chTOG is a conserved mitotic error correction factor
Figures
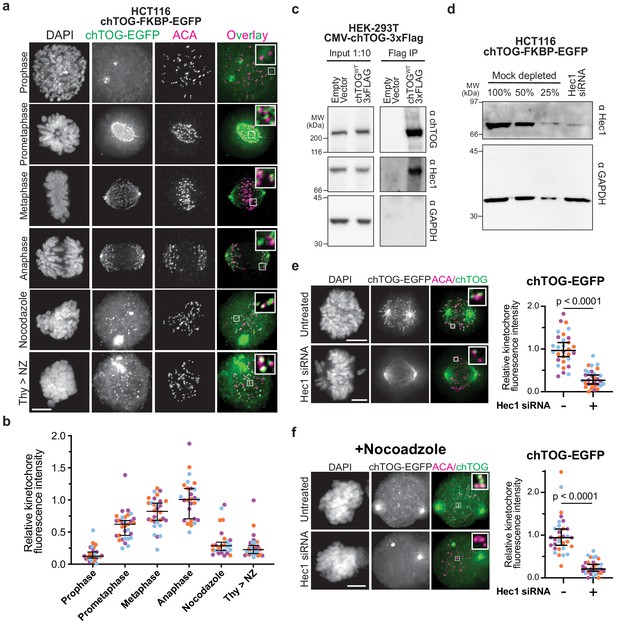
chTOG localizes to kinetochores during mitosis.
(a) Immunofluorescence images of chTOG subcellular localization during mitosis, as visualized in HCT116 cells expressing endogenously epitope-tagged chTOG-EGFP. Anti-centromere protein antibody (ACA) staining marks the centromere-binding proteins and representative images are shown with inlays of kinetochore proximal chTOG at each stage of mitosis. (b) Quantification of chTOG kinetochore association. Each data point represents mean chTOG-EGFP fluorescence intensity at all kinetochores in a single cell normalized to the mean value of the anaphase population (c) HEK-293T cells with either an empty vector control or overexpressed chTOG-3Flag were immunoprecipitated using anti-Flag antibodies. Immunoblots of the input (left) or Flag IP (right) show that the endogenous Hec1/Ndc80 protein specifically co-purified with chTOG. Endogenous and epitope tagged chTOG cannot be individually resolved by anti-chTOG immunoblotting because the 3Flag tag increases the protein’s predicted MW by only 3%. Anti-GAPDH served as a non-specific control. (d) Immunoblotting with anti-Hec1 antibodies was performed on samples of mock-depleted lysate that were diluted to contain the indicated percent of total protein and compared to a lysate prepared from a population of HCT116 cells treated with Hec1 siRNA. Greater than 75% of Hec1 protein was depleted in the siRNA-treated cells. Anti-GAPDH is a loading control. (e) Kinetochore localization of chTOG-EGFP in Hec1 depleted HCT116 cells was determined by fluorescence microscopy. Representative images are shown and were quantified on the right to show endogenously tagged chTOG-EGFP signal at kinetochores decreased by ~70% in siRNA-treated HCT116 cells. Each data point represents mean fluorescence intensity at all kinetochores in a single cell normalized to the mean value of the mock depleted population. (f) Same as (e) but cells were treated with 10 µM nocodazole for 1 hr prior to fixation to preclude any chTOG bound to the microtubule tips from this analysis. All scale bars are 5 µm; contrast on inlays was adjusted independently; data points on graphs are grouped from three experimental replicates and colored according to each replicate with median and 95% confidence intervals displayed in black. p-Values determined by an unpaired Mann-Whitney test.
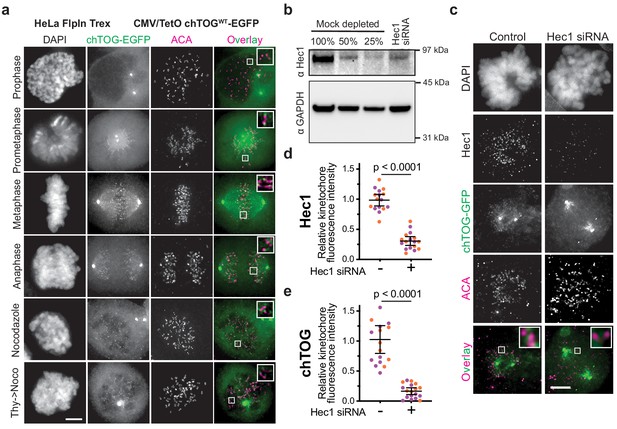
Exogenously expressed chTOG-EGFP localizes to kinetochores in HeLa cells.
(a) Representative immunofluorescence images of chTOG subcellular localization during mitosis, as visualized in HeLa cells where exogenous chTOG-EGFP was expressed by addition of 1 µg/mL doxycycline. Anti-centromere antibody (ACA) staining marks the centromere-binding proteins and inlays of kinetochore proximal chTOG are shown in the boxes. (b) siRNA targeting Hec1 depleted more than 50% of the protein in a population of HeLa cells. Immunoblotting with anti-Hec1 antibodies was performed on samples of mock-depleted lysate that were diluted to contain the indicated percent of total protein and compared to a lysate prepared from a population of HeLa cells treated with Hec1 siRNA. (c) Representative images from immunofluorescence microscopy quantified in (d and e) showing Hec1 siRNA depleted 80% of the Hec1 and chTOG-EGFP proteins from kinetochores in HeLa cells. Each data point represents the mean fluorescence intensity at all kinetochores in a single cell normalized to the mean value of the mock depleted population. All scale bars are 5 µm; contrast on inlays was adjusted independently; data points on graphs are grouped from two experimental replicates and colored according to each replicate with mean and 95% confidence intervals displayed in black. p-Values determined by an unpaired t test.
