Cohesin mutations are synthetic lethal with stimulation of WNT signaling
Figures
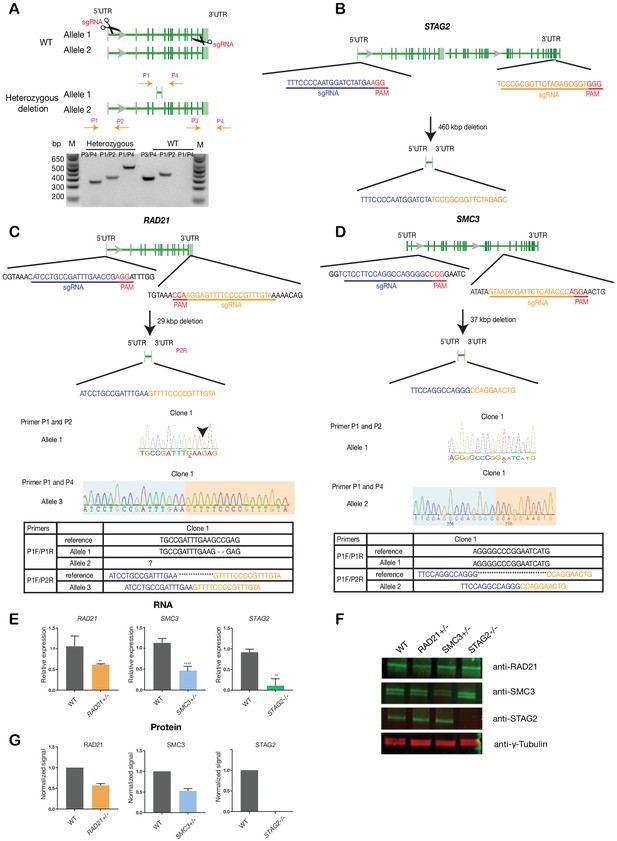
Creation of MCF10A isogenic cell lines with cohesin gene deletions.
(A) Top, schematic diagram shows the deletion strategy for genes encoding cohesin subunits RAD21, SMC3, and STAG2 using two sgRNAs targeting the 5'UTR and the 3'UTR of each gene. Bottom, heterozygous clones were identified by PCR using specific primer pairs flanking the deletion region. Representative DNA gel shows the PCR products yielded using specific primer pairs for MCF10A parental and RAD21+/- deletion clone. M, ladder marker. (B,C,D) Schematic deletion strategy and summary of the allele sequences for the STAG2 homozygous deletion clone, and the RAD21 and SMC3 heterozygous deletion clones. (E) RNA levels of the targeted genes in MCF10A cohesin-deficient clones. (F) Representative immunoblot and (G) quantification of cohesin protein levels. γ-tubulin was used as loading control. n = 3 independent experiments, mean ±s.d., one-tailed student t test: **p≤0.01; ****p≤0.0001. Guide RNAs and PCR primers can be found in Supplementary file 1.
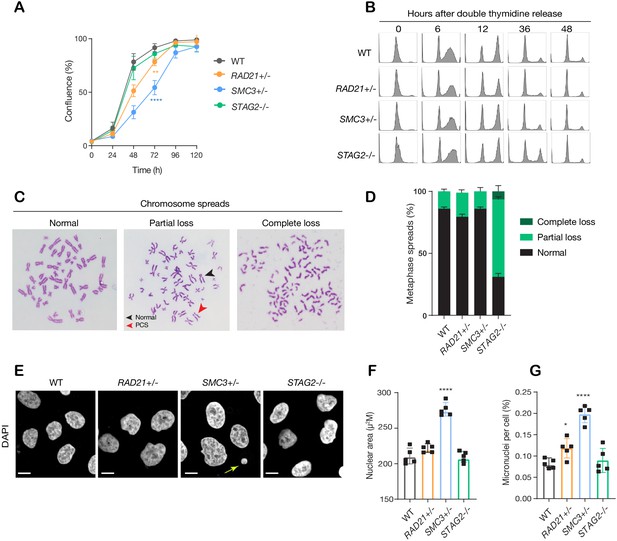
minor deficiencies in cell cycle progression, chromosome segregation and nuclear morphology in cohesin-deficient isogenic cell lines.
(A) Proliferation curves of MCF10A parental and cohesin-deficient clones. n = 3 independent experiments, mean ± sd, two-way ANOVA: **p≤0.01; ****p≤0.0001. Doubling times in hours of MCF10A parental cells are 15.5 ± 0.1; RAD21+/-, 17.1 ± 0.2; SMC3+/-, 21.4 ± 0.9; STAG2-/-, 16.4 ± 0.2 respectively. (B) Flow cytometry analysis of cell cycle progression. (C) Representative metaphase spread images of cohesin-deficient cells. Black arrow indicates a normal chromosome; Red arrow indicates premature chromatid separation (PCS). (D) Quantification of chromosome cohesion defects. A minimum of 20 metaphase spreads were examined per individual experiment, n = 2 independent experiments, mean ±s.d. (E) Representative confocal images of nuclear morphology. Scale bar, 15 μM. Yellow arrow indicates a micronucleus. (F) Quantification of nuclear area. (G) Quantification of micronuclei (MN). A minimum of 1000 cells was examined per individual experiment. n = 5 independent experiments (square symbols), mean ±s.d., one-way ANOVA: *p≤0.05; ****p≤0.0001. Source data is available for A,D,F,G in Figure 2—source data 1.
-
Figure 2—source data 1
Raw data for Figure 2a,d,f,g.
