Integron activity accelerates the evolution of antibiotic resistance
Figures
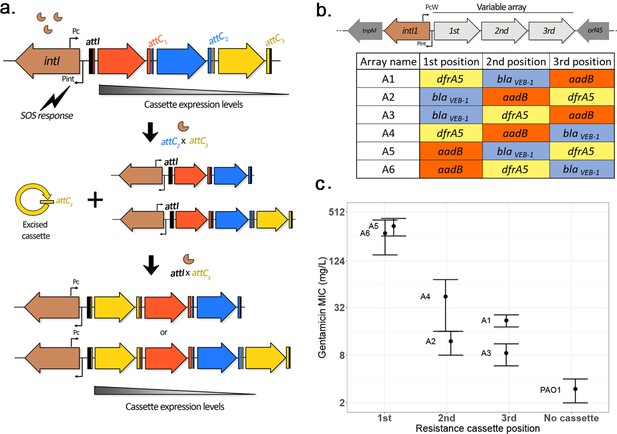
Overview of integron system.
(a) Diagram of the integron mechanism: the integron consists of an integrase gene, intI, followed by an array of promoterless gene cassettes (represented here by arrows). Cassettes are expressed from the Pc promoter within the integrase gene, with decreasing cassette expression along the array. Following the induction of the SOS response, the integrase enzyme promotes cassette excision (recombination between a cassette attC site and the attC of the preceding cassette, causing excision of the cassette into its circular form). Due to the presence of a replication step in the excision process, a copy of the original array is conserved. Re-integration of the circular cassette can then occur through recombination between the cassette attC site and the attI site located at the start of either array, leading to an apparent ‘cut-and-paste’ recombination if the cassette integrates in the excised array, or can be assimilated to a ‘copy-and-paste’ outcome if it integrates in a conserved copy of the array. By shuffling and duplicating cassettes, the integron has the potential to quickly modulate cassette expression levels. (b) Custom integron arrays: the native integron array of the R388 plasmid was replaced by the custom integron arrays WTA1 to WTA6 containing three integron cassettes in every possible order. (c) Effect of position of the aadB cassette on gentamicin resistance levels. Error bars represent standard error (n = 2–4).
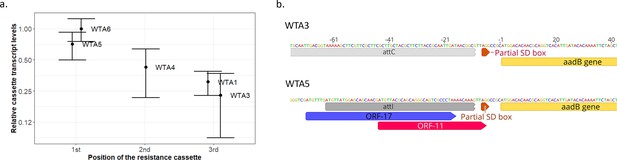
Transcriptional and translational origin of the aadB expression gradient.
(A) Transcript levels of the aadB cassette in the different arrays. Transcription levels are normalized relatively to the best transcribed array (WTA6). Error bars represent the standard error of three independent biological replicates. (B) Representation of the genomic environment of the aadB cassette when aadB is in last (WTA3) or first position (WTA5) in our arrays. The two ORFs overlapping the attI, which have been shown to improve cassette translation (Hanau-Berçot et al., 2002; Papagiannitsis et al., 2017), are represented in color.
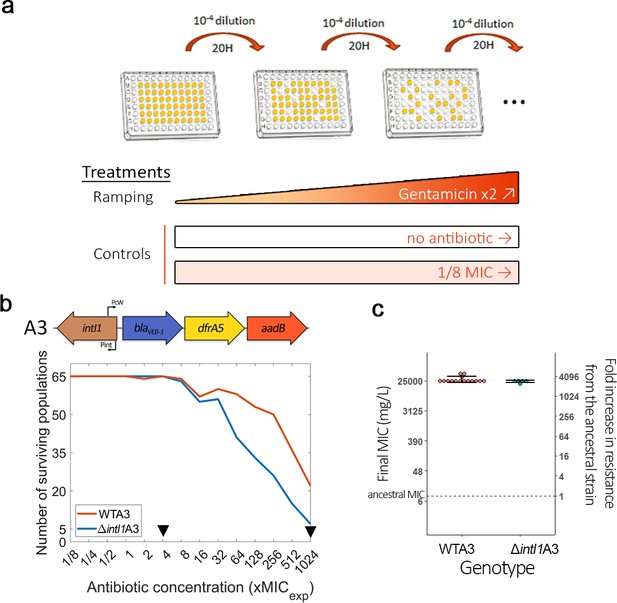
Integrase activity can increase bacteria evolvability against antibiotics.
