Characterization of the mechanism by which the RB/E2F pathway controls expression of the cancer genomic DNA deaminase APOBEC3B
Figures
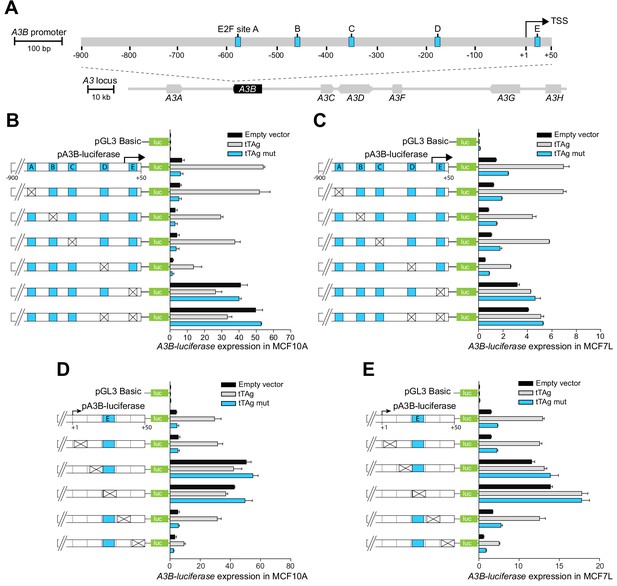
The A3B promoter harbors a repressive cis-element in the +1 to +50 region.
(A) Schematic of the 7-gene human APOBEC3 locus with the A3B promoter magnified to depict five predicted E2F binding sites (A-E in blue) relative to the TSS at +1 (scales indicated). (B–E) Relative luciferase activity of MCF10A or MCF7 cells expressing the indicated firefly luciferase construct (pGL3-basic, pA3B-luciferase, or mutant pA3B-luciferase), a renilla luciferase internal control plasmid (not shown), and a tTAg plasmid (empty, wildtype, or LxCxE mutant). Mutation clusters are depicted by X’s (mutant sequences in Supplementary file 4). Experiments report mean ± SD of n ≥ 2 technical replicates and are representative of n = 3 biologically independent replicates.
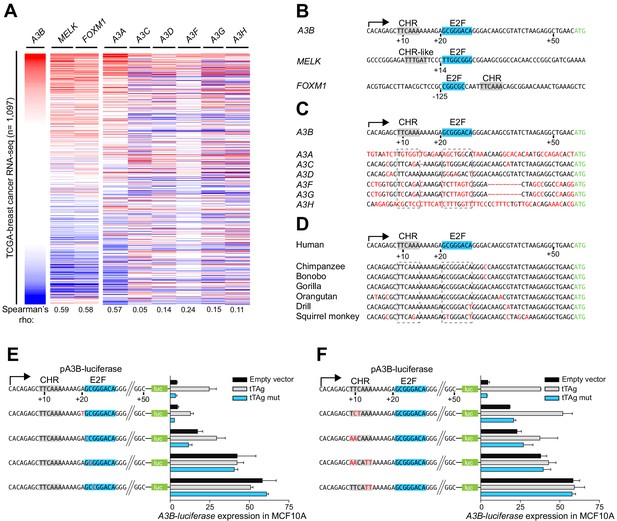
A3B repression requires both CHR and E2F cis-elements.
(A) Heatmap depicting high-to-low A3B expression levels in TCGA breast cancer specimens (n = 1,097) and correlations with two known RB/E2F target genes, MELK and FOXM1, and related APOBEC3 genes (Spearman’s rho indicated). (B) Comparison of the A3B promoter and analogous regions of MELK and FOXM1. Known and predicted E2F and CHR elements are indicated in blue and light gray, respectively. (C–D) Alignments of the A3B promoter sequence and corresponding promoter sequences of related human APOBEC3 genes and representative non-human primate A3B genes. (E–F) Relative luciferase activity of MCF10A cells expressing the indicated firefly luciferase construct (pA3B-luciferase or mutant pA3B-luciferase), a renilla luciferase internal control plasmid (not shown), and a tTAg plasmid (empty, wildtype, or LxCxE mutant). Panel (E) reports data for E2F mutants and (F) for CHR mutants. Experiments report mean ± SD of n ≥ 2 technical replicates and are representative of n = 3 biologically independent replicates.
