Mapping endothelial-cell diversity in cerebral cavernous malformations at single-cell resolution
Figures
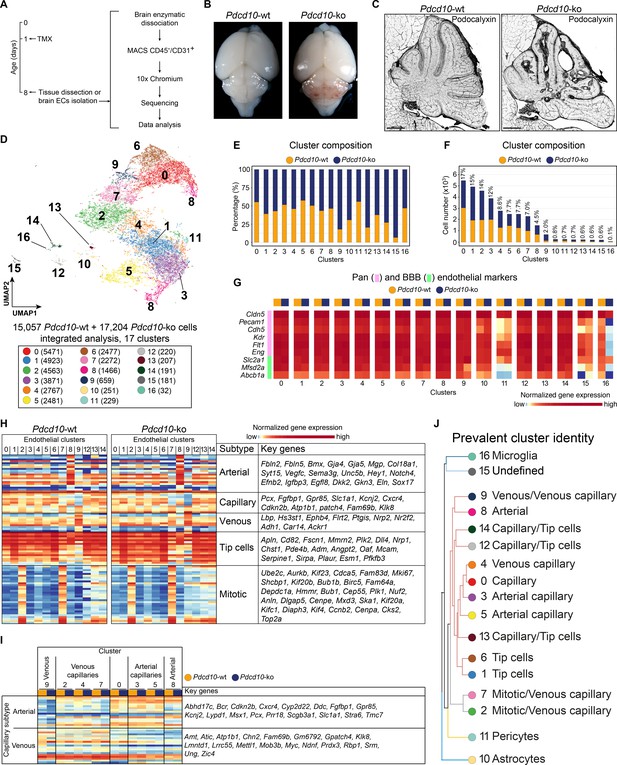
scRNA-sequencing of Pdcd10-wt and Pdcd10-ko endothelial cells.
(A) Experimental scheme (see Materials and methods for details). (B) Representative photographs of Pdcd10-wt (left) and Pdcd10-ko (right) whole brains at P8. (C) Representative confocal microscopy of the vasculature of Pdcd10-wt (left) and Pdcd10-ko (right) cerebella at P8, stained for Podocalyxin (black; see also Figure 1—figure supplement 1). Scale bars: 1 mm. (D) UMAP plot showing detected cell subpopulations in the Pdcd10-wt and Pdcd10-ko integrated analysis. The total numbers of cells within each cluster are shown in brackets in the color legend (bottom panel). (E) Plot of the percentages of Pdcd10-wt (orange) and Pdcd10-ko (blue) cells in each of the cluster. (F) Plot of the numbers of Pdcd10-wt (orange) and Pdcd10-ko (blue) cells in each cluster. The percentages of total cells in each cluster (%) is reported above each bar. (G) Heatmap of the selected pan- (pink) and blood–brain barrier- (BBB; green) endothelial cell markers (normalized expression shown; see Materials and methods) for the Pdcd10-wt and Pdcd10-ko cells in each cluster. (H) Heatmap of the selected endothelial cell subtype markers (from top to bottom, as indicated: arterial, capillary, venous, tip cells, mitotic), to show the normalized expression levels of the Pdcd10-wt (left) and Pdcd10-ko (right) cells in each cluster (see also Supplementary file 1). For each subtype, the key genes are listed accordingly (top-to-bottom). (I) Heatmap of normalized expression of arterial and venous capillary markers in the capillary and mitotic/capillary clusters (C0, C2, C3, C4, C5, C7). The venous (C9; left) and arterial (C8; right) clusters are reported for reference. The Pdcd10-wt (orange) and Pdcd10-ko cells (blue) are shown separately. Key genes are listed according to the top-to-bottom order in the heatmap. (J) Summary of the prevalent identities of the 17 clusters based on endothelial cell subtype marker expression as in (H, I) and Figure 1—figure supplement 1. The dendrogram of the hierarchical clustering is shown on the left. Non/mixed-ECs clusters (C10, C11, C15, C16) show early segregation. Arterial/venous (C8, C9), capillary (C0, C3, C4, C5), tip (C1 and C6) and mitotic/capillary (C2, C7) cells segregate as distinct groups of the final branches. Clusters 12, 13, and 14 show features of both capillaries and tip cells, but only C12 and C14 segregate on neighbor final branches.
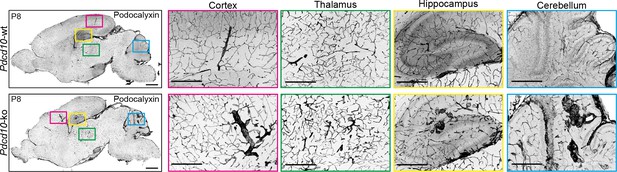
Representative confocal microscopy of Pdcd10-wt and Pdcd10-ko brains at P8 immunostained for Podocalyxin.
Higher magnification images of different brain regions are shown on the right (color coded from left panel) for cortex, thalamus, hippocampus, and cerebellum. Scale bars: 1 mm (main panel left), 500 μm (magnifications).
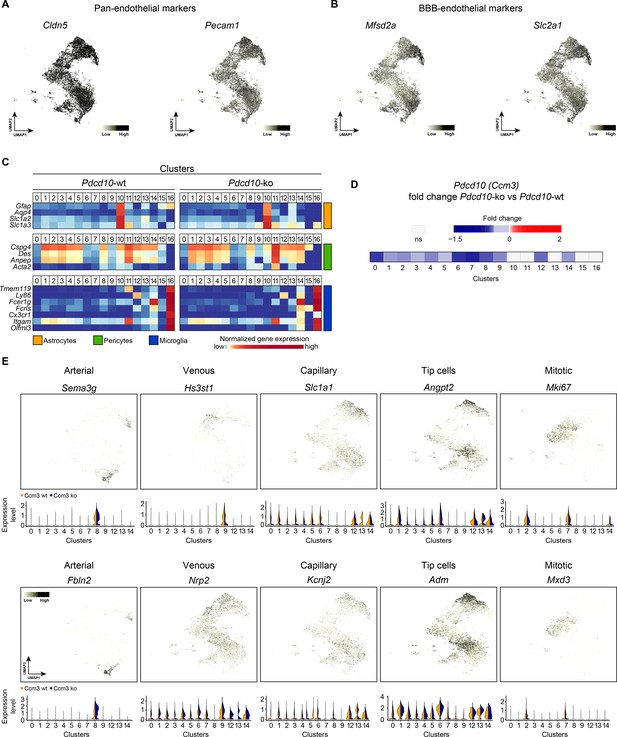
Expression of selected markers of endothelial and contaminant cells in the identified clusters.
(A, B) UMAP non-linear dimensional reduction method for the 32,261 cells examined (Pdcd10-wt, 15,057; Pdcd10-ko, 17,204), color coded for expression levels of pan-endothelial markers Cldn5 and Pecam1 (A) and blood–brain barrier (BBB) endothelial markers Mfsd2a, Slc2a1 (also known as Glut1) (B). Considering jointly Pdcd10-wt and Pdcd10-ko cells in the different clusters, the expression of the endothelial markers appears homogenously distributed in the different clusters. (C) Heatmap (for normalized expression; see Materials and methods) of selected contaminant markers (indicated left) for astrocytes, pericytes and microglia in Pdcd10-wt and Pdcd10-ko cells of each cluster (0–16) (see also Supplementary file 1). (D) Heatmap of average logFC (p<0.05, Pdcd10-ko vs. Pdcd10-wt) for Pdcd10 expression in the 17 clusters identified (0–16). (E) UMAP for the 32,261 cells examined (Pdcd10-wt, 15,057; Pdcd10-ko, 17,204), color coded for expression levels of the indicated EC subtype markers, for (left to right) arterial, venous, capillary, tip and mitotic cells. For each UMAP, the corresponding violin plots of Pdcd10-wt (orange) and Pdcd10-ko (blue) cells in each endothelial cluster is shown.