(a) Schematic representation of the experimental evolution protocol. (b) Top: Representation of the WTA3 integron. Bottom: Survival curves of the PA01:WTA3 and PA01:ΔintI1A3 populations during ramping treatment, monitored using OD595. The black triangles represent time points where populations were sequenced using whole genome sequencing. (c) Average minimum inhibitory concentration (MIC) of a random subset of the PA01:WTA3 (15 populations) and PA01:ΔintI1A3 populations (five populations) from the final ×1024 MIC time point. The MIC of the ancestral populations is represented by a dashed line. Error bars represent standard deviation. The MIC of each population was averaged from three biological replicates.
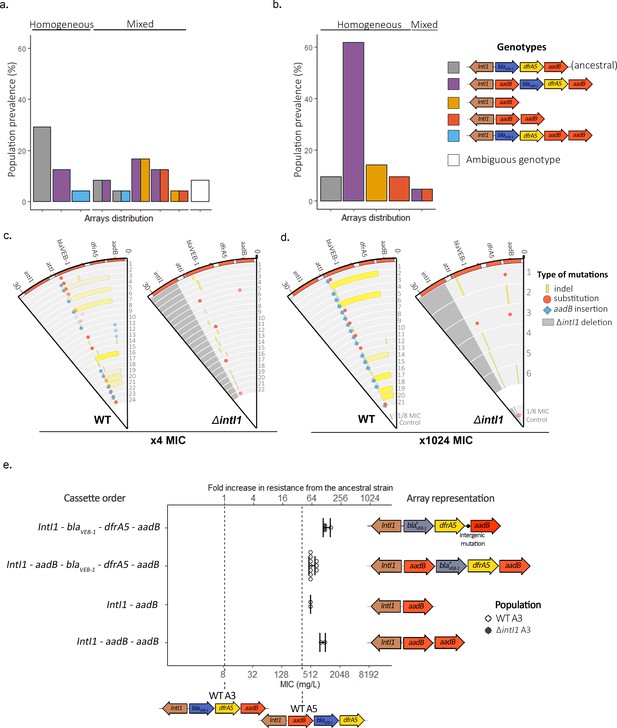
Extensive cassette re-arrangements are linked with integrase activity.
(a,b) Distribution of cassette re-arrangements at ×4 MIC (a) and ×1024 MIC (b) time points in the WTA3 populations. Homogeneous populations represent populations where only one type of array could be identified while mixed populations contain different arrays as indicated by the corresponding colors. Ambiguous populations correspond to re-arrangements that could not be identified with confidence from short-read data. No re-arrangement was found in the ΔintI1A3 populations. (c,d) Representation of the plasmid mutations and re-arrangements in the surviving PA01:WTA3 and PA01: ΔintI1A3 populations at ×4 MIC (c) and ×1024 MIC (d), mapped to the integron reference sequence. Each circle represents a separate population, with the inner circle representing the variants present in an equimolar pool of six 1/8 MIC control populations. Indels are represented in yellow and single-nucleotide substitutions in red. aadB insertions are represented by blue lozenges. The color intensity represents the frequency of the corresponding mutation/recombination. The dark gray area in the PA01:ΔintI1A3 populations represents the location of the intI1 deletion. (e) Resistance levels provided by evolved plasmids in the ancestral chromosomal background. The plasmids of 15 PA01:WTA3 and 2 PA01:ΔintI1A3 populations were extracted and transformed back into the ancestral PA01 strain. The populations are grouped by array cassette order. Other mutations in the arrays (blaVEB-1 mutations or intergenic mutations upstream of aadB) are indicated in the arrays representations on the right. Resistance levels of the less (WTA3) and most (WTA5) resistant unevolved custom arrays are represented by the dashed lines. Error bars represent standard deviation and the MIC of each plasmid was averaged from three biological replicates.
