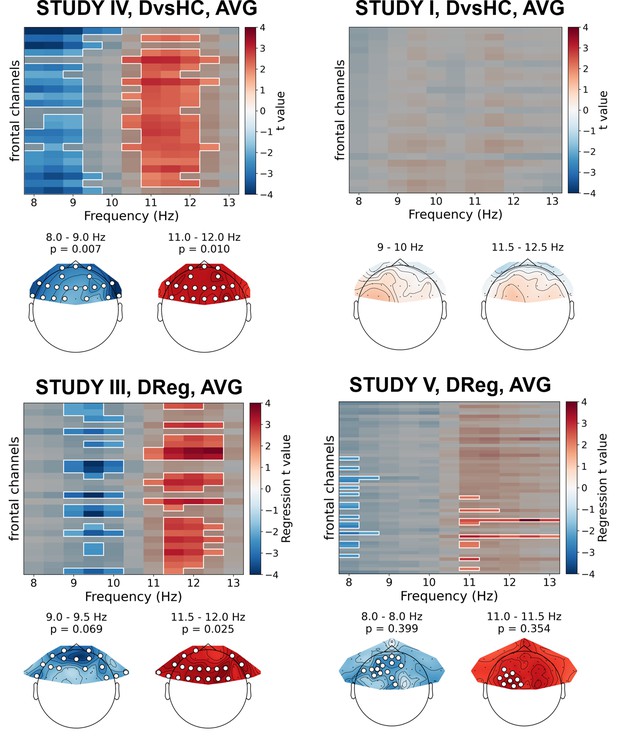
An example of a more detailed visualization of asymmetry effects: group-level averages for DvsHC, Study V, AVG from F.
First row shows topography of average alpha power for both groups; second row shows topography of average alpha asymmetry for both groups.
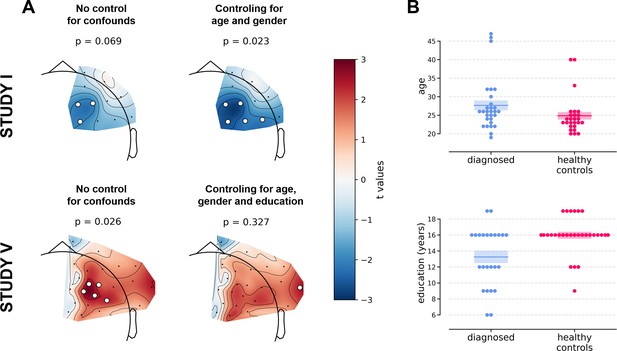
Selected results of cluster-based analyses showing the influence of statistical control for confounding variables like age, gender (Studies I and V), and education (Study V).
Selected results of cluster-based analyses on standardized data.
Heatmaps in the upper part of each panel represent regression t values for channel by frequency search space. More positive/negative t values indicate higher/lower power with higher BDI. Clusters are indicated in the heatmaps with white outline. In each panel we present two topographies below the heatmap: showing average effect for lower and higher frequency ranges determined by the positions of the clusters. Channels that are part of a cluster are marked with white dots in the topographical plots.
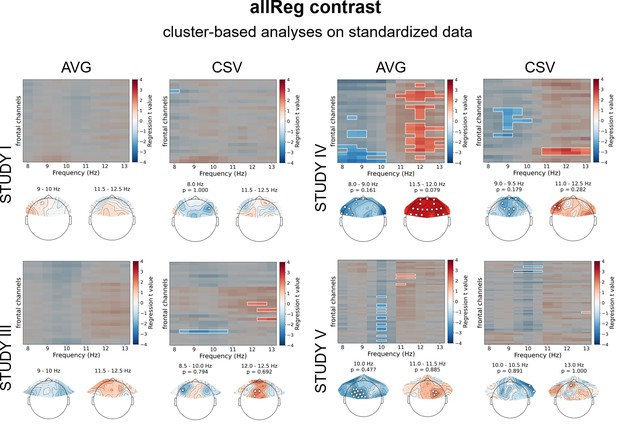
Results of cluster-based analyses on standardized data for allReg contrast (linear regression between FAA and BDI on all subjects together).
Heatmaps in the upper part of each panel represent regression t values for channel by frequency search space. More positive/negative t values indicate higher/lower power with higher BDI. Clusters are indicated in the heatmaps with white outline. In each panel we present two topographies below the heatmap: showing average effect for 9–10 Hz and 11.5–12.5 Hz range respectively. Channels that are part of a cluster are marked with white dots in the topographical plots.

Results of cluster-based analyses on standardized data for DReg contrast (linear regression between FAA and BDI restricted to the non-diagnosed subjects).
Heatmaps in the upper part of each panel represent regression t values for channel by frequency search space. More positive/negative t values indicate higher/lower power with higher BDI. Clusters are indicated in the heatmaps with white outline. In each panel we present two topographies below the heatmap: showing average effect for 9–10 Hz and 11.5–12.5 Hz range respectively. Channels that are part of a cluster are marked with white dots in the topographical plots.
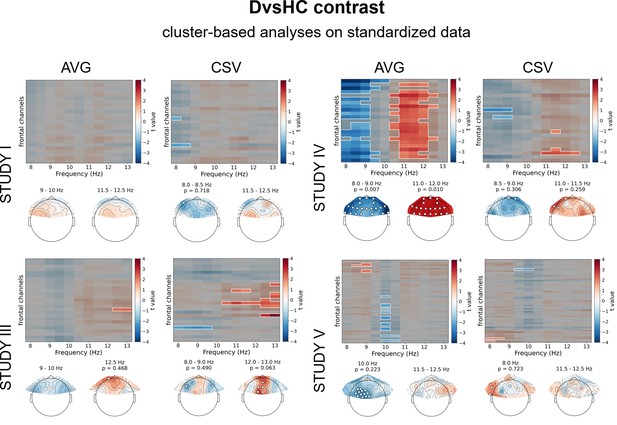
Results of cluster-based analyses on standardized data for DvsHC contrast (comparison between diagnosed and healthy controls).
Heatmaps in the upper part of each panel represent t values for channel by frequency search space. More positive (red) t values indicate higher power in given channel and frequency for diagnosed participants. More negative (blue) t values indicate higher power in given channel and frequency for healthy controls. Clusters are indicated in the heatmaps with white outline. In each panel we present two topographies below the heatmap: showing average effect for 9–10 Hz and 11.5–12.5 Hz range respectively. Channels that are part of a cluster are marked with white dots in the topographical plots.
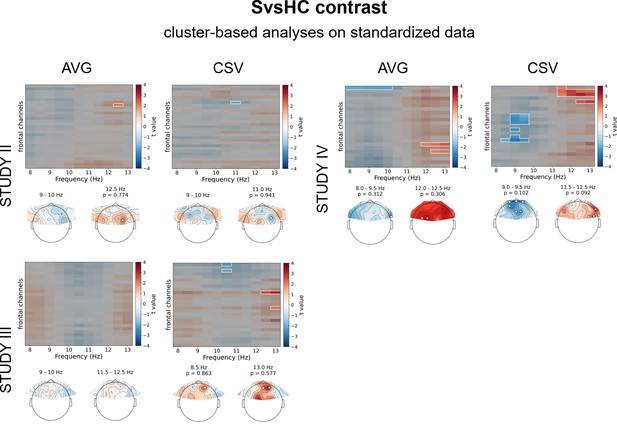
Results of cluster-based analyses on standardized data for SvsHC contrast (comparison between subclinical and healthy controls).
Heatmaps in the upper part of each panel represent t values for channel by frequency search space. More positive (red) t values indicate higher power in given channel and frequency for diagnosed participants. More negative (blue) t values indicate higher power in given channel and frequency for healthy controls. Clusters are indicated in the heatmaps with white outline. In each panel we present two topographies below the heatmap: showing average effect for 9–10 Hz and 11.5–12.5 Hz range respectively. Channels that are part of a cluster are marked with white dots in the topographical plots.
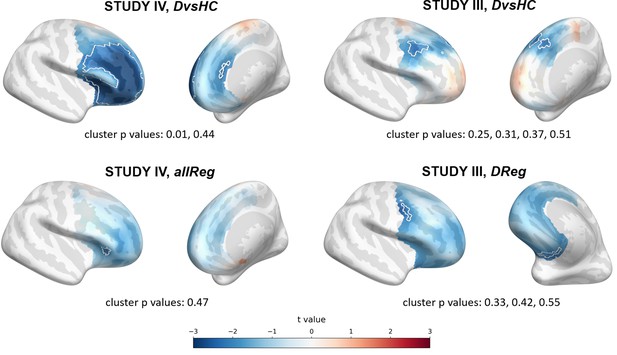
Selected results of source level analyses showing spatial t value maps for respective contrasts.