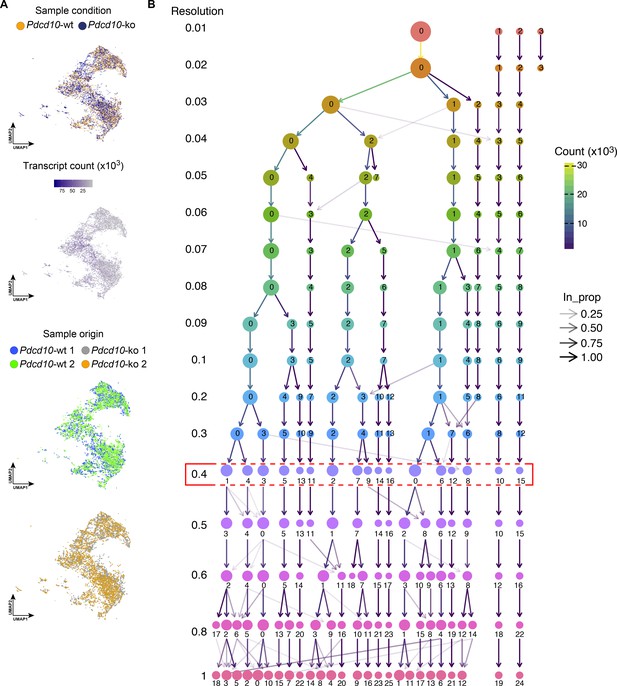
Sequencing quality control and clustering tree of examinedPdcd10-wt andPdcd10-ko ECs.
(A) UMAP non-linear dimensional reduction method for the 32,261 cells examined (Pdcd10-wt, 15,057; Pdcd10-ko, 17,204), color coded, from top to bottom, for sample conditions, transcript counts (number of unique molecular identifiers) and sample origin. Top: Homogenous distribution of Pdcd10-wt (orange dots) and Pdcd10-ko (blue dots) cells shown among the clusters. Middle: Number of unique transcript sequences shown for each cell. Bottom: Homogenous distribution of cells isolated from the four mice used in the analysis (blue and green; Pdcd10-wt, gray and orange: Pdcd10-ko). (B) Clustering tree of examined Pdcd10-wt and Pdcd10-ko endothelial cells. Results from clustering using Seurat with resolution parameters from 0.01 to 1. At a resolution of 0.01, we see the formation of four clusters, one of which continues to split up to form new clusters as the resolution increased. Resolution 0.09–0.1 and resolution 0.4–0.5 are two stable regions in this tree where no additional sub-branching happened in between and the number of clusters stayed the same. After resolution 0.5, many low in-proportion (in_prop) edges ad new clusters with multiple parent clusters showing up, indicating cluster instability. Red box highlights the resolution (0.4) selected for further analysis.
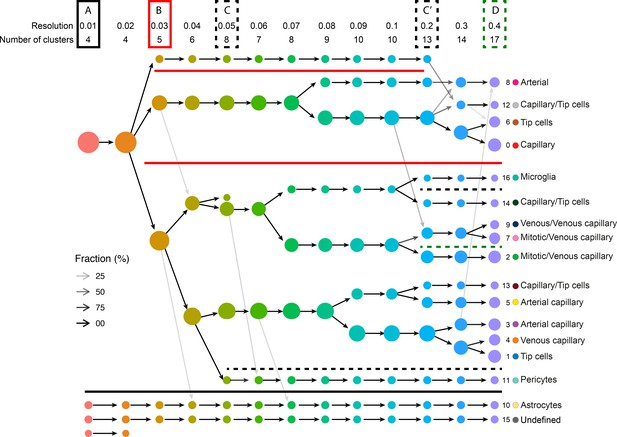
Clustering tree of examined Pdcd10-wt and Pdcd10-ko endothelial cells with overlaid prevalent cluster identity.
Resolutions from 0.01 to 0.4, which is part of the clustering tree shown in Figure 1—figure supplement 3. Box A (solid black line): at resolution 0.01, two of the four contaminant cell clusters (C10 and C15) are formed and remain unchanged to resolution 0.4. Box B (solid red line): at resolution 0.03, two major lineages are branched. It is clear at resolution 0.4 that arterial and venous clusters (C8 and C9, respectively) are derive from these two distinct lineages, so are the two major tip cell clusters (C1 and C6). It suggests that C1 and C6 are not the result of over-clustering, but rather represent two distinct tip cell sub-populations. Box (C & C’) (dashed black line): at resolution 0.05 (C) and 0.2 (C’), the remaining two contaminant cell clusters (C11 and C16) are formed and remain unchanged to resolution 0.04. Box D (dashed green line): at resolution 0.4, the mitotic/venous capillary and the venous/venous capillary clusters (C7 and C9, respectively) were formed from the same parent cluster.
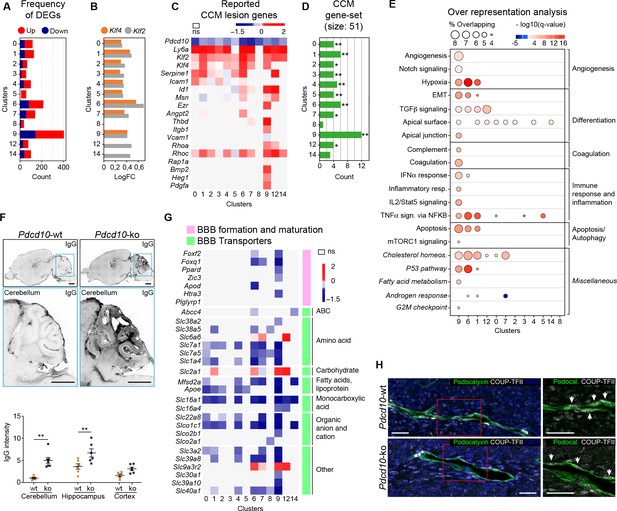
Pdcd10 deletion induces specific transcriptional profiles in distinct endothelial cell subpopulations.