- https://cdn.elifesciences.org/articles/61405/elife-61405-fig2-data1-v2.xlsx
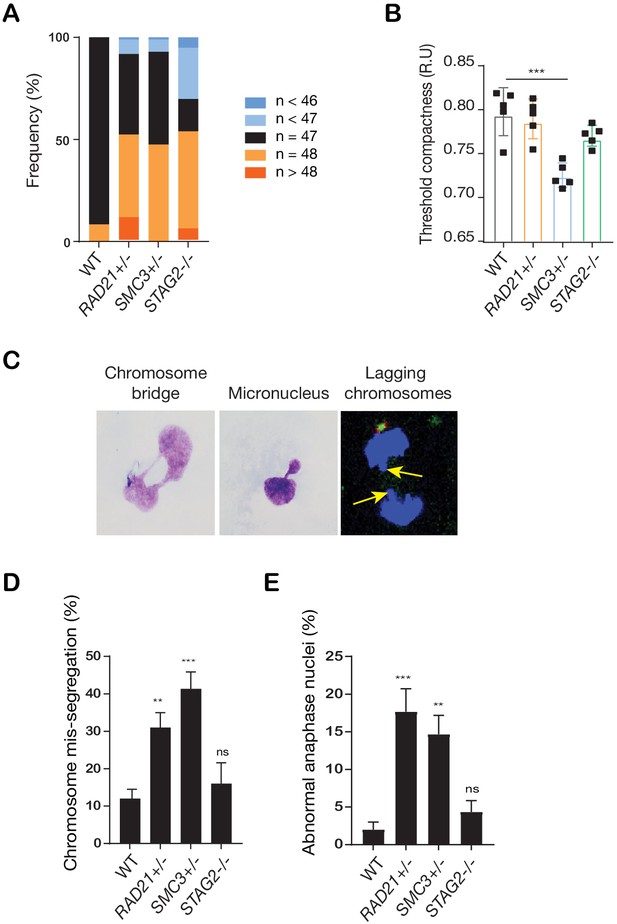
Cohesin deficiency causes a modest increase in chromosome mis-segregation events in MCF10A cells.
(A) Frequency of aneuploidy in cohesin-deficient MCF10A clones compared to parental MCF10A (WT). (B) Threshold compactness in the nuclei of cohesin-deficient clones compared to wild type MCF10A (R.U. = relative units). n = 5 independent experiments (square symbols), mean ± s.d., one-way ANOVA: ***p≤0.001. (C) Representative images of abnormal interphase nuclei showing chromosome bridges, micronuclei, and lagging chromosomes at anaphase. (D) Quantification of chromosome bridges and micronuclei events in MCF10A cohesin-deficient cells. (E) Quantification of chromosome mis-segregation observed in MCF10A cohesin-deficient clones. At least 100 mitotic cells were examined per condition, n = 3 independent experiments, mean ±s.d., one-way ANOVA **p≤0.01; ***p≤0.0005; ns, not significant.
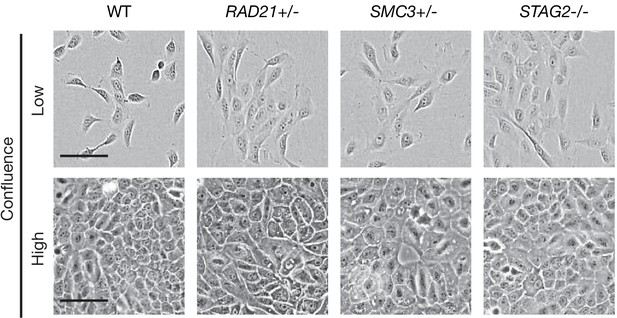
Cell morphology of MCF10A parental and cohesin-deficient cells grown at low and high confluence.
MCF10A cells were grown in low (1000 cells/well) and high density (4000 cells/well) in 96-well plates. Cohesin-deficient cells display a cobblestone-like morphology that is similar to MCF10A parental cells (WT). Scale bar, 250 μM.
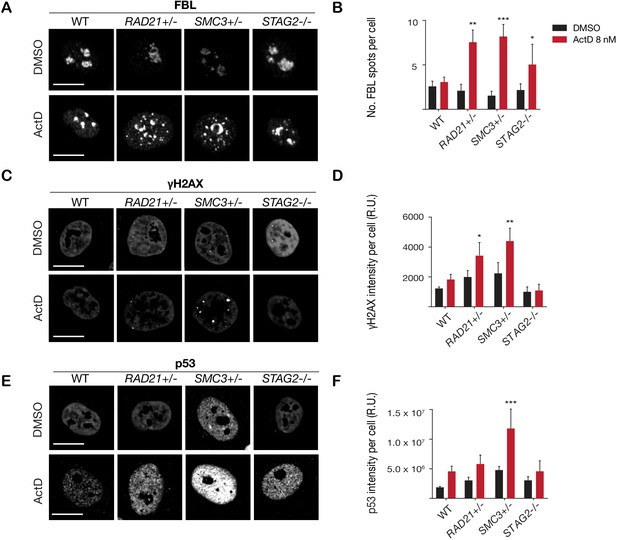
Cohesin-deficient cells have increased sensitivity to nucleolar stress and DNA damaging agents.
(A) Representative image and (B) quantification of nucleolar dispersal observed in parental (WT) MCF10A cells and cohesin-deficient clones after exposure to a DNA damaging agent, Actinomycin D (ActD) 8 nM. Fibrillarin (FBL) staining was used as a marker for nucleoli. (C) Representative image and (D) quantification of DNA damage foci observed in parental (WT) MCF10A cells and cohesin-deficient clones after exposure to ActD. An antibody detecting γH2AX was used to visualize foci of DNA double-strand breaks. (E) Representative image and (F) quantification of nuclear p53 in parental (WT) MCF10A cells and cohesin-deficient clones after exposure to ActD. A minimum of 500 cells was examined per individual experiment. n = 3 independent experiments, mean ± s.d., one-way ANOVA: *p≤0.05; **p≤0.01; ***p≤0.0005. Scale bar, 15 μM. Source data is available for Figure 2B,D,F in Figure 3—source data 1.