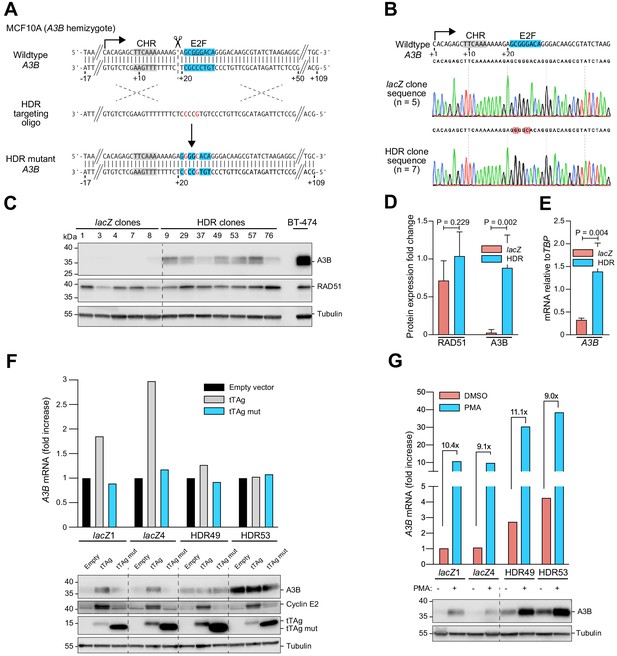
Single-base substitutions in the endogenous predicted E2F binding site induce A3B expression independent of activation by the PKC/ncNF-κB pathway.
Complementary supporting data are in Figure 3—figure supplement 1. (A) Schematic of CRISPR/Cas9-mediated HDR of the predicted E2F binding site in A3B hemizygous MCF10A cells. Top: CRISPR/Cas9 (scissors) introduces a DNA break (dashed line) adjacent to the predicted E2F binding site (blue). Middle: The ssDNA oligo used for HDR has two point mutations in the predicted E2F binding site including one that disrupts the PAM (underlined). Bottom: A3B promoter sequence of properly targeted clones. (B) Sanger DNA sequencing chromatograms of the E2F promoter region of a representative control clone (lacZ clones, n = 5) and a representative clone with the targeted E2F point mutations (HDR clones, n = 7). (C–D) A3B and RAD51 protein levels in control lacZ and HDR clones with tubulin as a loading control (representative immunoblots and quantification from n = 3 experiments). A3B-overexpressing BT-474 cells were used as a positive control. P-values from unpaired t-test. (E) A3B mRNA expression levels in control lacZ and HDR clones quantified by RT-qPCR (mean ± SD; p-value from unpaired t-test). (F) RT-qPCR (top) and immunoblot (bottom) results showing the effects of wildtype and LxCxE mutant tTAg on the A3B gene (top) and protein (bottom) expression in two representative lacZ and HDR49 clones. Cyclin E2 was used as a positive immunoblot control for tTAg-mediated induction of an RB/E2F-repressed gene. Tubulin was used as an immunoblot loading control. (G) Expression of A3B mRNA (top) and protein (bottom) upon PMA-treatment of the indicated lacZ control and HDR mutant clones. The magnitude of mRNA induction is indicated for each DMSO control and PMA-treated pair. Tubulin was used as an immunoblot loading control.