(A) Numbers of significant differentially expressed genes (DEGs) (padj <0.05) in each cluster, showing up-regulation (red) and down-regulation (blue). (B) Average log fold changes of Klf4 (orange) and Klf2 (gray) expression in each cluster (Pdcd10-ko vs. Pdcd10-wt; padj <0.05). (C) Heatmap of selected known CCM lesion markers, as average logFC (padj <0.05) of Pdcd10-ko versus Pdcd10-wt cells (see also Supplementary file 2). (D) Enrichment of CCM-associated genes (source: Rare Diseases GeneRIF Library) among DEGs (Pdcd10-ko vs. Pdcd10-wt) for each cluster (see Materials and methods). X-axis indicates the number of DEGs identified as CCM-associated genes in each cluster. Asterisks show the significance of enrichment: *p<0.05; **p<0.01. (E) Over-representated Molecular Signatures Databases hallmark gene sets in DEGs (average |logFC| ≥ 0.3; padj <0.05, see Materials and methods). The sizes of the dots reflect the proportion (%) of overlap between the DEGs and the reference gene sets, while the intensities of the colors show the -log10(q-value), color coded in red for up-regulated gene sets and in blue for down-regulated gene sets. Gene sets in italics have not been described previously for CCM. (F) Representative confocal microscopy of IgG leakage in Pdcd10-wt (left) and Pdcd10-ko (right) brain sections. Bottom images: higher magnification of the cerebellum (light blue boxed areas at top). Higher magnification of the hippocampus and cortex are shown in Figure 2—figure supplement 1. Scale bars: 1 mm. Bottom panel: Quantification of IgG leakage (mean ± SEM; **p<0.01; ANOVA followed by Sidak multiple comparisons). Pdcd10-wt, n = 6; Pdcd10-ko, n = 7. (G) Heatmap of log fold expression changes of selected genes (padj <0.05) important for BBB formation and maturation (pink), and typical BBB transporters (green) between Pdcd10-ko and Pdcd10-wt. (H) Representative confocal microscopy of the venous marker COUP-TFII (encoded by the Nr2f2 transcript, white), Podocalyxin (pan-endothelial, green) and DAPI (blue) of a Pdcd10-wt vessel (top) and a Pdcd10-ko lesion (bottom), both in the cerebellum. Arrows, COUP-TFII–positive endothelial nuclei. Scale bars: 25 μm.
-
Figure 2—source data 1
Source data file for Figure 2F.
- https://cdn.elifesciences.org/articles/61413/elife-61413-fig2-data1-v1.xlsx
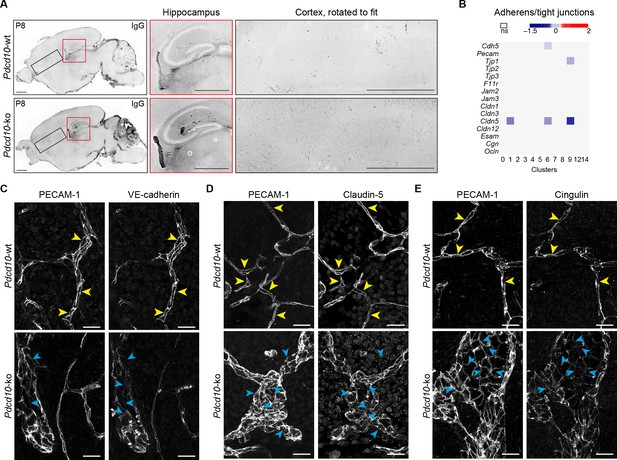
Pdcd10 deletion increases vessel permebility and impairs endothelial cell-to-cell junction organization.
(A) Representative confocal microscopy for IgG leakage in Pdcd10-wt (top) and Pdcd10-ko (bottom) brain sections, as in Figure 2F. Higher magnifications of hippocampus (red box) and cortex (black box) are shown on the right. Quantification of IgG leakage is shown in Figure 2F. (B) Heatmap of average logFC (Pdcd10-ko vs. Pdcd10-wt; p<0.05) of junctional molecule expression in the endothelial clusters (as indicated). (C–E) Representative confocal microscopy of adherens and tight junction markers VE-cadherin (C; encoded by Cdh5; right), Claudin-5 (D; right) and Cingulin (E; right) in Pdcd10-wt vessels and Pdcd10-ko lesions in the cerebellum. PECAM-1 staining is shown on left of each panel. Yellow arrowheads, regular junctions in Pdcd10-wt vessels; light blue arrows, altered junctions in Pdcd10-ko. Scale bars: 25 μm.
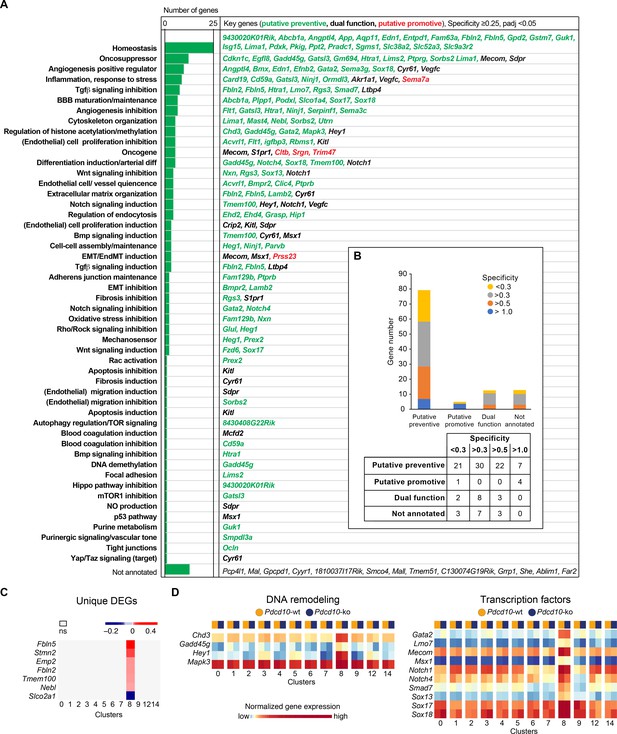
Unique functional markers suggest arterial Pdcd10-ko ECs may be protected from forming cavernomas.
(A) Annotation of C8 unique marker genes. 111 unique marker genes were identified using a log fold change (specificity) cutoff of 0.25 and padj <0.05. Mouse Genome Informatics (http://www.informatics.jax.org/) and PubMed (https://pubmed.ncbi.nlm.nih.gov/) were used for manual annotation (keywords: ‘gene name AND entothelial cells’ or ‘gene name’ if the previous search gave zero results). Based on the functions of these genes and the deregulation of these functions reported in Pdcd10-ko lesions, we tentatively categorized these unique marker genes into ‘preventive’ (green), ‘promotive’ (red) and ‘dual-function’ (black) (See details of annotation in Supplementary file 5 and 6). Here, 13 unique marker genes either lack clearly defined functionality or have non-EC related functions and could not be annotated. (B) Statistics of annotated unique marker genes in C8. Each gene was counted only once, although it can appear in more than one category in (A). (C) Heatmap of log fold changes of uniquely differentially expressed genes (unique DEGs) in C8. Unique DEGs were identified using thresholds: average |logFC| > 0.2 and padj <0.05. (D) Heatmap of normalized expression levels of the selected C8 unique marker genes (left: regulators of DNA remodeling; right: transcription factors) in the Pdcd10-wt and Pdcd10-ko cells in each cluster.