Cluster limits are marked with white outlines, and corresponding cluster p-values are shown below each panel. Color bar at the bottom presents color coding for the t values.
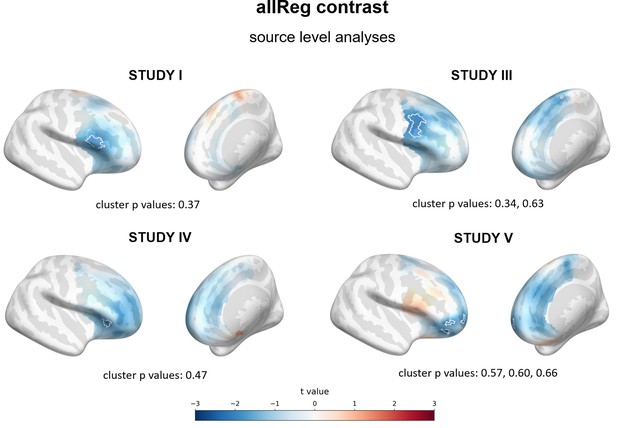
Results of source level analyses for allReg contrast (linear regression between FAA and BDI on all subjects) showing spatial t value maps for regression analyses.
Cluster limits are marked with white outlines, and corresponding cluster p-values are shown below each panel. Color bar at the bottom presents color coding for the t values: more positive (red)/negative (blue) relationship between FAA and BDI.
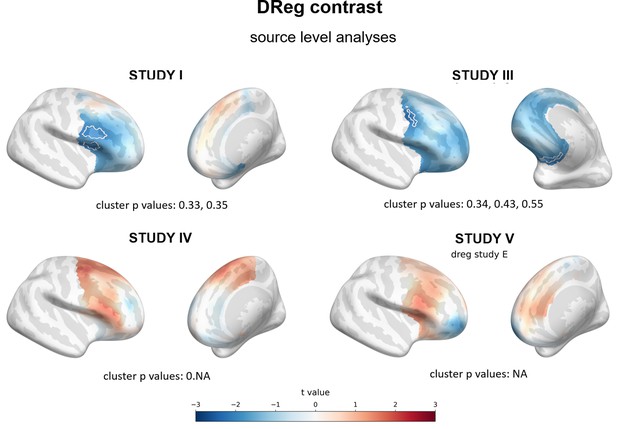
Results of source level analyses for DReg contrast (linear regression between FAA and BDI restricted to diagnosed subjects) showing spatial t value maps for regression analyses.
Cluster limits are marked with white outlines, and corresponding cluster p-values are shown below each panel. Color bar at the bottom presents color coding for the t values: more positive (red)/negative (blue) relationship between FAA and BDI. Linear regression restricted to diagnosed subjects; NA: no cluster with p<0.05 is present.
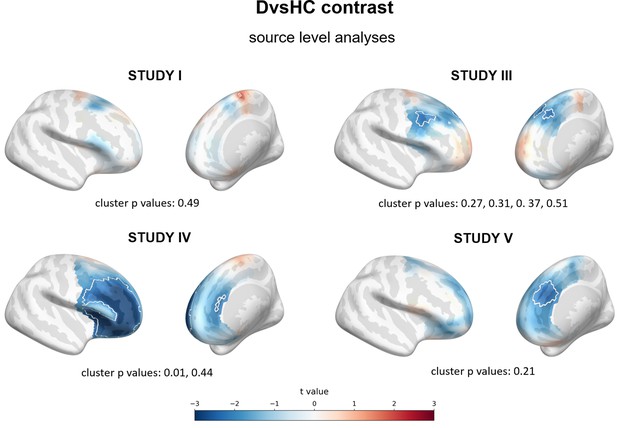
Results of source level analyses for DvsHC contrast (comparison between diagnosed and healthy controls) showing spatial t value maps for regression analyses.
Cluster limits are marked with white outlines, corresponding cluster p-values are shown below each panel. More positive (red) t values indicate more right-sided (less left-sided) alpha asymmetry for diagnosed participants. More negative (blue) t values indicate more right-sided (less left-sided) FAA for healthy controls.
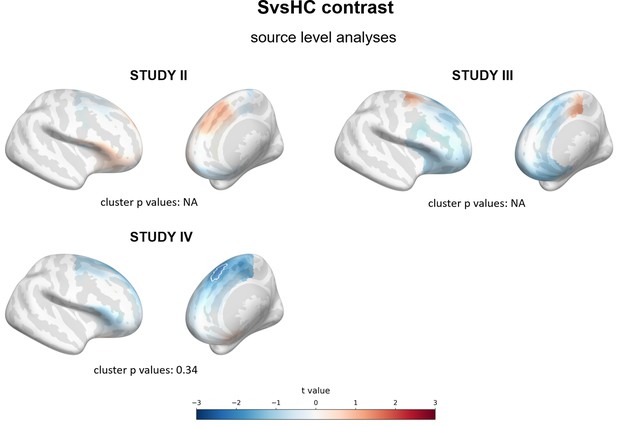
Results of source level analyses for SvsHC contrast (comparison between subclinical and healthy controls) showing spatial t value maps for regression analyses.
Cluster limits are marked with white outlines, corresponding cluster p-values are shown below each panel. More positive (red) t values indicate more right-sided (less left-sided) alpha asymmetry for subclinical participants. More negative (blue) t values indicate more right-sided (less left-sided) FAA for healthy controls. NA: no cluster with p<0.05 is present.
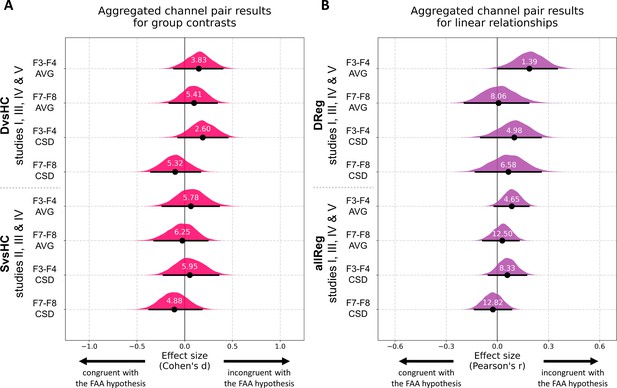
Results for channel pair analyses where studies including identical group contrasts (A) and linear contrasts (B) are combined.
Each row corresponds to one analysis on a single channel pair. The contrasts, studies, and channel pairs are labeled on the y axis. The black dots correspond to observed effect sizes in Cohen’s d/Pearson’s r, while the black lines indicate 95% confidence intervals for the effect size estimated using bias-corrected accelerated bootstrapping. The magenta/purple shapes represent bootstrap distributions and the white numbers printed on the distributions are Bayes factors for the null hypothesis (BF01). BF01 of 4 indicates that the data are four times more likely under the null than the alternative hypothesis. BF01 between 3 and 10 are considered moderate evidence for the null hypothesis.
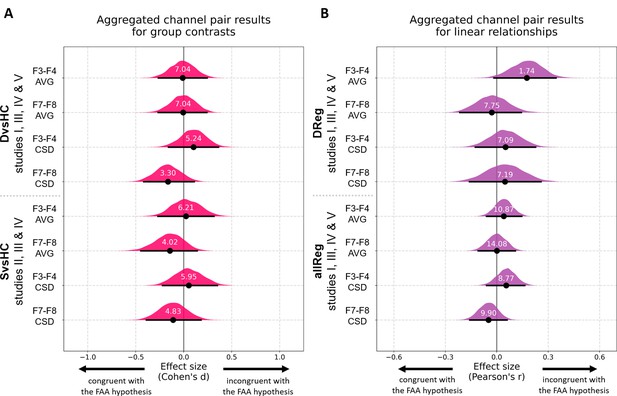
Results for channel pair analyses with control for confounds where studies including identical group contrasts (A) and linear contrasts (B) are combined.
Each row corresponds to one analysis on a single channel pair. The contrasts, studies, and channel pairs are labeled on the y axis. The black dots correspond to observed effect sizes in Cohen’s d/Pearson’s r, while the black lines indicate 95% confidence intervals for the effect size estimated using bias-corrected accelerated bootstrapping. The magenta/purple shapes represent bootstrap distributions and the white numbers printed on the distributions are Bayes factors for the null hypothesis (BF01). BF01 of 4 indicates that the data are four times more likely under the null than the alternative hypothesis. BF01 between 3 and 10 are considered moderate evidence for the null hypothesis.