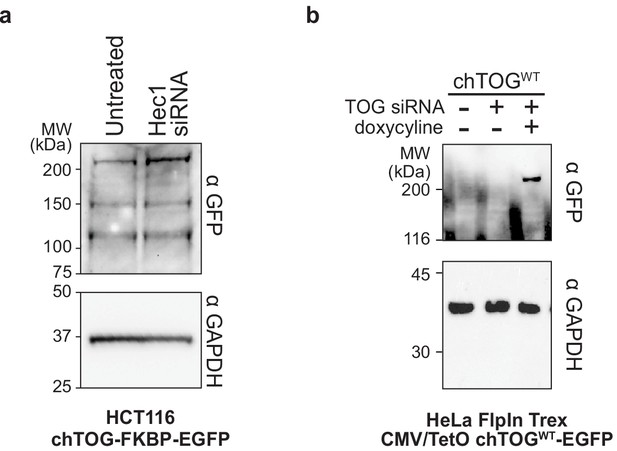
Hec1 siRNA does not affect chTOG protein levels.
(a) Immunoblotting with anti-EGFP antibodies was performed on samples of mock-depleted lysate and Hec1-depelted lysate from HCT116 cells expressing endogenously epitope-tagged chTOG-EGFP. Anti-GAPDH is a loading control. (b) Immunoblotting with anti-EGFP antibodies was performed on lysates from HeLa FlpIn Trex chTOGWT cells seen in Figure 1—figure supplement 1. Cells were treated with siRNA and or doxycycline to deplete endogenous chTOG and express ectopic chTOG-EGFP, suggesting the specificity of EGFP antibodies for the fusion protein.
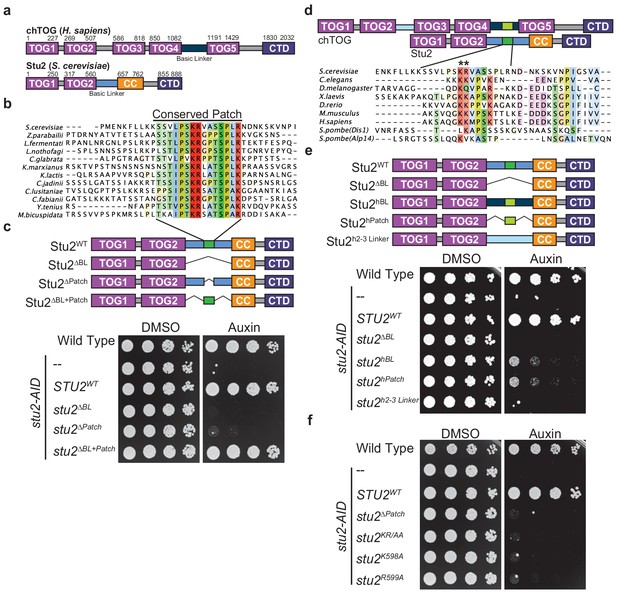
Two conserved basic residues are essential for Stu2 function.
(a) Schematic of the yeast Stu2 and human chTOG proteins describing the domains in each protein. Specific residues marking domains are indicated on the top of each protein. (b) ClustalO multiple sequence alignment generated from full-length Saccharomyces cerevisiae Stu2 and related proteins in Ascomycota. Fifteen amino acids within the otherwise divergent ~110 amino acid ‘basic linker’ are colored based on percent conservation and biochemical properties of the side chain. (c) Schematic of the Stu2 mutant proteins used to investigate the essential nature of the conserved patch in S. cerevisiae. Cell viability was analyzed in stu2-AID strains expressing the indicated Stu2 mutant proteins by plating fivefold serial dilutions in the presence of vehicle (left) and or auxin (right) to degrade the endogenous Stu2-AID protein. (d) Schematic of human chTOG and yeast Stu2 proteins showing how ClustalO multiple sequence alignments were performed specifically on basic linker regions (below) from metazoan and fungal species (colored similarly to (b)). (e) Schematic of the Stu2 mutant proteins used to investigate the orthologous behavior of the human basic linker and conserved patch (linkers are colored to match (d)). Cell viability determined as in (c) using a serial dilution growth assay. (f) The pair of conserved basic residues identified by asterisks in (d) were mutated individually or as a pair to alanine and found to be required for S. cerevisiae viability as assayed in (c).
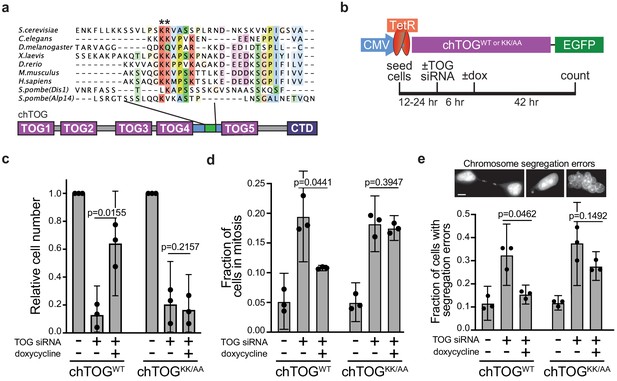
Mutating two conserved basic residues in chTOG causes mitotic delay and lethal chromosome segregation errors.
(a) ClustalO multiple sequence alignment generated from basic linker regions of metazoan and fungal species shown in Figure 2. The pair of basic residues mutated to alanine denoted with asterisks. (b) Assay to analyze the ability of doxycycline-inducible codon-optimized, chTOGWT and chTOGKK/AA to complement cellular activities after siRNA-mediated depletion of endogenous chTOG. Cells depleted of chTOG or expressing ectopic chTOG proteins were assayed for (c) proliferation, (d) mitotic index, and (e) chromosome segregation errors. Scale bar is 5 µm; mean values and 95% confidence intervals for three experimental replicates displayed; p-values determined by paired t test.