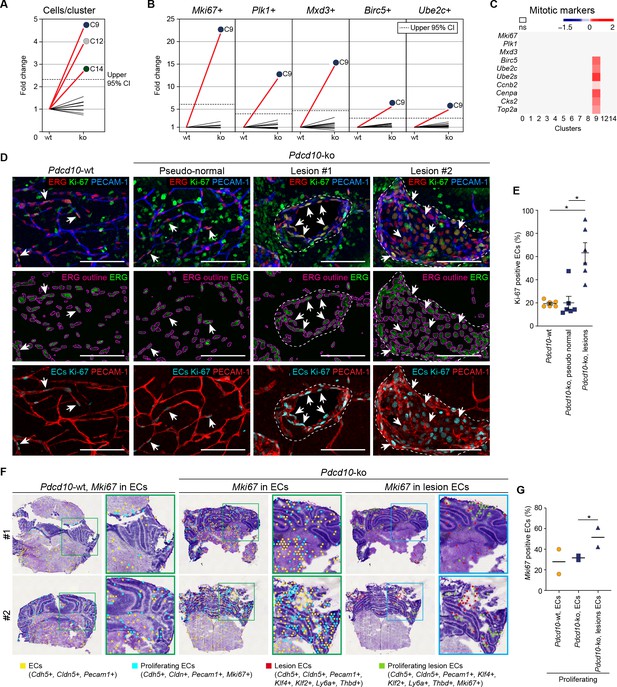
Pdcd10-ko cells of C9 express the same top mitotic phenotype as overt cavernomas.
(A) Plot of fold-change in numbers of Pdcd10-ko versus Pdcd10-wt cells in each cluster. (B) Plots of fold-change in the percentages (Pdcd10-ko vs. Pdcd10-wt) of cells positive for Mki67, Plk1, Mxd3, Birc5, and Ube2c in each cluster (as indicated). Dashed lines in (A) and (B) show upper limit of 95% confidence interval (CI), calculated from the mean fold-changes among the clusters. Red lines, clusters with fold-change >95% CI upper limit. (C) Heatmap of average logFC (padj <0.05) in the Pdcd10-ko vs. Pdcd10-wt for selected mitotic markers (as indicated). (D) Representative confocal microscopy of Pdcd10-wt and Pdcd10-ko mouse cerebellum at P8. Upper panels: Immunostained for PECAM-1 (pan-endothelial, blue), ERG (pan-endothelial nuclei, red) and Ki-67 (encoded by Mki67 transcript; mitotic nuclei, green). Central panels: Immunostained for ERG (green) with outlines of the segmented nuclei (magenta). Lower panels: Results of the filtering procedure showing EC-specific immunostaining of Ki-67 (ECs Ki-67, light blue) and PECAM-1 (red) (see Materials and methods). Arrows, mitotic ECs. Scale bars: 100 μm. (E) Quantification of mitotic ECs in the cerebellum, as normal vessels in Pdcd10-wt, Pdcd10-ko pseudo-normal vessels, and Pdcd10-ko lesions (mean ± SEM; *p<0.01; ANOVA followed by Tukey’s multiple comparisons). Pdcd10-wt, n = 6; Pdcd10-ko, n = 6. (F) Visium analysis showing spots positive for ECs (yellow) and lesion EC markers (red), identified as indicated, with co-expression of Mki67. Proliferating ECs (light blue) and proliferating lesion ECs (green) containing spots are shown. Higher magnifications of boxed areas are also shown. (G) Quantification of the percentages (%) of proliferating ECs (light blue) and proliferating lesion ECs (green) containing spots as in (F) (Mean; *p<0.01 Fisher’s exact test).
-
Figure 4—source data 1
Source data file for Figure 4E.
- https://cdn.elifesciences.org/articles/61413/elife-61413-fig4-data1-v1.xlsx
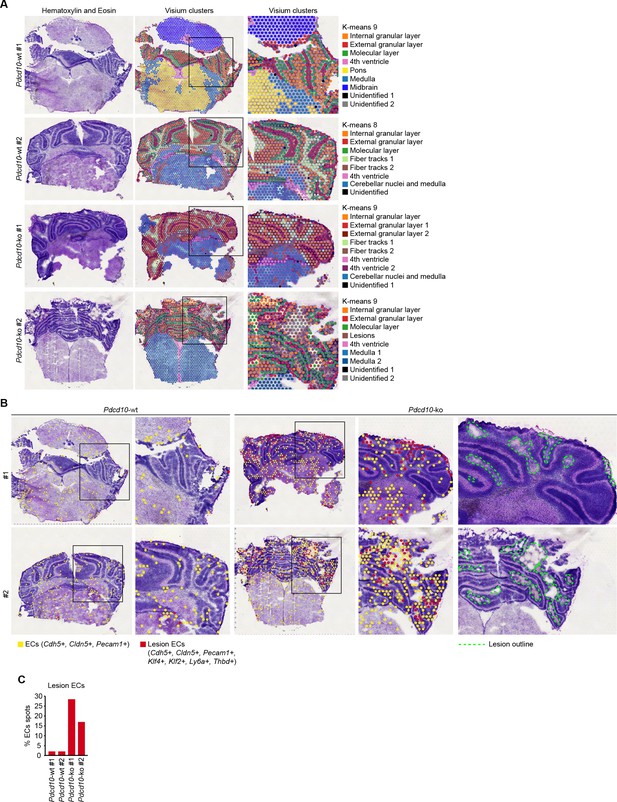
Validation of Visium analysis and identification of spots co-expressing endothelial and CCM lesion markers.
(A) Quality control of Visium analysis showing identification of the distinct areas of the representative Pdcd10-wt and Pdcd10-ko brain sections, as indicated. Identification was carried out using the Allen Brain Atlas (http://atlas.brain-map.org/) (see Material and methods for details). With the exception of Pdcd10-wt mouse #1 (second row), 9 (K-means) Visium clusters were necessary to obtain correct identification. Color coding of Visium clusters is shown superimposed on hematoxylin and eosin staining. (B) Representative Visium analysis for positive spots for ECs (yellow) and lesion ECs (red), identified at low (left) and high magnification (boxed areas), as indicated. Green dashed line, lesions in hematoxylin and eosin staining (right panel). (C) Quantification of proportions (%) of endothelial cell positive spots co-expressing lesion endothelial cell markers, as illustrated in (B).
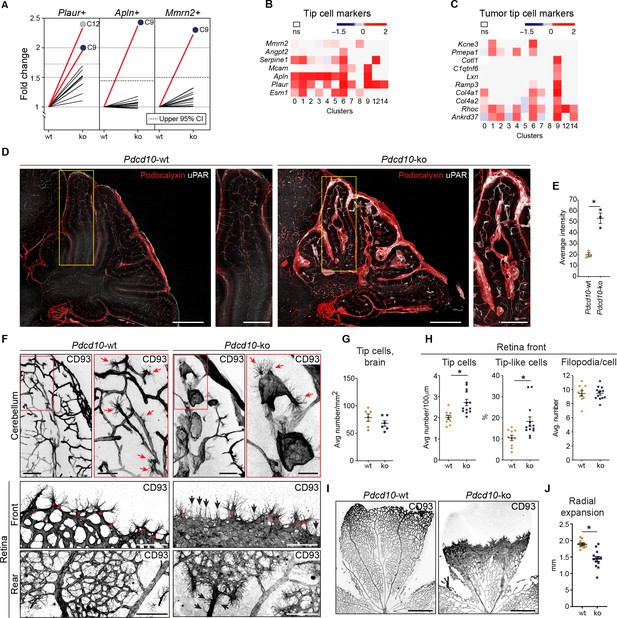
Pdcd10-ko cells widely enhance the tip cell phenotype with greatest expression for cluster C9.