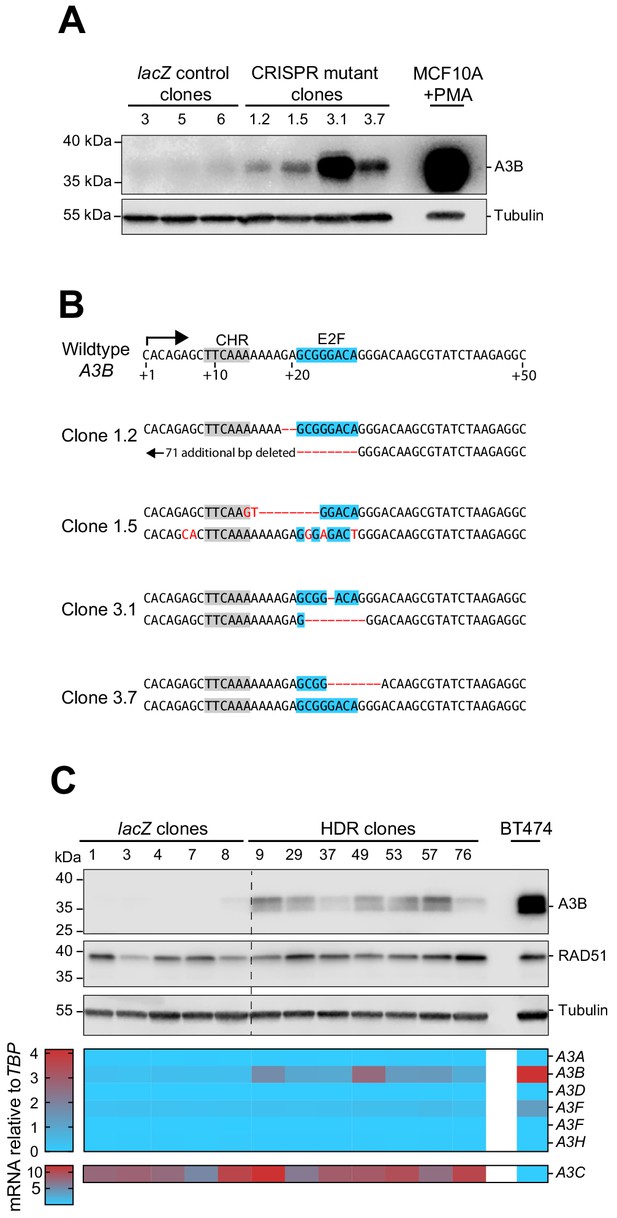
CRISPR/Cas9 disruption of the endogenous E2F site in the A3B promoter.
(A) Four independent MCF10A clones bearing mutations within the predicted E2F binding site were obtained using two different gRNAs in parallel to three clones with a control lacZ gRNA. Immunoblot showing A3B protein levels in the indicated control or targeted clones. PMA-treated MCF10A (25 ng/mL, 6 hr) was used as a positive control (representative of n = 3 blots). (B) DNA sequences of the E2F region of the A3B promoter of the four CRISPR/Cas9-targeted clones in panel (A). Clones 1.2 and 1.5 were obtained using gRNA_50UTR#1, and clones 3.1 and 3.7 were obtained using gRNA_50UTR#3 (gRNA sequences in Supplementary file 4). (C) Immunoblots from Figure 3C are shown again here for comparison with a heatmap summary of APOBEC3 family member mRNA levels in the same control and HDR-targeted clones. APOBEC3 mRNA levels were determined by RT-qPCR, normalized to TBP, and presented as a heatmap for comparison. A3C is shown on a separate scale due to relatively high expression in MCF10A.
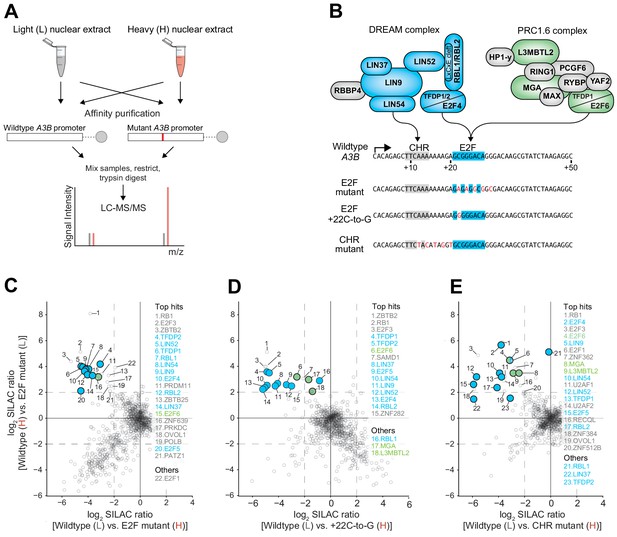
The DREAM and PRC1.6 repressive complexes bind to the CHR-E2F region of the A3B promoter.
Immunoblot validations of representative binding proteins from proteomic experiments are in Figure 4—figure supplement 1. (A) Schematic of the SILAC DNA pull-down strategy used to identify proteins from MCF7 cells capable of interacting with A3B promoter sequences. (B) Illustration of DREAM and PRC1.6 complexes positioned over the indicated A3B promoter elements (proteomic hits shaded blue and green, respectively). (C–E) Log2-transformed SILAC ratios of proteins purified using the indicated promoter sequences and identified through LC-MS/MS (dashed line, SILAC ratio threshold >2.0 [log2]). ‘Top hits’ are proteins surpassing the >2.0 log2 SILAC ratio threshold in both datasets (rank based on heavy versus light SILAC ratio). ‘Others’ are proteins of interest surpassing the >2.0 log2 SILAC ratio threshold in at least one dataset. Data for DREAM and PRC1.6 components are shaded blue and green, respectively.