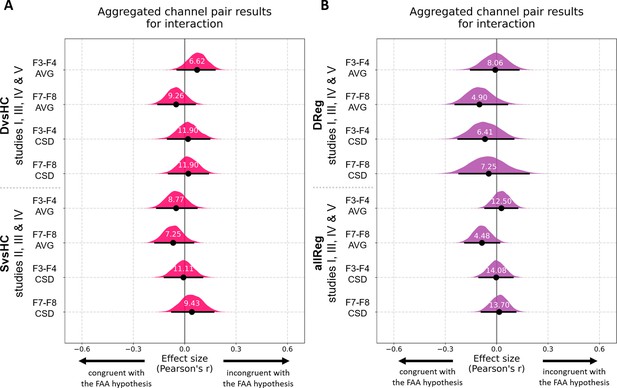
Results for channel pair gender × contrast interaction analyses on aggregated data with control for confounds.
In group contrasts (A) the interaction term is female * diagnosed; in linear contrasts (B) it is female * depression score. Each row corresponds to one analysis on a single channel pair. The contrasts, studies, and channel pairs are labeled on the y axis. The black dots correspond to observed effect sizes in Pearson’s r, while the black lines indicate 95% confidence intervals for the effect size estimated using bias-corrected accelerated bootstrapping. The magenta/purple shapes represent bootstrap distributions and the white numbers printed on the distributions are Bayes factors for the null hypothesis (BF01). BF01 of 4 indicates that the data are four times more likely under the null than the alternative hypothesis. BF01 between 3 and 10 are considered moderate evidence for the null hypothesis.
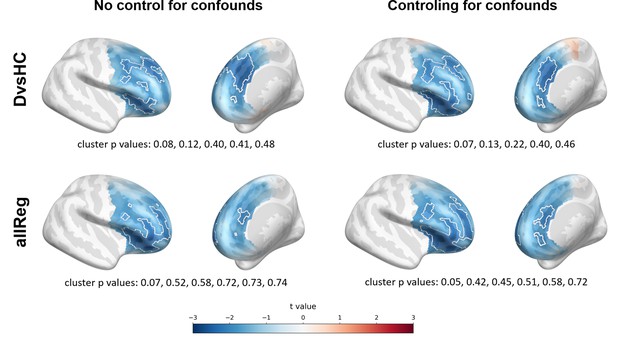
Selected results for aggregated source space analyses showing spatial t value maps for respective contrasts.
Cluster limits are marked with white outlines, and corresponding cluster p-values are shown below each panel. Color bar at the bottom presents color coding for the t values.

Selected results for aggregated source space analyses showing spatial t value maps for respective contrasts.
Cluster limits are marked with white outlines, and corresponding cluster p-values are shown below each panel. Color bar at the bottom presents color coding for the t values.
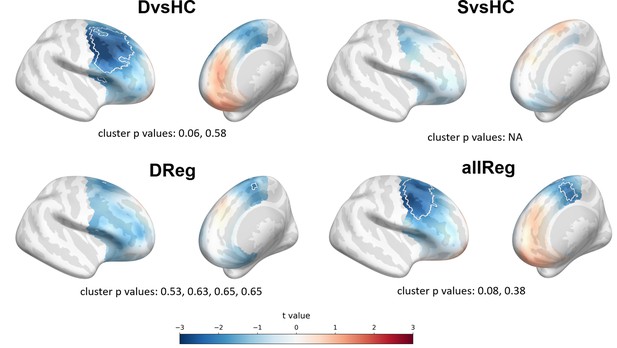
Results for interaction analyses (gender × contrast) for aggregated studies in source space.
In group contrasts (DvsHC and SvsHC) the interaction term is female * diagnosed; in linear contrasts (DReg, allReg) it is female * depression score. Cluster limits are marked with white outlines, and corresponding cluster p-values are shown below each panel. Color bar at the bottom presents color coding for the t values.
Tables
Results for all channel-pair analyses.
Each row represents two channel-pair results for a given contrast, study, and space combination; uncorrected for multiple comparisons. Electrode placement for each study is shown in Figure 1C. (N: number of participants included in given contrast; ES: effect size; Cohen’s d for group comparison and Pearson’s r for regression; CI: bootstrap 95% confidence interval for the effect size).
| No. | Contrast | Study | Space | N | Selected electrodes without correction | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Pair 1 (F3–F4) | Pair 2 (F7–F8) | |||||||||||
| t | p | ES | CI | t | p | ES | CI | |||||
| 1 | DvsHC | I | avg | 29 vs 22 | −2.073 | 0.043 | −0.573 | [−1.135,–0.024] | −0.365 | 0.717 | −0.101 | [−0.644, 0.465] |
| 2 | DvsHC | I | csd | 29 vs 22 | 0.132 | 0.896 | 0.038 | [−0.550, 0.608] | 0.553 | 0.583 | 0.153 | [−0.415, 0.689] |
| 3 | DvsHC | III | avg | 27 vs 21 | 0.904 | 0.371 | 0.247 | [−0.316, 0.689] | −0.536 | 0.595 | −0.145 | [−0.760, 0.452] |
| 4 | DvsHC | III | csd | 27 vs 21 | 0.849 | 0.401 | 0.226 | [−0.307, 0.721] | −0.129 | 0.898 | −0.035 | [−0.590, 0.536] |
| 5 | DvsHC | IV | avg | 22 vs 72 | 0.450 | 0.654 | 0.094 | [−0.310, 0.510] | 0.277 | 0.783 | 0.059 | [−0.413, 0.463] |
| 6 | DvsHC | IV | csd | 22 vs 72 | 0.767 | 0.449 | 0.212 | [−0.287, 0.771] | −1.396 | 0.172 | −0.345 | [−0.808, 0.167] |
| 7 | DvsHC | V | avg | 24 vs 29 | 2.823 | 0.007 | 0.743 | [0.218, 1.255] | 1.927 | 0.061 | 0.501 | [−0.023, 0.998] |
| 8 | DvsHC | V | csd | 24 vs 29 | 0.748 | 0.458 | 0.208 | [−0.380, 0.775] | −0.727 | 0.471 | −0.202 | [−0.766, 0.370] |
| 9 | SvsHC | II | avg | 23 vs 28 | 0.201 | 0.841 | 0.056 | [−0.502, 0.614] | −0.662 | 0.511 | −0.179 | [−0.780, 0.381] |
| 10 | SvsHC | II | csd | 23 vs 28 | 1.144 | 0.258 | 0.318 | [−0.221, 0.827] | −0.199 | 0.843 | −0.054 | [−0.611, 0.508] |
| 11 | SvsHC | III | avg | 33 vs 21 | −0.852 | 0.398 | −0.209 | [−0.640, 0.306] | −1.328 | 0.190 | −0.332 | [−0.804, 0.216] |
| 12 | SvsHC | III | csd | 33 vs 21 | −1.181 | 0.244 | −0.280 | [−0.730, 0.219] | −1.094 | 0.280 | −0.302 | [−0.798, 0.219] |
| 13 | SvsHC | IV | avg | 21 vs 72 | 1.359 | 0.184 | 0.346 | [−0.198, 0.816] | 1.247 | 0.219 | 0.254 | [−0.138, 0.646] |
| 14 | SvsHC | IV | csd | 21 vs 72 | 0.558 | 0.581 | 0.147 | [−0.393, 0.655] | 0.141 | 0.889 | 0.035 | [−0.388, 0.605] |
| 15 | allReg | I | avg | 54 | −1.138 | 0.260 | −0.156 | [−0.397, 0.088] | −0.180 | 0.858 | −0.025 | [−0.394, 0.258] |
| 16 | allReg | I | csd | 54 | −0.540 | 0.591 | −0.075 | [−0.309, 0.167] | 0.077 | 0.939 | 0.011 | [−0.335, 0.280] |
| 17 | allReg | III | avg | 91 | 0.545 | 0.587 | 0.058 | [−0.105, 0.263] | −0.781 | 0.437 | −0.083 | [−0.271, 0.101] |
| 18 | allReg | III | csd | 91 | 0.209 | 0.835 | 0.022 | [−0.169, 0.207] | 0.422 | 0.674 | 0.045 | [−0.168, 0.225] |
| 19 | allReg | IV | avg | 117 | 1.138 | 0.258 | 0.106 | [−0.076, 0.264] | 0.675 | 0.501 | 0.063 | [−0.111, 0.212] |
| 20 | allReg | IV | csd | 117 | 1.307 | 0.194 | 0.121 | [−0.096, 0.312] | −1.024 | 0.308 | −0.095 | [−0.265, 0.093] |
| 21 | allReg | V | avg | 53 | 2.489 | 0.016 | 0.329 | [0.106, 0.517] | 1.463 | 0.150 | 0.201 | [−0.023, 0.409] |
| 22 | allReg | V | csd | 53 | 0.857 | 0.395 | 0.119 | [−0.127, 0.361] | −0.220 | 0.827 | −0.031 | [−0.262, 0.210] |
| 23 | DReg | I | avg | 29 | 0.980 | 0.336 | 0.185 | [−0.212, 0.531] | 0.320 | 0.751 | 0.061 | [−0.470, 0.522] |
| 24 | DReg | I | csd | 29 | −1.303 | 0.204 | −0.243 | [−0.540, 0.091] | −0.504 | 0.618 | −0.097 | [−0.501, 0.337] |
| 25 | DReg | III | avg | 27 | 0.278 | 0.784 | 0.055 | [−0.268, 0.426] | 0.055 | 0.957 | 0.011 | [−0.273, 0.238] |
| 26 | DReg | III | csd | 27 | 0.407 | 0.688 | 0.081 | [−0.254, 0.413] | 1.347 | 0.190 | 0.260 | [−0.047, 0.498] |
| 27 | DReg | IV | avg | 22 | 1.754 | 0.095 | 0.365 | [0.012, 0.643] | −0.114 | 0.910 | −0.026 | [−0.459, 0.427] |
| 28 | DReg | IV | csd | 22 | 1.938 | 0.067 | 0.398 | [−0.043, 0.632] | −0.415 | 0.683 | −0.092 | [−0.519, 0.351] |
| 29 | DReg | V | avg | 24 | 1.497 | 0.149 | 0.304 | [0.045, 0.563] | −0.367 | 0.717 | −0.078 | [−0.444, 0.285] |
| 30 | DReg | V | csd | 24 | 0.789 | 0.438 | 0.166 | [−0.264, 0.501] | 0.635 | 0.532 | 0.134 | [−0.320, 0.615] |
-
DvsHC – diagnosed and healthy controls; SvsHC – sub-clinical and healthy controls; allReg – linear regression for all subjects together; DReg – linear regression for only diagnosed subjects; avg – average reference; csd – current source density.
