A sulfur-aromatic gate latch is essential for opening of the Orai1 channel pore
Figures
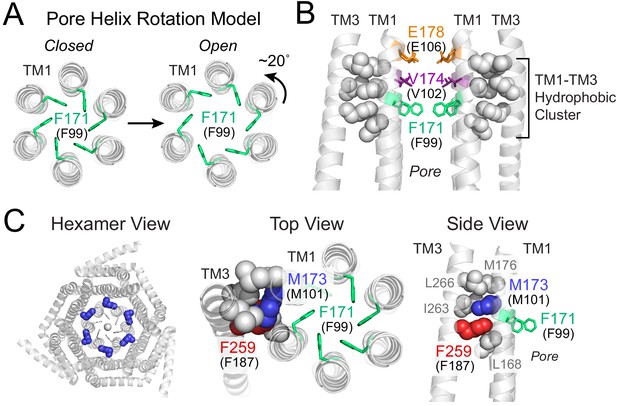
Gating model of Orai1 involving reorientation of the F99 gate.
(A) Schematic cartoon of the pore helix rotation model. Gating occurs through a modest rotation of TM1 and dilation of the outer pore (Yeung et al., 2018), which moves F171 (hOrai1 F99) away from the pore axis and decreases the free energy barrier in the hydrophobic stretch for ion permeation. For simplicity, TMs 2–4 are not shown. (B) A cluster of hydrophobic residues (shown as gray spheres) between the TM1 pore helices and the TM3 segment. The selectivity filter E178 (hOrai1 E106, orange) and hydrophobic gate residues V174 (hOrai1 V102, purple) and F171 (hOrai1 F99, green) in the pore are also shown. Four TM1 helices and two TM3 helices are shown for clarity. (C) Position of M101 within the hydrophobic cluster. F171 (hOrai1 F99) is also shown within the context of the hexameric channel and relative to the hydrophobic cluster residues. Residues M173 (hOrai1 M101) and F259 (hOrai1 F187) are depicted as blue and red spheres, respectively. The F171 gate is shown as green sticks. hOrai1 numbering is shown in parentheses.
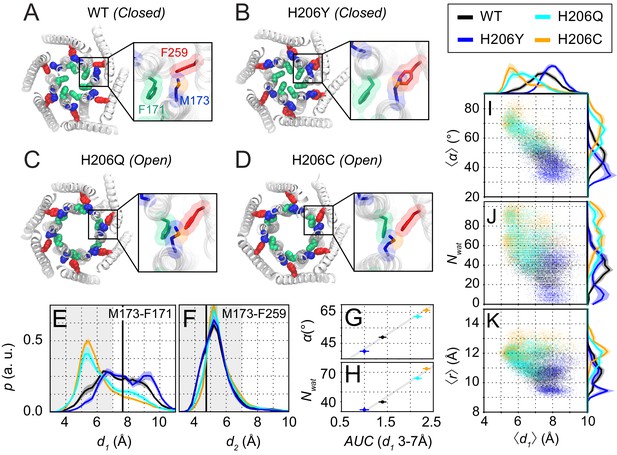
Molecular dynamics simulations reveal sulfur-aromatic interactions between M173 and F171 in open but not closed channel states.
(A–D) Snapshots of WT, H206Y (hOrai1 H134Y), H206Q (hOrai1 H134Q) and H206C (hOrai1 H134C) dOrai mutants from the MD simulation runs showing positions of key residues in the TM1 and TM3 helices. F171 (hOrai1 F99, green), M173 (hOrai1 M101, blue), and F259 (hOrai1 F187, red) are represented as spheres. Insets: Enlarged views of the F171-M173-F259 locus (hOrai1 F99-M101-F187) with sulfur atoms of M173 shown in orange. (E–F) Distribution of distances between M173 with F171 (d1) and M173 with F259 (d2), respectively. Distances were measured from the sulfur of the methionine and the center of mass of the phenylalanine ring. The mean and standard error of mean of M173-F171 distances over simulation repeats are 7.4 ± 0.1 Å for WT, 6.4 ± 0.2 Å for H206C, 6.6 ± 0.1 Å for H206Q, and 7.9 ± 0.1 Å for H206Y. Compared to closed WT and H206Y channels, constitutively active dOrai mutants H206Q/C show a greater proportion of M173-F171 interactions within the sulfur-aromatic interaction distance of 3–7 Å. By contrast, M173-F259 distance was constant across the different dOrai variants. Black solid lines in (E) and (F) represents distances observed in the closed dOrai crystal structure 4HKR. (G–H) Area under the curve (AUC) of M173-F171 interaction distances within 7 Å in panel E plotted against pore helix rotation (G) and number of pore waters (H) (Pearson correlation coefficient of 0.98 and 0.99, respectively). (I–K) Scatter plots of the hexameric average of M173-F171 distances plotted against corresponding average pore helix rotation (I), pore hydration (J) and average pore diameter (K). The distributions of the x and y parameters are displayed on the periphery of the graphs. In general, shorter M173-F171 distances were associated with increased pore helix rotation, pore hydration and pore diameter across the Orai variants. These dynamical channel properties are anti-correlated within simulations of each system (Pearson correlation coefficient of −0.65 (WT), –0.68 (H206C), −0.72 (H206Q), and −0.17 (H206Y) for panel I). Similarly, these properties were anti-correlated across our mutational landscape (Pearson correlation coefficients of −0.79 (panel I), –0.39 (panel J), and −0.56 (panel K)). All analysis was done after the equilibration window of 100 ns.