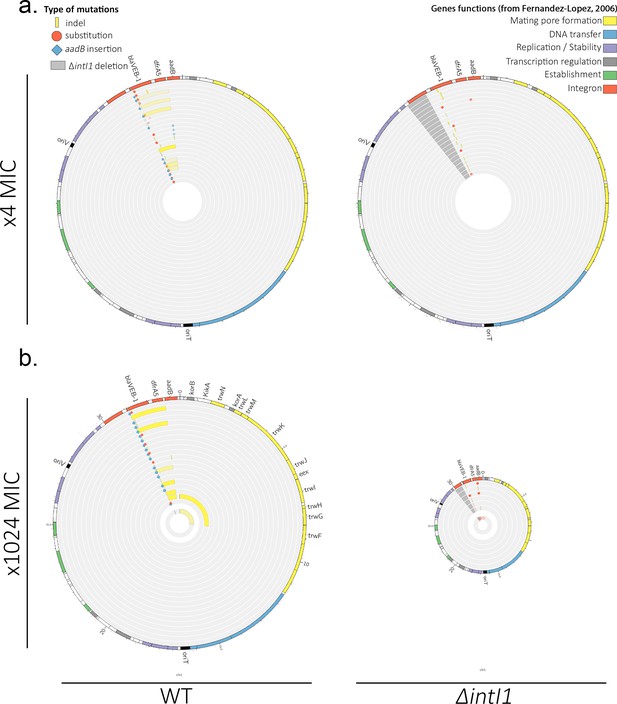
Plasmid mutations and re-arrangements at (a) ×4 MIC and (b) ×1024 MIC.
Representation of the plasmid mutations and re-arrangements in PA01:WTA3 and PA01: ΔintI1A3 populations, mapped to the plasmid reference sequence. Each circle represents a separate population. Indels are represented in yellow and single-nucleotide substitutions in red. aadB insertions are represented by blue lozenges. The color intensity represents the frequency of the corresponding mutation/recombination. The dark gray area in the PA01:ΔintI1 populations represents the location of the intI1 deletion. The function of each R388 gene as described in Fernández-López et al., 2006 is indicated by a specific color in the outer circle. Apart from oriT and oriV, given as reference, only the name of genes where mutations are located is indicated. In (b), the inner circles correspond to the equimolar ⅛ MIC control populations.
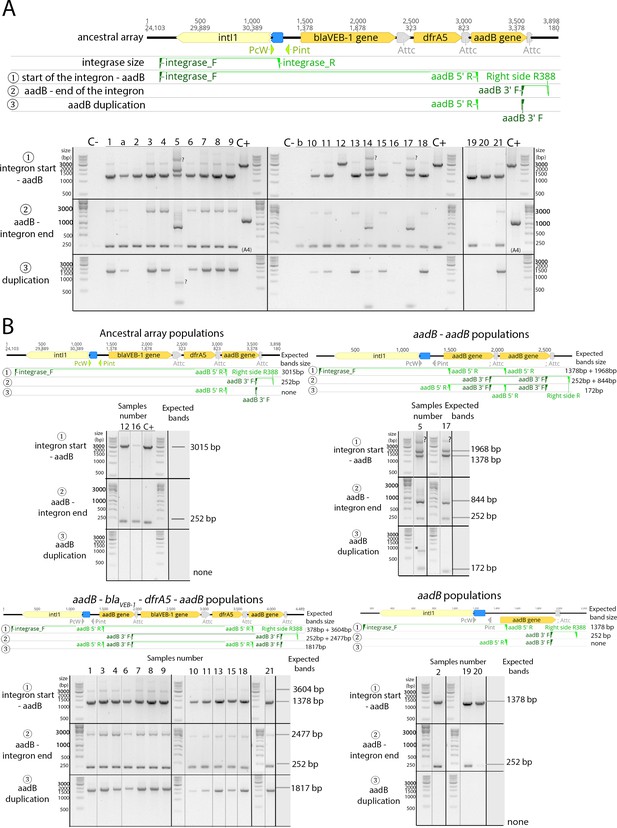
Re-arrangements detection by PCR in ×1024 MIC WTA3 populations.