Results for all channel-pair analyses corrected for confounds.
Each row represents two channel-pair results for a given contrast, study, and space combination; uncorrected for multiple comparisons. Electrode placement for each study is shown in Figure 1C. (N: number of participants included in given contrast; ES: effect size; Cohen’s d for group comparison and Pearson’s r for regression; CI: bootstrap 95% confidence interval for the effect size).
| No. | Contrast | Study | Space | N | Selected electrodes corrected for confounds | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Pair 1 (F3–F4) | Pair 2 (F7–F8) | |||||||||||
| t | p | ES | CI | t | p | ES | CI | |||||
| 1 | DvsHC | I | avg | 29 vs 22 | −2.679 | 0.010 | −0.789 | [−1.413,–0.218] | −0.782 | 0.438 | −0.230 | [−0.691, 0.288] |
| 2 | DvsHC | I | csd | 29 vs 22 | 0.269 | 0.789 | 0.079 | [−0.501, 0.681] | 0.305 | 0.762 | 0.090 | [−0.404, 0.622] |
| 3 | DvsHC | III | avg | 27 vs 21 | 0.204 | 0.839 | 0.064 | [−0.642, 0.684] | −1.161 | 0.252 | −0.366 | [−0.994, 0.310] |
| 4 | DvsHC | III | csd | 27 vs 21 | −0.162 | 0.872 | −0.051 | [−0.691, 0.547] | −0.703 | 0.486 | −0.221 | [−0.998, 0.558] |
| 5 | DvsHC | IV | avg | 22 vs 71 | 0.310 | 0.757 | 0.077 | [−0.338, 0.504] | 0.085 | 0.933 | 0.021 | [−0.412, 0.450] |
| 6 | DvsHC | IV | csd | 22 vs 71 | 0.978 | 0.331 | 0.244 | [−0.250, 0.816] | −1.484 | 0.141 | −0.370 | [−0.842, 0.135] |
| 7 | DvsHC | V | avg | 24 vs 29 | 1.862 | 0.069 | 0.540 | [0.033, 1.044] | 1.817 | 0.075 | 0.527 | [−0.008, 1.101] |
| 8 | DvsHC | V | csd | 24 vs 29 | 0.518 | 0.607 | 0.150 | [−0.415, 0.742] | −0.722 | 0.474 | −0.209 | [−0.689, 0.235] |
| 9 | SvsHC | II | avg | 23 vs 28 | 0.351 | 0.728 | 0.105 | [−0.444, 0.746] | −0.654 | 0.516 | −0.196 | [−0.883, 0.450] |
| 10 | SvsHC | II | csd | 23 vs 28 | 1.293 | 0.203 | 0.387 | [−0.152, 0.943] | 0.035 | 0.972 | 0.010 | [−0.653, 0.532] |
| 11 | SvsHC | III | avg | 33 vs 21 | −1.169 | 0.248 | −0.350 | [−0.946, 0.285] | −1.768 | 0.084 | −0.529 | [−1.071, 0.029] |
| 12 | SvsHC | III | csd | 33 vs 21 | −1.382 | 0.173 | −0.414 | [−1.086, 0.160] | −1.197 | 0.237 | −0.358 | [−0.997, 0.212] |
| 13 | SvsHC | IV | avg | 21 vs 71 | 1.058 | 0.293 | 0.269 | [−0.232, 0.776] | 0.466 | 0.642 | 0.118 | [−0.293, 0.515] |
| 14 | SvsHC | IV | csd | 21 vs 71 | 0.739 | 0.462 | 0.188 | [−0.302, 0.649] | −0.263 | 0.793 | −0.067 | [−0.524, 0.483] |
| 15 | allReg | I | avg | 54 | −1.352 | 0.182 | −0.188 | [−0.440, 0.078] | −0.349 | 0.728 | −0.049 | [−0.369, 0.240] |
| 16 | allReg | I | csd | 54 | −0.431 | 0.668 | −0.061 | [−0.313, 0.187] | −0.019 | 0.985 | −0.003 | [−0.335, 0.304] |
| 17 | allReg | III | avg | 91 | 0.233 | 0.816 | 0.025 | [−0.156, 0.263] | −1.170 | 0.245 | −0.127 | [−0.313, 0.092] |
| 18 | allReg | III | csd | 91 | −0.005 | 0.996 | −0.001 | [−0.190, 0.204] | −0.060 | 0.953 | −0.007 | [−0.238, 0.224] |
| 19 | allReg | IV | avg | 116 | 0.904 | 0.368 | 0.085 | [−0.088, 0.237] | 0.355 | 0.723 | 0.034 | [−0.139, 0.197] |
| 20 | allReg | IV | csd | 116 | 1.461 | 0.147 | 0.137 | [−0.064, 0.316] | −1.300 | 0.196 | −0.122 | [−0.296, 0.059] |
| 21 | allReg | V | avg | 53 | 1.685 | 0.099 | 0.236 | [−0.014, 0.449] | 1.544 | 0.129 | 0.218 | [−0.034, 0.433] |
| 22 | allReg | V | csd | 53 | 0.769 | 0.446 | 0.110 | [−0.148, 0.339] | −0.058 | 0.954 | −0.008 | [−0.214, 0.185] |
| 23 | DReg | I | avg | 29 | 0.767 | 0.450 | 0.152 | [−0.324, 0.503] | 0.265 | 0.793 | 0.053 | [−0.452, 0.564] |
| 24 | DReg | I | csd | 29 | −1.273 | 0.215 | −0.247 | [−0.588, 0.156] | −0.506 | 0.617 | −0.101 | [−0.504, 0.437] |
| 25 | DReg | III | avg | 27 | −0.054 | 0.958 | −0.012 | [−0.434, 0.485] | −0.079 | 0.937 | −0.018 | [−0.425, 0.346] |
| 26 | DReg | III | csd | 27 | 0.055 | 0.957 | 0.012 | [−0.407, 0.408] | 1.375 | 0.184 | 0.294 | [−0.096, 0.614] |
| 27 | DReg | IV | avg | 22 | 1.979 | 0.063 | 0.423 | [−0.001, 0.719] | −0.062 | 0.951 | −0.015 | [−0.409, 0.473] |
| 28 | DReg | IV | csd | 22 | 1.761 | 0.095 | 0.383 | [0.004, 0.674] | −0.269 | 0.791 | −0.063 | [−0.538, 0.418] |
| 29 | DReg | V | avg | 24 | 0.926 | 0.366 | 0.208 | [−0.209, 0.565] | −0.707 | 0.488 | −0.160 | [−0.571, 0.278] |
| 30 | DReg | V | csd | 24 | 0.723 | 0.478 | 0.164 | [−0.345, 0.554] | 0.298 | 0.769 | 0.068 | [−0.569, 0.735] |
-
DvsHC – diagnosed and healthy controls; SvsHC – sub-clinical and healthy controls; allReg – linear regression for all subjects together; DReg – linear regression for only diagnosed subjects; avg – average reference; csd – current source density.
Results for cluster-based permutation test on frontal asymmetry space.