-
Figure 3—source data 1
Raw data for Figure 3.
- https://cdn.elifesciences.org/articles/61405/elife-61405-fig3-data1-v2.xlsx
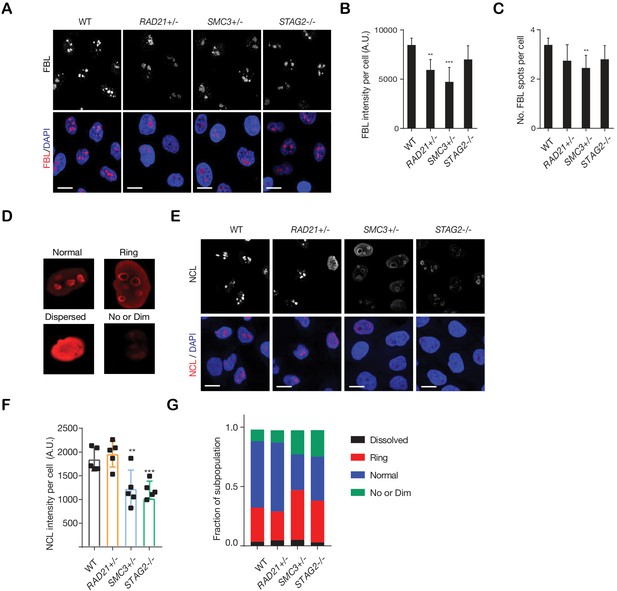
Cohesin-deficient cells show altered nucleolar morphology.
(A) Representative images of nucleolar morphology as determined using fibrillarin (FBL) as a marker of nucleoli. (B,C) Quantification of FBL intensity and number of FBL spots per cell in each genotype. A minimum of 1000 cells was examined per individual experiment, n = 3 independent experiments, mean ± s.d., one-way ANOVA: *p≤0.05; **p≤0.01; ***p≤0.0005. (D) Types of nucleolar morphology observed in MCF10A parental cells. (E) Representative images of nucleolar morphology determined by using nucleolin (NCL) as a marker. (F) Quantification of NCL intensity per cell in each genotype. At least 500 cells were examined per individual experiment, n = 3 independent experiments, mean ±s.d., one-way ANOVA: **p≤0.01; ***p≤0.0005. (G) Quantification of NCL morphology classes present in cohesin-deficient cells. A.U., Arbitrary unit.
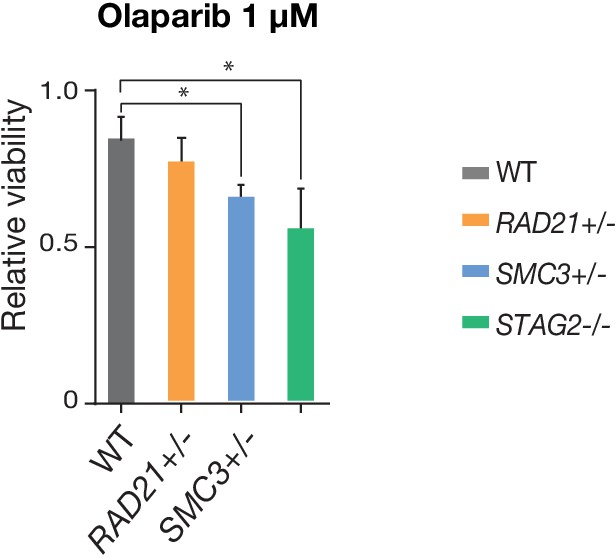
PARP sensitivity of cohesin-deficient cells.
STAG2- and SMC3-deficient MCF10A cells show reduced viability following 48 hr of treatment with the PARP inhibitor, Olaparib, relative to parental cells (WT). n = 2 independent experiments, mean ±s.d., one-way ANOVA: *p≤0.05.
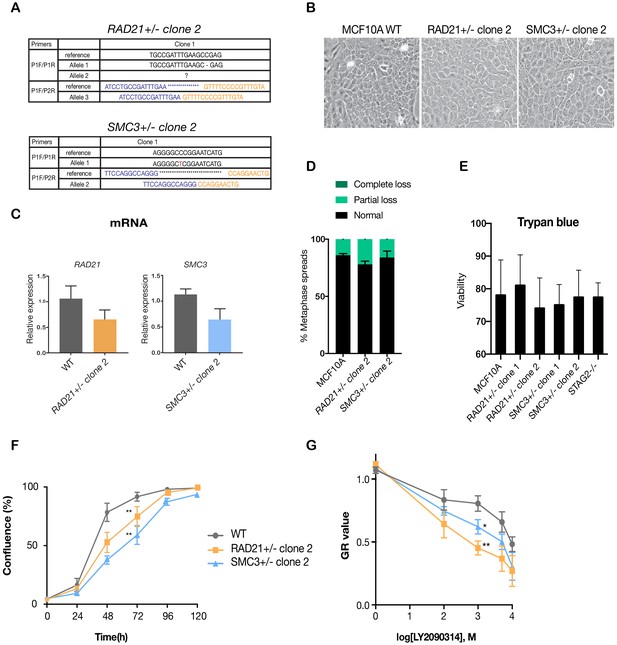
Data replication with additional MCF10A isogenic cell lines with cohesin gene deletions.