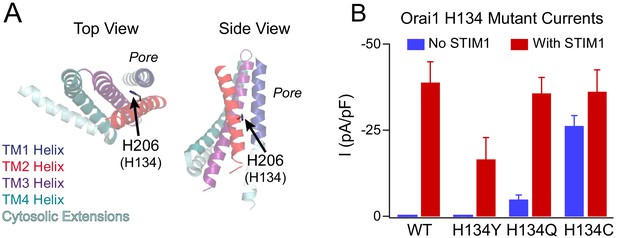
Phenotypes of the loss- and gain-of-function mutants at the Orai1 H134 gating locus.
(A) Side and top views of TM2 H206 residue (hOrai1 H134) in a monomer in the dOrai crystal structure (PDB ID: 4HKR). Numbering hOrai1 is shown in parentheses. (B) Current densities of WT and H134Y/Q/C Orai1 mutants without and with STIM1 co-expression (Yeung et al., 2018). The Orai1 H134Y variant conducts smaller, non-selective currents in the presence of STIM1 compared with WT channels, and H134Q/C Orai1 channels are constitutively active at baseline. The current amplitude of H134C in the absence of STIM1 is significantly larger than that of H134Q.
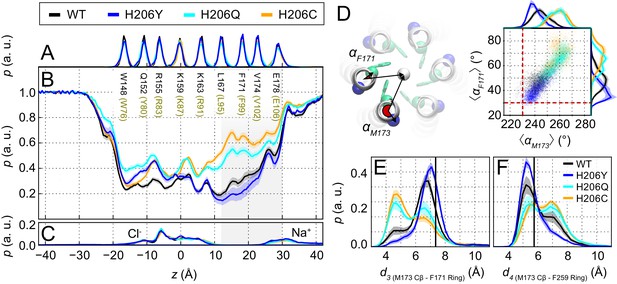
Pore hydration and M173 orientation in MD simulations H206 mutants.
(A) Relative probability distributions of the axial position of Cα atoms for all pore-lining residues (human Orai1 numbering in brown). Average distribution of water oxygen atoms (B) and Na+ and Cl− ions (C) along the pore axis. The water occupancies of H206Q/C mutants deviate from those of WT and H206Y dOrai most significantly in the hydrophobic stretch (V102-L95) of the pore. Data in panels (A–C) have been previously reported (Yeung et al., 2018). (D) Molecular rendering of the geometric positions involved in computing pore helix rotation angles αF171 and αM173. These angles are computed between the center of mass of all TM1 helices (white sphere), the center of mass of each TM1 helix, and the Cα atom of the residue of interest, shown for F171 (green) and M173 (blue), respectively. Scatterplot of the hexameric average of pore helix rotation computed at position F171 with respect to pore helix rotation at position M173. The relationship between these variables is highly correlated (Pearson correlation coefficient of 0.88). The pore rotation angle of F171 (30.6°) and M173 (230.1°) measured from the crystal structure of dOrai (4HKR) is shown as a red dashed lines. (E–F) Distribution of distances between M173 with F171 (d3) and M173 with F259 (d4), respectively. Distances were measured from the Cβ of the methionine and the center of mass of the phenylalanine ring. The mean and standard error of mean of M173-F171 distances over simulation repeats are 6.5 ± 0.1 Å for WT, 7.0 ± 0.1 Å for H206Y, 6.0 ± 0.1 Å for H206Q, and 5.8 ± 0.2 Å for H206C. In (F), M173-F259 distances over simulation repeats are 6.1 ± 0.1 Å for WT, 5.8 ± 0.1 Å for H206Y, 6.6 ± 0.1 Å for H206Q, and 6.6 ± 0.1 Å for H206C. Compared to closed WT and H206Y channels, constitutively active dOrai mutants H206Q/C show a greater proportion of M173-F171 directed orientations. By contrast, M173-F259 distance reflected this trend in reverse. Black solid lines in (E) and (F) represent distances observed in the crystal structure (4HKR) of closed dOrai (d3 = 7.3 Å and d4 = 5.8 Å).