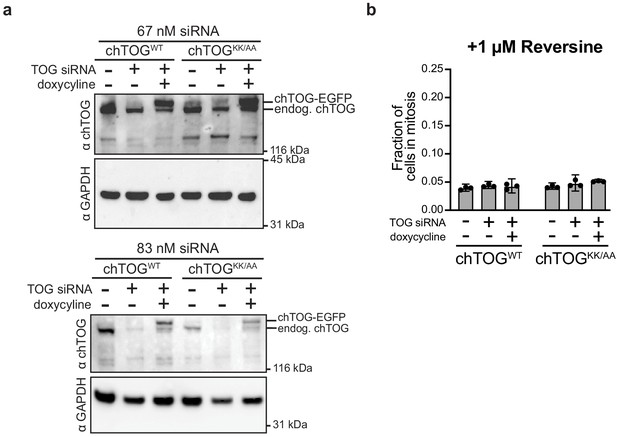
chTOGWT and chTOGKK/AA are expressed at equivalent levels; mitotic delays from chTOG depletion or mutation require Mps1 activity.
(a) Immunoblots were performed on lysates of HeLa cells expressing EGFP epitope-tagged chTOGWT and chTOGKK/AA proteins after depletion of the endogenous chTOG by siRNA with the indicated concentration of siRNA. All future experiments used 83 nM siRNA; GAPDH served as a loading control. Unlike 3FLAG epitope tag, both endogenous and EGFP tagged chTOG can be resolved with anti-chTOG antibodies. (b) Cells depleted of chTOG or expressing ectopic chTOG proteins were assayed for mitotic index after 1-hr treatment with Mps1 inhibitor, Reversine. Mean values and 95% confidence intervals for three experimental replicates displayed.
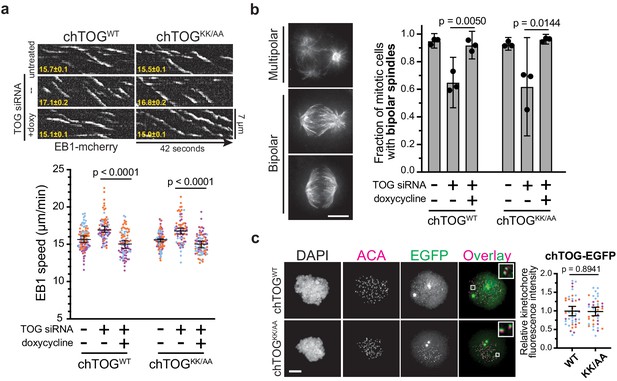
The chTOG basic pair mutant regulates microtubule dynamics, spindle structure, and localizes to kinetochores.
(a) Images isolated from live-cell TIRF microscopy were used to generate kymographs of EB1-mCherry in chTOGWT and chTOGKK/AA-expressing cells with average EB1 track speed (µm/min) in bottom left and quantifications shown below. Each data point represents the mean EB1 track speed per cell and is grouped from three experimental replicates and colored according to each replicate with median and 95% confidence intervals displayed in black. p-Values were determined with an unpaired Mann-Whitney test. (b) Representative images of each spindle phenotype observed in mitotic chTOG-depleted, chTOGWT, or chTOGKK/AA expressing cells. While bipolar spindles exhibited two distinct phenotypes, we first quantified the fraction of cells exhibiting multipolar or bipolar spindles. Mean values and 95% confidence interval for three replicates were reported, p values calculated with a paired t test. (c) Representative images (left) and quantification (right) of chTOGWT and chTOGKK/AA localization to kinetochores in the absence of endogenous chTOG. Cells were treated with nocodazole to eliminate microtubules. Each data point represents the mean chTOG-EGFP fluorescence intensity at all kinetochores in a single cell normalized to the mean value of chTOGWT-expressing cells. Data are grouped from three experimental replicates and colored according to each replicate with mean values and 95% confidence displayed in black. p-Value was determined with an unpaired t test. All scale bars are 5 µm.
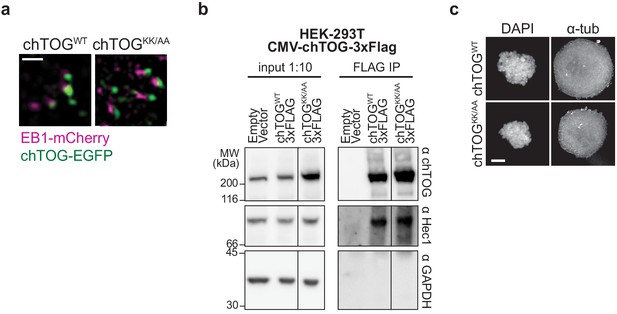
The basic pair mutant localizes to microtubule plus-ends and interacts with Hec1.
(a) Representative live-cell TIRF microscopy image of chTOGWT and chTOGKK/AA binding to microtubule plus-ends as marked by EB1-mCherry. Scale bar is 1 µm. (c) chTOG was immunoprecipitated from HEK-293T cells expressing either an empty vector control, chTOGWT-3Flag, or chTOGKK/AA-3FLAG proteins, using anti-Flag antibodies. Immunoblotting of the input lysates (left) and immunoprecipitations (right) show that the WT and mutant chTOG proteins co-purify endogenous Hec1. Anti-GAPDH is shown as a non-specific control for the immunoprecipitation. This is the same immunoblot from Figure 1, and the line denotes where a single, non-relevant lane was cropped from the image. (c) Representative image of nocodazole-treated cells from Figure 4c demonstrating the complete depolymerization of microtubules in these cells at this dose of drug; the scale bar is 5 µm.
