Autophagy compensates for Lkb1 loss to maintain adult mice homeostasis and survival
Figures
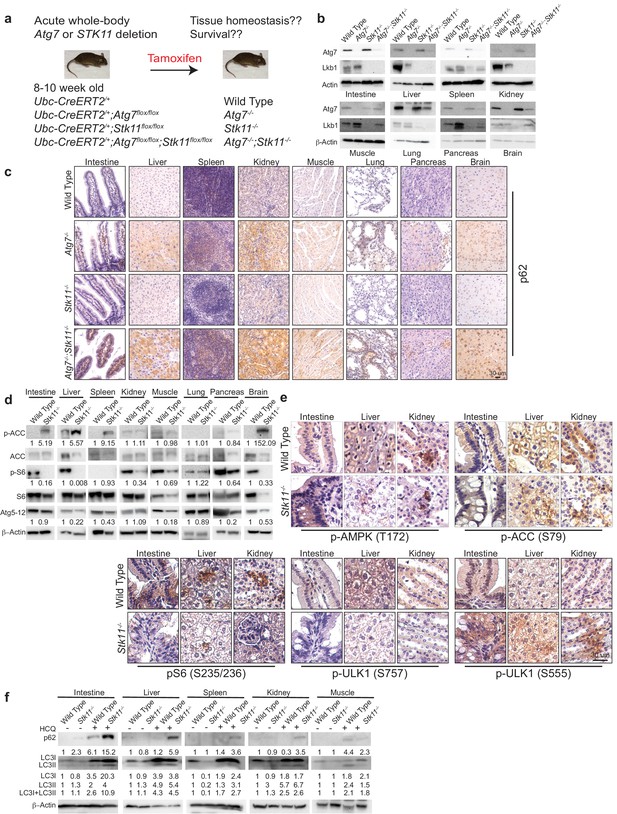
Autophagy is upregulated in tissues of Stk11-deficient mice.
(a) Experimental design for generation of Atg7-/-, Stk11-/-, and Atg7-/-;Stk11-/- adult mice. (b) Western blotting for Atg7 and Lkb1 of the indicated tissues from WT control, Atg7-/-, Stk11-/-, and Atg7-/-;Stk11-/- adult mice. β-actin serves as a protein loading control. (c) Representative IHC for p62 of different tissues from WT control, Atg7-/-, Stk11-/-, and Atg7-/-;Stk11-/- adult mice. (d) Western blotting for pACC (S79), total ACC, pS6 (S235/236), total S6, and Atg5-Atg12 from different tissues of WT control and Stk11-/- mice. β-actin serves as a protein loading control. Numbers indicate the quantification of phospho-protein levels normalized to total levels of protein, β-actin and WT control. (e) Representative IHC for pAMPK (Th172), pACC (S79), pS6 (S235/236), and pULK1 (S555 and S757) in different tissues of WT control and Stk11-/- adult mice. (f) Western blotting for p62, LC3I and LC3II in different tissues of WT control and Stk11-/- mice with or without HCQ treatment. β-actin serves as a protein loading control. Numbers indicate the quantification of protein levels normalized to β-actin and WT control.
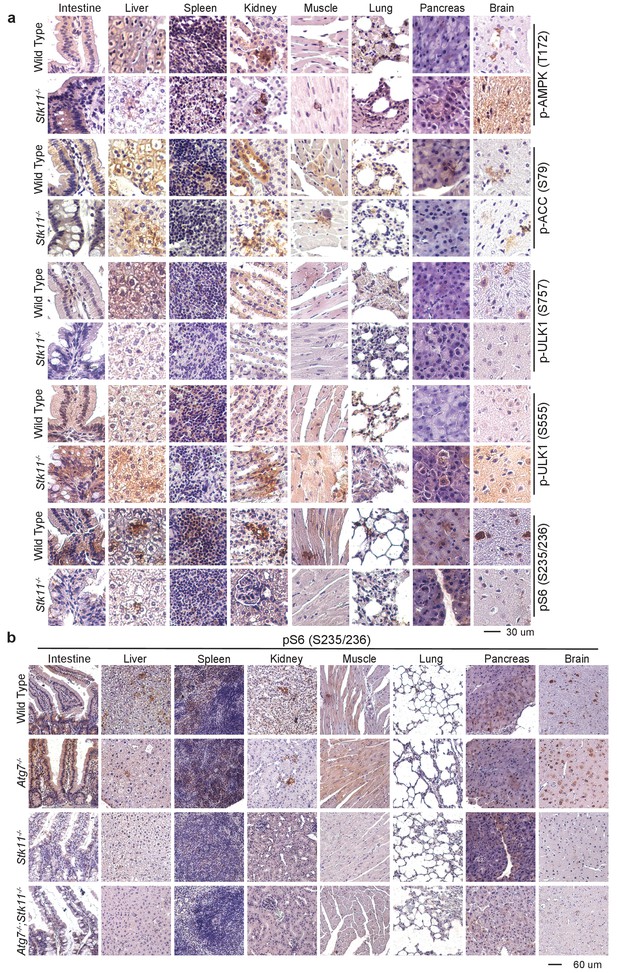
AMPK is active in Lkb1-deleted tissues.