(A) Top: Primer binding sites and amplicons used in the screen for integrase size and cassette re-arrangements. Bottom: Full gels of the PCR screen for cassettes re-arrangements of the ×1024 MIC WTA3 populations. The population number identifier is indicated at the top. Genomic DNA from the WTA3 ancestor was used as positive control, unless indicated. Samples with a letter corresponds to populations latter excluded from the analysis due to detected cross-contamination. The former location of non-relevant lanes excluded from the gels images is indicated by the gray vertical line, and care was taken to keep the vertical alignment within the gels pictures during the figure composition. (B) Predicted amplicons and corresponding populations for each of the cassette array arrangements identified at the ×1024 MIC time point by full genome sequencing. Bands that cannot be explained by the genomic data are indicated by a question mark (?). Black borders separate different gels images. Gray lines indicate where the full gel images were spliced to keep only the corresponding populations while keeping the vertical alignment to the size ladder. Alignments figures created using Geneious version 10.2.
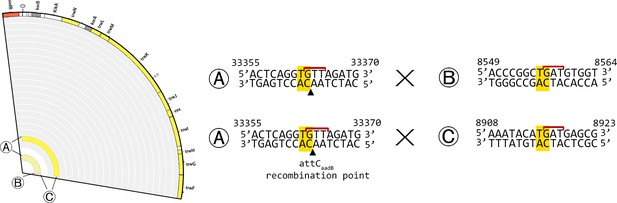
Re-arrangements in the plasmid backbone of the WT populations at ×1024 MIC.
Left: Re-arrangements in the plasmid backbone of the WT populations at ×1024 MIC. Each junction site is indicated by a letter. Right: junction sequences for each re-arrangement. The junction site is indicated in yellow (as the crossover point is unclear due to sequence homology between each junction, the entire homology is highlighted). The GNT integron secondary motif is indicated by a red line. The recombination point of the aadB attC site is indicated by a black arrow.
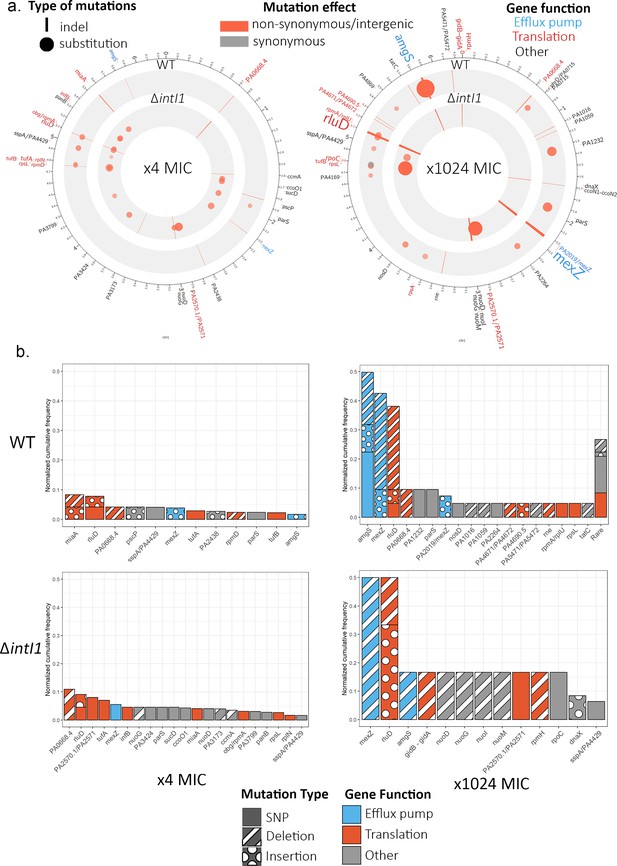
Chromosome evolution.
(a) Summary of the chromosomal mutations at ×4 MIC (left) and at ×1024 MIC (right) mapped to the PAO1 reference sequence. Each circle represents a summary of each genotype. The type (indel, substitution) of mutation for each gene is represented by the shape of the marker (line, circle), while the marker color represents the effect of the mutations (nonsynonymous/intergenic vs. synonymous), and its color intensity and size represent its normalized cumulative frequency per gene. The size of the gene labels on the outer ring represents the overall cumulative frequency of mutations present in this gene across all populations from this time point. (b) Cumulative frequency of mutations for each gene normalized by the number of populations within each genotype and time point. Genes are colored by resistance mechanism, and the type of the mutations (single-nucleotide substitution, insertion, deletion) is indicated by the patterning.