(A) Summary of the allele sequences for additional RAD21 and SMC3 heterozygous clones, RAD21+/- clone two and SMC3+/- clone 2. (B) Cohesin-deficient clone 2 cells display a cobblestone-like morphology that is similar to MCF10A parental cells (WT). (C) RNA levels of the targeted genes in the additional MCF10A cohesin-deficient clones. (D) Quantification of chromosome cohesion defects in RAD21+/- clone two and SMC3+/- clone 2. A minimum of 20 metaphase spreads were examined per individual experiment, n = 2 independent experiments, mean ±s.d. (E) Trypan blue measurement of cell viability for all cohesin-deficient clones in this study. Survival rates of mutant clones under standard culture conditions are not significantly different from wild type, n = 2 independent experiments, mean ±s.d. (F) Proliferation curves of MCF10A parental and RAD21+/- clone two and SMC3+/- clone 2. (G) Dose-response curves were generated for validation of LY2090314 in RAD21+/- clone two and SMC3+/- clone two compared with MCF10A cells. For (F) and (G), n = 3 independent experiments, mean ± sd, two-way ANOVA: *p≤0.05; **p≤0.01.
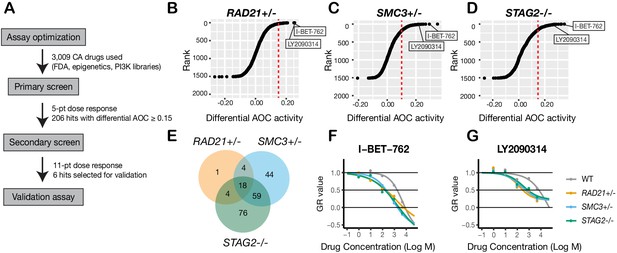
A synthetic lethal screen identifies common sensitivity of cohesin-deficient cells to WNT activation and BET inhibition.
(A) Schematic overview of the synthetic lethal screen. (B,C,D) Overview of the differential area over the curve (AOC) activity of all compounds tested in cohesin-deficient cell lines relative to parental MCF10A cells in the primary screen. A threshold of differential AOC ≥ 0.15 (red dashed lines) was used to filter candidate compounds of interest. (E) Venn diagram showing the number of common and unique compounds that inhibited RAD21+/-, SMC3+/-, and STAG2-/- in the primary screen. (F,G) Dose-response curves of I-BET-762 and LY2090314. Source data is available for B–E in Figure 4—source data 1.
-
Figure 4—source data 1
Raw cell counts for SL screen compound treatments.
- https://cdn.elifesciences.org/articles/61405/elife-61405-fig4-data1-v2.xlsx
-
Figure 4—source data 2
Table and AOC measurements of all hit compounds from the screen.
- https://cdn.elifesciences.org/articles/61405/elife-61405-fig4-data2-v2.xlsx
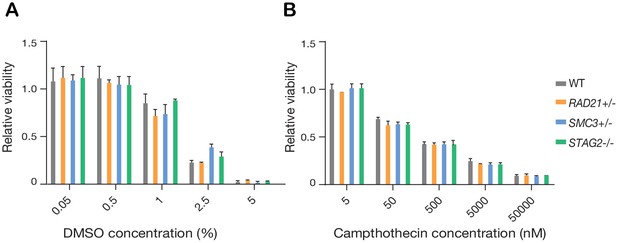
Synthetic lethal screen controls.
(A) DMSO (negative control) and (B) Camptothecin (positive control) titrations were used to establish a cytotoxicity range in MCF10A cells and cohesin-deleted derivatives. n = 2 independent experiments, mean ±s.d.
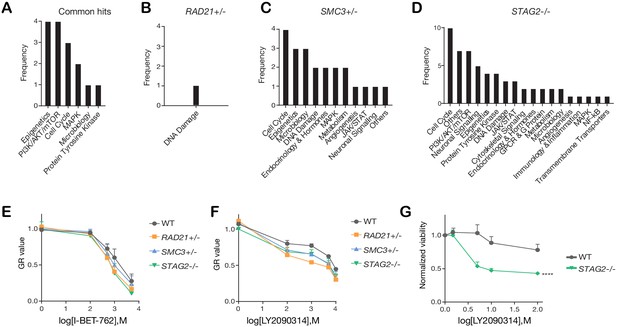
Categories of compounds that differentially inhibit cohesin-deficient cells.
(A) Common and (B,C,D) unique categories of compounds identified in the primary screen. See Figure 4—source data 1 for details. (E,F) Dose-response curves were generated for validation of I-BET-762 and LY2090314 in MCF10A cells. (G) Dose-response curve of LY2090314 in the K562 STAG2R614* mutant cell line. Two-way ANOVA: ****p≤0.0001 Source data is available for Figure 4A–D in Figure 4—source data 2.
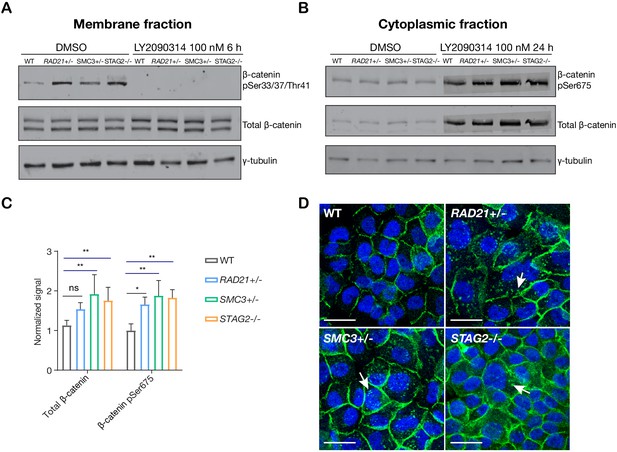
LY2090314-mediated WNT stimulation leads to β-catenin stabilization in cohesin-deficient MCF10A cells.
