Glucose restriction drives spatial reorganization of mevalonate metabolism
Figures
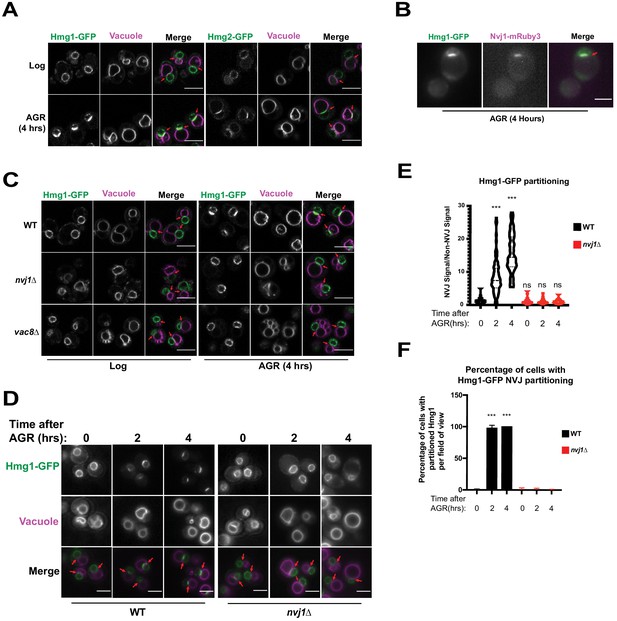
The HMG-CoA Reductases, Hmg1 and Hmg2, enrich at the NVJ in an Nvj1-dependent manner during AGR.
(A) Confocal microscopy images of yeast expressing endogenously tagged Hmg1-GFP or Hmg2-GFP (green) grown in SC media with 2% glucose to an OD of 0.5 (Log), or in 0.001% glucose for 4 hr (AGR). Vacuoles (magenta) were visualized by staining with FM4-64. Red arrows represent the relative location of the Nucleus-Vacuole Junction (NVJ), or where the nuclear envelope is closely apposed to the vacuole in nvj1Δ cells. Scale bars = 5 μm. (B) Epifluorescence microscopy images showing the overlap of Hmg1-GFP with Nvj1-mRuby3 after cells were exposed to four hours of AGR. Red arrows represent the location of the NVJ. Scale bars = 5 μm (C) Confocal microscopy images of yeast expressing endogenously tagged Hmg1-GFP (green) in wild type (WT), nvj1Δ, or vac8Δ yeast. Red arrows represent the location of NVJ. (D) Epifluorescence microscopy images of WT or nvj1Δ yeast expressing endogenously tagged Hmg1-GFP growing in 2% glucose to an OD of 0.5 (Log) or in 0.001% glucose for 2 or 4 hr (AGR). (E) Quantification of Hmg1 partitioning measured by line scan across the nuclear envelope (NE) and plotted as the ratio of Hmg1-GFP intensity at the NVJ relative to Hmg1-GFP intensity opposite the NVJ. Scale bars represent 5 μm. (Brown-Forsyth and Welch ANOVA. N > 51 cells. ***p-value<0.001). (F) Quantification of percentage of cells displaying Hmg1 NVJ partitioning per field of view. Any cell with partitioning >2.0 was considered as displaying Hmg1 NVJ partitioning. (Brown-Forsyth and Welch ANOVA. N > 51 cells. ***p-value<0.001).
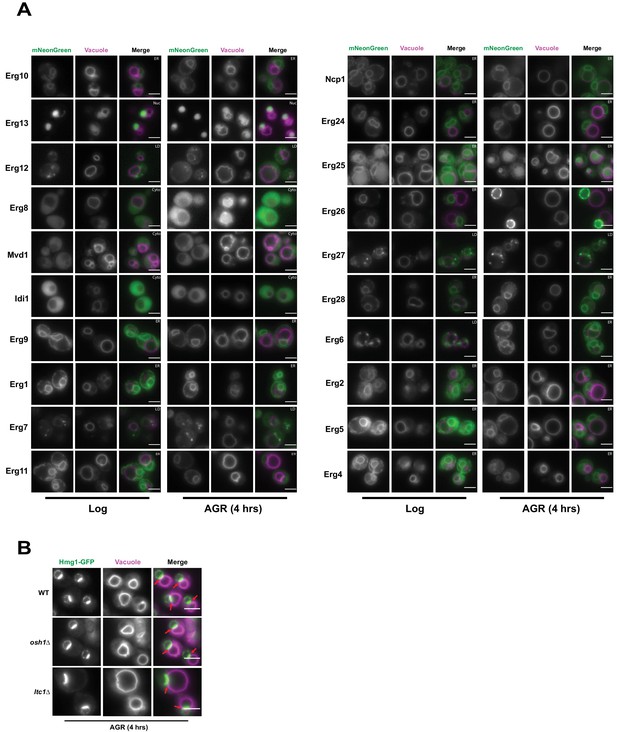
Hmg1 NVJ partitioning is specific amongst ergosterol biosynthesis proteins and independent of NVJ-associated sterol transport proteins.