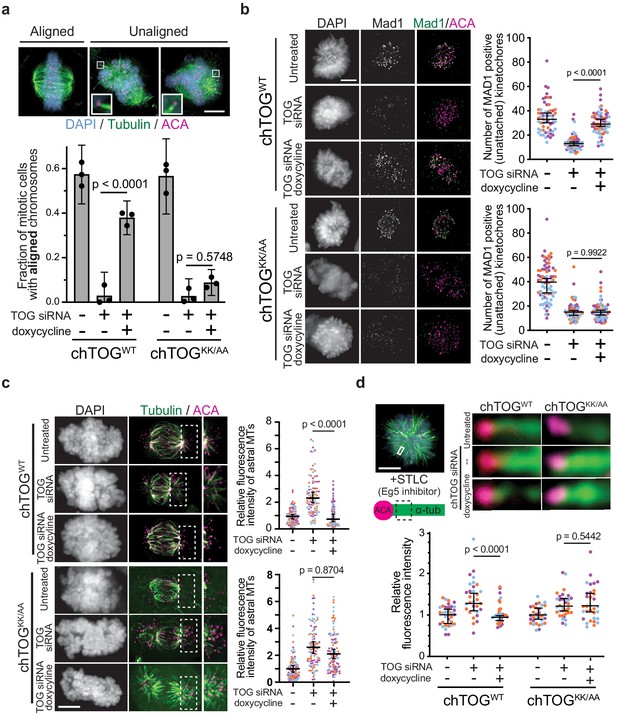
The chTOG basic pair is required to regulate kinetochore-microtubule attachment stability.
(a) Representative images of each chromosome alignment phenotype observed in mitotic chTOG-depleted, chTOGWT, or chTOGKK/AA show a large fraction of chTOGKK/AA expressing cells form bipolar spindles with excessive astral microtubules that attach to kinetochores (image inlays) and prevent chromosome alignment. Phenotypes are quantified below where mean values and 95% confidence intervals for three replicates are reported. p-Values were determined with a paired t test and contrast on inlays was adjusted independently. (b–d) Data points on graphs are grouped from three experimental replicates and colored according to each replicate with median and 95% confidence intervals displayed in black. p-Values determined by an unpaired Mann-Whitney test. (b) Representative images (left) and quantification (right) of Mad1 immunostaining as a marker for kinetochore-microtubule attachment state in chTOG depleted, chTOGWT (top), or chTOGKK/AA (bottom) expressing cells. Each data point represents the number of kinetochores with Mad1 puncta per cell. (c) Representative images (left) and quantification (right) of cold-stable astral (erroneous) kinetochore-microtubule attachments in chTOG depleted, chTOGWT (top), or chTOGKK/AA (bottom) expressing cells. Each data point represents the tubulin fluorescence intensity of all astral microtubules on one half of the mitotic spindle. (d) Monopolar spindles (top left) were formed by inhibiting Eg5/KIF11 with STLC to allow the fluorescence intensity quantification of kinetochore-bound microtubule bundles at low-tension, erroneous attachments in control cells or chTOG-depleted cells expressing chTOGWT or chTOGKK/AA (right). Each data point is relative intensity normalized to the mean of uninduced, mock-depleted cells. All scale bars are 5 µm.
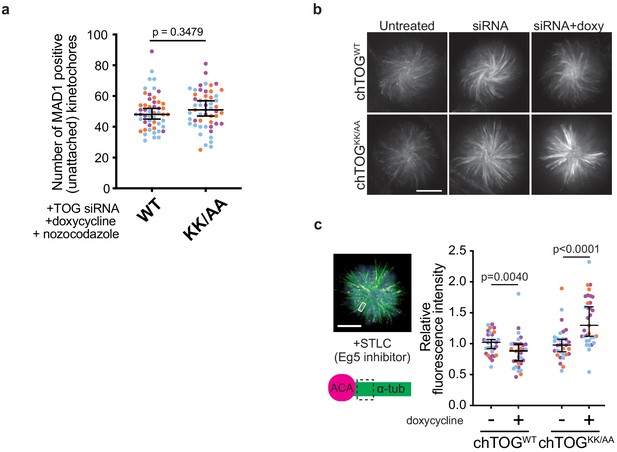
The basic pair mutant does not affect Mad1 recruitment to kinetochores and competes with endogenous chTOG.
(a) Cells depleted of chTOG and expressing chTOGWT or chTOGKK/AA were immunostained for Mad1 as in Figure 5b; however, cells were first treated with nocodazole to ensure the basic pair did not contribute to Mad1 kinetochore recruitment. (b) Representative images of whole cells from the experiment in Figure 5d show the anti-tubulin immunostaining for STLC-treated monopoles. (c) HeLa cells expressing chTOGWT and chTOGKK/AA in the presence of endogenous chTOG were arrested with STLC to form monopolar spindles and erroneous low-tension kinetochore-microtubule attachments. Immunofluorescence was performed with anti-tubulin and anti-ACA antibodies and the intensity of tubulin staining was quantified near kinetochores as an indirect measure of the number of microtubules within an attached bundle. The basic pair mutant prematurely stabilizes erroneous attachments even in the presence of endogenous chTOG. Data points on graphs are grouped from three experimental replicates and colored according to each replicate with median and 95% confidence intervals displayed in black. p-Values determined by an unpaired Mann-Whitney test.
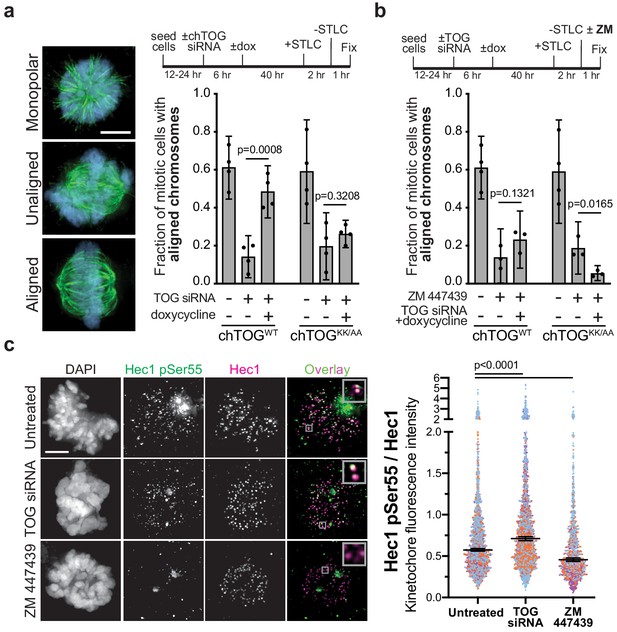
chTOG- and Aurora-B-dependent error pathways likely function independently.