MD simulation trajectories of WT dOrai reveal interaction of M173 with F259.
Multiple synchronized molecular renderings of a single simulation repeat (0 to 440 ns). Molecular renderings are shown in three views; top down view of gate latch interaction site formed by two subunits, top down hexamer view, side view of two pairs of diagonal subunits TM1 and TM3, from left to right, respectively. Across all panels, key residues involved in the gate latch are M173 (blue), F171 (green), and, F259 (red), in agreement with the molecular renderings in Figure 2A–E. Only TM1 and TM3 are shown for simplicity. The subunits selected in the gate latch detail view are identified in the hexamer view by a black box in the hexamer view. The side view of two pairs of diagonal subunits depicts the counter-clockwise subunits in a ribbon representation. In all four subunits, F171 (green), V174 (purple), and E178 (orange), are shown in stick representation, in addition to other residues of the TM1-TM3 hydrophobic cluster described in Figure 1B (white spheres). The pore lining residues W148, Q152, R155, K159, K163, and L167 are all shown in two of the subunits (white sticks), as well as Cl- atoms which bind transiently within the basic region of the pore (small white spheres).
MD simulation trajectories of the open mutant H206C dOrai reveal interactions of M173 with F171 and F259.
Multiple synchronized molecular renderings of a single simulation repeat (0 to 556 ns) of the constitutively active dOrai H206C mutant. The other details of the simulation and rendering are same as in Figure 2—video 1.
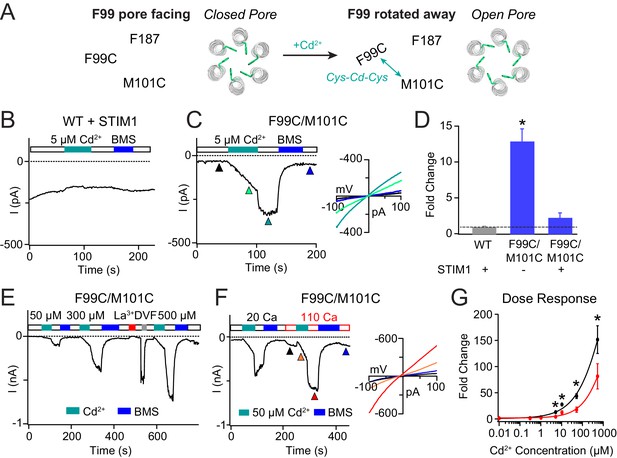
Enhancing the M101-F99 interaction via a metal-ion bridge activates F99C/M101C Orai1 in the absence of STIM1.
(A) Schematic of proposed mechanism of action of Cd2+ on F99C/M101C channels. (B) Cd2+ (5 µM) has no effect on WT Orai1 channels gated by STIM1. (C) By contrast, the activity of the F99C/M101C Orai1 variant without STIM1 is significantly boosted by Cd2+ (5 µM) and can be reversed by BMS (5 mM). Inset shows the current-voltage relationships of F99C/M101C at the time points indicated by the arrowheads. (D) Summary of Cd2+ potentiation (5 µM) on F99C/M101C channels without and with STIM1. (E) Dose-dependence of Cd2+ potentiation on F99C/M101C channels (40 s of Cd2+ application). (F) Application of Cd2+ in 110 mM Ca2+-containing external solution significantly decreases the relative extent of current potentiation. Note the larger pre-Cd2+ baseline current amplitude in the 110 mM Ca2+ solution. Inset: Current-voltage relationships of F99C/M101C in 110 mM Ca2+ solution at the indicated time points. (G) Summary of the fold increase in current amplitude of F99C/M101C channels following Cd2+ application in 20 mM versus 110 mM Ca2+ external solutions. Black and red lines represent polynomial fits to the data to enable visualization of the overall trends. Less relative potentiation is seen in 110 mM Ca2+ solution, suggesting that permeating Ca2+can compete with Cd2+ for a binding site within the pore. Values are mean ± S.E.M. N = 4–11 cells *p<0.05 by Student’s T-test. Numerical data for this figure can be found in Figure 3—source data 1.