However, such tip cell phenotype is functionally defective. (A) Plots of the fold-changes in the proportions (Pdcd10-ko vs. Pdcd10-wt) of cells positive for Plaur, Apln, and Mmrn2 in each cluster. Dashed line shows upper limit of 95% confidence interval (CI), calculated on the mean fold-change among the clusters. Red lines highlight clusters with fold-changes > 95% CI upper limit. (B) Heatmap of average logFC (padj <0.05) for Pdcd10-ko vs. Pdcd10-wt for selected tip cell markers (as indicated). (C) Heatmap of average logFC (padj <0.05) for Pdcd10-ko vs. Pdcd10-wt for selected tumor tip cell markers (as indicated). (D) Representative confocal microscopy of Pdcd10-wt and Pdcd10-ko mouse cerebellum at P8, immunostained for Podocalyxin (pan-endothelial, red) and uPAR (tip cell marker, encoded by Plaur, white). Scale bars: main, 500 μm; magnifications, 250 μm. (E) Quantification of uPAR staining as in (D) (mean ± SEM; *p<0.01; unpaired t-test). Pdcd10-wt, n = 3; Pdcd10-ko, n = 3. (F) Representative confocal microscopy of Pdcd10-wt and Pdcd10-ko mouse cerebellum (top) and retina (bottom) at P8, immunostained for CD93 (pan-ECs membrane and filopodia, at P8). Retina vessels are shown for the migrating front (Front) and proximal to the optic nerve (Rear). Red arrows and asterisks (in magnifications), tip cells (filopodia-rich cells); black arrows, tip-like cells at the migrating front in the Pdcd10-ko retina. Tip-like cells were present in the vein proximal to the optic nerve (Rear, black arrows) exclusively for the Pdcd10-ko. Scale bars: main, 100 μm. magnifications, 50 μm. (G) Quantification of tip cells in the cerebellum (mean ± SEM; ns; unpaired t-test). Pdcd10-wt, n = 6; Pdcd10-ko, n = 6. (H) Quantification of tip cells and tip-like cells at the retina front, for tip cell density (mean number/100 μm; left), proportion of tip-like cells (%; middle), and number of filopodia/cell (mean ± SEM; right; *p<0.01; unpaired t-test with Welch’s correction or Mann-Whitney for tip-like cells). Pdcd10-wt, n = 10; Pdcd10-ko, n = 13. (I) Representative lower magnification images of whole-mount retina preparations immunostained for CD93. Scale bars: 500 μm. (J) Quantification of radial expansion (mm; mean ± SEM; *p<0.01; unpaired t-test). Pdcd10-wt, n = 10; Pdcd10-ko, n = 13.
-
Figure 5—source data 1
Source data file for Figure 5E, G, H, and J.
- https://cdn.elifesciences.org/articles/61413/elife-61413-fig5-data1-v1.xlsx
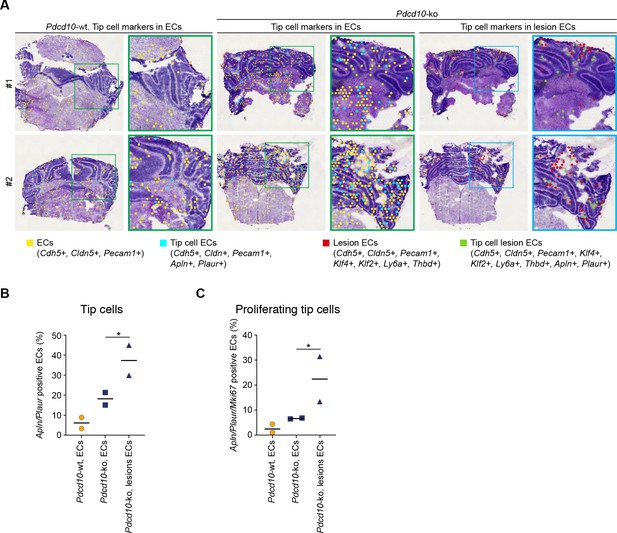
Lesion endothelial cells co-express mitotic and tip cell markers.
(A) Representative Visium analysis of endothelial cell (yellow) and lesion endothelial cell marker (red) positive spots, identified as indicated, showing co-expression of tip cell markers Plaur and Apln in Pdcd10-wt and Pdcd10-ko brains. Spots containing tip cell ECs (orange) and tip cell lesion ECs (green) are shown at low (left panels) and high (boxed areas) magnification. (B, C) Quantification of the proportions (%) of spots containing tip cell ECs (orange) and tip cell lesion ECs (green) (B), as illustrated in (A), and co-expressing the proliferation marker Mki67 (C) (Mean; *p<0.01; Fisher’s exact test).
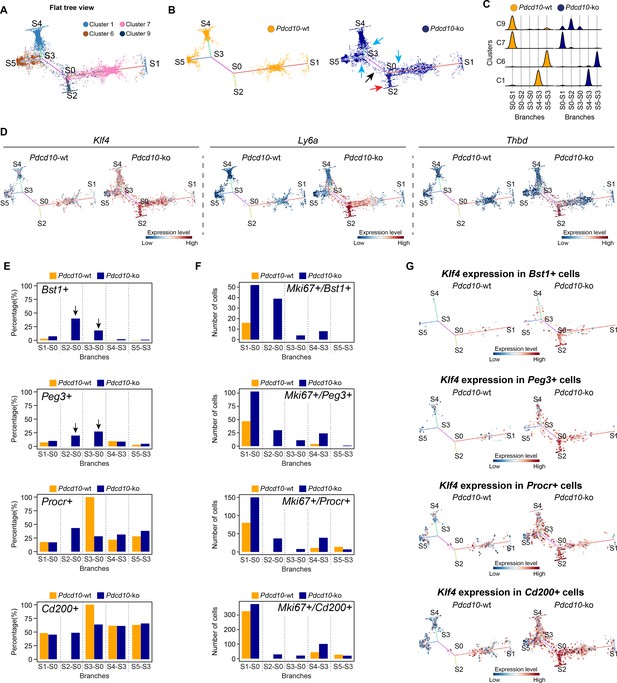
STREAM trajectory analysis shows Pdcd10-ko venous cluster 9 concentrated in a distinctive branch with mitotic and abnormal tip cell functions.