(a) Representative IHC for pAMPK (T172), pACC (S79), pS6 (S235/236), and pULK1 (S555 and S757) in the tissues of WT control and Stk11-/- mice. (b) Representative IHC for pS6 (S235/236) at lower magnification in the tissues of WT control, Atg7-/-, Stk11-/-, and Atg7-/-;Stk11-/- adult mice.
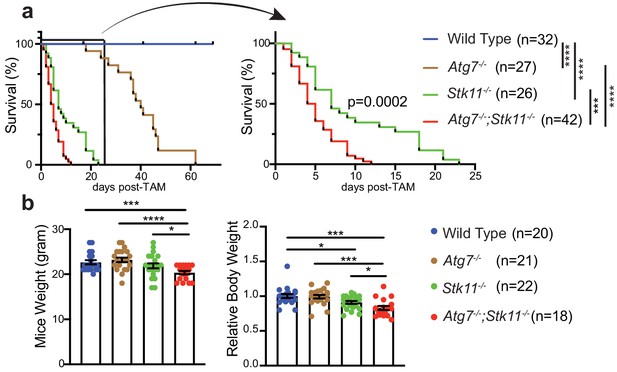
Autophagy compensates for acute Lkb1 loss to support the survival of adult mice.
(a) Kaplan-Meier survival curve of WT control, Atg7-/-, Stk11-/-, and Atg7-/-;Stk11-/- adult mice. ***p<0.001, and ****p<0.0001 (log-rank Mantel-Cox test). (b) Left: Body weight was obtained at 10 days post-TAM administration. For the actual weight, 8–10 weeks old mice with original weight range between 20 and 25 g were used. Right: For relative mice weight, each final weight was normalized to its original weight before TAM administration, subsequently normalized to the WT control. Data are mean ± s.e.m. *p<0.05, **p<0.01, and ***p<0.001.
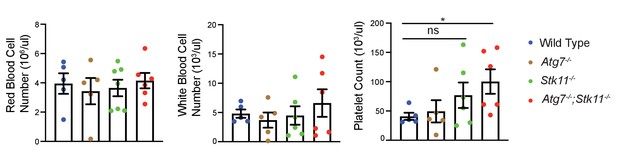
Autophagy and Lkb1 inhibitions do not lead to pancytopenia.
Quantification of red blood cells, white blood cells, and platelets of WT control, Atg7-/-, Stk11-/-, and Atg7-/-;Stk11-/- adult mice. Data are mean ± s.e.m. *p<0.05, ns: non-significant.

Autophagy ablation deteriorates impaired intestinal structure and function caused by acute Stk11 deletion.