(A) Immunoblot of the membrane fraction of parental (WT) and cohesin-deficient MCF10A cells shows increased basal level of β-catenin phosphorylation at Ser33/37/Thr41. (B) Immunoblot of the cytoplasmic fraction shows increased level of both total and phosphorylated β-catenin at Ser675 after parental (WT) and cohesin-deficient MCF10A cells were treated with LY2090314 at 100 nM for 24 hr. (C) Quantification of protein levels for total and phosphorylated β-catenin at Ser675. n = 3 independent experiments, mean ± s.d., one-way ANOVA: *p≤0.05, **p≤0.01. (D) Immunofluorescence images show cytosolic accumulation of active β-catenin in cohesin-deficient MCF10A cells treated with LY2090314 100 nM for 24 hr, relative to parental (WT) MCF10A cells. White arrows indicate puncta of β-catenin (pSer675). Scale bar = 25 μM. Full length blots and molecular size markers are available for A,B in Figure 5—source data 1. Source quantification data is available for C in Figure 5—source data 2.
-
Figure 5—source data 1
Untrimmed blots for Figure 5A, B.
- https://cdn.elifesciences.org/articles/61405/elife-61405-fig5-data1-v2.pdf
-
Figure 5—source data 2
Quantitation of blots in Figure 5A, B.
- https://cdn.elifesciences.org/articles/61405/elife-61405-fig5-data2-v2.xlsx
-
Figure 5—source data 3
Untrimmed blots for Figure 5A, B.
- https://cdn.elifesciences.org/articles/61405/elife-61405-fig5-data3-v2.pdf
-
Figure 5—source data 4
Untrimmed blots for Figure 5A, B.
- https://cdn.elifesciences.org/articles/61405/elife-61405-fig5-data4-v2.pdf
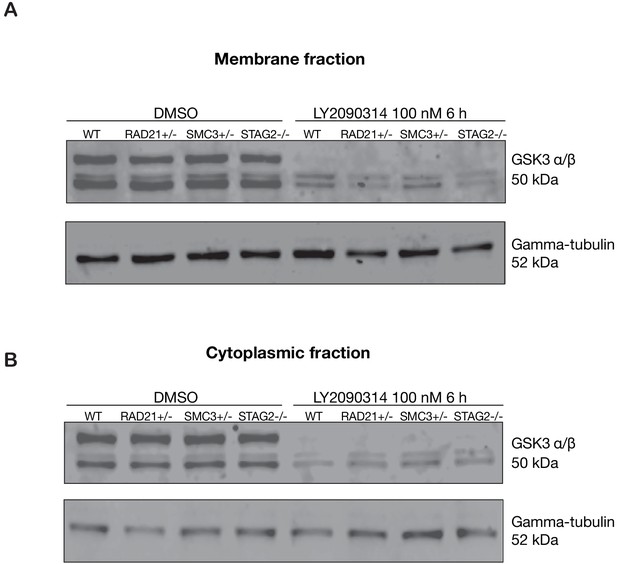
GSK3 levels are unaffected in cohesin-deficient MCF10A cells.
Anti-GSK3 immunoblot of membrane fraction from parental MCF10A (WT) or cohesin-deficient cells treated with either DMSO or 100 nM LY2090314. (A) Membrane fraction after 6 hr of treatment with DMSO, or with LY2090314. (B) Cytoplasmic fraction after 6 hr of treatment with DMSO, or with LY2090314. Untrimmed blots and molecular size markers are available for A,B in Figure 5—source data 3.
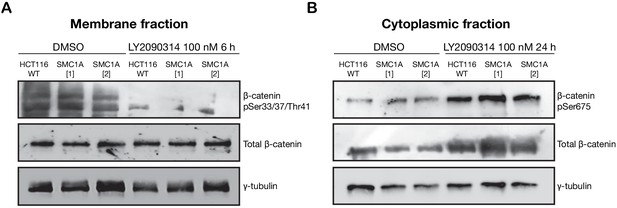
LY2090314-mediated WNT stimulation leads to increased β-catenin stabilization in SMC1A mutant HCT116 cells.
Unmodified HCT116 cells (WT), and cells modified to contain SMC1A mutations: SMC1A (1), c.2027A > G; SMC1A (2), c.2479 C > T, were treated with DMSO or LY2090314 at 100 nM for 24 hr. (A) Immunoblot of the membrane fraction of HCT116 wild type (WT) and SMC1A mutant-expressing HCT116 cells. The proteasome-targeted form of β-catenin phosphorylated at Ser33/37/Thr41 is slightly less abundant in the SMC1A mutants, and this phospho isoform is degraded in all cells following treatment with LY2090314. (B) Immunoblot of the cytoplasmic fraction shows increased basal level of phosphorylated β-catenin at Ser675 in the SMC1A mutant-expressing cells compared with HCT116 WT, and LY2090314 treatment markedly increased total β-catenin in the SMC1A mutants compared with HCT116 WT. Untrimmed blots and molecular size markers are available for A,B in Figure 5—source data 4.