(A) Flat tree view of the STREAM trajectory inferred for Pdcd10-wt and Pdcd10-ko cells of the most affected clusters (C1, C6, C7, C8), color coded as indicated. Branching and end points are numbered sequentially (S0, S1, S2, S3, S4, S5) and each branch is named after the points connected (S0–S1, S0–S2, S4–S3, S3–S0, S5–S3). For corresponding three-dimensional views of the STREAM trajectories see Figure 6—figure supplement 1A and B. (B) Flat tree view of the trajectory as in (A) showing separately Pdcd10-wt (orange) and Pdcd10-ko (blue) cells. Red and black arrows, Pdcd10-ko specific branches (S0–S2, S3–S0, respectively). Light-blue arrows, Pdcd10-ko cells showing altered distributions within the other branches. (C) Distributions of Pdcd10-wt (orange) and Pdcd10- ko (blue) cells according to cluster, for the five branches of the trajectory. Pdcd10-ko cells of Cluster 9 (top row, blue peaks) show the most heterogeneous distributions with the cells in branches S0–S1, S0–S2, and S3–S0. (D) Flat tree view showing expression levels in Pdcd10-wt (left) and Pdcd10-ko (right) cells for three representative lesion markers: Klf4, Ly6a, and Thbd. (E) Plots showing the percentage of cells positive for the progenitor cell markers Bst1, Peg3, Procr, and Cd200 (as indicated) for Pdcd10-wt (orange) and Pdcd10-ko (blue) in each branch of the trajectories in (A) and (B). See Materials and Methods for definition of positive cells. (F) Plots showing numbers of cells double positive for Mki67 and for indicated progenitor cell markers for Pdcd10-wt (orange) and Pdcd10-ko (blue) in each branch of the trajectories in (A) and (B). See Materials and methods for definition of positive cells. (G) Flat tree view showing expression levels of Klf4 for Pdcd10-wt (left) and Pdcd10-ko (right), for each of the progenitor cell subpopulations investigated. In (D and G) dots are color coded for expression levels, as blue, low; red, high.
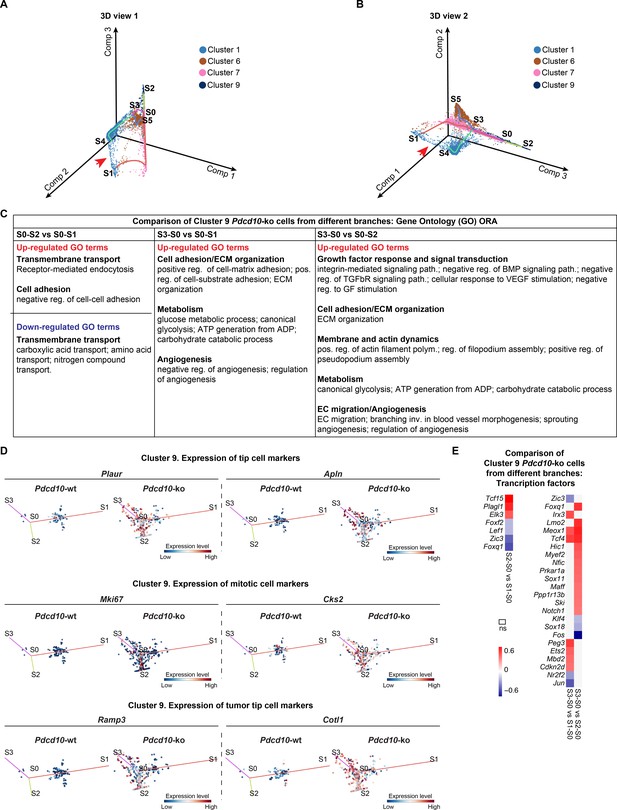
STREAM trajectory analysis of C9 Pdcd10-ko cells from different branches for the expression of functions (GO) and tip, mitotic and transcription factor genes.
(A, B) STREAM trajectory inferred for Pdcd10-wt and Pdcd10-ko cells of the most affected clusters of C1, C6, C7, and C8, color coded as indicated. Two three-dimensional views (A, B) of the trajectories generated by STREAM are shown. Red arrows, small subset of cells belonging to C1 that bridge the ends of the S0–S1 and S4–S3 branches. Comp: principal components. (C) Pdcd10-ko cells of cluster 9 were distributed over different branches of the trajectory (S3–S0, S0–S2, S0–S1), as shown in Figure 6B,C. Multiple comparisons were performed, as indicated, to calculate differentially expressed genes between these three branches. Over-representation analysis (ORA) of the up-regulated (red text) and down-regulated (blue text) differentially expressed genes (average |logFC| ≥ 0.25, padj <0.05) for the Gene Ontology (GO) biological processes. GO-terms were functionally grouped as shown in bold text. (D) Flat tree view showing expression levels for Pdcd10-wt (left) and Pdcd10-ko (right) cells from cluster nine only, for representative tip, mitotic, and tumor tip cell markers. Dots are color coded for expression levels (blue, low; red, high). (E) Heatmaps of average logFC (p<0.05) for transcription factors calculated from the comparisons in (C). Left: comparison of S0–S2 versus S0–S1; right: comparison of S3–S0 versus S1-S0 (left) and S2-S0 (right).
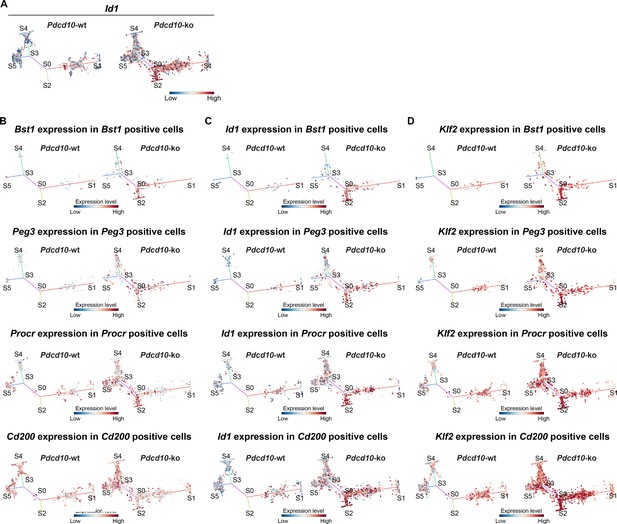
STREAM trajectory analysis of resident endothelial progenitor cells from different branches for the expression of the CCM-related genes Id1 and Klf2.
(A) Flat tree views showing expression levels for Pdcd10-wt (left) and Pdcd10-ko (right) cells of Id1. (B–D) Flat tree views showing expression levels for Pdcd10-wt (left) and Pdcd10-ko (right) cells for each progenitor subpopulation investigated (B-D, top to bottom: Bst1+, Peg3+, Procr+, Cd200+) for the expression levels of the corresponding identifier genes (B), and for Id1 (C) and Klf2 (D) in each of the progenitor subpopulations investigated. Dots are color coded for expression levels (blue, low; red, high).
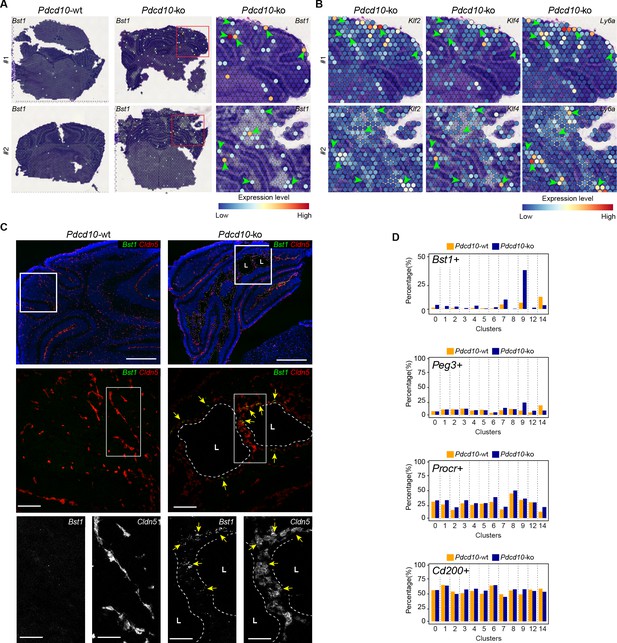
Venous-resident Pdcd10-ko endothelial progenitor cells support the formation of cavernomas.