-
Figure 3—source data 1
Numerical data for cadmium-mediated current potentiation in F99C/M101C Orai1.
- https://cdn.elifesciences.org/articles/60751/elife-60751-fig3-data1-v2.docx

Cd2+-mediated Orai1 current potentiation and inhibition is replicated by Zn2+.
(A) In addition to Cd2+, the transition metal-ion Zn2+ also interacts with cysteine residues. Like Cd2+, Zn2+ (50 µM) potentiates Orai1 F99C/M101C current. (B) Zn2+ (5 µM) inhibits M101C/F187C channels to a similar extent as an equivalent dose of Cd2+. Thus, the potentiation and inhibition effects observed are generalizable to other divalent ion bridges.
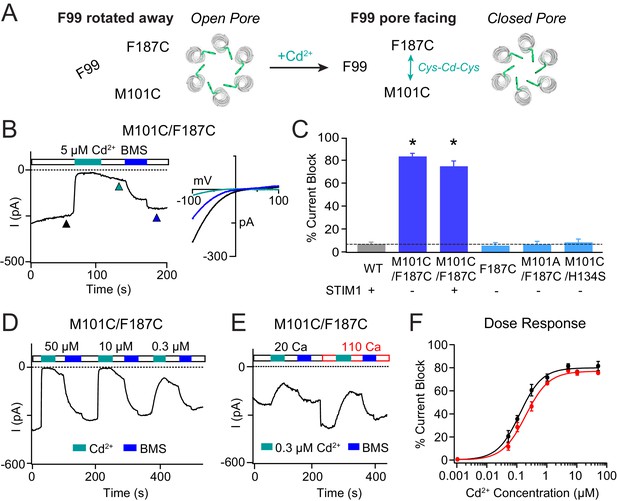
Stabilizing the M101-F187 interaction via a metal-ion bridge promotes pore closure.
(A) Schematic of proposed mechanism of action of Cd2+ on M101C/F187C channels. The addition of Cd2+ induces a bridge between M101C and F187C which releases the F99 gate returning to the ‘closed’ orientation. (B) Cd2+ (5 µM) strongly inhibits the current of M101C/F187C channels, which can be reversed by BMS (5 mM). Inset: Current-voltage relationships of M101C/F187C at the time points indicated by the arrowheads. (C) Summary of Cd2+ inhibition (5 µM) on M101C/F187C channels without and with STIM1. Double mutants with only one cysteine at positions M101C or F187C do not exhibit block by Cd2+, indicating that the coordination of Cd2+ requires cysteines at both residues. (D) M101C/F187C channels display a dose-dependent increase in current inhibition (40 s of Cd2+ application). (E) Application of Cd2+ in 110 mM Ca2+-containing external solution does not notably affect the extent of current inhibition by Cd2+. (F) Summary of the fold increase in current amplitude of M101C/F187C channels following Cd2+ application in 20 mM versus 110 mM Ca2+ external solutions. The lack of difference in effect suggests that the Cd2+ binding site is not within the pore, and instead at the TM1-TM3 interface as implied by the crystal structure. Values are mean ± S.E.M. N = 4–8 cells for each point; *p<0.05 by Student’s T-test. Numerical data for this figure can be found in Figure 4—source data 1.
-
Figure 4—source data 1
Numerical data for cadmium-mediated current inhibition in M101C/F187C Orai1.