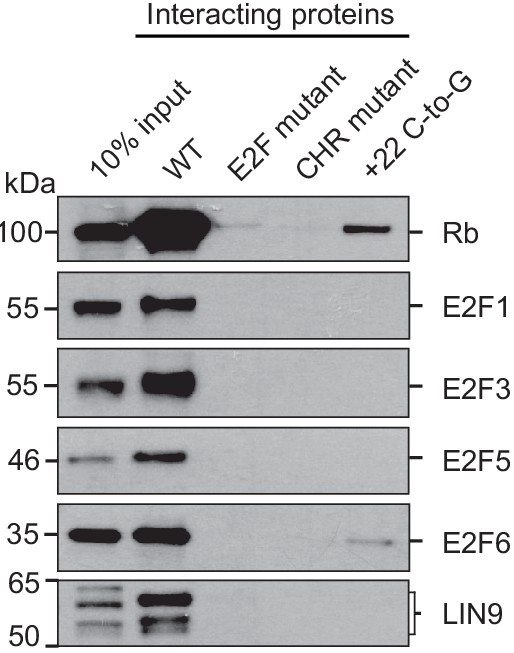
Immunoblot validations of representative A3B promoter-binding proteins identified in proteomics experiments.
Immunoblots were done for the indicated proteins, which were pulled-down by wildtype A3B promoter sequence but not by the indicated mutant sequences.
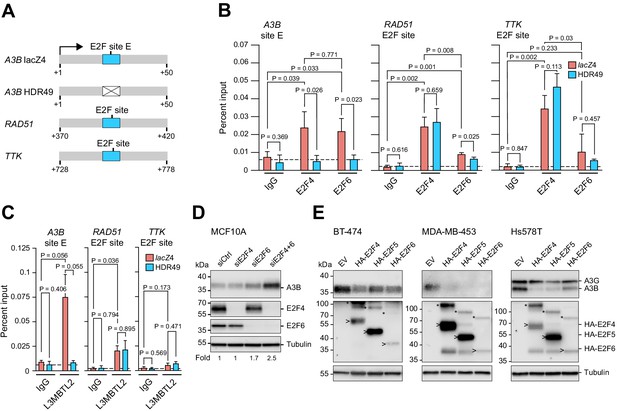
Endogenous A3B regulation by both E2F4/DREAM and E2F6/PRC1.6 complexes.
(A) Schematics of the promoter regions interrogated by ChIP experiments. Wildtype E2F sites are depicted by blue boxes and the mutant E2F site in the A3B promoter by a white X-box. (B–C) E2F4, E2F6, and L3MBTL2 occupancy at the indicated E2F sites in A3B, RAD51, and TTK, as analyzed by ChIP-qPCR using lacZ4 and HDR49 cells. Experiments in (B) report mean ± SD of n = 3 biologically independent replicates and in (C) of n = 2 biologically independent replicates (p values from unpaired t-test). Dashed lines indicate the average IgG background. (D) Immunoblots of A3B, E2F4, and E2F6 in MCF10A cells treated 24 hr with the indicated siRNAs. Tubulin was used as a loading control. Representative blots are shown and fold-changes below are based on the average values from n = 3 biologically independent replicates. (E) Immunoblots of A3B and the indicated HA-tagged E2F proteins in BT-474, MDA-MB-453, and Hs578T cells transduced with lentiviral constructs encoding mCherry-T2A-HA-E2F4, -HA-E2F5, -HA-E2F6, or -EV (empty vector control). Representative blots of n = 2 experiments are shown. Asterisks indicate E2Fs still fused with mCherry due to incomplete ribosome skipping at the T2A site, and arrowheads indicate bands for free E2F. Tubulin was used as a loading control.