(A) Epifluorescence microscopy images of twenty ergosterol biosynthesis proteins endogenously tagged with mNeonGreen (mNg) imaged in log phase and after 4 hr of AGR. Cells were co-stained with FM4-64 (magenta) to visualize vacuoles. Cellular compartments occupied by each protein are designated in white text in the top right corner of each image. Scale bars = 5 μm. (B) Epifluorescence microscopy images of cells endogenously tagged with Hmg1-GFP in wild type (WT), osh1Δ, or ltc1Δ cells exposed to 4 hr of AGR. Scale bars = 5 μm.
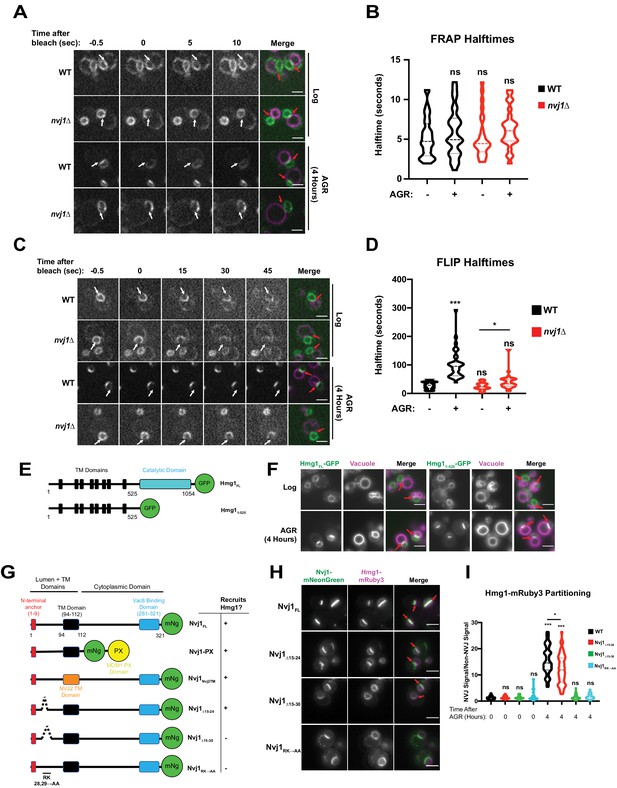
Hmg1 is selectively retained at the NVJ and requires a luminal domain of Nvj1.
(A) Spinning disk confocal microscopy images from FRAP movies of yeast expressing endogenously tagged Hmg1-GFP and stained with FM4-64 (magenta). Portions of the NE corresponding to the NVJ were photobleached (white arrows) and recovery was monitored in both WT and nvj1Δ cells grown to log phase or after 4 hr of AGR. Red arrows represent the location of the NVJ. (B) Quantification of Hmg1-GFP FRAP halftimes after photobleaching. (Brown-Forsyth and Welch ANOVA. N > 26 cells) (C) Spinning disk confocal microscopy images taken from FLIP movies. Conditions and strains were the same as in (A). One region of the NE that lies opposite the NVJ was photobleached every 5 s (white arrows) and Hmg1-GFP signal at the NVJ was monitored for loss of fluorescence. Time above the images indicates seconds after the first bleach pulse. Red arrows represent the location of the NVJ (D) Quantification of FLIP halftimes. (Brown-Forsyth and Welch ANOVA. N > 29 cells. *p-value>0.05 ***p-value<0.001). (E) Cartoon of Hmg1-GFP constructs endogenously expressed in yeast with both the cytosolic catalytic domain (blue) and TM/luminal domains (black) (Hmg1FL) or the TM/luminal domains alone (Hmg11-525). (F) Epifluorescence microscopy images of yeast expressing endogenously tagged Hmg1FL-GFP or Hmg11-525-GFP. (G) Cartoon of Nvj1-mNeonGreen constructs expressed in nvj1Δ cells. Nvj1FL-mNg contains all domains of Nvj1 including the N-terminal luminal anchor (red), the TM domain (black), and the Vac8-binding domain (VBD) (blue). Chimeric constructs either replaced the VBD with a vacuole-binding PX domain (Nvj1PX) or replaced the endogenous TM domain with the TM domain of Nvj2 (Nvj1NVJ2TM). Images of these constructs can be found in Figure 3—figure supplement 1A. Truncations of Nvj1 removed either residues 15–24 (Nvj115-24Δ), 15–30 (Nvj115-30Δ), or mutated residues 28 and 29 to alanine (Nvj1RK→AA). (H) Epifluorescence microscopy images of yeast expressing truncation constructs depicted in (G). Cells were co-expressing endogenously tagged Hmg1-mRuby3 (magenta). Scale bars represent 5 μm. (I) Quantification of Hmg1 partitioning at the NVJ for yeast expressing truncations of Nvj1-mNg and Hmg1-mRuby3. (Brown-Forsyth and Welch ANOVA. N > 61 cells. *p-value<0.05 ***p-value<0.001).