- https://cdn.elifesciences.org/articles/60751/elife-60751-fig4-data1-v2.docx
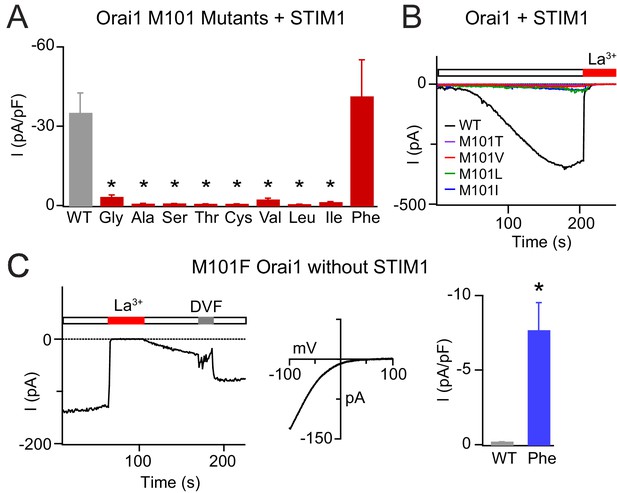
Mutations at M101 expected to disrupt M101-F99 interactions abrogate Orai1 gating.
(A) Current densities of Orai1 M101 mutants in the presence of STIM1. The majority of M101 mutations, even those to large residues similar in hydrophobicity to the native Met, abolish Orai1 activation by STIM1. (B) Time course traces of LOF M101 mutants with co-expressed with STIM1. Unlike WT Orai1 channels, these mutants do not conduct current after store-depletion by 8 mM BAPTA in the internal solution. (C) M101F evokes a GOF effect. This mutant is constitutively active even in the absence of STIM1. The constitutively active M101F variant is blocked by La3+ (150 µM) and its current-voltage relationship indicates a highly Ca2+-selective current. Values are mean ± S.E.M. N = 4–9 cells *p<0.05 by Student’s T-test. Numerical data for this figure can be found in Figure 5—source data 1.
-
Figure 5—source data 1
Numerical data for M101 Orai1 mutants.
- https://cdn.elifesciences.org/articles/60751/elife-60751-fig5-data1-v2.docx
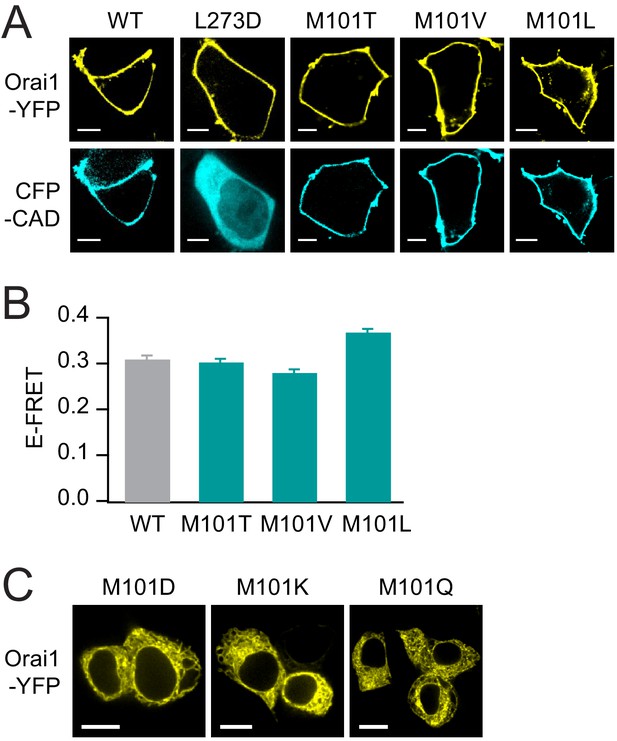
The LOF M101 mutants retain normal levels of binding to STIM1.
(A) M101 mutants recruit CFP-CAD to the plasma membrane indicating robust binding between Orai1 and CAD. L273D, which cannot bind to STIM1 (Li et al., 2011), is shown as a negative control. Scale bars: 5 Å. (B) The loss-of-function M101 mutants exhibit normal levels of E-FRET between Orai1-YFP and CFP-CAD, comparable to WT Orai1. Values are mean ± S.E.M. N = 51–70 cells. M101 mutants bind STIM1 yet cannot be gated, underscoring the importance of the methionine at this position for channel gating. (C) Confocal images of Orai1-YFP M101D/K/Q showing that these mutants do not express to the plasma membrane. Scale bars:10 Å. Numerical data for this figure can be found in Figure 5—source data 1.
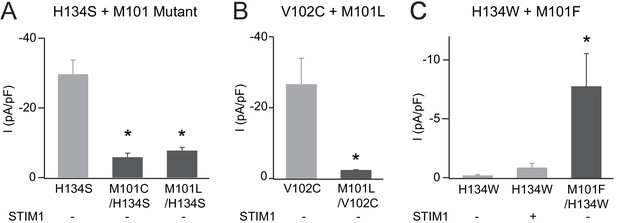
Additional analysis of M101 mutations.