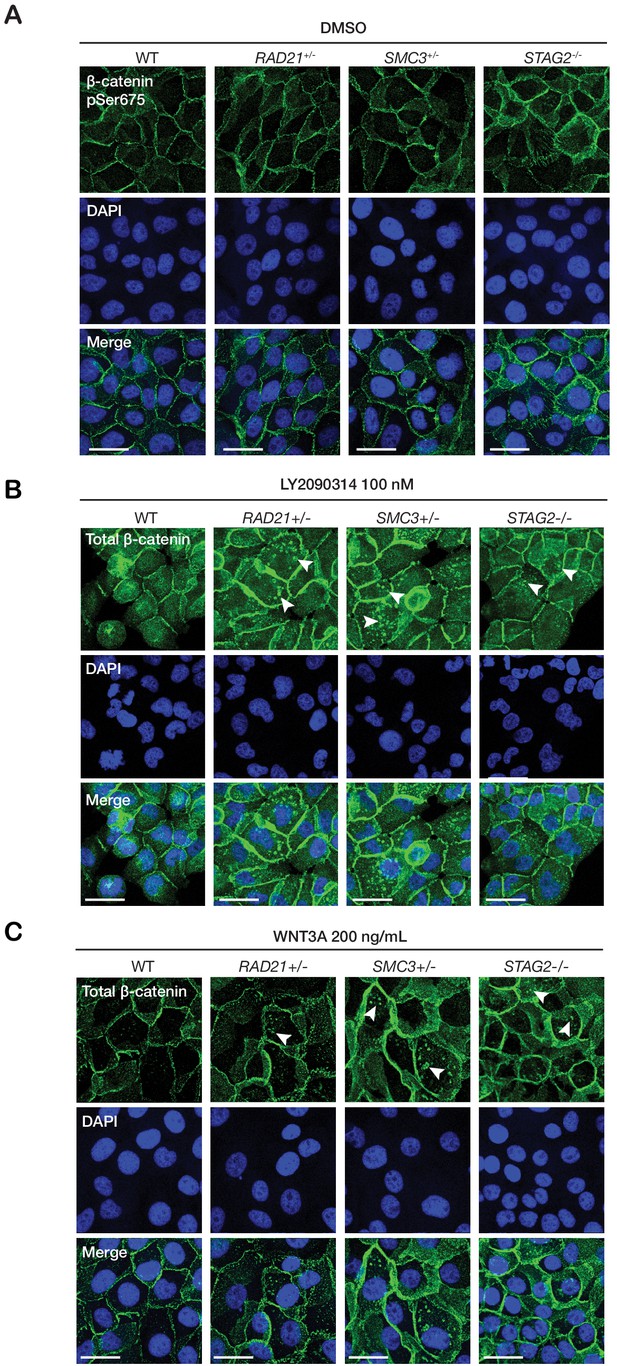
WNT3A phenocopies LY2090314-mediated β-catenin accumulation in the cytoplasm.
Immunofluorescence images show that (A) parental MCF10A (WT) or cohesin-deficient cells treated with 0.5% DMSO have no β-catenin puncta. In contrast, 24 hr of treatment with both (B) LY2090314 and (C) WNT3A leads to formation of β-catenin puncta in the cytoplasm of cohesin-deficient cells (arrows), but not in parental MCF10A (WT) cells. Scale bar = 25 µm.
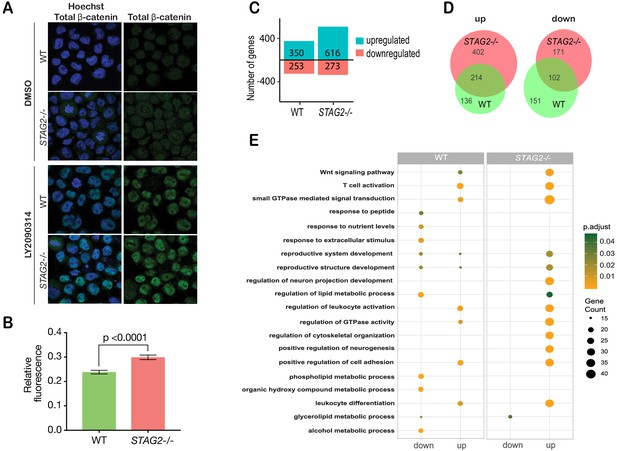
Cohesin-STAG2 mutant CMK cells show increased sensitivity to Wnt signaling.
(A) Immunofluorescence images showing slightly increased nuclear accumulation of β-catenin in STAG2-CMK cells (STAG2-/-) compared to parental (WT) following treatment with LY2090314 at 100 nM for 24 hr. (B) Quantification of nuclear total β-catenin in parental (WT) and STAG2-CMK cells (STAG2-/-) cells. Fluorescence of nuclear total β-catenin was determined relative to the nuclear area. Image J was used quantify cells from 10 different confocal fields. The graph depicts s.e.m. from analyses of 170–188 cell nuclei, and the p value was calculated using a student’s t test. (C) Histogram showing the number of genes upregulated or downregulated at FDR ≤ 0.05 upon WNT3A treatment in parental (WT) and STAG2-/- CMK cells. WNT3A stimulation was performed on three biological replicates, from three independent experiments. (D) Overlap of genes significantly upregulated or downregulated (FDR ≤ 0.05) upon WNT3A treatment between parental (WT) and STAG2-/- CMK cells. (E) Top enriched pathways (ranked by gene count) from the significantly downregulated and upregulated genes (FDR ≤ 0.05) following WNT3A treatment in STAG2-/- and parental (WT) CMK cells using the ClusterProfiler R package on the Gene Ontology Biological Process dataset modeling for both cell types (WT or STAG2-/-) and regulation pattern (up- or downregulation). Source data is available for C,D in Figure 6—source data 1.
-
Figure 6—source data 1
Gene expression data for Figure 6.