(a) Representative H&E staining of duodenum, jejunum, and ileum for WT control, Atg7-/-, Stk11-/-, and Atg7-/-;Stk11-/- adult mice. (b) Left: Representative alcian blue staining of adult mouse intestine shows the enlargement of mucin-secreting cells in Stk11-/- and Atg7-/-;Stk11-/- mice. Right: Quantification of alcian blue positive cells in duodenum, jejunum, and ileum. Data are mean ± s.e.m. *p<0.05, **p<0.01, ***p<0.001, and ****p<0.0001. (c) Left: Representative IHC for intestinal lysozyme shows decrease of Paneth cell population in Stk11-/- and Atg7-/-;Stk11-/- crypts. Right: Quantification of lysozyme-positive cells in duodenum, jejunum, and ileum. Data are mean ± s.e.m. **p<0.01, ***p<0.001, ****p<0.0001, and ns means non-significant. (d) Left: Representative IHC for OLFM4 of intestine shows the decrease of stem cells with greater extent in Atg7-/-;Stk11-/- crypts compared with WT control and Stk11-/- mice. Right: Quantification of OLFM4-positive cells in duodenum, jejunum, and ileum. Data are mean ± s.e.m. *p<0.05, **p<0.01, and ns means non-significant. (e) Left: Representative IHC for cleaved caspase-3 of intestine delineates increase of cell death in intestine villi and tips of villi in Atg7-/-;Stk11-/- compared with WT control and Stk11-/- mice. Right: Quantification of cleaved caspase-3. Data are mean ± s.e.m. *p<0.05. (f) Left: Representative TUNEL assay in intestine sections shows increase of cell death in intestine villi and tips of villi in Atg7-/-;Stk11-/- mice compared with WT control, Atg7-/- and Stk11-/- mice. Quantification of TUNEL-positive cells in intestine. Data are mean ± s.e.m. ***p<0.001. (g) Representative relative FITC-dextran levels in sera of WT control, Atg7-/-, Stk11-/-, and Atg7-/-;Stk11-/- adult mice at 4 hr post-oral gavage of FITC-dextran. Data are mean ± s.e.m. *p<0.05, **p<0.01, ***p<0.001. (h) The level of serum D-lyxose in WT control, Atg7-/-, Stk11-/-, and Atg7-/-;Stk11-/- adult mice measured by LC-MS shows an increase of D-lyxose in Atg7-/-;Stk11-/- sera compared with WT control mice. Data are mean ± s.e.m. *p<0.05, **p<0.01. (i) Kaplan-Meier survival curve of Stk11-/-, and Atg7-/-;Stk11-/- mice treated with or without broad-spectrum antibiotics (log-rank Mantel-Cox test).
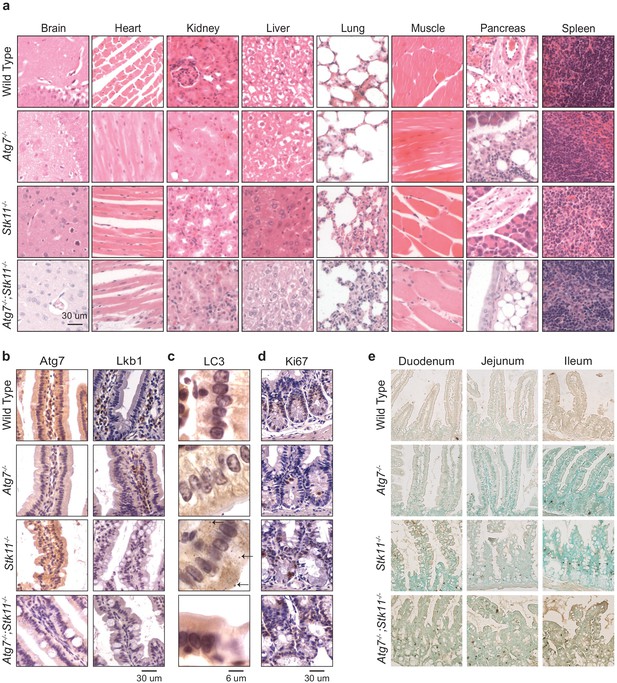
The histology of most mouse tissues and intestinal cell proliferation are not impacted by short-term deletion of Atg7 and Lkb1.
(a) Representative brain, heart, kidney, liver, lung, muscle, pancreas, and spleen histology (H&E staining) of WT control, Atg7-/-, Stk11-/-, and Atg7-/-; Stk11-/- adult mice. (b) Representative IHC for Atg7 and Lkb1 from intestine of WT control, Atg7-/-, Stk11-/- , and Atg7-/-;Stk11-/- adult mice. (c) Representative IHC for LC3 from intestine of WT control, Atg7-/-, Stk11-/-, and Atg7-/-; Stk11-/- adult mice shows accumulation of LC3II puncta (arrows) in Stk11-deficient mice. (d) Representative IHC for Ki67 from intestine of WT control, Atg7-/-, Stk11-/-, and Atg7-/-;Stk11-/- adult mice shows no significant difference in the rate of proliferation within the groups. (e) Representative lower magnification of TUNEL assay in intestine sections shows increase of cell death in intestine villi and tips of villi in Atg7-/-;Stk11-/- mice compared with WT control, Atg7-/-, and Stk11-/-.