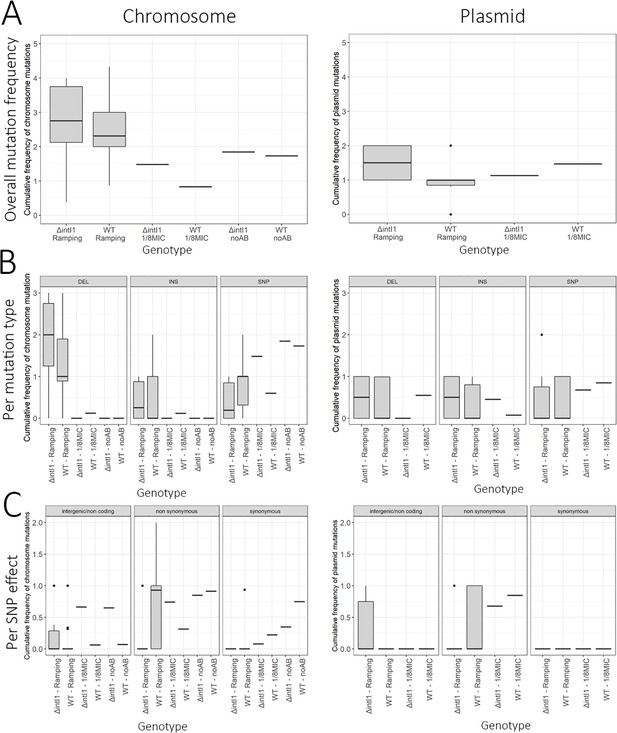
Summary statistics of mutations in the ×1024 MIC populations.
Box plots representing the average cumulative mutation frequencies for each population at the ×1024 MIC time point for (A) all mutations (B) per mutation type (C) per mutation effect (SNP only). The lower and upper hinges correspond to the first and third quartiles, while the middle line corresponds to the median. Cassette re-arrangements (duplications and deletions) are not included.
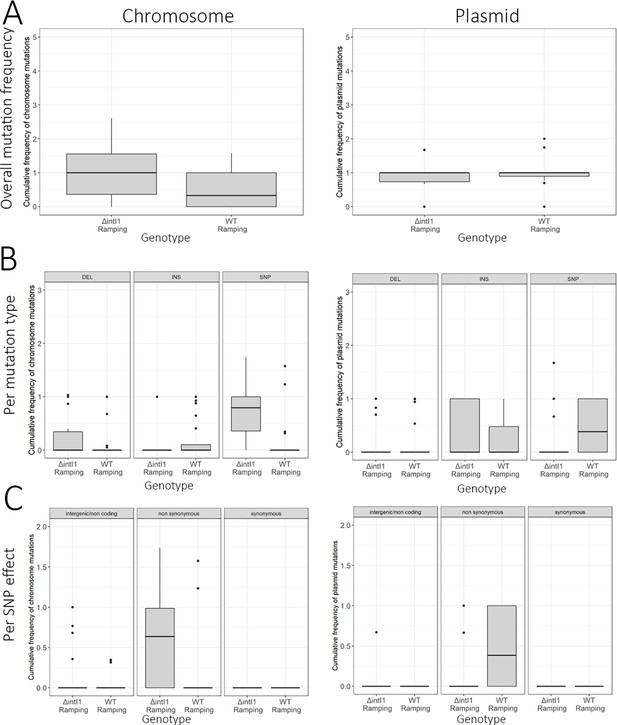
Summary statistics of mutations in the ×4 MIC populations.
Box plots representing the average cumulative mutation frequencies for each population at the ×4 MIC time point for (A) all mutations (B) per mutation type (C) per mutation effect (SNP only). The lower and upper hinges correspond to the first and third quartiles while the middle line corresponds to the median. Cassette re-arrangements (duplications and deletions) are not included. Supplementary Informations.