(A) The addition of M101 mutation to the strongly active H134S mutant significantly abrogates channel current amplitude without STIM1. (B) Similarly, M101L/V102C channels have significantly smaller current amplitudes compared to V102C alone. (C) M101F mutation partially rescues the LOF phenotype of the Orai1 H134W mutant. Values are mean ± S.E.M. N = 4–6 cells *p<0.05 by Student’s T-test. Numerical data for this figure can be found in Figure 5—source data 1.
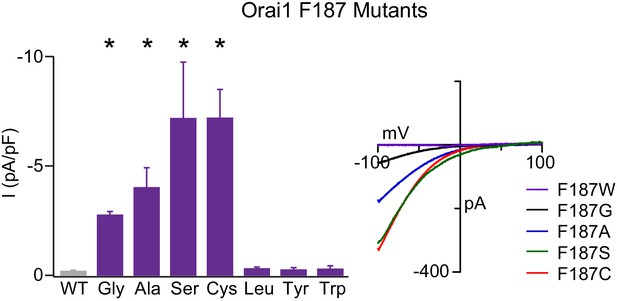
F187 is required for stabilizing the closed channel state.
Current densities of F187 mutants in the absence of STIM1. Smaller substitutions such as F187G/A/S/C produced GOF channels whereas larger substitutions remained closes in the absence of TSIM1. The right plot shows the current-voltage relationships of the GOF mutants. Values are mean ± S.E.M. N = 4–6 cells *p<0.05 by Student’s T-test. Numerical data for this figure can be found in Figure 5—source data 1.
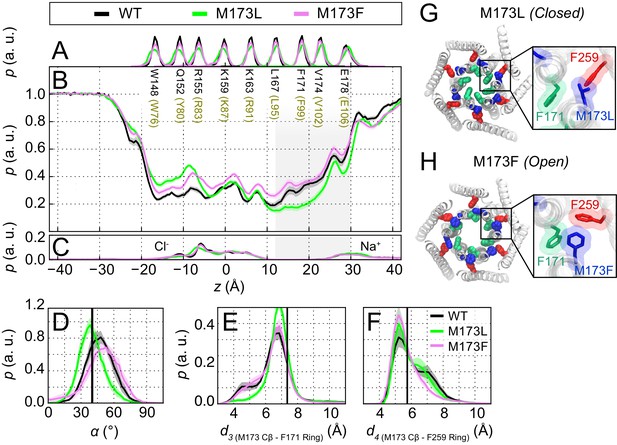
The loss-of-function M173L mutation decreases pore hydration and stabilizes closure of the hydrophobic gate.
(A) Relative probability distributions of the axial position of Cα atoms for all pore-lining residues (hOrai1 numbering in brown). Average distribution of water oxygen atoms (B) and Na+ and Cl− ions (C) along the pore axis. The P values from a two-sided Welch’s t-test of mean water oxygen count at z = 15 Å across all simulation repeats indicate a significant difference with respect to WT for M173L (1.2 × 10−2, p<0.05) but not for M173F (1.1 × 10−1). (D) Relative distributions of the radial angle of residue 171 defined as the angle between the pore axis, the center of mass of the two helical turns centered at residue 171, and the Cα atom of residue 171 in the different mutants. The mean and standard error of mean of F171 radial angle over simulation repeats are 41 ± 1 ° for M173L and 48 ± 1 ° for M173F. The P values from a two-sided Welch’s t-test for each system indicate significant differences (p<0.05) between the mean of these distributions with respect to WT simulations (6.1 × 10−24, 5.2 × 10−3). The black solid line represents the angle observed in the closed dOrai crystal structure 4HKR. (E–F) Distribution of distances between M173 with F171 (d3) and M173 with F259 (d4), respectively. Distances were measured from the Cβ of the methionine and the center of mass of the phenylalanine ring. The mean and standard error of mean of M173-F171 distances over simulation repeats are 6.5 ± 0.1 Å for WT, 6.9 ± 0.1 Å for M173L, and 6.6 ± 0.1 Å for M173F. In (F), M173-F259 distances over simulation repeats are 6.1 ± 0.1 Å for WT, 6.0 ± 0.1 Å for M173L, and 5.8 ± 0.1 Å for M173F. Black solid lines in (A) and (B) represent distances observed in the crystal structure of closed dOrai (d3 = 7.3 Å and d4 = 5.8 Å). (G) Snapshot of the positions of M173L, F171, and F259 in the closed M173L mutant. M173L decreases pore hydration and pore helix rotation, thereby evoking closure of the F171 hydrophobic gate. (H) Snapshot of the positions of M173F, F171, and F259 in the constitutively active M173F mutant. Only TM1 and TM3 helices are shown in (G–H) for simplicity.