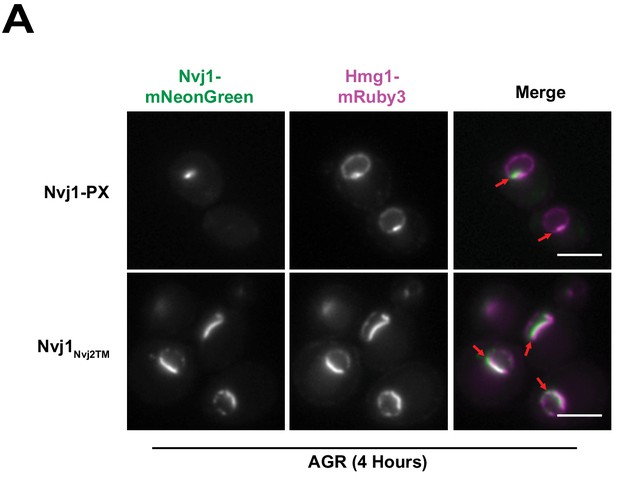
Hmg1 NVJ partitioning is not dependent on the cytoplasmic or transmembrane regions of Nvj1.
(A) Epifluorescence microscopy images of cells expressing ADH promoter-driven Nvj1-mNeonGreen (Nvj1-mNg) chimeric constructs (green) either substituting the endogenous transmembrane region with that of Nvj2 (Nvj1Nvj2TM) or substituting the endogenous Vac8 binding domain with a vacuole binding PX domain (Nvj1-PX). Chimera strains co-expressed endogenously tagged Hmg1-mRuby3 (magenta) and lacked endogenous Nvj1. Images were taken after exposing cells to four hours of AGR. Scale bars represent 5μm.
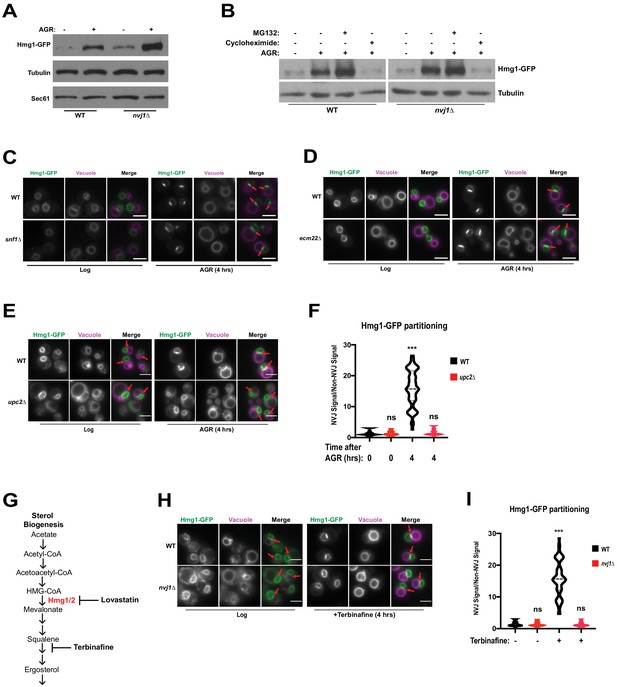
Hmg1 protein accumulates in nvj1Δ yeast via enhanced synthesis, which coincides with Upc2-dependent Hmg1 clustering at the NVJ.