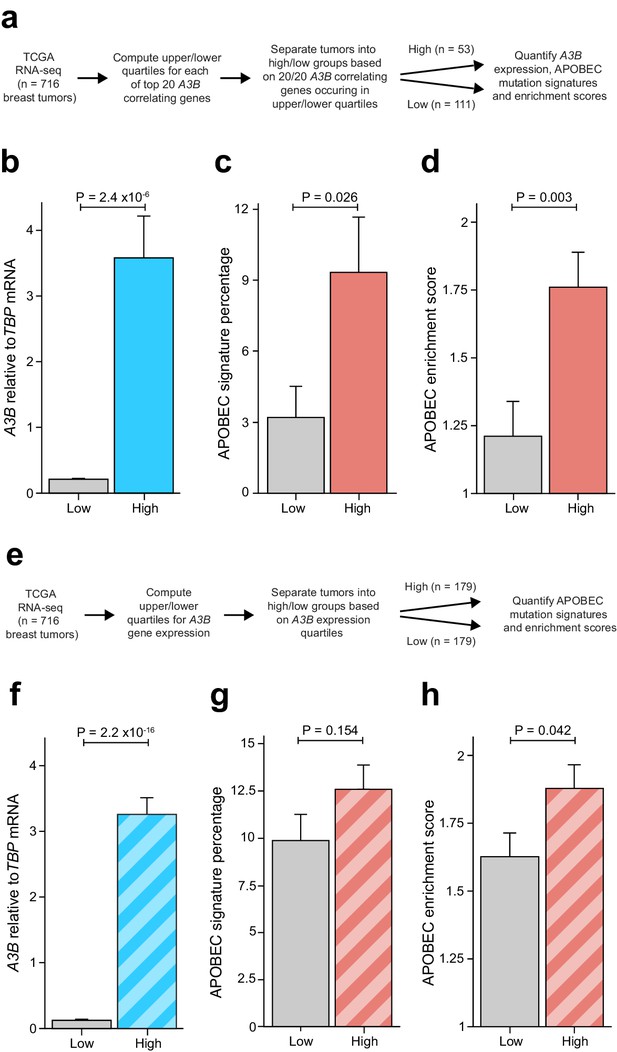
Elevated levels of APOBEC signature mutations in breast tumors with coordinated overexpression of an E2F-repressed gene set.
Complementary analyses are presented in Figure 6—figure supplement 1. (A) Schematic depicting the bioinformatics workflow of TCGA breast tumor data sets based on the 20 genes most strongly associated with A3B expression (Figure 2A and Supplementary file 1). (B–D) The mean A3B mRNA levels, mean APOBEC mutation percentages, and mean APOBEC enrichment scores in breast tumors with coordinated overexpression (high) or repression (low) of the 20 gene set (mean ± SD; n = 53 tumors in the high group and n = 111 in the low group; p values from Welch’s t-test). (E) Schematic depicting the bioinformatics workflow of TCGA breast tumor data sets based solely on A3B mRNA expression levels. (F–H) The mean A3B mRNA levels, mean APOBEC mutation percentages, and mean APOBEC enrichment scores in breast tumors with high or low A3B mRNA levels (mean ± SD of top and bottom quartiles; n = 179 tumors in each group; p values from Welch’s t-test).
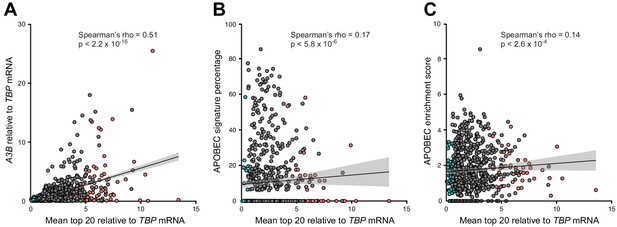
Global pairwise comparisons of the mean mRNA levels of the top 20 E2F-repressed/A3B-associated genes and A3B mRNA levels and APOBEC mutation signature prevalence in primary breast tumors.
(A–C) Reanalysis of the data sets in Figure 6, panels B-D. To facilitate pairwise comparisons, we first calculated a single relative mean expression value to represent the top 20 E2F-repressed/A3B-associated genes from each breast tumor RNA-seq data set. Then, the mean values from the top 20 E2F-repressed/A3B-associated genes were compared with A3B mRNA expression levels (panel A), APOBEC signature percentages (panel B), and APOBEC signature enrichment values for each tumor (panel C). Non-parametric Spearman’s correlations and non-exact p values are show with each plot, in addition to a linear model (black line) with corresponding 95% confidence interval (gray shading). Data points from tumors in the high (n = 53) and low (n = 111) groups are highlighted in red and blue, respectively, for comparison with Figure 6B–D.