MD simulation trajectory of the LOF mutant M173L dOrai.
Molecular rendering showing a simulation of the LOF M173L mutant (0 to 426 ns). Residues M173L (blue), the hydrophobic gate residue F171 (green), and TM3 residue F259 (red) are highlighted. Only TM1 and TM3 are shown for simplicity. The middle panel shows the top view and the right panel a side view. The black box is enlarged in the left panel. The video was smoothened over three frames to remove small amplitude, high-frequency fluctuations. Mutation of the latch (M173L) results in F171 stabilization into a pore-facing orientation and diminishes pore hydration and channel opening (Figure 6B,G and Figure 5A).
MD simulation trajectory of the GOF mutant M173F dOrai.
Molecular renderings of a single simulation repeat smoothened (over three frames) (0 to 520 ns). Residues M173F (blue), the hydrophobic gate residue F171 (green), and TM3 residue F259 (red) are highlighted. Only TM1 and TM3 are shown for simplicity. F171 is displaced away from the pore due to π-π, cation-π, or other interactions with M173F and F259 residues resulting in a GOF phenotype.
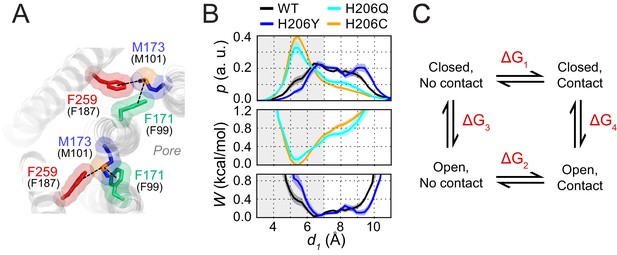
Free energy analysis of M173-F171 interactions.
(A) A snapshot of the H206C dOrai GOF mutant showing two subunits in which the M173 latch is engaged with the F171 hydrophobic gate and F259 to evoke gate opening. hOrai1 numbering is shown in parentheses. (B) Top: M173-F171 distances for the various H206 mutants replotted from Figure 2E. The gray shaded area corresponding to d1 <7 Å represents the range of distances used to infer the occurrence of Met-Phe contact. Middle and bottom: The potential of mean-force (W) for the M173-F171 distance for the mutants simulated as indicated in Figure 2. (C) Schematic of thermodynamic cycle associated with the M173-F171 interaction in closed and open channel states.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Cell line (Homo-sapiens) | HEK293-H | Thermo Fisher Scientific | 11631017 | RRID:CVCL_6643 |
| Commercial assay or kit | QuikChange II XL Site-Directed Mutagenesis Kit | Agilent | 200522 | |
| Transfected construct (human) | Orai1-YFP | Clontech Navarro-Borelly et al., 2008 | ||
| Transfected construct (human) | mCherry-STIM1 | Richard Lewis (Stanford) | ||
| Transfected construct (human) | CFP-CAD | Richard Lewis (Stanford) | ||
| Chemical compound, drug | Lipofectamine 2000 | Thermo Fisher Scientific | 11668019 | |
| Chemical compound, drug | cadmium chloride | Sigma-Aldrich | 202908 | |
| Chemical compound, drug | BMS, bis(2-mercaptoethylsulfone) | Calbiochem | 145626-87-5 | |
| Sequenced-based reagent | mutagenesis primers for Orai1 F99C | IDT Yamashita et al., 2017 | accatggcgcagccggagagcagagcc ggctctgctctccggctgcgccatggt | |
| Sequenced-based reagent | mutagenesis primers for Orai1 M101A | IDT | This paper | ccaccattgccaccgcggcgaagccggaga tctccggcttcgccgcggtggcaatggtgg |
| Sequenced-based reagent | mutagenesis primers for Orai1 M101C | IDT McNally et al., 2009 | ctccaccattgccacgcaggcgaagccggagag ctctccggcttcgcctgcgtggcaatggtggag | |
| Sequenced-based reagent | mutagenesis primers for Orai1 M101F | IDT | This paper | ccaccattgccacgaaggcgaagccggag ctccggcttcgccttcgtggcaatggtgg |