(A) Immunoblot of cells expressing endogenously tagged Hmg1-GFP grown to log-phase or treated with AGR for 4 hr. Tubulin and Sec61 antibodies were used as loading controls. (B) Immunoblot of Hmg1-GFP expressing cells grown to log-phase or treated with AGR for 4 hr. AGR-treated cells were also co-treated with either 100 μg/mL cycloheximide or 25 μM MG132. Tubulin was used as a loading control. (C) Epifluorescence microscopy images of cells expressing endogenously tagged Hmg1-GFP in either a wild type (WT) or Snf1 knock-out (snf1Δ) background. (D) Epifluorescence microscopy images of cells expressing endogenously tagged Hmg1-GFP in either a wild type (WT) or ecm22Δ background. Scale bars = 5 μm. (E) Epifluorescence microscopy images of endogenously tagged Hmg1-GFP (green) with FM4-64-stained vacuoles (magenta) grown to log phase or after 4 hr of AGR in wild type (WT) and upc2Δ yeast. Scale bars = 5 μm. Red arrows indicate relative position of the NVJ. (F) Quantification of Hmg1-GFP partitioning at NVJ from images represented in (E). (Brown-Forsyth and Welch ANOVA. N > 49 cells. ***p-value<0.001). (G) Schematic representing abbreviated ergosterol biogenesis pathway. (H) Epifluorescence microscopy images of wild type and nvj1Δ cells expressing endogenously tagged Hmg1-GFP treated with 10 μg/mL of terbinafine for 4 hr. Scale bars represent 5 μm. (I) Quantification of Hmg1-GFP partitioning at the NVJ from images shown in (H). (Brown-Forsyth and Welch ANOVA. N > 51 cells. ***p-value<0.001).
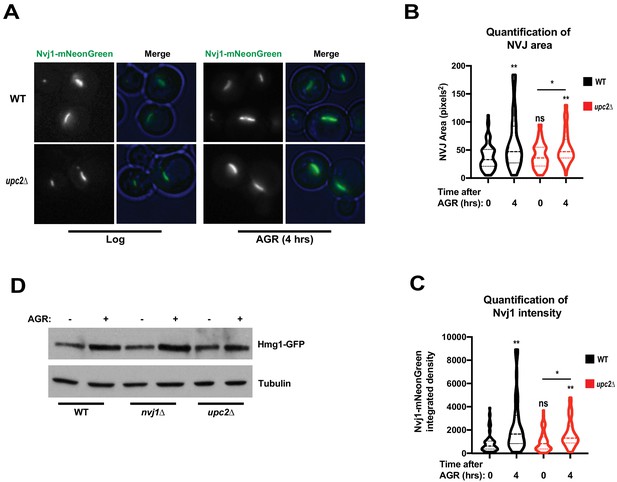
Loss of Hmg1 partitioning in upc2Δ cells does not function through Nvj1 or Hmg1 expression.
(A) Epifluorescence images of yeast expressing Nvj1-mNeonGreen in either a WT or upc2Δ background. Blue outline was generated in Fiji by converting brightfield images to blue images and using ‘adjust color balance’ function. (B) Quantification of NVJ area, as measured by the length of Nvj1-mNeonGreen signal in each cell. (Brown-Forsyth and Welch ANOVA. N > 68. *p-value<0.05 **p-value<0.01). (C) Quantification of Nvj1-mNeonGreen signal intensity in either WT or upc2Δ cells. (Brown-Forsyth and Welch ANOVA. N > 68. *p-value<0.05 **p-value<0.01). (D) Immunoblot analysis of cells expressing endogenously tagged Hmg1-GFP in either WT, nvj1Δ, or upc2Δ backgrounds grown to either log phase or treated with 4 hr of AGR. Tubulin was used as a loading control.
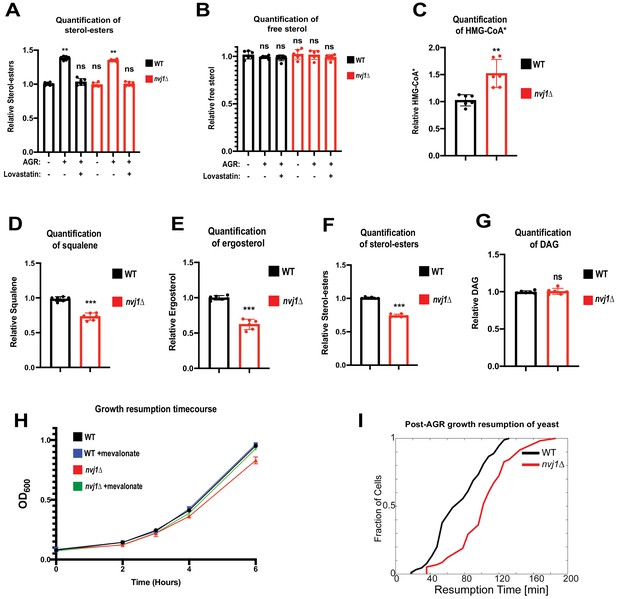
Hmg1 NVJ partitioning coincides with de novo sterol-ester production and increased Hmg1 catalytic efficiency.