(a) Mitotic error correction was assayed by inducing errors with STLC to inhibit Eg5 and then washing out the inhibitor in control cells or chTOG-depleted cells expressing chTOGWT or chTOGKK/AA. The chromosome alignment phenotype (left) was quantified 60 min after inhibitor washout. (b) The same error correction assay was performed as in (a) but was supplemented with a low dose of Aurora B kinase inhibitor ZM 447439. Untreated populations in (a and b) are the same and both display mean values and the 95% confidence interval of three or four experimental replicates; p values were determined with paired t tests. (c) Representative immunofluorescence images (left) and quantifications (right) of the relative fluorescence intensity of phosphorylated Hec1 analyzed with a phospho-specific antibody to Ser55. Each data point represents individual kinetochore intensities of Hec1 pSer55 antibody ratioed to Hec1 antibody. Data points on graphs are grouped from three experimental replicates and colored according to each replicate with median and 95% confidence intervals displayed in black. p-Values were determined by an unpaired Mann-Whitney test. All scale bars are 5 µm.
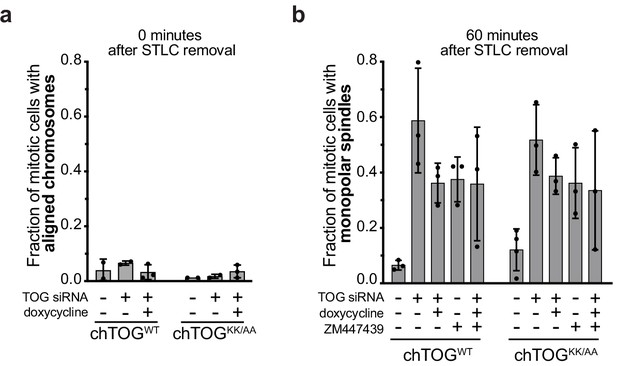
Formation of bipolar or monopolar spindles in STLC washout experiments.
(a) Cells under all experimental conditions in Figure 4 were quantified for the fraction of mitotic cells with aligned chromosome immediately after STLC removal. In every condition, cells responded equivalently to STLC treatment and washout. (b) Fraction of cells with monopolar spindles 60 min after STLC removal show that chTOG-depleted cells struggle to form bipolar spindles, while all other experimental conditions form bipolar spindles equivalently. For all experiments, mean values and 95% confidence intervals for three experimental replicates displayed; p values determined by paired t test.
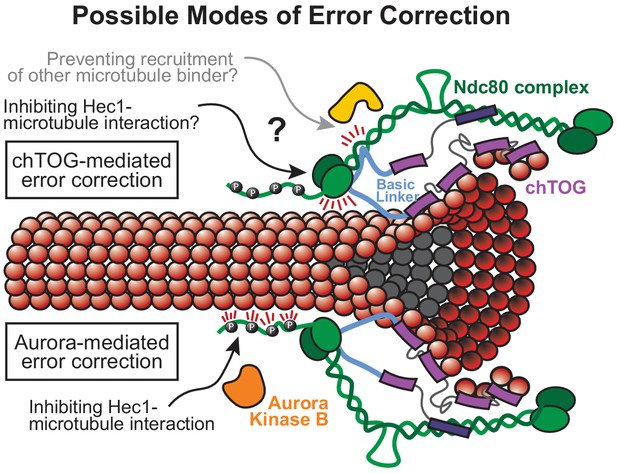
Possible model for chTOG-mediated error correction, independent of Aurora B kinase activity.
Possible models of chTOG-mediated error correction (top) where chTOG kinetochore localization depends on the C-terminus while the TOG domains are still capable of binding tubulin at the microtubule tip in the ‘bent’ conformation. This may position the basic linker near the Ndc80/Hec1 CH domain to regulate Ndc80 complex activity. We favor the basic linker directly modulating Hec1 microtubule binding behavior, but chTOG could also prevent recruitment of other microtubule binding proteins. This activity is independent of Aurora-B-mediated error correction (bottom) that also recognizes low-tension attachments in the absence of chTOG, phosphorylates the Ndc80 tail domain, but on average cannot fully destabilize the bond.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Chemical compound, drug | Nocodazole | Sigma-Aldrich | M1404 | |
| Chemical compound, drug | Thymidine | Sigma-Aldrich | T9250 | |
| Chemical compound, drug | S-trityl-L-cysteine (STLC) | Sigma-Aldrich | 164739 | |
| Chemical compound, drug | Reversine | Sigma-Aldrich | R3904 | |
| Chemical compound, drug | ZM447439 | Selleckchem | S1103 | |
| Chemical compound, drug | Puromycin Dihydrochloride | Sigma-Aldrich | A11138-03 | |
| Chemical compound, drug | Hygromycin B | Thermo Fisher | 10687010 | |
| Chemical compound, drug | Doxycycline | Sigma-Aldrich | D9891 | |
| Chemical compound, drug | Indole-3-acetic acid (Auxin) | Sigma-Aldrich | I3750 | |
| Chemical compound, drug | Polyethyleneimine (PEI) | Polysciences | 23966–1 | Linear, MW 25000 |
| Chemical compound, drug | Lipfectamine 2000 | Thermo Fisher | 11668027 | |
| Chemical compound, drug | Lipofectamine RNAiMax | Thermo Fisher | 13778075 | |
| Other | Protein G Dynabeads | Thermo Fisher | 10009D | |
| Other | DAPI stain | Thermo Fisher | D1306 | IF: 60 ng/mL |
| Antibody | Anti-Flag (M2) [mouse monoclonal] | Millipore-Sigma | Cat# F3165; RRID:AB_259529 | IP (1 µg / 60 µL Prot G bead) |
| Antibody | Anti-GAPDH (6C5) [mouse monoclonal] | Millipore-Sigma | Cat# MAB374; RRID:AB_2107445 | WB (1 µg/mL) |
| Antibody | Anti-CKAP5(chTOG) [rabbit polyclonal] | GeneTex | Cat# GTX30693; RRID:AB_625852 | WB (1:1000) |
| Antibody | Anti-Hec1 (9G3) [mouse monoclonal] | Thermo Fisher | Cat# MA1-23308; RRID:AB_2149871 | WB (2 µg/mL) IF (1 µg/mL) |
| Antibody | Anti-GFP (JL-8) [mouse monoclonal] | Takara | Cat# 632381; RRID:AB_2313808 | WB (0.5 µg/mL) |