Each row represents cluster-based results for a given contrast, study, and space combination (N: number of participants included in given contrast; min t, max t: lowest and highest t value in the search space, respectively; n significant points: total number of significant points in the search space before cluster-based correction; n clusters: number of clusters found in given analysis; largest cluster size: number of channels participating in the cluster; largest cluster p: p-value for the largest cluster, NA means that no cluster was found in given analysis).
| No. | Contrast | Study | Space | N | Cluster-based permutation test on frontal asymmetry space | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| Min t | Max t | n significant points | n clusters | Largest cluster size | Largest cluster p | |||||
| 1 | DvsHC | I | avg | 29 vs 22 | −2.110 | 0.023 | 3 | 1 | 3 | 0.069 |
| 2 | DvsHC | I | csd | 29 vs 22 | −0.720 | 1.983 | 0 | 0 | NA | NA |
| 3 | DvsHC | III | avg | 27 vs 21 | −1.172 | 1.058 | 0 | 0 | NA | NA |
| 4 | DvsHC | III | csd | 27 vs 21 | −1.258 | 1.875 | 0 | 0 | NA | NA |
| 5 | DvsHC | IV | avg | 22 vs 72 | −0.751 | 2.069 | 1 | 1 | 1 | 0.345 |
| 6 | DvsHC | IV | csd | 22 vs 72 | −1.600 | 1.142 | 0 | 0 | NA | NA |
| 7 | DvsHC | V | avg | 24 vs 29 | −1.260 | 2.823 | 6 | 2 | 5 | 0.026 |
| 8 | DvsHC | V | csd | 24 vs 29 | −2.901 | 1.425 | 2 | 2 | 1 | 0.156 |
| 9 | SvsHC | II | avg | 23 vs 28 | −2.581 | 0.201 | 1 | 1 | 1 | 0.164z |
| 10 | SvsHC | II | csd | 23 vs 28 | −0.855 | 1.254 | 0 | 0 | NA | NA |
| 11 | SvsHC | III | avg | 33 vs 21 | −1.410 | 1.021 | 0 | 0 | NA | NA |
| 12 | SvsHC | III | csd | 33 vs 21 | −2.315 | 2.101 | 2 | 2 | 1 | 0.227 |
| 13 | SvsHC | IV | avg | 21 vs 72 | −1.356 | 2.760 | 2 | 1 | 2 | 0.052 |
| 14 | SvsHC | IV | csd | 21 vs 72 | −1.193 | 1.296 | 0 | 0 | NA | NA |
| 15 | allReg | I | avg | 54 | −1.519 | 0.022 | 0 | 0 | NA | NA |
| 16 | allReg | I | csd | 54 | −0.727 | 1.052 | 0 | 0 | NA | NA |
| 17 | allReg | III | avg | 91 | −1.207 | 0.906 | 0 | 0 | NA | NA |
| 18 | allReg | III | csd | 91 | −1.290 | 2.210 | 1 | 1 | 1 | 0.287 |
| 19 | allReg | IV | avg | 117 | −1.807 | 2.153 | 1 | 1 | 1 | 0.202 |
| 20 | allReg | IV | csd | 117 | −1.173 | 1.353 | 0 | 0 | NA | NA |
| 21 | allReg | V | avg | 53 | −1.001 | 2.489 | 3 | 1 | 3 | 0.077 |
| 22 | allReg | V | csd | 53 | −3.291 | 1.552 | 2 | 2 | 1 | 0.187 |
| 23 | DReg | I | avg | 29 | −0.159 | 1.552 | 0 | 0 | NA | NA |
| 24 | DReg | I | csd | 29 | −1.303 | 0.487 | 0 | 0 | NA | NA |
| 25 | DReg | III | avg | 27 | −2.041 | 0.976 | 0 | 0 | NA | NA |
| 26 | DReg | III | csd | 27 | −1.420 | 1.662 | 0 | 0 | NA | NA |
| 27 | DReg | IV | avg | 22 | −1.155 | 1.754 | 0 | 0 | NA | NA |
| 28 | DReg | IV | csd | 22 | −0.415 | 1.938 | 0 | 0 | NA | NA |
| 29 | DReg | V | avg | 24 | −1.482 | 1.497 | 0 | 0 | NA | NA |
| 30 | DReg | V | csd | 24 | −2.254 | 1.554 | 1 | 1 | 1 | 0.555 |
-
DvsHC – diagnosed and healthy controls; SvsHC – sub-clinical and healthy controls; allReg – linear regression for all subjects together; DReg – linear regression for only diagnosed subjects; avg – average reference; csd – current source density.
Results for cluster-based permutation test on frontal asymmetry space corrected for confounds.
Each row represents cluster-based results for a given contrast, study and space combination (N: number of participants included in given contrast; min t, max t: lowest and highest t value in the search space, respectively; n significant points: total number of significant points in the search space before cluster-based correction; n clusters: number of clusters found in given analysis; largest cluster size: number of channels participating in the cluster; largest cluster p: p-value for the largest cluster, NA means that no cluster was found in given analysis).
| No. | Contrast | Study | Space | N | Cluster-based permutation test on frontal asymmetry space corrected for confounds | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| Min t | Max t | n significant points | n clusters | Largest cluster size | Largest cluster p | |||||
| 1 | DvsHC | I | avg | 29 vs 22 | −2.679 | −0.149 | 5 | 1 | 5 | 0.023 |
| 2 | DvsHC | I | csd | 29 vs 22 | −1.020 | 1.591 | 0 | 0 | NA | NA |
| 3 | DvsHC | III | avg | 27 vs 21 | −1.506 | 1.361 | 0 | 0 | NA | NA |
| 4 | DvsHC | III | csd | 27 vs 21 | −1.310 | 1.378 | 0 | 0 | NA | NA |
| 5 | DvsHC | IV | avg | 22 vs 71 | −0.814 | 1.812 | 0 | 0 | NA | NA |
| 6 | DvsHC | IV | csd | 22 vs 71 | −1.812 | 0.978 | 0 | 0 | NA | NA |
| 7 | DvsHC | V | avg | 24 vs 29 | −1.618 | 2.341 | 3 | 3 | 1 | 0.327 |
| 8 | DvsHC | V | csd | 24 vs 29 | −3.017 | 0.738 | 2 | 2 | 1 | 0.205 |
| 9 | SvsHC | II | avg | 23 vs 28 | −2.622 | 0.351 | 1 | 1 | 1 | 0.177 |
| 10 | SvsHC | II | csd | 23 vs 28 | −0.838 | 1.742 | 0 | 0 | NA | NA |
| 11 | SvsHC | III | avg | 33 vs 21 | −1.768 | 1.218 | 0 | 0 | NA | NA |
| 12 | SvsHC | III | csd | 33 vs 21 | −2.090 | 1.899 | 1 | 1 | 1 | 0.358 |
| 13 | SvsHC | IV | avg | 21 vs 71 | −1.584 | 1.490 | 0 | 0 | NA | NA |
| 14 | SvsHC | IV | csd | 21 vs 71 | −1.400 | 0.739 | 0 | 0 | NA | NA |
| 15 | allReg | I | avg | 54 | −1.726 | 0.039 | 0 | 0 | NA | NA |
| 16 | allReg | I | csd | 54 | −0.816 | 0.987 | 0 | 0 | NA | NA |
| 17 | allReg | III | avg | 91 | −1.231 | 0.722 | 0 | 0 | NA | NA |
| 18 | allReg | III | csd | 91 | −1.540 | 2.119 | 1 | 1 | 1 | 0.335 |
| 19 | allReg | IV | avg | 116 | −1.804 | 1.852 | 0 | 0 | NA | NA |
| 20 | allReg | IV | csd | 116 | −1.300 | 1.461 | 0 | 0 | NA | NA |
| 21 | allReg | V | avg | 53 | −1.286 | 2.350 | 1 | 1 | 1 | 0.324 |
| 22 | allReg | V | csd | 53 | −3.541 | 0.832 | 2 | 2 | 1 | 0.173 |
| 23 | DReg | I | avg | 29 | −0.244 | 1.516 | 0 | 0 | NA | NA |
| 24 | DReg | I | csd | 29 | −1.291 | 0.565 | 0 | 0 | NA | NA |
| 25 | DReg | III | avg | 27 | −1.700 | 0.852 | 0 | 0 | NA | NA |
| 26 | DReg | III | csd | 27 | −1.311 | 1.574 | 0 | 0 | NA | NA |
| 27 | DReg | IV | avg | 22 | −1.173 | 2.728 | 1 | 1 | 1 | 0.121 |
| 28 | DReg | IV | csd | 22 | −0.269 | 2.235 | 1 | 1 | 1 | 0.260 |
| 29 | DReg | V | avg | 24 | −1.593 | 0.926 | 0 | 0 | NA | NA |
| 30 | DReg | V | csd | 24 | −2.075 | 2.586 | 2 | 2 | 1 | 0.376 |
-
DvsHC – diagnosed and healthy controls; SvsHC – sub-clinical and healthy controls; allReg – linear regression for all subjects together; DReg – linear regression for only diagnosed subjects; avg – average reference; csd – current source density.