- https://cdn.elifesciences.org/articles/61405/elife-61405-fig6-data1-v2.xlsx
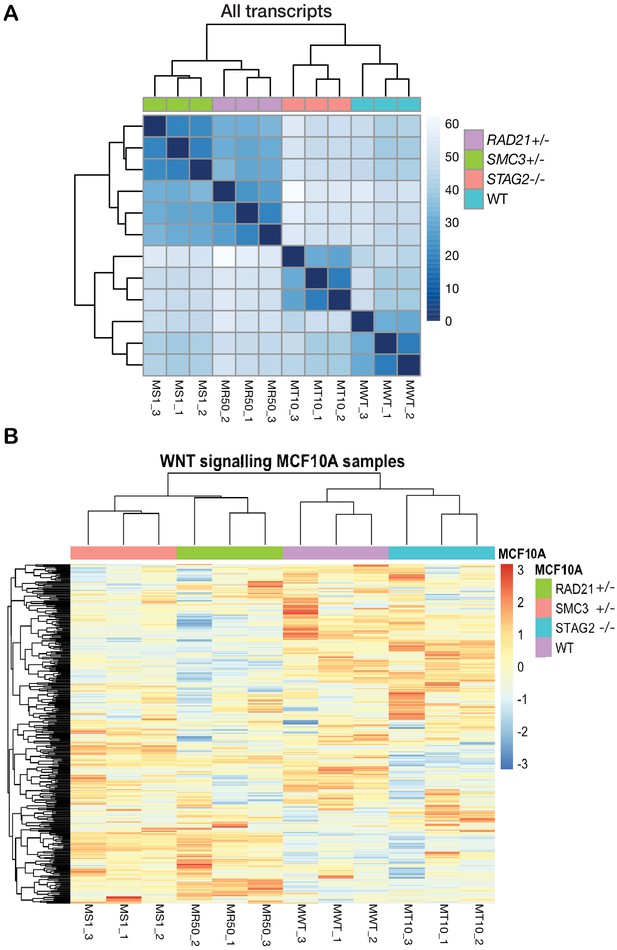
RNA-sequencing profiling of cohesin-deficient MCF10A cells.
(A) Whole genome correlation matrix of MCF10A cells. The dendrogram shows a common clustering of the biological replicates. There are two main clusters. The STAG2-/- (MT10_1–3) transcriptomes are more similar to the WT (MWT_1–3). SMC3+/- (MS_1–3) and RAD21+/- (MR50_1–3) populate the second cluster. (B) MCF10A gene expression profile of genes involved in WNT signaling pathways according to Gene Ontology annotation (GO:0016055). The heatmap shows the clustering of the scaled (z-score) normalized count (log2RPKM) for each replicate. RNA-sequencing data is available in the GEO database under GSE154086.
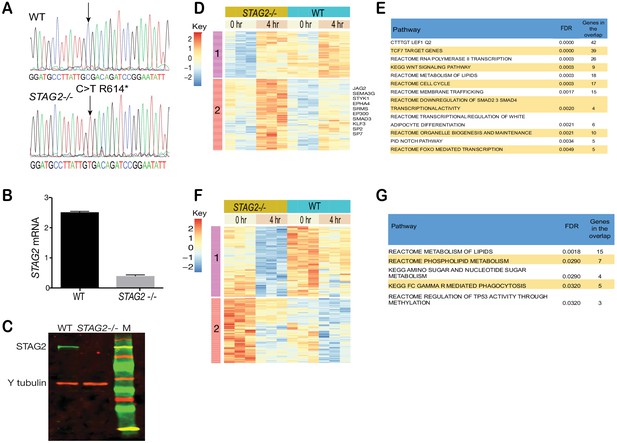
Enhanced sensitivity of cohesin-mutant CMK cells to Wnt stimulation.
(A) Sanger sequencing shows successful CRISPR-CAS9 editing introducing the STAG2 R614* C > T mutation site into CMK cells. (B) Reduced STAG2 mRNA and (C) protein in STAG2-/- CMK cells. (D) Heatmap showing log2 RPKM values of 402 genes upregulated only in STAG2 mutant cells (FDR ≤ 0.05) following 4 hr of WNT3A treatment. (E) GSEA analyses of the 402 upregulated genes. (F) Heatmap showing log2 RPKM values of 171 genes downregulated only in STAG2 mutant cells (FDR ≤ 0.05) following 4 hr of WNT3A treatment. (G) GSEA analyses of the 171 genes significantly downregulated only in the STAG2 mutant line. Gene lists are provided in Figure 6—source data 1.
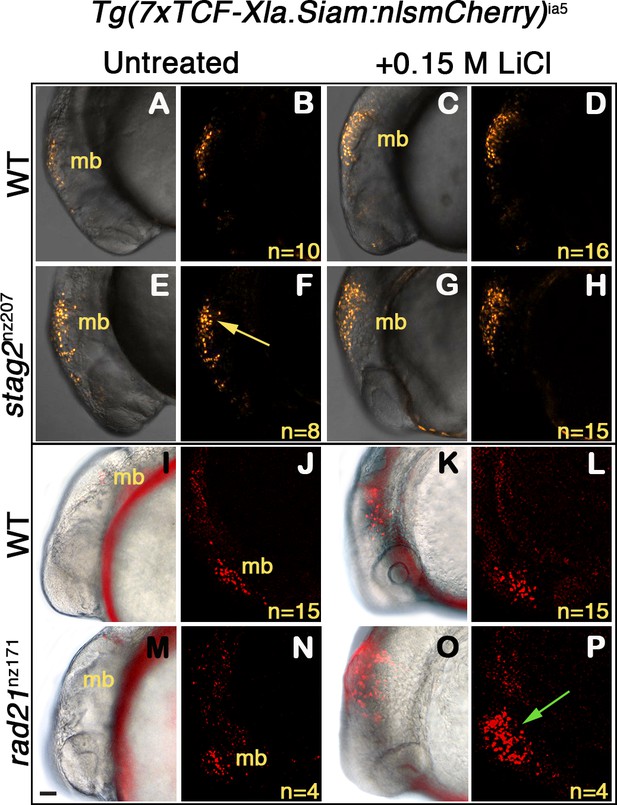
Zebrafish stag2b and rad21 cohesin mutants show increased sensitivity to Wnt signaling.