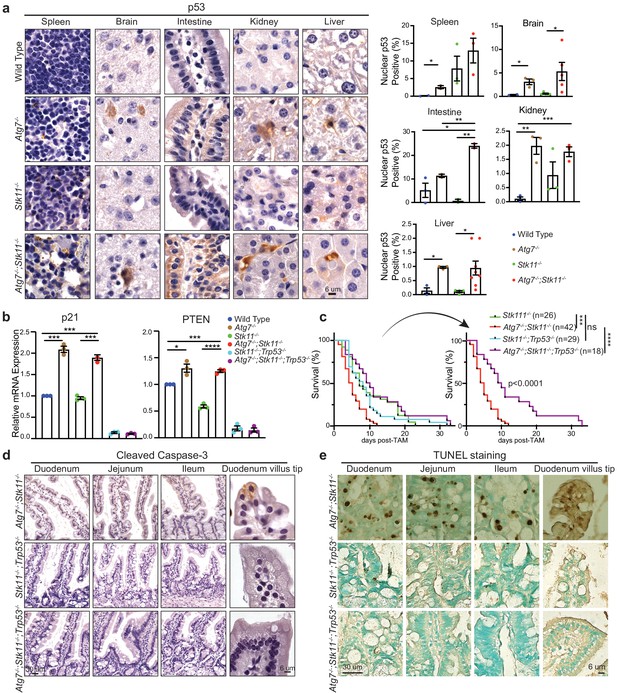
p53 deficiency extends the life span of Atg7-/-;Stk11-/- mice.
(a) Left: Representative IHC for p53 in different tissues of WT control, Atg7-/-, Stk11-/-, and Atg7-/-;Stk11-/- adult mice shows an increase of nuclear p53 in Atg7-ablated tissues. Right: Bar graphs represent the quantification of nuclear p53 in different tissues. Data are mean ± s.e.m. *p<0.05, **p<0.01, and ***p<0.001. (b) Quantitative real-time PCR of Cdkn1a (p21) and PTEN for intestine tissues of WT control, Atg7-/-, Stk11-/- and Atg7-/-;Stk11-/- adult mice. Data are mean ± s.e.m. *p<0.05, ***p<0.001, and ****p<0.0001. (c) Kaplan-Meier survival curve of Stk11-/-, Atg7-/-;Stk11-/-, Stk11-/-;Trp53-/-, and Atg7-/-;Stk11-/-;Trp53-/- adult mice. ***p<0.001, ****p<0.0001 and ns: non-significant (log-rank Mantel-Cox test). (d) Representative IHC for cleaved caspase-3 of intestine from Atg7-/-;Stk11-/-, Atg7+/+;Stk11-/-;Trp53-/-, and Atg7-/-;Stk11-/-;Trp53-/- adult mice. (e) Representative TUNEL assay of intestine from Atg7-/-;Stk11-/-, Atg7+/+;Stk11-/-;Trp53-/- and Atg7-/-;Stk11-/-;Trp53-/- adult mice.
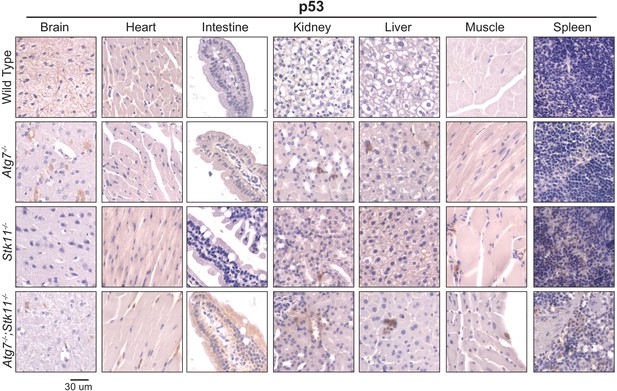
p53 is activated in the absence of autophagy.
Representative IHC of p53 with lower magnification in brain, heart, intestine, kidney, liver, muscle, and spleen of WT control, Atg7-/-, Stk11-/-, and Atg7-/-;Stk11-/- adult mice shows the increase of nuclear p53 in Atg7-ablated tissues.