Results for all cluster-based analyses on standardized data.
Each row represents cluster-based results for given contrast, study, and space (N: number of participants included in given contrast). Results for cluster-based permutation test on frontal asymmetry space (min t, max t: lowest and highest t value in the search space, respectively; n significant points: total number of significant points in the search space before cluster-based correction; n clusters: number of clusters found in given analysis; largest cluster size: number of channels by frequency points participating in the cluster; largest cluster p: p-value for the largest cluster, NA means that no cluster was found in given analysis).
| No. | Contrast | Study | Space | N | Cluster-based analyses on standardized data | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| Min t | Max t | n significant points | n clusters | Largest cluster size | Largest cluster p | |||||
| 1 | DvsHC | I | avg | 29 vs 22 | −1.453 | 1.434 | 0 | 0 | NA | NA |
| 2 | DvsHC | I | csd | 29 vs 22 | −2.227 | 1.879 | 3 | 2 | 2 | 0.718 |
| 3 | DvsHC | III | avg | 27 vs 21 | −1.999 | 2.326 | 2 | 1 | 2 | 0.468 |
| 4 | DvsHC | III | csd | 27 vs 21 | −2.917 | 3.882 | 23 | 3 | 16 | 0.063 |
| 5 | DvsHC | IV | avg | 22 vs 72 | −4.256 | 3.053 | 129 | 2 | 66 | 0.007 |
| 6 | DvsHC | IV | csd | 22 vs 72 | −3.180 | 2.582 | 22 | 7 | 6 | 0.259 |
| 7 | DvsHC | V | avg | 24 vs 29 | −2.516 | 2.267 | 21 | 3 | 15 | 0.223 |
| 8 | DvsHC | V | csd | 24 vs 29 | −2.703 | 2.420 | 10 | 6 | 3 | 0.723 |
| 9 | SvsHC | II | avg | 23 vs 28 | −1.685 | 2.185 | 1 | 1 | 1 | 0.774 |
| 10 | SvsHC | II | csd | 23 vs 28 | −2.059 | 1.714 | 1 | 1 | 1 | 0.941 |
| 11 | SvsHC | III | avg | 33 vs 21 | −1.906 | 1.946 | 0 | 0 | NA | NA |
| 12 | SvsHC | III | csd | 33 vs 21 | −2.284 | 3.091 | 7 | 4 | 3 | 0.577 |
| 13 | SvsHC | IV | avg | 21 vs 72 | −2.319 | 2.519 | 19 | 5 | 5 | 0.306 |
| 14 | SvsHC | IV | csd | 21 vs 72 | −2.950 | 3.130 | 25 | 4 | 11 | 0.092 |
| 15 | allReg | I | avg | 54 | −1.295 | 1.450 | 0 | 0 | NA | NA |
| 16 | allReg | I | csd | 54 | −2.082 | 1.899 | 1 | 1 | 1 | 1.000 |
| 17 | allReg | III | avg | 91 | −1.814 | 1.949 | 0 | 0 | NA | NA |
| 18 | allReg | III | csd | 91 | −2.626 | 2.547 | 14 | 4 | 6 | 0.692 |
| 19 | allReg | IV | avg | 117 | −3.406 | 2.701 | 75 | 4 | 40 | 0.079 |
| 20 | allReg | IV | csd | 117 | −2.644 | 2.918 | 32 | 5 | 13 | 0.179 |
| 21 | allReg | V | avg | 53 | −2.752 | 2.321 | 22 | 6 | 13 | 0.477 |
| 22 | allReg | V | csd | 53 | −3.733 | 2.023 | 15 | 6 | 5 | 0.891 |
| 23 | DReg | I | avg | 29 | −2.011 | 1.936 | 0 | 0 | NA | NA |
| 24 | DReg | I | csd | 29 | −2.721 | 2.093 | 8 | 5 | 3 | 1.000 |
| 25 | DReg | III | avg | 27 | −4.167 | 3.824 | 89 | 2 | 53 | 0.025 |
| 26 | DReg | III | csd | 27 | −3.525 | 3.285 | 34 | 4 | 17 | 0.105 |
| 27 | DReg | IV | avg | 22 | −1.802 | 2.213 | 1 | 1 | 1 | 0.888 |
| 28 | DReg | IV | csd | 22 | −2.219 | 2.890 | 12 | 4 | 9 | 0.316 |
| 29 | DReg | V | avg | 24 | −2.926 | 4.155 | 58 | 7 | 18 | 0.354 |
| 30 | DReg | V | csd | 24 | −3.551 | 3.084 | 46 | 9 | 14 | 0.387 |
-
DvsHC – diagnosed and healthy controls; SvsHC – sub-clinical and healthy controls; allReg – linear regression for all subjects together; DReg – linear regression for only diagnosed subjects; avg – average reference; csd – current source density.
Results for all cluster-based analyses on standardized data corrected for confounds.
Each row represents cluster-based results for given contrast, study, and space (N: number of participants included in given contrast). Results for cluster-based permutation test on frontal asymmetry space (min t, max t: lowest and highest t value in the search space, respectively; n significant points: total number of significant points in the search space before cluster-based correction; n clusters: number of clusters found in given analysis; largest cluster size: number of channels by frequency points participating in the cluster; largest cluster p: p-value for the largest cluster, NA means that no cluster was found in given analysis).
| No. | Contrast | Study | Space | N | Cluster-based analyses on standardized data corrected for confounds | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| Min t | Max t | n significant points | n clusters | Largest cluster size | Largest cluster p | |||||
| 1 | DvsHC | I | avg | 29 vs 22 | −1.766 | 1.716 | 0 | 0 | NA | NA |
| 2 | DvsHC | I | csd | 29 vs 22 | −2.007 | 1.614 | 0 | 0 | NA | NA |
| 3 | DvsHC | III | avg | 27 vs 21 | −1.387 | 1.896 | 0 | 0 | NA | NA |
| 4 | DvsHC | III | csd | 27 vs 21 | −3.345 | 3.054 | 17 | 4 | 6 | 0.567 |
| 5 | DvsHC | IV | avg | 22 vs 71 | −4.295 | 3.304 | 143 | 2 | 72 | 0.006 |
| 6 | DvsHC | IV | csd | 22 vs 71 | −3.095 | 2.854 | 33 | 5 | 12 | 0.179 |
| 7 | DvsHC | V | avg | 24 vs 29 | −1.982 | 1.976 | 0 | 0 | NA | NA |
| 8 | DvsHC | V | csd | 24 vs 29 | −2.026 | 2.798 | 10 | 4 | 7 | 0.772 |
| 9 | SvsHC | II | avg | 23 vs 28 | −1.511 | 1.841 | 0 | 0 | NA | NA |
| 10 | SvsHC | II | csd | 23 vs 28 | −2.059 | 1.876 | 1 | 1 | 1 | 1.000 |
| 11 | SvsHC | III | avg | 33 vs 21 | −2.964 | 2.422 | 22 | 5 | 10 | 0.374 |
| 12 | SvsHC | III | csd | 33 vs 21 | −2.424 | 3.562 | 12 | 4 | 4 | 0.865 |
| 13 | SvsHC | IV | avg | 21 vs 71 | −2.667 | 2.858 | 34 | 3 | 31 | 0.113 |
| 14 | SvsHC | IV | csd | 21 vs 71 | −2.389 | 3.317 | 23 | 3 | 10 | 0.273 |
| 15 | allReg | I | avg | 54 | −1.404 | 1.614 | 0 | 0 | NA | NA |
| 16 | allReg | I | csd | 54 | −1.752 | 2.155 | 4 | 2 | 2 | 1.000 |
| 17 | allReg | III | avg | 91 | −2.119 | 1.963 | 5 | 1 | 5 | 0.612 |
| 18 | allReg | III | csd | 91 | −2.558 | 2.825 | 25 | 3 | 17 | 0.112 |
| 19 | allReg | IV | avg | 116 | −3.414 | 3.009 | 92 | 4 | 54 | 0.038 |
| 20 | allReg | IV | csd | 116 | −2.571 | 3.112 | 32 | 6 | 11 | 0.216 |
| 21 | allReg | V | avg | 53 | −1.942 | 2.057 | 1 | 1 | 1 | 1.000 |
| 22 | allReg | V | csd | 53 | −3.005 | 2.362 | 11 | 5 | 5 | 0.945 |
| 23 | DReg | I | avg | 29 | −1.902 | 1.990 | 0 | 0 | NA | NA |
| 24 | DReg | I | csd | 29 | −2.583 | 2.088 | 9 | 5 | 4 | 0.955 |
| 25 | DReg | III | avg | 27 | −3.938 | 3.530 | 70 | 4 | 36 | 0.073 |
| 26 | DReg | III | csd | 27 | −3.198 | 4.098 | 39 | 7 | 21 | 0.035 |
| 27 | DReg | IV | avg | 22 | −2.213 | 2.662 | 4 | 2 | 2 | 0.659 |
| 28 | DReg | IV | csd | 22 | −3.056 | 3.114 | 16 | 4 | 10 | 0.265 |
| 29 | DReg | V | avg | 24 | −3.200 | 4.246 | 84 | 7 | 27 | 0.266 |
| 30 | DReg | V | csd | 24 | −3.533 | 3.378 | 78 | 7 | 36 | 0.092 |
-
DvsHC – diagnosed and healthy controls; SvsHC – sub-clinical and healthy controls; allReg – linear regression for all subjects together; DReg – linear regression for only diagnosed subjects; avg – average reference; csd – current source density.