(A) Expression levels of Bst1 as measured according to Visium Spatial Transcriptomics. All of the sequenced dots are shown, color coded for expression, as blue, low; red, high. Negative spot outlines are shown without any fill color to allow the visualization of underlying H and E staining (dark and light violet). Both complete sections and magnified boxed areas are shown. Arrowheads, examples of positive spots. (B) Expression levels of representative lesion markers (as indicated) from Visium analysis. For each gene and for each sample, the same magnified areas are shown as in (A). Arrowheads, same spots as indicted in (A). (C) Representative in-situ hybridization for Bst1 (green) and Cldn5 (red) using the RNA-Scope assay for Pdcd10-wt and Pdcd10-ko mouse cerebellum at P8. Three magnifications are shown (increasing top to bottom). Top: DAPI staining (blue) is also shown. Bottom: Cldn5 and Bst1 shown separately (white). Yellow arrows points to colocalization of Cldn5 and Bst1 signals. Lesion lumens (L) are outlined by white dashed lines. Scale bars: 500 μm (left), 100 μm (middle), 50 μm (right). (D) Plots of percentages (%) of cells positive for progenitor cell markers Bst1, Peg3, Procr, and Cd200 (as indicated) for Pdcd10-wt (orange) and Pdcd10-ko (blue) cells of each cluster. See Materials and Methods for definition of positive cells.
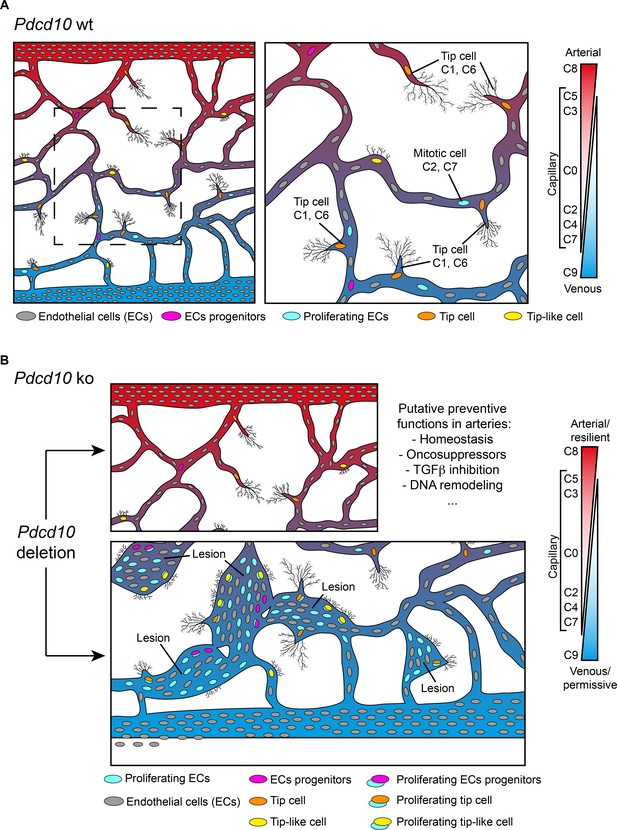
Model of brain cavernoma development based on the scRNA-seq analysis of Pdcd10-ko brain endothelial cells complemented with the findings obtained by immunofluorescence microscopy and spatial transcriptomics (Visium).
Brain ECs (gray) are a heterogeneous population distributed in different clusters on the basis of their gene expression in both Pdcd10-wt (A) and Pdcd10-ko (B). Besides arterial, arterial capillary, venous, venous-capillary also tip cell and mitotic endothelial clusters could be detected at the developmental stage examined, P8 (orange and light blue , respectively). Besides, tip-like cells could be recognized morphologically (yellow). They are significantly increased in Pdcd10-ko, but their transcriptomic features remain to be studied. Resident endothelial cell progenitor cells (pink) were detected in all clusters, although in different proportions, which indicates that they are heterogeneous and share several transcriptional features with the ECs of the vessels where they reside. In (A) dashed square on the left is enlarged for the image on the right. (B) Although Pdcd10 deletion takes place comparably in arterial and venous ECs, only venous ECs can respond to Pdcd10 deletion. Pdcd10-ko venous cells follow defective differentiation and angiogenic programs, with abnormal tip cell traits and increased mitosis, giving rise to cavernous branching and mulberry lesions. Moreover, the venous compartment responds to Pdcd10 deletion with increased numbers and expression levels of venous resident endothelial cell progenitors. Arterial ECs are instead resilient to Pdcd10 deletion. It appears that the resistance of these cells is maintained by a relatively large set of putative defensive genes, some of which modulated by Pdcd10 deletion. In parallel, endothelial progenitor cells within arterial cluster C8 neither increased in number nor were transcriptionally modified by Pdcd10 deletion.
Videos
Three-dimensional rendering of a whole section from a Pdcd10-ko cerebellum (100 μm thick) after confocal microscopy.
Staining for ERG (green), Ki67 (red), and Podocalyxin (blue) is shown. At 4 s and 20 s, the movie focuses on two examples of mulberry lesions. Ki67 staining in the ERG-positive nuclei is clearly visible (white arrows); to enhance the visibility of Ki67 staining, at each stop the ERG staining was hidden for a few seconds. Two Ki67 positive nuclei in pseudo-normal vessels are visible close to the second lesion shown (yellow arrows, top left and bottom right). Scale bars: as indicated on the bars, as the size changes during the movie.
Three-dimensional rendering of three retinal regions after confocal microscopy.