| Antibody | HRP-conjugated anti-mouse [sheep polyclonal] | GE Healthcare | Cat# NA931; RRID:AB_772210 | WB (1:10,0000) |
| Antibody | HRP-conjugated anti-rabbit [sheep polyclonal] | GE Healthcare | Cat# NA934; RRID:AB_2722659 | WB (1:10,0000) |
| Antibody | Anti-centromere antibody (ACA) [human polyclonal] | Antibodies Inc | Cat# 15–235; RRID:AB_2797146 | IF (1:600) |
| Antibody | Anti-alpha tubulin (DM1A) [mouse monoclonal] | Millipore-Sigma | Cat# T6199; RRID:AB_477583 | IF (2 µg/mL) |
| Antibody | Anti-Mad1 [rabbit polyclonal] | GeneTex | Cat# GTX109519; RRID:AB_1950847 | IF (1:1000) |
| Antibody | Anti-pSer55 Hec1 [rabbit purified polyclonal] | DeLuca et al., 2011 PMID:21266467 | IF (1:1000) | |
| Antibody | Alexa Fluor 594 conjugated anti-mouse [goat polyclonal] | Thermo Fisher | Cat# A11005; RRID:AB_2534073; | IF (1:300; 1:600 for Tubulin) |
| Antibody | Alexa Fluor 647 conjugated anti-mouse [goat polyclonal] | Thermo Fisher | Cat# A21235; RRID:AB_2535804 | IF (1:300; 1:600 for Tubulin) |
| Antibody | Alexa Fluor 594 conjugated anti-rabbit [goat polyclonal] | Thermo Fisher | Cat# A11037; RRID:AB_2534095 | IF (1:300) |
| Antibody | Alexa Fluor 647 conjugated anti-rabbit [goat polyclonal] | Thermo Fisher | Cat # A21244; RRID:AB_ 2535812 | IF (1:300) |
| Antibody | Alexa Fluor 594 conjugated anti-human [goat polyclonal] | Thermo Fisher | Cat# A11014; RRID:AB_2534081 | IF (1:300) |
| Antibody | AlexFluor 647 conjugated anti-human [goat polyclonal] | Thermo Fisher | Cat# A21445; RRID:AB_2535862 | IF (1:300) |
| Transfected construct | siRNA to Hec1 (custom sequence) | Qiagen | CCCUGGGUCGUGUCAGGAA | |
| Transfected construct | siRNA to chTOG (flexitube) | Qiagen | Cat# SI02653588 | AAGGGTCGACTCAATGATTCA |
| Recombinant DNA reagent | Stu2WT-3V5 | Miller et al., 2016 PMID:27156448 | pSB2232 | See Table 1 for more details |
| Recombinant DNA reagent | Stu2∆BL-3V5 | Miller et al., 2019 PMID:31584935 | pSB2260 | See Table 1 for more details |
| Recombinant DNA reagent | Stu2∆Patch-3V5 | This study | pSB2634 | See Table 1 for more details |
| Recombinant DNA reagent | Cloning intermediate | This study | pSB2820 | See Table 1 for more details |
| Recombinant DNA reagent | Stu2∆BL+Patch-3V5 | This study | pSB2852 | See Table 1 for more details |
| Recombinant DNA reagent | Stu2∆K598A-3V5 | This study | pSB2818 | See Table 1 for more details |
| Recombinant DNA reagent | Stu2∆R599A-3V5 | This study | pSB2819 | See Table 1 for more details |
| Recombinant DNA reagent | Stu2∆KR598AA-3V5 | This study | pSB2781 | See Table 1 for more details |
| Recombinant DNA reagent | Stu2hBL-3V5 | This study | pSB3076 | See Table 1 for more details |
| Recombinant DNA reagent | Stu2hPatch-3V5 | This study | pSB3075 | See Table 1 for more details |
| Recombinant DNA reagent | Stu2h2-3 Linker-3V5 | This study | pSB3089 | See Table 1 for more details |
| Recombinant DNA reagent | pCDNA3_chTOGWT-EGFP | This study | pSB2822 | See Table 1 for more details |
| Recombinant DNA reagent | pCDNA3_chTOGKK1142AA-EGFP | This study | pSB2823 | See Table 1 for more details |
| Recombinant DNA reagent | FRT/TO | Etemad et al., 2015 | pSB2353 | See Table 1 for more details |
| Recombinant DNA reagent | FRT/TO_chTOGWT-EGFP | This Study | pSB2860 | See Table 1 for more details |
| Recombinant DNA reagent | FRT/TO_chTOGKK1142AA-EGFP | This Study | pSB2863 | See Table 1 for more details |
| Recombinant DNA reagent | FRT/TO_ chTOGWT-6His-3Flag | This Study | pSB2976 | See Table 1 for more details |
| Recombinant DNA reagent | FRT/TO_chTOGKK1142AA-6His-3Flag | This Study | pSB2977 | See Table 1 for more details |
| Recombinant DNA reagent | EB1-mCherry | Addgene | RRID:Addgene_55035 | See Table 1 for more details |
| Recombinant DNA reagent | pLPH2 | This Study | pSB2998 | See Table 1 for more details |
| Recombinant DNA reagent | pLPH2_EB1-mCherry | This Study | pSB3217 | See Table 1 for more details |
| Strain, strain background (Saccharomyces cerevisiae) | W303 | Miller et al., 2016 PMID:27156448 | SBY3 | See Table 2 for more details |
| Genetic reagent (S. cerevisiae) | STU2-IAA7; TIR1 | Miller et al., 2016 PMID:27156448 | SBY13772 | See Table 2 for more details |
| Genetic reagent (S. cerevisiae) | STU2-IAA7; TIR1; Stu2WT | Miller et al., 2019 PMID:31584935 | SBY13901 | See Table 2 for more details |
| Genetic reagent (S. cerevisiae) | STU2-IAA7; TIR1; Stu2∆BL | This study | SBY17069 | See Table 2 for more details |
| Genetic reagent (S. cerevisiae) | STU2-IAA7; TIR1; Stu2KR/AA | This study | SBY17206 | See Table 2 for more details |