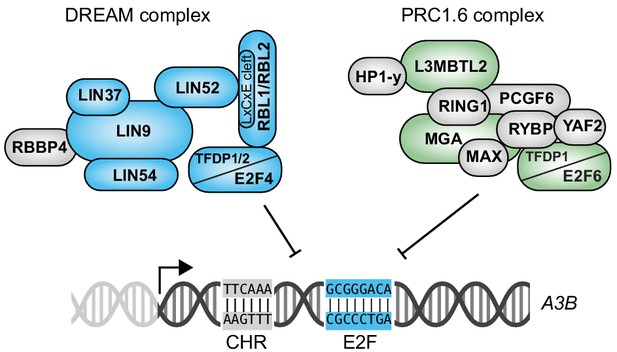
Model for coordinated repression of A3B transcription by both E2F4/DREAM and E2F6/PRC1.6 complexes.
Transcriptional repression of A3B through the combined activities of E2F4/DREAM and E2F6/PRC1.6 complexes. Other regulatory mechanisms including A3B transcriptional activation by PKC/ncNF-κB signal transduction are not shown. See text for details and discussion.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Cell line (Homo sapiens, female) | MCF10A | ATCC | Cat#:CRL-10317 RRID:CVCL_0598 | |
| Cell line (Homo sapiens, female) | MCF10A-4C10 | This study | Hemizygous for A3B | Request by contacting RSH |
| Cell line (Homo sapiens, female) | MCF7 | ATCC | Cat#:HTB-22 RRID:CVCL_0031 | |
| Cell line (Homo sapiens, female) | BT474 | ATCC | Cat#:CLR-7913 RRID:CVCL_0179 | |
| Cell line (Homo sapiens, female) | Hs578T | ATCC | Cat#:HTB-126 RRID:CVCL_0332 | |
| Cell line (Homo sapiens, female) | MDA-MB-453 | ATCC | Cat#:HTB-131 RRID:CVCL_0418 | |
| Cell line (Homo sapiens, female) | HEK 293T | ATCC | Cat#:CRL-3216 RRID:CVCL_0063 | |
| Antibody | Anti-RAD51 (rabbit monoclonal) | Abcam | Cat#:ab133534 RRID:AB_2722613 | WB (1:10,000) |
| Antibody | Anti-E2F4 (mouse monoclonal) | Santa Cruz | Cat#:sc-398543 | WB (1:250) ChIP: 5 µg per 20 µg Dynabeads |
| Antibody | Anti-E2F6 (rabbit polyclonal) | Abcam | Cat#:ab53061 RRID:AB_2097254 | WB (1:500) ChIP: 5 µg per 20 µg Dynabeads |
| Antibody | Anti-HA (rabbit monoclonal) | Cell Signaling | Cat#:3724 RRID:AB_1549585 | WB (1:5000) |
| Antibody | Anti-Rb (mouse monoclonal) | Santa Cruz | Cat#:sc-102 RRID:AB_628209 | WB (1:300) |
| Antibody | Anti-E2F1 (mouse monoclonal) | Santa Cruz | Cat#:sc-251 RRID:AB_627476 | WB (1:1,000) |
| Antibody | Anti-E2F3 (mouse monoclonal) | Santa Cruz | Cat#:sc-56665 RRID:AB_1122397 | WB (1:800) |
| Antibody | Anti-E2F5 (mouse monoclonal) | Santa Cruz | Cat#:sc-374268 RRID:AB_10988935 | WB (1:800) |
| Antibody | Anti-E2F6 (mouse monoclonal) | Santa Cruz | Cat#:sc-53273 RRID:AB_783163 | WB (1:300) |
| Antibody | Anti-LIN9 (mouse monoclonal) | Santa Cruz | Cat#:sc-398234 | WB (1:300) |
| Antibody | Anti-tubulin (mouse monoclonal) | Sigma-Aldrich | Cat#:T5168 RRID:AB_477579 | WB (1:20,000) |
| Antibody | Anti-A3B (rabbit monoclonal) | NIH AIDS Reagent Program | Cat#:12397 RRID:AB_2721202 | WB (1:1,000) |
| Antibody | Anti-L3MBTL2 (rabbit polyclonal) | Active Motif | Cat#:39569 RRID:AB_2615062 | ChIP: 5 µg per 20 µg Dynabeads |
| Recombinant DNA reagent | pGL4.74 TK-RL renilla control (plasmid) | Promega | Cat#:E692A | Internal control for luciferase assays |
| Recombinant DNA reagent | pGL3 basic (plasmid) | Promega | Cat#:E1751 | Base vector for luciferase assays |
| Recombinant DNA reagent | pA3B-luciferase (plasmid) | This study | Wildtype A3B promoter + luciferase | Request by contacting RSH |