(A) Quantification of SEs in wild type and nvj1Δ cells collected at log phase, after 4 hr of AGR, or after a 4 hr co-treatment of AGR in the presence of 20 μg/mL lovastatin. Neutral lipids were extracted, separated by TLC, and visualized using Cu(II) Sulfate spray and charring. Quantification performed by densitometry. (Brown-Forsyth and Welch ANOVA. N = 6. **p-value<0.01) (B) Quantification of free sterols using the same strains and methodology as in (A). (Brown-Forsyth and Welch ANOVA. N = 6. ***p-value<0.001). (C) Scintillation counting quantification of radiolabeled HMG-CoA* produced after a 15 min pulse with 14C-acetate in intact WT or nvj1Δ cells grown for 4 hr in AGR. After quenching the radio-labeling reaction, soluble metabolites were isolated and endogenous HMG-CoA was converted into mevalonate using an in vitro HMGCR reaction. Mevalonate was subsequently separated and isolated by TLC, visualized by autoradiography, and quantified by scintillation counting. (Brown-Forsyth and Welch ANOVA. N = 6. **p-value<0.01). (D–G) Quantifications of radiolabeled DAG and mevalonate-derived lipids. Radio-labeling was performed as described in (C). Lipids were extracted and separated by TLC and visualized by autoradiography. Squalene and SE bands were quantified by scintillation counting. Ergosterol and DAG bands were quantified by densitometry. (Brown-Forsyth and Welch ANOVA. N = 6. ***p-value<0.001). (H) Growth curves of WT and nvj1Δ cells. Cells were treated with AGR for 10 hr in the absence or presence of 10 μg/mL mevalonate, and subsequently diluted to OD600 = 0.1 in SC media containing 2% glucose lacking mevalonate. OD600 measurements were taken every hour following dilution in fresh glucose-containing media. (I) Resumption probability graph depicting probability of cells producing a daughter bud as a function of time following an SC to SC+glucose transition (N > 85 cells).
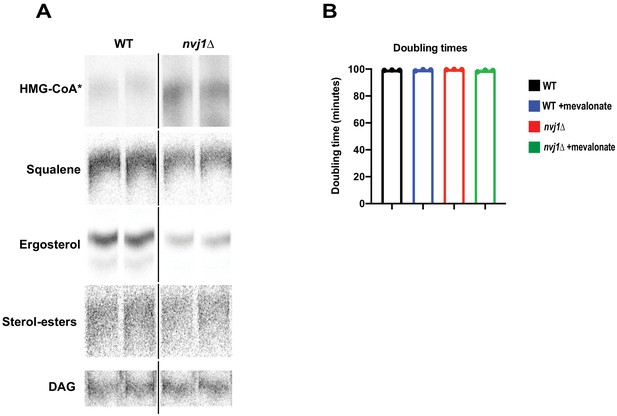
Cells lacking endogenous Nvj1 accumulate Hmg1 substrates and have decreased mevalonate-derived pathway products.
(A) Autoradiograms taken of TLC plates to visualize HMG-CoA* (visualized as mevalonate, see Figure 3 legend or supplemental methods) or squalene, ergosterol, SE, and DAG in wild type (WT) or nvj1Δ cells. Squalene, ergosterol, SE, and DAG were visualized on the same plate. DAG was used as a control band for neutral lipid species. Brightness/contrast was adjusted linearly in ImageJ to visualize bands. (B) Quantification of growth resumption in WT and nvj1Δ cells after ten hours in AGR. Resumption of growth was scored using single-cell time lapse imaging. Cells lacking Nvj1 exhibit approximately a 30 min delay in growth resumption after re-introduction to glucose.
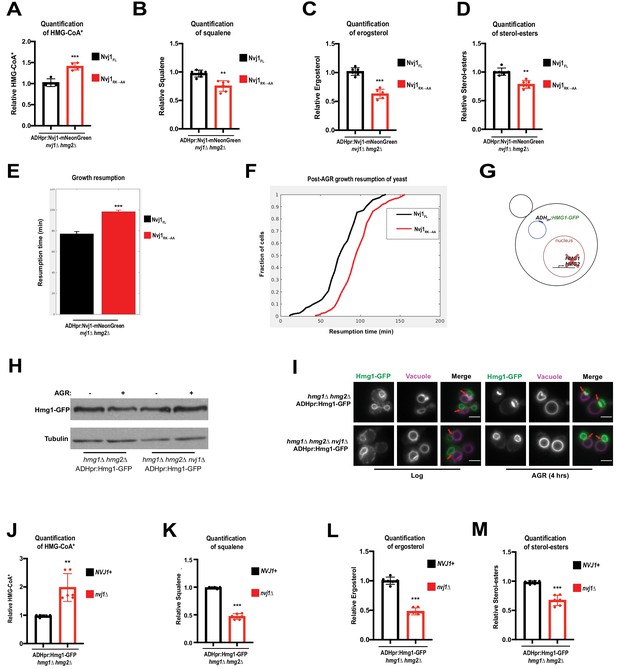
AGR-induced compartmentalization of Hmg1 increases enzymatic efficiency.