Wnt reporter Tg(7xTCF-Xla.Siam:nlsmCherry)ia5 control embryos, Tg(7xTCF-Xla.Siam:nlsmCherry)ia5;stag2bnz207 and Tg(7xTCF-Xla.Siam:nlsmCherry)ia5;rad21nz171 cohesin-mutant embryos were treated with 0.15 M LiCl from 4 hpf to 20 hpf. (A–H) max projections of 4 (10 μm) optical sections. (A,C,E,G) TD (transmitted light detector) images merged with confocal images. (B,D,F,H) confocal images alone. (I,K,M,O) Brightfield/fluorescent and (J,N,L,P) confocal images of the same embryos in I,K,M,O. (A–D) and (I–J) Tg(7xTCF-Xla.Siam:nlsmCherry)ia5 control embryos (WT) have low level fluorescence (Wnt reporter activity) in the midbrain that is increased following treatment. (E–H) Tg(7xTCF-Xla.Siam:nlsmCherry)ia5; stag2bnz207 embryos have elevated baseline levels of fluorescence (Wnt reporter activity) relative to controls (yellow arrow) with not much further increase upon LiCl treatment. (M–P) Tg(7xTCF-Xla.Siam:nlsmCherry)ia5; rad21nz171 cohesin-mutant embryos show enhanced Wnt reporter activity in the midbrain (green arrow) upon LiCl treatment, relative to controls. hpf, hours post-fertilization. mb, midbrain. Scale bar, 50 µm.
Tables
Significant inhibitors of all three cohesin-deficient clones.
| Compound | Rank 1° screen | Rank 2° screen | Target | Pathway | 1° differential AOC activity | ||
|---|---|---|---|---|---|---|---|
| RAD21+/- | SMC3+/- | STAG2-/- | |||||
| WAY-600 | 1 | 3 | mTORC1/2 | PI3K/AKT/mTOR | 0.30 | 0.37 | 0.32 |
| I-BET-762 | 2 | 8 | BET proteins | Epigenetics | 0.25 | 0.27 | 0.29 |
| LY2090314 | 3 | 6 | GSK3 | WNT | 0.25 | 0.22 | 0.25 |
| Vistusertib (AZD2014) | 4 | 1 | mTORC1/2 | PI3K/AKT/mTOR | 0.18 | 0.33 | 0.26 |
| P276-00 | 5 | 17 | CDK1/4/9 | Cell Cycle | 0.17 | 0.29 | 0.28 |
| MK-8745 | 6 | 22 | Aurora A | Cell Cycle | 0.18 | 0.30 | 0.26 |
| Ethacridine lactate | 7 | 28 | Anti-infection | Microbiology | 0.18 | 0.25 | 0.27 |
| CUDC-101 | 8 | 36 | EGFR, HDAC, HER2 | Epigenetics | 0.16 | 0.25 | 0.24 |
| Dabrafenib (GSK2118436) | 9 | 9 | BRAF | MAPK | 0.18 | 0.27 | 0.18 |
| SAR131675 | 10 | 34 | VEGFR3 | Protein Tyrosine Kinase | 0.15 | 0.17 | 0.33 |
| ZM 447439 | 11 | 41 | Aurora A/B | Cell Cycle | 0.16 | 0.22 | 0.24 |
| Gitoxigenin Diacetate | 12 | 32 | NA | Other | 0.17 | 0.24 | 0.20 |
| UNC669 | 13 | 62 | Epigenetic Reader Domain | Epigenetics | 0.18 | 0.23 | 0.20 |
| 4-Phenylbutyric Acid | 14 | 29 | Endoplasmic reticulum stress | Other | 0.20 | 0.17 | 0.20 |
| Ipatasertib (GDC-0068) | 15 | 12 | AKT | PI3K/AKT/mTOR | 0.21 | 0.19 | 0.16 |
| VX-702 | 16 | 43 | P38 MAPK | MAPK | 0.15 | 0.19 | 0.21 |
| RVX-208 | 17 | 58 | BET proteins | Epigenetics | 0.16 | 0.20 | 0.17 |
| Dihydroergotamine mesylate | 18 | 49 | NA | Other | 0.16 | 0.19 | 0.16 |
| Olaparib | 351 | PARP1/2 | DNA Damage | 0.11 | 0.13 | 0.22 | |
-
Table 1—source data 1
Compounds with growth inhibitory activity (AOC) ranked to produce Table 1.
- https://cdn.elifesciences.org/articles/61405/elife-61405-table1-data1-v2.xlsx
-
Table 1—source data 2
Compounds effective in the secondary screen ranked to produce Table 1.
- https://cdn.elifesciences.org/articles/61405/elife-61405-table1-data2-v2.xlsx
Additional files
-
Supplementary file 1
List of sgRNA sequences and PCR primers.
- https://cdn.elifesciences.org/articles/61405/elife-61405-supp1-v2.docx
-
Supplementary file 2
TCGA analysis of STAG2 mutant vs wild type cancers.
- https://cdn.elifesciences.org/articles/61405/elife-61405-supp2-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/61405/elife-61405-transrepform-v2.docx