| Genetic reagent (S. cerevisiae) | STU2-IAA7; TIR1; Stu2K598A | This study | SBY17477 | See Table 2 for more details |
| Genetic reagent (S. cerevisiae) | STU2-IAA7; TIR1; Stu2R599A | This study | SBY17479 | See Table 2 for more details |
| Genetic reagent (S. cerevisiae) | STU2-IAA7; TIR1; Stu2∆Patch | This study | SBY17519 | See Table 2 for more details |
| Genetic reagent (S. cerevisiae) | STU2-IAA7; TIR1; Stu2∆BL+Patch | This study | SBY17593 | See Table 2 for more details |
| Genetic reagent (S. cerevisiae) | STU2-IAA7; TIR1; Stu2hBL | This study | SBY18799 | See Table 2 for more details |
| Genetic reagent (S. cerevisiae) | STU2-IAA7; TIR1; Stu2hPatch | This study | SBY18797 | See Table 2 for more details |
| Genetic reagent (S. cerevisiae) | STU2-IAA7; TIR1; Stu2h2-3 Linker | This study | SBY19023 | See Table 2 for more details |
| Genetic reagent (Homo sapiens) | HCT116 chTOG-FKBP-EGFP | Cherry et al., 2019 PMID:31058365 | SBM004 | See Table 3 for more details |
| Cell line (H. sapiens) | HEK 293T | Ding et al., 2013 PMID:23154965 | SBM033 | See Table 3 for more details |
| Genetic reagent (H. sapiens) | HeLa FlpIn Trex | Etemad et al., 2015 PMID:26621779 | SBM001 | See Table 3 for more details |
| Genetic reagent (H. sapiens) | HeLa FlpIn Trex; chTOGWT-EGFP | This study | SBM044 | See Table 3 for more details |
| Genetic reagent (H. sapiens) | HeLa FlpIn Trex; chTOGKK/AA-EGFP | This study | SBM046 | See Table 3 for more details |
| Genetic reagent (H. sapiens) | HeLa FlpIn Trex; chTOGWT-EGFP; EB1-mCherry | This study | SBM045 | See Table 3 for more details |
| Genetic reagent (H. sapiens) | HeLa FlpIn Trex; chTOGKK/AA-EGFP; EB1-mCherry | This study | SBM047 | See Table 3 for more details |
| Gene (S. cerevisiae) | STU2 | Saccharomyces Genome Database | SGD:S000004035 | |
| Gene (H. sapiens) | chTOG; CKAP5 | Consensus Coding DNA Sequence Database | CCDS: 31477.1 | |
| Gene (H. sapiens) | EB1; MAPRE1 | Consensus Coding DNA Sequence Database | CCDS: 13208.1 | |
| Software, algorithm | Prism 9 | GraphPad Software | Version 9.0.0 (86) | |
| Software, algorithm | TrackMate | Tinevez et al., 2017 PMID:27713081 | Version 4.0.0 |
Plasmids used in this study.
| Plasmid | Vector backbone | Gene of interest | Mutation description | Selection marker | Primers | Source |
|---|---|---|---|---|---|---|
| pSB2232 | pSB2223/pL300 | Stu2WT-3V5 | LEU2 | SB4372, SB4374 | Miller et al., 2016 PMID:27156448 | |
| pSB2260 | pSB2223/pL300 | Stu2∆BL-3V5 | ∆560–657: : GDGAG | LEU2 | SB4411, SB4413 | Miller et al., 2019 PMID:31584935 |
| pSB2634 | pSB2223/pL300 | Stu2∆Patch-3V5 | ∆592–607: : GDGAG | LEU2 | SB5248, SB4413 | This study |
| pSB2820 | pSB2223/pL300 | Cloning intermediate | ∆569–657:: GDGAG+592–607+GDGAG | LEU2 | SB5447 | This study |
| pSB2852 | pSB2223/pL300 | Stu2∆BL+Patch-3V5 | ∆560–657:: GDGAG+592–607+GDGAG | LEU2 | SB5519, SB5520 | This study |
| pSB2818 | pSB2223/pL300 | Stu2∆K598A-3V5 | K598A | LEU2 | SB5458, SB4413 | This study |
| pSB2819 | pSB2223/pL300 | Stu2∆R599A-3V5 | R599A | LEU2 | SB5459, SB4413 | This study |
| pSB2781 | pSB2223/pL300 | Stu2∆KR598AA-3V5 | K598A, R599A | LEU2 | SB5349, SB4413 | This study |
| pSB3076 | pSB2223/pL300 | Stu2hBL-3V5 | ∆560–657::chTOG1081-1167 | LEU2 | SB5248, SB4413 | This study |
| pSB3075 | pSB2223/pL300 | Stu2hPatch-3V5 | ∆560–657::GDGG+chTOG1137-1150+GDGAG | LEU2 | SB5447 | This study |
| pSB3089 | pSB2223/pL300 | Stu2h2-3 Linker-3V5 | ∆560–657::chTOG500-585 | LEU2 | SB5519, SB5520 | This study |
| pSB2822 | pCDNA3.1 | chTOGWT-EGFP | Neomycin | N/A | This study | |
| pSB2823 | pCDNA3.1 | chTOGKK1142AA-EGFP | K1142A, K1143A | Neomycin | N/A | This study |
| pSB2353 | pCDNA5 FRT/TO | N/A | Puromycin | N/A | Etemad et al., 2015 PMID:26621779 | |
| pSB2860 | pCDNA5 FRT/TO | chTOGWT-EGFP | Puromycin | SB5536, SB5537 | This Study | |
| pSB2863 | pCDNA5 FRT/TO | chTOGKK1142AA-EGFP | K1142A, K1143A | Puromycin | SB5536, SB5537 | This Study |
| pSB2976 | pCDNA5 FRT/TO | chTOGWT-6His-3Flag | Puromycin | SB5774, SB5775 | This Study | |
| pSB2977 | pCDNA5 FRT/TO | chTOGKK1142AA-6His-3Flag | K1142A, K1143A | Puromycin | SB5774, SB5775 | This Study |
| pSB3062 | EB1-mCherry | Neomycin | N/A | Davidson Lab (Addgene: 55035) | ||
| pSB2998 | pLPH2 | N/A | Hygromycin | N/A | This Study | |
| pSB3217 | pLPH2 | EB1-mCherry | Hygromycin | SB5938, SB5939, SB5940, SB5941 | This Study |
Yeast strains used in this study.