(A) Quantification of radiolabeled HMG-CoA* from nvj1Δhmg2Δ yeast expressing either Nvj1FL or Nvj1RK→AA. Growth and radiolabeling was performed as described above. (Brown-Forsyth and Welch ANOVA. N = 6. ***p-value<0.001). (B–D) Quantification of radiolabeled squalene, ergosterol, and SE produced from a 15 min radio-labeling pulse in nvj1Δhmg2Δ yeast expressing either Nvj1FL or Nvj1RK→AA. (Brown-Forsyth and Welch ANOVA. N = 6. **p-value<0.01, ***p-value<0.001). (E) Quantification of growth resumption times during SC to SC+glucose transition in nvj1Δhmg2Δ yeast expressing either Nvj1FL or Nvj1RK→AA (N > 82 cells, ***p<0.0001, Kolmogorov-Smirnov test). (F) Resumption probability graph depicting probability of cells producing a daughter bud as a function of time following an SC to SC+glucose transition (N > 82 cells). (G) Cartoon representation of ADHpr:Hmg1-GFP strains. Strains were generated by removal of endogenous Hmg1 and Hmg2, and replacing with ADH promoter-driven Hmg1-GFP (ADHpr:Hmg1-GFP). (H) Immunoblot of Hmg1-GFP isolated from ADHpr:Hmg1-GFP strains before or after treatment with AGR. Tubulin was used as a loading control. (I) Epifluorescence imaging of ADHpr:Hmg1-GFP strains showing that Hmg1 NVJ partitioning can be uncoupled from endogenous protein expression. Red arrows indicate relative position of the NVJ. Scale bars = 0.5 μm. (J) Quantification of radiolabeled HMG-CoA* from yeast expressing ADHpr:Hmg1-GFP in wildtype (NVJ1+) or nvj1Δ yeast. Growth and radiolabeling was performed as described above. (Brown-Forsyth and Welch ANOVA. N = 6. **p-value<0.01). (K–M) Quantification of squalene, ergosterol, and SE produced from a 15 min radio-labeling pulse in ADHpr:Hmg1-GFP strains. (Brown-Forsyth and Welch ANOVA. N = 6. ***p-value<0.001).
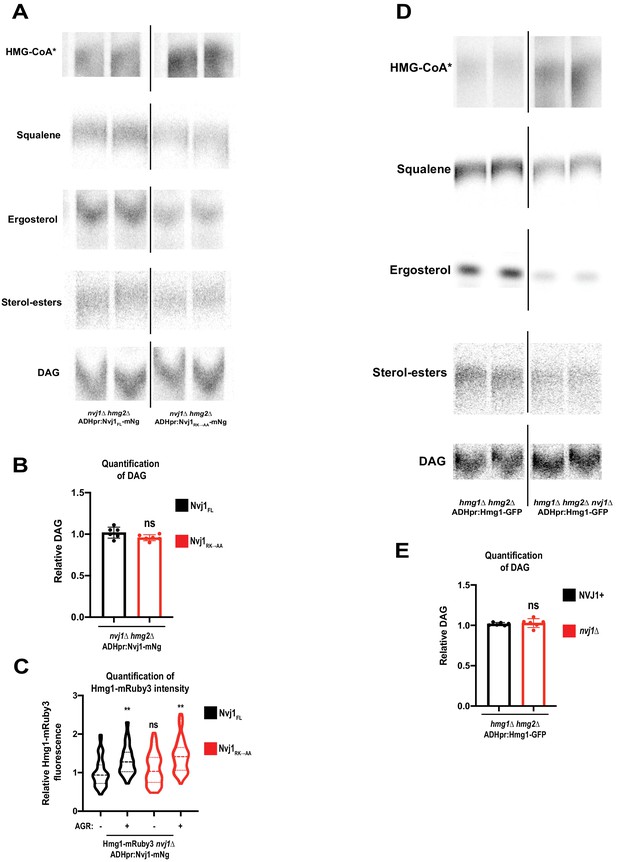
Cells expressing ADHpr:Hmg1-GFP in the absence of endogenous Nvj1 have exacerbated Hmg1 catalytic deficiency phenotype.