| Recombinant DNA reagent | pLenti-lox-empty vector (plasmid) | Carpenter et al., 2019 | ||
| Recombinant DNA reagent | pLenti-lox-BKPyV tTAg (plasmid) | Starrett et al., 2019 | ||
| Recombinant DNA reagent | pLenti-lox-BKPyV tTAg LxCxE mutant (plasmid) | Starrett et al., 2019 | ||
| Recombinant DNA reagent | pLenti4/TO-mCherry-T2A-MCS (plasmid) | This study | Base vector for E2F expression | Request by contacting RSH |
| Recombinant DNA reagent | pLenti4/TO-mCherry-T2A-HA-E2F4 (plasmid) | This study | Lentiviral vector for expression of E2F4 | Request by contacting RSH |
| Recombinant DNA reagent | pLenti4/TO-mCherry-T2A-HA-E2F5 (plasmid) | This study | Lentiviral vector for expression of E2F5 | Request by contacting RSH |
| Recombinant DNA reagent | pLenti4/TO-mCherry-T2A-HA-E2F6 (plasmid) | This study | Lentiviral vector for expression of E2F6 | Request by contacting RSH |
| Recombinant DNA reagent | pLentiCRISPR-LoxP-A3B-gRNA#1 (plasmid) | This study | Lentiviral vector for expression of gRNA targeting E2F site E | Request by contacting RSH |
| Recombinant DNA reagent | pLentiCRISPR-LoxP-A3B-gRNA#3 (plasmid) | This study | Lentiviral vector for expression of gRNA targeting E2F site E | Request by contacting RSH |
| Sequence-based reagent | Cas9-encoding modified RNA | TriLink Biotech | Cat#:L7206-100 | |
| Commercial assay or kit | Dual Luciferase Reporter Assay | Promega | Cat#:E1960 | |
| Commercial assay or kit | Neon Transfection System 100 µL Kit | ThermoFisher | Cat#:MPK10025 | |
| Software, algorithm | MaxQuant version 1.5.2.8 | MaxQuant | RRID:SCR_014485 | |
| Software, algorithm | Fiji | Fiji | RRID:SCR_002285 | |
| Software, algorithm | GraphPad Prism 6 | GraphPad | RRID:SCR_002798 | |
| Software, algorithm | Image Studio | LI-COR Biosciences | RRID:SCR_015795 | |
| Other | Spark Multimode Microplate Reader | Tecan | ||
| Other | Neon Transfection System | ThermoFisher | Cat#:MPK5000 | |
| Other | LI-COR Odyssey FC | LI-COR | Cat#:2800 | |
| Other | LightCycler 480 Instrument | Roche | Cat#:04640268001 | |
| Other | EASY-nLC 1200 system | ThermoFisher | Cat#:LC140 | |
| Other | Q Exactive HF mass spectrometer | ThermoFisher |
Additional files
-
Supplementary file 1
Genes associating positively with A3B expression in breast cancer.
- https://cdn.elifesciences.org/articles/61287/elife-61287-supp1-v2.xlsx
-
Supplementary file 2
Log-transformed SILAC values from proteomics experiments.
- https://cdn.elifesciences.org/articles/61287/elife-61287-supp2-v2.xlsx
-
Supplementary file 3
Gene expression values relative to TBP mRNA in TCGA breast tumors.
- https://cdn.elifesciences.org/articles/61287/elife-61287-supp3-v2.xls
-
Supplementary file 4
Sequences of oligonucleotides used in this study.
- https://cdn.elifesciences.org/articles/61287/elife-61287-supp4-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/61287/elife-61287-transrepform-v2.docx