| All strains are derivatives of SBY3 (W303) | |
|---|---|
| Strain | Relevant genotype |
| SBY3 (W303) | MATa ura3-1 leu2-3,112 his3-11 trp1-1 can1-100 ade2-1 bar1-1 |
| SBY13772 | MATa STU2-3HA-IAA7:KanMX DSN1-6His-3Flag:URA3 his3::pGPD1-TIR1:HIS3 |
| SBY13901 | MATa STU2-3HA-IAA7:KanMX DSN1-6His-3Flag:URA3 his3::pGPD1-TIR1:HIS3 leu2::pSTU2-STU2-3V5:LEU2 |
| SBY17069 | MATa STU2-3HA-IAA7:KanMX DSN1-6His-3Flag:URA3 his3::pGPD1-TIR1:HIS3 leu2::pSTU2-stu2(∆592–607::GDGAGLlinker)−3 V5:LEU2 |
| SBY17206 | MATa STU2-3HA-IAA7:KanMX DSN1-6His-3Flag:URA3 his3::pGPD1-TIR1:HIS3 leu2::pSTU2-stu2(K598A R599A)−3 V5:LEU2 |
| SBY17477 | MATa STU2-3HA-IAA7:KanMX DSN1-6His-3Flag:URA3 his3::pGPD1-TIR1:HIS3 leu2::pSTU2-stu2(K598A)−3 V5:LEU2 |
| SBY17479 | MATa STU2-3HA-IAA7:KanMX DSN1-6His-3Flag:URA3 his3::pGPD1-TIR1:HIS3 leu2::pSTU2-stu2(R599A)−3 V5:LEU2 |
| SBY17519 | MATa STU2-3HA-IAA7:KanMX DSN1-6His-3Flag:URA3 his3::pGPD1-TIR1:HIS3 leu2::pSTU2-stu2(∆560–657::GDGAGLlinker)−3 V5:LEU2 |
| SBY17593 | MATa STU2-3HA-IAA7:KanMX DSN1-6His-3Flag:URA3 his3::pGPD1-TIR1:HIS3 leu2::pSTU2-stu2(∆560–657::GDGAGLlinker:592–607:GDGAGLlinker)−3 V5:LEU2 |
| SBY18799 | MATa STU2-3HA-IAA7:KanMX DSN1-6His-3Flag:URA3 his3::pGPD1-TIR1:HIS3 leu2::pSTU2-Stu2(∆560–657::chTOG(1081–1167))−3 V5:LEU2 |
| SBY18797 | MATa STU2-3HA-IAA7:KanMX DSN1-6His-3Flag:URA3 his3::pGPD1-TIR1:HIS3 leu2::pSTU2-Stu2(∆560–657::linker-chTOG(1137–1150)-linker)−3 V5:LEU2 |
| SBY19023 | MATa STU2-3HA-IAA7:KanMX DSN1-6His-3Flag:URA3 his3::pGPD1-TIR1:HIS3 leu2::pSTU2-stu2(∆560–657::chTOG(500-585))−3 V5:LEU2 |
Human cell lines used in this study.
| Cell line | Parental | Modification 1 | Modification 2 | Source |
|---|---|---|---|---|
| SBM004 | HCT116 | CKAP5e44-FKBP- EGFP/CKAP5e44-FKBP-EGFP | Cherry et al., 2019 PMID:31058365 | |
| SBM033 | 293T | Ding et al., 2013 PMID:23154965 | ||
| SBM001 | HeLa FlpIn Trex | SV40: LacZ-ZeocinR | Etemad et al., 2015 PMID:26621779 | |
| SBM044 | HeLa FlpIn Trex | TRE: chTOGWT-EGFP SV40: PuromycinR | This study | |
| SBM046 | HeLa FlpIn Trex | TRE: chTOGKK/AA-EGFP SV40: PuromycinR | This study | |
| SBM045 | HeLa FlpIn Trex | TRE: chTOGWT-EGFP SV40: PuromycinR | hPGK1: EB1-mCherry IRES hygromycinR | This study |
| SBM047 | HeLa FlpIn Trex | TRE: chTOGKK/AA-EGFP SV40: PuromycinR | hPGK1: EB1-mCherry IRES hygromycinR | This study |