(A) Autoradiograms taken of TLC plates to visualize HMG-CoA*, squalene, ergosterol, SE, and DAG in nvj1Δhmg2Δ cells expressing either Nvj1FL or Nvj1RK→AA. Bands were visualized as described in Figure 5—figure supplement 1. (B) Densitometry quantification of DAG bands as shown in (A). (Brown-Forsyth and Welch ANOVA. N = 6). (C) Quantification of Hmg1-mRuby3 signal in log phase or after four hours of AGR treatment in nvj1Δhmg2Δ cells expressing either Nvj1FL or Nvj1RK→AA. (Brown-Forsyth and Welch ANOVA. N = 88. **p-value<0.01). (D) Autoradiograms taken of TLC plates to visualize HMG-CoA*, squalene, ergosterol, SE, and DAG in ADHpr:Hmg1-GFP lines in the presence or absence of Nvj1. Bands were visualized as described in Figure 5—figure supplement 1. (E) Densitometry quantification of DAG bands as shown in (D). (Brown-Forsyth and Welch ANOVA. N = 6).
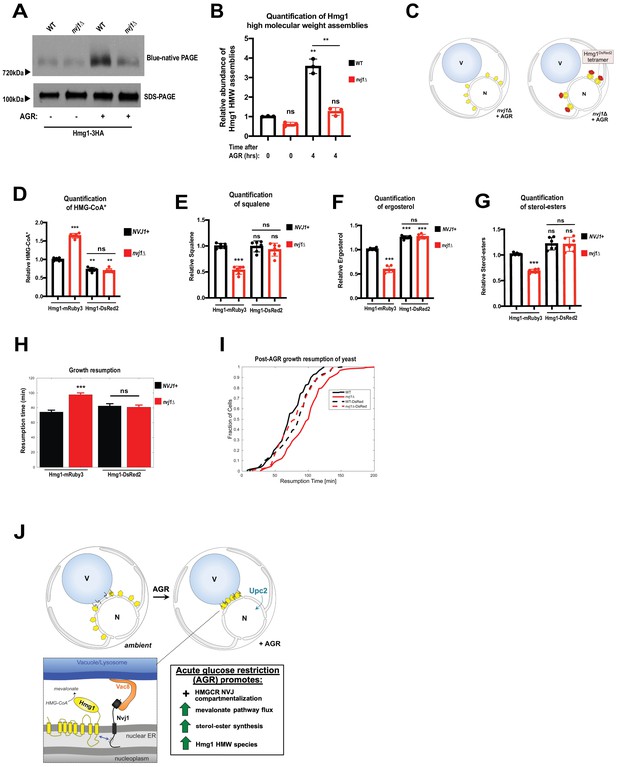
Artificial multimerization of Hmg1 rescues enzymatic deficiencies of nvj1Δ cells.
(A) Blue native PAGE (BN-PAGE) and SDS-PAGE analysis of Hmg1-3HA expressed in either WT or nvj1Δ cells under log-phase or AGR growth conditions. Samples were loaded to mitigate differences in total Hmg1 loading such that differences in the BN-PAGE could be easily interpreted. Arrows on the side indicate relative molecular mass in corresponding region as determined by molecular weight standards. (B) Quantification of BN-PAGE and SDS-PAGE analysis as shown in (A). For quantification of relative Hmg1-3HA abundance in HMW species, fold change relative to WT log samples were first calculated for BN-PAGE species and subsequently normalized by total expression of Hmg1 in each sample/condition as determined by SDS-PAGE. (Brown-Forsyth and Welch ANOVA. N = 3. **p-value<0.01). (C) Cartoon representation of Hmg1 artificial tetramerization by DsRed2 tagging in nvj1Δ cells. (D) Scintillation counting quantification of radiolabeled HMG-CoA* from yeast expressing endogenously tagged monomeric Hmg1-mRuby3 or tetrameric Hmg1-DsRed2 in wildtype (NVJ1+) or nvj1Δ yeast. Quantification performed as in (Figure 6A). (Brown-Forsyth and Welch ANOVA. N = 6. **p-value<0.01 ***p-value<0.001). (E–G) Quantification of radiolabeled squalene, ergosterol, and SE produced in Hmg1-mRuby3 or Hmg1-DsRed2 yeast. Radio labeling and quantifications performed as in (Figure 6B–D). (Brown-Forsyth and Welch ANOVA. N = 6. ***p-value<0.001). (H) Average growth resumption times measured by single-cell time lapse microscopy. (WT: N = 110, nvj1Δ: N = 175, WT Hmg1-DsRed2: N = 94, nvj1Δ Hmg1-DsRed2: N = 96). (N > 94 cells, **p<0.001, Kolmogorov-Smirnov test). (I) Resumption probability graph depicting probability of cells producing a daughter bud as a function of time following an SC to SC+glucose transition. (N > 94 cells). (J) Cartoon representation of Nvj1 and Upc2-mediated Hmg1 partitioning at the NVJ. Spatial partitioning of Hmg1 at the NVJ coincides with increased production of mevalonate-derived metabolites.