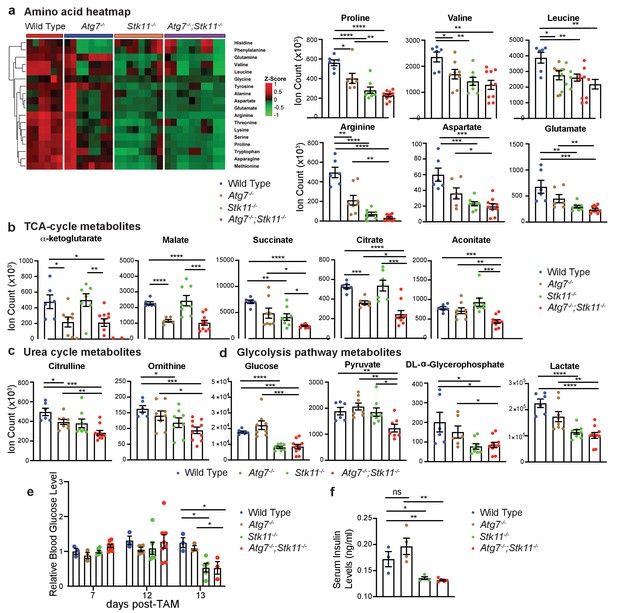
Major metabolic pathways are disturbed in Stk11-/- and Atg7-/-;Stk11-/- mice.
(a) Left: Representative heat map of all the amino acids in sera of WT control, Atg7-/-, Stk11-/-, and Atg7-/-;Stk11-/- adult mice compared with WT control mice at fasting state. Right: Bar graphs show the levels of amino acids that are significantly decreased in Stk11-/- and Atg7-/-;Stk11-/- mice sera compared with WT control mice. Data are mean ± s.e.m. *p<0.05, **p<0.01, ***p<0.001, and ****p<0.0001. (b-d). Metabolites that are significantly decreased in the sera of Stk11-/- and Atg7-/-;Stk11-/- mice compared with WT control mice at fasting state. Data are mean ± s.e.m. *p<0.05, **p<0.01, ***p<0.001, and ****p<0.0001. (e) Relative blood glucose levels of WT control, Atg7-/-, Stk11-/- and Atg7-/-;Stk11-/- adult mice normalized to WT control mice at fed state for the indicated time course after first TAM injection. Data are mean ± s.e.m. *p<0.05. (f) Quantification of serum insulin levels of WT control, Atg7-/-, Stk11-/- and Atg7-/-;Stk11-/- adult mice at fasted state at 10 days post-deletion. Data are mean ± s.e.m. *p<0.05, **p<0.01, ns: non-significant.
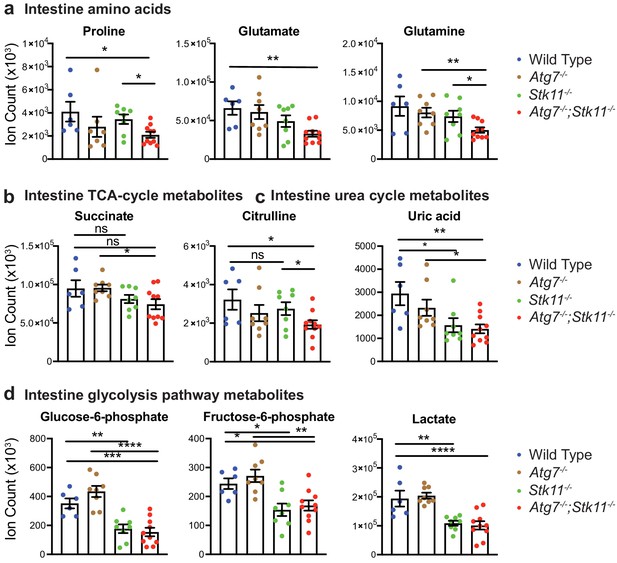
Acute loss of Lkb1 alone or together with Atg7 altered the levels of metabolites in intestine.
(a–d) Metabolites that are significantly decreased in the intestine of Stk11-/- and Atg7-/-;Stk11-/- mice compared with WT control mice. Data are mean ± s.e.m. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001.
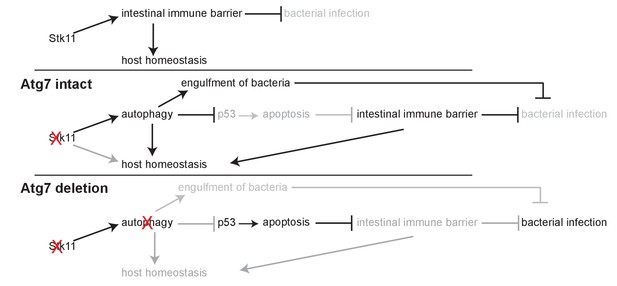
Mechanism by which Lkb1 interacts with autophagy to support adult mice homeostasis and survival.