Results for all source level analyses.
Each row represents source level results for given contrast, study, and space (N: number of participants included in given contrast). Results for cluster-based permutation test on frontal asymmetry source space (min t, max t: lowest and highest t value in the search space, respectively; n significant points: total number of significant points in the search space before cluster-based correction; n clusters: number of clusters found in given analysis; largest cluster p: p-value for the largest cluster, NA means that no cluster was found in given analysis).
| No. | Contrast | Study | N | Source level analysis | |||||
|---|---|---|---|---|---|---|---|---|---|
| Min t | Max t | n significant points | n clusters | Largest cluster size | Largest cluster p | ||||
| 1 | DvsHC | I | 29 vs 22 | −1.906 | 2.404 | 2 | 1 | 2 | 0.489 |
| 2 | DvsHC | III | 27 vs 21 | −2.557 | 1.158 | 29 | 4 | 14 | 0.267 |
| 3 | DvsHC | IV | 22 vs 72 | −3.921 | 1.151 | 320 | 2 | 316 | 0.010 |
| 4 | DvsHC | V | 24 vs 29 | −2.612 | 0.943 | 24 | 1 | 24 | 0.207 |
| 5 | SvsHC | II | 23 vs 28 | −0.909 | 1.321 | 0 | 0 | NA | NA |
| 6 | SvsHC | III | 34 vs 21 | −1.809 | 1.341 | 0 | 0 | NA | NA |
| 7 | SvsHC | IV | 21 vs 72 | −2.339 | 0.679 | 11 | 1 | 11 | 0.340 |
| 8 | allReg | I | 54 | −2.319 | 1.549 | 20 | 1 | 20 | 0.367 |
| 9 | allReg | III | 92 | −2.232 | 0.290 | 21 | 2 | 20 | 0.343 |
| 10 | allReg | IV | 117 | −2.220 | 1.211 | 10 | 1 | 10 | 0.466 |
| 11 | allReg | V | 53 | −2.644 | 1.201 | 15 | 3 | 6 | 0.574 |
| 12 | DReg | I | 29 | −2.292 | 1.150 | 41 | 2 | 22 | 0.328 |
| 13 | DReg | III | 27 | −2.624 | −0.375 | 26 | 3 | 18 | 0.335 |
| 14 | DReg | IV | 22 | −1.216 | 2.064 | 0 | 0 | NA | NA |
| 15 | DReg | V | 24 | −2.068 | 1.758 | 0 | 0 | NA | NA |
-
DvsHC – diagnosed and healthy controls; SvsHC – sub-clinical and healthy controls; allReg – linear regression for all subjects together; DReg – linear regression for only diagnosed subjects.
Results for all source level analyses corrected for confounds.
Each row represents source level results for given contrast, study, and space (N: number of participants included in given contrast) Results for cluster-based permutation test on frontal asymmetry source space (min t, max t: lowest and highest t value in the search space, respectively; n significant points: total number of significant points in the search space before cluster-based correction; n clusters: number of clusters found in given analysis; largest cluster p: p-value for the largest cluster, NA means that no cluster was found in given analysis).
| No. | Contrast | Study | N | Source level analysis corrected for confounds | |||||
|---|---|---|---|---|---|---|---|---|---|
| Min t | Max t | n significant points | n clusters | Largest cluster size | Largest cluster p | ||||
| 1 | DvsHC | I | 29 vs 22 | −1.416 | 2.057 | 1 | 1 | 1 | 0.708 |
| 2 | DvsHC | III | 27 vs 21 | −2.161 | 1.704 | 3 | 2 | 2 | 0.555 |
| 3 | DvsHC | IV | 22 vs 71 | −3.685 | 1.066 | 365 | 2 | 346 | 0.011 |
| 4 | DvsHC | V | 24 vs 29 | −2.630 | 0.287 | 64 | 3 | 46 | 0.231 |
| 5 | SvsHC | II | 23 vs 28 | −0.814 | 1.911 | 0 | 0 | NA | NA |
| 6 | SvsHC | III | 34 vs 21 | −1.770 | 1.526 | 0 | 0 | NA | NA |
| 7 | SvsHC | IV | 21 vs 71 | −2.732 | −0.083 | 58 | 1 | 58 | 0.192 |
| 8 | allReg | I | 54 | −1.846 | 1.143 | 0 | 0 | NA | NA |
| 9 | allReg | III | 92 | −2.195 | 0.698 | 15 | 1 | 15 | 0.391 |
| 10 | allReg | IV | 116 | −2.313 | 0.910 | 44 | 4 | 19 | 0.375 |
| 11 | allReg | V | 53 | −2.901 | 0.317 | 79 | 4 | 36 | 0.264 |
| 12 | DReg | I | 29 | −2.164 | 1.010 | 15 | 2 | 11 | 0.428 |
| 13 | DReg | III | 27 | −2.991 | 0.175 | 66 | 3 | 63 | 0.186 |
| 14 | DReg | IV | 22 | −1.397 | 1.818 | 0 | 0 | NA | NA |
| 15 | DReg | V | 24 | −2.893 | 1.784 | 22 | 1 | 22 | 0.352 |
-
DvsHC – diagnosed and healthy controls; SvsHC – sub-clinical and healthy controls; allReg – linear regression for all subjects together; DReg – linear regression for only diagnosed subjects; nonDReg – linear regression for only the non-diagnosed subjects.
Results for analyses on data aggregated across studies: tests on frontal asymmetry on selected channel pairs.
Each row represents a given contrast × reference × control for confounds combination. N: number of participants included in given contrast, control for confounds: whether the FAA data was residualized with respect to confounding variables (age, gender and education); ES: effect size, measured as Cohen’s d for DvsHC and SvsHC contrasts and Spearman’s r for Dreg and allReg contrasts; CI: bootstrap confidence interval for the effect size; BF01: Bayes Factor for the null hypothesis.
| No. | Contrast | Space | N | Control for confounds | Aggregated channel pair analyses | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Pair 1 (F3–F4) | Pair 2 (F7–F8) | |||||||||||
| ES | CI | BF01 | ES | CI | BF01 | |||||||
| 1 | DvsHC | avg | 102 vs 144 | − | 0.147 | [−0.111, 0.396] | 3.831 | 0.098 | [−0.161, 0.338] | 5.405 | ||
| 2 | DvsHC | avg | 102 vs 143 | + | −0.011 | [−0.264, 0.244] | 7.042 | −0.006 | [−0.266, 0.240] | 7.042 | ||
| 3 | DvsHC | csd | 102 vs 144 | − | 0.188 | [−0.069, 0.449] | 2.597 | −0.100 | [−0.354, 0.164] | 5.319 | ||
| 4 | DvsHC | csd | 102 vs 143 | + | 0.103 | [−0.159, 0.362] | 5.236 | −0.164 | [−0.417, 0.108] | 3.300 | ||
| 5 | SvsHC | avg | 77 vs 121 | − | 0.065 | [−0.236, 0.359] | 5.780 | −0.025 | [−0.320, 0.240] | 6.250 | ||
| 6 | SvsHC | avg | 77 vs 120 | + | 0.024 | [−0.269, 0.315] | 6.211 | −0.144 | [−0.447, 0.140] | 4.016 | ||
| 7 | SvsHC | csd | 77 vs 121 | − | 0.053 | [−0.223, 0.355] | 5.952 | −0.108 | [−0.372, 0.179] | 4.878 | ||
| 8 | SvsHC | csd | 77 vs 120 | + | 0.053 | [−0.222, 0.350] | 5.952 | −0.110 | [−0.389, 0.179] | 4.831 | ||
| 9 | DReg | avg | 102 | − | 0.188 | [0.006, 0.354] | 1.387 | 0.007 | [−0.186, 0.188] | 8.065 | ||
| 10 | DReg | avg | 102 | + | 0.175 | [−0.018, 0.348] | 1.742 | −0.029 | [−0.211, 0.155] | 7.752 | ||
| 11 | DReg | csd | 102 | − | 0.099 | [−0.072, 0.260] | 4.975 | 0.065 | [−0.136, 0.252] | 6.579 | ||
| 12 | DReg | csd | 102 | + | 0.051 | [−0.131, 0.216] | 7.092 | 0.048 | [−0.157, 0.255] | 7.194 | ||
| 13 | allReg | avg | 315 | − | 0.085 | [−0.017, 0.184] | 4.651 | 0.029 | [−0.080, 0.126] | 12.5 | ||
| 14 | allReg | avg | 314 | + | 0.041 | [−0.063, 0.143] | 10.87 | 0.001 | [−0.120, 0.104] | 14.085 | ||
| 15 | allReg | csd | 315 | − | 0.059 | [−0.054, 0.166] | 8.333 | −0.026 | [−0.141, 0.082] | 12.821 | ||
| 16 | allReg | csd | 314 | + | 0.055 | [−0.056, 0.168] | 8.772 | −0.048 | [−0.158, 0.059] | 9.901 | ||
-
DvsHC – diagnosed and healthy controls; SvsHC – sub-clinical and healthy controls; allReg – linear regression for all subjects together; DReg – linear regression for only diagnosed subjects; avg – average reference; csd – current source density.