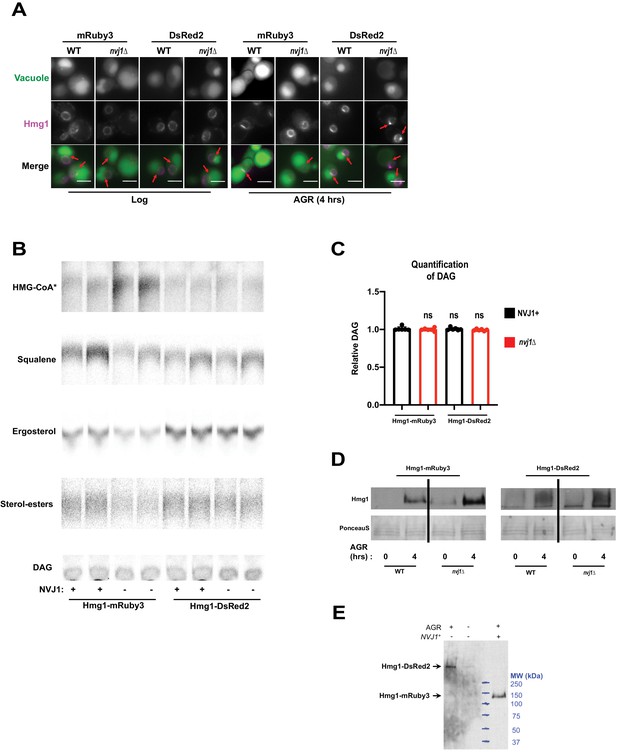
Artificial inter-enzyme association via DsRed2 tagging rescues nvj1Δ associated Hmg1 catalytic deficiency.
(A) Epifluorescence imaging of yeast expressing endogenously tagged Hmg1-mRuby3 or Hmg1-DsRed2 (magenta) in log-phase and after 4 hr of AGR treatment. Vacuoles (green) were stained by incubating yeast with 5 μg/mL CMAC dye for 2 hr prior to imaging. Red arrows in merged images indicate the NVJ. Red arrows in the Hmg1-DsRed2 nvj1Δ AGR-treated greyscale image indicates prominent Hmg1-positive foci on the NE. Scale bars = 0.5 μm. (B) Autoradiograms taken of TLC plates to visualize HMG-CoA* (as previously described) or squalene, ergosterol, SE, and DAG in cells expressing endogenously tagged monomeric Hmg1-mRuby3 or tetrameric Hmg1-DsRed2 in the presence or absence of endogenous Nvj1. Bands were visualized as described in Figure 5—figure supplement 1. (C) Densitometry quantification of DAG bands as shown in (A). (Brown-Forsyth and Welch ANOVA. N = 6). (D) Immunoblot analysis of Hmg1-mRuby3 and Hmg1-DsRed2 strains expressed in either WT or nvj1Δ yeast. PonceauS total protein stain was used as a loading control. (E) Immunoblot analysis of Hmg1-DsRed2 and Hmg1-mRuby3 strains indicating that Hmg1-DsRed2 runs higher in the SDS-PAGE gel than expected. Blue lines indicate position of molecular weight standards. One lane of Hmg1-mRuby3 is included for comparison.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Saccharomyces cervisiae) | W303 | This paper | W303 | See Supplementary file 1 for all yeast strains |
| Transfected construct (Saccharomyces cervisiae) | pRS305 vector backbone (Leu selection) | This paper | pRS305 | See Supplementary file 2 for all yeast expression plasmids |
| Antibody | Anti-GFP (rabbit polyclonal) | Abcam | Cat# abcam290 | WB (1:10,000) |
| Antibody | anti-mRuby3 (rabbit polyclonal) | Invitrogen | Cat# R10367 | WB (1:1,000) |
| Antibody | anti-DsRed2 (mouse monoclonal) | Origene | Cat#: TA180005 | WB: (1:1,000) |
| Antibody | Anti-tubulin (rat monoclonal) | Abcam | Cat#: ab6160 | WB: (1:15,000) |
| Antibody | Sec61 (rabbit poly) | J. Friedman lab (UTSW) | N/A | WB: (1:15,000) |
| Software, algorithm | Segmentation and tracking code for single-cell time-lapse imaging of yeast budding | from N. E. Wood, A. Doncic, A fully-automated, robust, and versatile algorithm for long-term budding yeast segmentation and tracking. PLoS ONE. 14, e0206395 (2019). Folch et al., 1957 | --- |
Additional files
-
Supplementary file 1
Table S1: Yeast strains used in this study.
- https://cdn.elifesciences.org/articles/62591/elife-62591-supp1-v2.docx
-
Supplementary file 2
Table S2: Plasmids used in this study.
- https://cdn.elifesciences.org/articles/62591/elife-62591-supp2-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/62591/elife-62591-transrepform-v2.docx