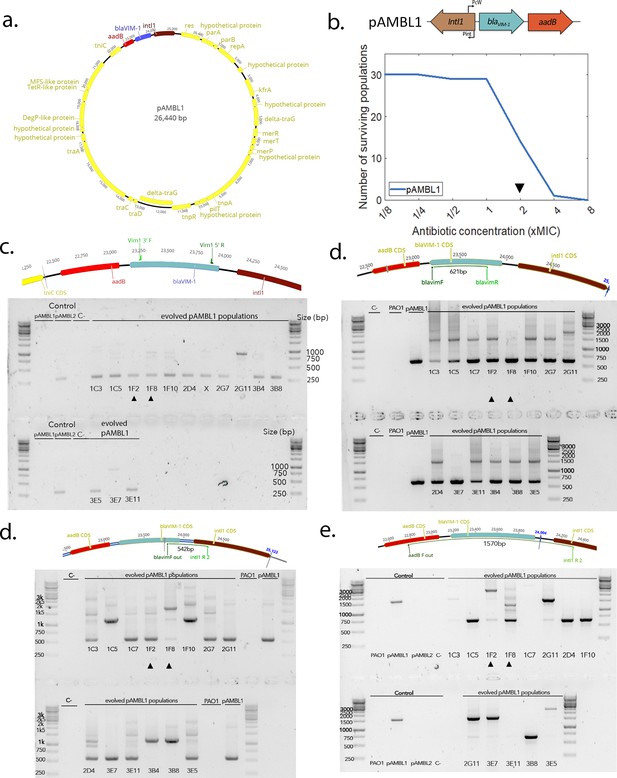
pAMBL1 re-arrangements.
(a) Representation of the pAMBL1 plasmid. The integron is highlighted in color. (b) Survival curve of the 30 PA01:pAMBL1 populations under ramping treatment. The black arrow indicates the time point at which populations were plated and duplications amplified by PCR/NGS. (c–e) Detection of cassette re-arrangements by PCR. The expected positions of the primers on the ancestral pAMBL1 and the size of the corresponding amplicon are indicated on top.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Pseudomonas aeruginosa) | PA01 | Lab strain | NC_002516 | |
| Strain, strain background (Escherichia coli) | MG-1 | Poirel et al., 1999 | AF205943 | E. coli clinical isolate containing a qacI–aadB–aacA1/orfG–blaVEB1–aadB–arr2–cmIA5–blaOXA–10/aadA1 integron array |
| Strain, strain background (Escherichia coli) | EIEC-4 | Gassama et al., 2004 | E. coli clinical isolate containing a dfrA5 integron cassette | |
| Recombinant DNA reagent | R388 | Avila and de la Cruz, 1988 | NC_028464.1 | |
| Recombinant DNA reagent | WTA1 | This study | Custom integron array drfA5–blaVEB1–aadB on R388 plasmid backbone | |
| Recombinant DNA reagent | WTA2 | This study | Custom integron array blaVEB1– aadB–dfrA5 on R388 plasmid backbone | |
| Recombinant DNA reagent | WTA3 | This study | Custom integron array blaVEB1–dfrA5–aadB on R388 plasmid backbone | |
| Recombinant DNA reagent | WTA4 | This study | Custom integron array dfrA5–aadB–blaVEB1 on R388 plasmid backbone | |
| Recombinant DNA reagent | WTA5 | This study | Custom integron array aadB–blaVEB1– dfrA5 on R388 plasmid backbone | |
| Recombinant DNA reagent | WTA6 | This study | Custom integron array aadB–dfrA5–blaVEB1 on R388 plasmid backbone | |
| Recombinant DNA reagent | Δint1A3 | This study | Array WTA3 with a 948 bp deletion of the integrase intI1 | |
| Recombinant DNA reagent | pAMBL1 | San Millan et al., 2015a | KP873172.1 | Clinical plasmid containing a blaVIM-1–aadB integron array |
| Software, algorithm | breseq | Barrick et al., 2014 | RRID:SCR_010810 | Version 0.33.2 |
| Software, algorithm | CNOGpro | Brynildsrud, 2018 |
Additional files
-
Supplementary file 1
Primers used in this study.
- https://cdn.elifesciences.org/articles/62474/elife-62474-supp1-v2.xlsx
-
Supplementary file 2
List of mutations / recombinations at x1024 MIC.
- https://cdn.elifesciences.org/articles/62474/elife-62474-supp2-v2.xlsx
-
Supplementary file 3
List of mutations / recombinations at x4 MIC.
- https://cdn.elifesciences.org/articles/62474/elife-62474-supp3-v2.xlsx
-
Supplementary file 4
List of duplicated cassettes from the INTEGRALL database.
- https://cdn.elifesciences.org/articles/62474/elife-62474-supp4-v2.doc
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/62474/elife-62474-transrepform-v2.pdf