Results for analyses on data aggregated across studies, cluster-based permutation tests on frontal asymmetry in the source space.
Each row represents the result for given contrast × control for confounds combination. N: number of participants included in given contrast, control for confounds: whether the FAA data was residualized with respect to confounding variables (age, gender and education), min t, max t: lowest and highest t value in the search space, respectively; n significant points: total number of significant points in the search space before cluster-based correction; n clusters: number of clusters found in given analysis; largest cluster p: p-value for the largest cluster, NA means that no cluster was found in given analysis.
| No. | Contrast | N | Control for confounds | Aggregated source level analyses | |||||
|---|---|---|---|---|---|---|---|---|---|
| Min t | Max t | n significant points | n clusters | Largest cluster size | Largest cluster p | ||||
| 1 | DvsHC | 102 vs 144 | − | −2.719 | 0.956 | 178 | 5 | 100 | 0.077 |
| 2 | DvsHC | 102 vs 143 | + | −2.846 | 0.908 | 211 | 5 | 116 | 0.066 |
| 3 | SvsHC | 77 vs 121 | − | −1.831 | −0.11 | 0 | 0 | NA | NA |
| 4 | SvsHC | 77 vs 120 | + | −1.871 | 0.182 | 0 | 0 | NA | NA |
| 5 | DReg | 102 | − | −2.387 | 1.376 | 31 | 3 | 25 | 0.328 |
| 6 | DReg | 102 | + | −3.095 | 1.354 | 112 | 3 | 68 | 0.155 |
| 7 | allReg | 315 | − | −3.044 | 0.411 | 174 | 6 | 160 | 0.067 |
| 8 | allReg | 314 | + | −3.107 | 0.282 | 235 | 6 | 195 | 0.054 |
-
DvsHC – diagnosed and healthy controls; SvsHC – sub-clinical and healthy controls; allReg – linear regression for all subjects together; DReg – linear regression for only diagnosed subjects.
Number of significant results compatible with the traditional FAA effect partitioned into analyses and contrasts.
If one or more such results have been found for a given cell, then the p-value for the binomial test is also shown.
| Number of significant results congruent with the FAA effect | |||||
|---|---|---|---|---|---|
| Channel pairs N = 120 | Cluster correction N = 60 | Source space N = 30 | |||
| DvsHC | 2/32, p=0.48 | 1/16, p=0.56 | 2/8, p=0.057 | ||
| SvsHC | 0/24 | 0/12 | 0/6 | ||
| DReg | 0/32 | 0/16 | 0/8 | ||
| allReg | 0/32 | 0/16 | 0/8 | ||
Descriptive statistics for each study presented in the article (N – number of participants, M – mean score, SD – standard deviation, BDI-I and BDI-II – Beck Depression Inventory I and II, PHQ-9 – Patient Health Questionnaire-9).
| Study I (N=51) | ||||
|---|---|---|---|---|
| Diagnosed | Healthy Controls, BDI-I ≤ 5 | Subclinical | Unclassified | |
| N | 29 | 22 | - | - |
| Age | M = 27.66, SD = 7.13 | M = 23.68, SD = 2.83 | - | - |
| 19 - 47 range | 20 - 33 range | - | - | |
| Gender | 9 male, 20 female | 11 male, 11 female | - | - |
| BDI-I score | M = 20.93, SD = 8.21 | M = 2.00, SD = 1.48 | - | - |
| Study II (N=76) | ||||
| Undiagnosed (N=76) | ||||
| Diagnosed | Healthy Controls, BDI-I ≤ 5 | Subclinical, BDI-I ≥ 10 | Unclassified, 5 < BDI-I < 10 | |
| N | - | 28 | 25 | 23 |
| Age | - | M = 25.32, SD = 6.46 | M = 24.44, SD = 5.08 | M = 25.22, SD = 6.78 |
| - | 18 - 43 range | 19 - 38 range | 18 - 40 range | |
| Gender | - | 8 males, 20 females | 4 males, 21 females | 9 males, 14 females |
| BDI-I score | - | M = 2.29, SD = 1.72 | M = 17.56, SD = 8.13 | M = 7.91, SD = 1.16 |
| Study III (N=91) | ||||
| Diagnosed | Undiagnosed (N=64) | |||
| Healthy Controls, BDI-II ≤ 5 | Subclinical, BDI-II ≥ 10 | Unclassified, 5 < BDI-II < 10 | ||
| N | 27 | 21 | 34 | 9 |
| Age | M = 27.19, SD = 7.23 | M = 24.29, SD = 4.99 | M = 25.06, SD = 6.58 | M = 26.78, SD = 8.74 |
| 19 - 42 range | 19 - 41 range | 18 - 44 range | 22 - 49 range | |
| Gender | 6 males, 21 females | 7 males, 14 females | 10 males, 24 females | 2 males, 7 females |
| BDI-II score | M = 34.26, SD = 9.18 | M = 2.24, SD = 1.70 | M = 24.06, SD = 10.08 | M = 6.78, SD = 0.97 |
| Study IV (N=117) | ||||
| Diagnosed | Undiagnosed (N=95) | |||
| Healthy Controls, BDI-II ≤ 5 | Subclinical, BDI-II ≥ 10 | Unclassified, 5 < BDI-II < 10 | ||
| N | 22 | 72 | 21 | 2 |
| (12 past MDD, 10 present MDD) | ||||
| Age | M = 18.91, SD = 1.34 | M = 19.00, SD = 1.23 | M = 18.43, SD = 0.81 | M = 18.00 |
| 18 - 24 range | 18 - 23 range | 18 - 21 range | ages 18, 18 | |
| Gender | 8 males, 14 females | 33 males, 39 females | 3 males, 18 females | 1 males, 1 females |
| BDI-II score | M = 21.82, SD = 5.70 | M = 1.60, SD = 1.48 | M = 22.95, SD = 4.25 | M = 6.50, SD = 0.71 |
| Study V (N=53) | ||||
| Diagnosed | Healthy Controls, PHQ-9 ≤ 5 | Subclinical | Unclassified | |
| N | 24 | 29 | - | - |
| Age | M = 30.88, SD = 10.37 | M = 31.45, SD = 9.15 | - | - |
| 16 - 52 range | 19 - 52 range | - | - | |
| Gender | 13 males, 11 females | 11 males, 13 females | - | - |
| PHQ-9 score | M = 18.33, SD = 3.50 | M = 2.66, SD = 1.80 | - | - |
Summary of the contrasts (DvsHC – diagnosed vs healthy controls; SvsHC – sub-clinical vs healthy controls; allReg – linear regression between FAA and depression questionnaire score for all subjects together; DReg – linear regression only for the diagnosed subjects) and confounds (age, gender, and education) used in each study.
| STUDY | ||||||
|---|---|---|---|---|---|---|
| I | II | III | IV | V | ||
| Contrast type | ||||||
| DvsHC | + | + | + | + | ||
| SvsHC | + | + | + | |||
| allReg | + | + | + | + | ||
| DReg | + | + | + | + | ||
| Control for confounds | ||||||
| gender | + | + | + | + | + | |
| age | + | + | + | + | + | |
| education | + | + | + | |||