Loss of Lkb1 causes hypoglycemia, impaired intestinal epithelium barrier integrity, increased general infection, and disturbed host sera metabolism. When autophagy is intact, these dysfunctions are temporarily compensated by autophagy upregulation, partly through preventing p53 activation. However, autophagy deficiency further exacerbates the dysfunctions induced by Stk11 deletion, thereby accelerating mouse death.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Antibody | Atg7 (Rabbit polyclonal) | Sigma Aldrich | Cat# A2856 | IHC: (1:400) WB: (1:2000) |
| Antibody | Lkb1 (mouse monoclonal) | Sant Cruz Biotechnology | Cat# sc-32245 | IHC: (1:50) WB: (1:500) |
| Antibody | P62 (Guinea pig)/ (Rabbit polyclonal) | Enzo Life Sciences/ American Research Products | Cat# PW9860-0100/ 03-GP62-C | IHC: (1:1000) WB: (1:2500) |
| Antibody | p-AMPKTh172(Rabbit polyclonal) | Cell Signaling Technology | Cat#: 2535S | IHC: (1:100) |
| Antibody | p-ACCS79(Rabbit polyclonal) | Cell Signaling Technology | Cat#: 3661 | IHC: (1:800) WB: (1:1000) |
| Antibody | ACC (Rabbit polyclonal) | Cell Signaling Technology | Cat#: 3676 | WB: (1:1000) |
| Antibody | p-S6S235/236(Rabbit polyclonal) | Cell Signaling Technology | Cat#: 4858 | IHC: (1:300) WB: (1:2000) |
| Antibody | S6 (Rabbit polyclonal) | Cell Signaling Technology | Cat#: 2217 | WB: (1:1000) |
| Antibody | p-ULK1S555(Rabbit polyclonal) | Cell Signaling Technology | Cat#: 5869 | IHC: (1:100) |
| Antibody | p-ULK1S757(Rabbit polyclonal) | Cell Signaling Technology | Cat. #: 14202 | IHC: (1:800) |
| Antibody | LC3 (Rabbit polyclonal) | Nano Tolls | Cat. #: LC3-5F10 | IHC: (1:100) WB: (1:4000) |
| Antibody | Ki67 (Rabbit polyclonal) | Abcam | Cat. #: ab-15580 | IHC: (1:400) |
| Antibody | Cleaved Caspase3 (Rabbit polyclonal) | Cell Signaling Technology | Cat. #: 9661S | IHC: (1:250) |
| Antibody | OLFM4 (Rabbit polyclonal) | Cell Signaling Technology | Cat. #:39141 | IHC: (1:2000) |
| Antibody | Lysozyme (Rabbit polyclonal) | Aligent | Cat. #: A0099 | IHC: (1:2000) |
| Antibody | P53 (mouse monoclonal) | Novus Biologicals | Cat. #: NB200-103SS | IHC: (1:300) |
| Antibody | Atg5 (Rabbit polyclonal) | Abcam | Cat. #: ab108327 | WB: (1:500) |
| Antibody | β-actin (mouse monoclonal) | Sigma Aldrich | Cat. #: A1978 | WB: (1:2000) |
| Commercial assay or kit | TUNEL assay | Abcam | ab206386 | |
| Software, algorithm | Prism GraphPad | Prism 8 | RRID:SCR_002798 | |
| Software, algorithm | Adobe Illustrator | CC2020 | RRID:SCR_010279 | |
| Mouse genotype | Ubc-CreERT2 | Jackson Laboratory | ||
| Mouse genotype | Stk11flox/flox | Jackson Laboratory | ||
| Mouse genotype | Trp53flox/flox | Jackson Laboratory | ||
| Mouse genotype | Atg7flox/flox | Komatsu et al., 2005 |
Additional files
-
Source data 1
Levels of metabolites in the sera of mice measured by LC_MS.
- https://cdn.elifesciences.org/articles/62377/elife-62377-data1-v2.xlsx
-
Source data 2
Levels of metabolites in the intestine of mice measured by LC_MS.
- https://cdn.elifesciences.org/articles/62377/elife-62377-data2-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/62377/elife-62377-transrepform-v2.pdf