| Sequenced-based reagent | mutagenesis primers for Orai1 M101G | IDT | This paper | ccaccattgccaccccggcgaagccggaga tctccggcttcgccggggtggcaatggtgg |
| Sequenced-based reagent | mutagenesis primers for Orai1 M101I | IDT | This paper | accattgccactatggcgaagccggagag ctctccggcttcgccatagtggcaatggt |
| Sequenced-based reagent | mutagenesis primers for Orai1 M101L | IDT | This paper | accattgccaccaaggcgaagccggag ctccggcttcgccttggtggcaatggt |
| Sequenced-based reagent | mutagenesis primers for Orai1 M101S | IDT | This paper | tccaccattgccacgctggcgaagccggag ctccggcttcgccagcgtggcaatggtgga |
| Sequenced-based reagent | mutagenesis primers for Orai1 M101T | IDT | This paper | ccattgccaccgtggcgaagccggag ctccggcttcgccacggtggcaatgg |
| Sequenced-based reagent | mutagenesis primers for Orai1 M101V | IDT | This paper | accattgccaccacggcgaagccggag ctccggcttcgccgtggtggcaatggt |
| Sequenced-based reagent | mutagenesis primers for Orai1 V102C | IDT McNally et al., 2012 | gcacctccaccattgcgcacatggcgaagccggag ctccggcttcgccatgtgcgcaatggtggaggtgc | |
| Sequenced-based reagent | mutagenesis primers for Orai1 H134C | IDT Yeung et al., 2018 | catgagcgcaaacaggcacacagccaccagcact agtgctggtggctgtgtgcctgtttgcgctcatg | |
| Sequenced-based reagent | mutagenesis primers for Orai1 H134Q | IDT Yeung et al., 2018 | atgagcgcaaacagctgcacagccaccag ctggtggctgtgcagctgtttgcgctcat | |
| Sequenced-based reagent | mutagenesis primers for Orai1 H134S | IDT Yeung et al., 2018 | catgagcgcaaacaggctcacagccaccagcact agtgctggtggctgtgagcctgtttgcgctcatg | |
| Sequenced-based reagent | mutagenesis primers for Orai1 H134W | IDT Yeung et al., 2018 | gatcatgagcgcaaacagccacacagccaccagcactgt acagtgctggtggctgtgtggctgtttgcgctcatgatc | |
| Sequenced-based reagent | mutagenesis primers for Orai1 H134Y | IDT Yeung et al., 2018 | atcatgagcgcaaacagatacacagccaccagcactg cagtgctggtggctgtgtatctgtttgcgctcatgat | |
| Sequenced-based reagent | mutagenesis primers for Orai1 F187A | IDT Yeung et al., 2018 | ccacctcagctagggcgagcagcgtgccga tcggcacgctgctcgccctagctgaggtgg | |
| Sequenced-based reagent | mutagenesis primers for Orai1 F187C | IDT Yeung et al., 2018 | cctcagctaggcagagcagcgtgccg cggcacgctgctctgcctagctgagg | |
| Sequenced-based reagent | mutagenesis primers for Orai1 F187G | IDT | This paper | ccacctcagctaggccgagcagcgtgccga tcggcacgctgctcggcctagctgaggtgg |
| Sequenced-based reagent | mutagenesis primers for Orai1 F187L | IDT | This paper | accacctcagctagtaagagcagcgtgcc ggcacgctgctcttactagctgaggtggt |
| Sequenced-based reagent | mutagenesis primers for Orai1 F187S | IDT | This paper | ccacctcagctaggctgagcagcgtgccga tcggcacgctgctcagcctagctgaggtgg |
| Sequenced-based reagent | mutagenesis primers for Orai1 F187W | IDT | This paper | accacctcagctagccagagcagcgtgccg cggcacgctgctctggctagctgaggtggt |
| Sequenced-based reagent | mutagenesis primers for Orai1 F187Y | IDT | This paper | caccacctcagctagatagagcagcgtgccga tcggcacgctgctctatctagctgaggtggtg |
| Sequenced-based reagent | mutagenesis primers for Orai1 L273D | IDT Li et al., 2011 | ccgccagctcgttgtcctcctggaactgtc gacagttccaggaggacaacgagctggcgg |