Pdcd10-wt (top) and Pdcd10-ko (bottom) retinas were stained for the membrane marker CD93 (white). Rear and front regions were acquired in the same areas shown in Figure 5F. The central area was from the same retina and shows high density vascular plexus for Pdcd10-ko. Increased numbers of tip cells were observed in all of the vascular regions of Pdcd10-ko retina.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strains, strain backgrounds (mice) | Cdh5(PAC)-Cre-ERT2, C57BL/6 background | Wang et al., 2010 | ||
| Strains, strain backgrounds (mice) | Cdh5(PAC)-Cre- ERT2/Ccm3f/f, C57BL/6 background | Bravi et al., 2015 | ||
| Strains, strain backgrounds (mice) | Cldn5(BAC)-GFP (Tg(Cldn5-GFP)Cbet/U), C57BL/6 background | Honkura et al., 2018 | ||
| Strains, strain backgrounds (mice) | Cdh5(PAC)-Cre-ERT2/Ccm3f/f/Cldn5(BAC)-GFP, C57BL/6 background | This paper | ||
| Antibody | Anti-CD93 (sheep polyclonal) | R and D | AF1696, RRID:AB_354937 | IF (1:500) |
| Antibody | Anti-Erg (rabbit monoclonal) | Abcam | Ab92513, RRID:AB_2630401 | IF (1:400) |
| Antibody | Anti-Ki67 (rat monoclonal) | Invitrogen | 14-5698-82, RRID:AB_10854564 | IF (1:200) |
| Antibody | Anti-Podocalyxin (goat polyclonal) | R and D | AF1556, RRID:AB_354858 | IF (1:400) |
| Antibody | Anti-CoupTFII (rabbit polyclonal) | LSBio | LS-C356225 | IF (1:200) |
| Antibody | Anti-PECAM-1 (rat monoclonal) | BD | 55369, RRID:AB_394815 | IF (1:200) |
| Antibody | Anti-PECAM-1 (armenian hamster monoclonal) | Millipore | MAB1398z, RRID:AB_94207 | IF (1:500) |
| Antibody | Anti-Claudin-5 (mouse monoclonal, Alexa fluor 488-conjugated) | Invitrogen | 352588, RRID:AB_2532189 | IF (1:200) |
| Antibody | Anti-VE-cadherin (rat monoclonal) | BD | 550548, RRID:AB_2244723 | IF (1:200) |
| Antibody | Anti-Cingulin (rabbit polyclonal) | Invitrogen | 36–4401 | IF (1:500) |
| Antibody | Alexa Fluor 488, 555 and 647 donkey secondary antibodies | Jackson laboratories | IF (1:400) | |
| Antibody | Alexa Fluor 488 conjugated donkey anti-mouse | Jackson laboratories | IF (1:400) | |
| Antibody | Anti-uPAR antibody | Tjwa et al., 2009 | IF (1:200) | |
| Commercial assays or kits | 3-Plex positive controls | ACD | #320881 | for detecting Polr2a, PPIB, UBC mRNA |
| Commercial assays or kits | 3-Plex negative controls | ACD | #320871 | for detecting DapB mRNA of Bacillus subtilis |
| Commercial assays or kits | Multiplex fluorescent Reagent Kit v2 | ACD | 323100 | |
| Commercial assays or kits | Cldn5 | ACD | 491611-C2 | |
| Commercial assays or kits | Bst1 | ACD | 559841-C3 | |
| Commercial assays or kits | CD45 MicroBeads | Miltenyi Biotech | 130-052-301 | |
| Commercial assays or kits | CD31 MicroBeads | Miltenyi Biotech | 130-097-418, RRID:AB_2814657 | |
| Commercial assays or kits | Adult Brain Dissociation kits | Miltenyi Biotech | 130-107-677 | |
| Commercial assays or kits | Single Cell 3’ Reagent kits v2 | 10 × genomics | PN-120237 | |
| Instruments | GentleMACS Octo Dissociator | Miltenyi Biotech | 130-095-937 | |
| Instruments | Chromium controller | 10 × genomics | NA | |
| Instruments | NovaSeq 6000 system | Illumina | RRID:SCR_016387 | |
| Instruments | Vibratome | Leica | VT1200s, RRID:SCR_018453 | |
| Instruments | Sp8 confocal microscope | Leica | ||
| Instruments | Cryostat | Thermo Scientific | CryoStar NX50 | |
| Instruments | High Sensitivity RNA ScreenTape | Agilent | ||
| Others | Visium Spatial Gene Expression slides | 10 × genomics | PN-2000233 | |
| Others | Visium Tissue Optimization Slides and reagents | 10 × genomics | PN-1000193 | |
| Software, algorithms | Fiji | open source, http://fiji.sc/ | RRID:SCR_002285 | |
| Software, algorithms | Cell Ranger | 10 × genomics | v2.1.0, RRID:SCR_017344 | |
| Software, algorithms | R package Seurat | Butler et al., 2018; Stuart et al., 2019 | v3.1, RRID:SCR_016341 | |
| Software, algorithms | R package clustree | Zappia and Oshlack, 2018 | v0.4.2, RRID:SCR_016293 | |
| Software, algorithms | R package EnrichR | Chen et al., 2013; Kuleshov et al., 2016 | v2.1, RRID:SCR_001575 | https://amp.pharm.mssm.edu/Enrichr/ |
| Software, algorithms | GSEA | Liberzon et al., 2015; Subramanian et al., 2005 | v2.2.3, RRID:SCR_003199 | https://www.gsea-msigdb.org/gsea/msigdb/annotate.jsp |
| Software, algorithms | STREAM | Chen et al., 2019 | v0.4.1 | |
| Software, algorithms | Space Ranger | 10 × genomics | v1.0.0 | |
| Software, algorithms | Loupe cell browser | 10 × genomics | v4.0, RRID:SCR_018555 |
Additional files
-
Supplementary file 1
Marker genes used in Figure 1H, with supporting references.
- https://cdn.elifesciences.org/articles/61413/elife-61413-supp1-v1.xlsx
-
Supplementary file 2
Known lesion marker genes, showing average logFC in each cluster and bibliographic references.
The table refers to Figure 2C and shows, for the genes included in the heatmap: (1) gene symbol, with basic gene informations; (2) the average logFC observed in each cluster (Pdcd10-ko vs. Pdcd10-wt); (3) a selected list of bibliographic references supporting the choice of each marker.
- https://cdn.elifesciences.org/articles/61413/elife-61413-supp2-v1.xlsx
-
Supplementary file 3
EnrichR results from RareDisease GeneRif dataset.
The table refers to Figure 2D and show the results of the over-representation analysis. The table is showing, for cerebral cavernous malformation term: (1) the cluster that showed significant enrichment; (2) the DEGs contributing to the enrichment, with basic gene informations; (3) the average logFC observed in each cluster (Pdcd10-ko vs. Pdcd10-wt).
- https://cdn.elifesciences.org/articles/61413/elife-61413-supp3-v1.xlsx
-
Supplementary file 4
Over-representation analysis results.
The table refers to Figure 2E and show the results of the over-representation analysis. Hallmarks are divided in categories, as in the figure. For each hallmark is shown: (1) the cluster that showed significant enrichment; (2) the DEGs contributing to the enrichment, with basic gene informations; (3) the average logFC observed in each cluster (Pdcd10-ko vs. Pdcd10-wt).
- https://cdn.elifesciences.org/articles/61413/elife-61413-supp4-v1.xlsx
-
Supplementary file 5
References supporting the annotation of unique marker genes of cluster eight set in alphabetical order and summarized in Figure 3A.
- https://cdn.elifesciences.org/articles/61413/elife-61413-supp5-v1.xlsx
-
Supplementary file 6
References supporting the annotation of DEGs (Pdcd10-ko vs. Pdcd10-wt) of cluster eight set in alphabetical order and summarized in Figure 3D.
- https://cdn.elifesciences.org/articles/61413/elife-61413-supp6-v1.xlsx
-
Supplementary file 7
Canonical marker genes of clusters.
The differentially expressed genes with avg_logFC > 0 in both genotypes and minimump_p_val < 0.05 are considered as canonical markers of clusters.
- https://cdn.elifesciences.org/articles/61413/elife-61413-supp7-v1.xlsx
-
Supplementary file 8
Differentially expressed genes between Pdcd10 (Ccm3) ko and Pdcd10 (Ccm3) wt in each cluster.
The genes are considered differentially expressed if p_val_adj < 0.05.
- https://cdn.elifesciences.org/articles/61413/elife-61413-supp8-v1.xlsx
-
Supplementary file 9
Details of statistical analysis.
- https://cdn.elifesciences.org/articles/61413/elife-61413-supp9-v1.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/61413/elife-61413-transrepform-v1.docx





