The alternative sigma factor σX mediates competence shut-off at the cell pole in Streptococcus pneumoniae
Figures
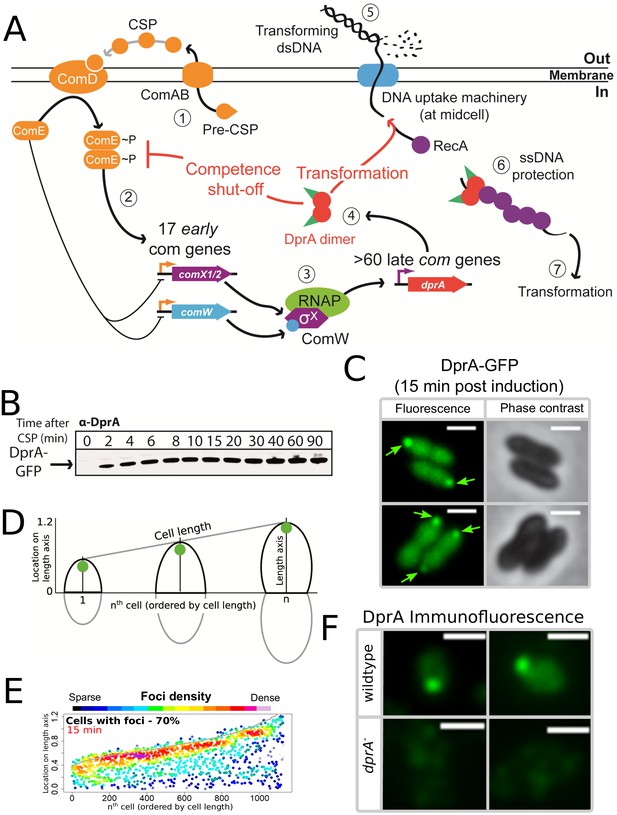
DprA: Localization and roles in competence and transformation.
(A) (1) Pre-CSP is exported and matured by the ComAB transporter, and then promotes phosphorylation of the histidine kinase ComD. (2) ComD transphosphorylates ComE, which then stimulates the expression of 17 early com genes, including two copies of comX. (3) These encode an alternative sigma factor σX, which controls late com genes including dprA. (4) DprA dimers load RecA onto ssDNA to mediate transformation and interact with ComE~P to shut-off competence. (5) Transforming DNA is internalized in single strand form and is protected from degradation by DprA and RecA (6), which then mediate transformation (7). Orange arrows, early com promoters; purple arrows, late com promoters. (B) Western blot tracking cellular levels of DprA-GFP after competence induction in strain R3728. α-DprA antibody used. (C) Sample fluorescence microscopy images of R3728 strain producing DprA-GFP 15 min after competence induction. Scale bars, 1 µm. (D) Schematic representation of focus density maps with half cells represented as vertical lines in ascending size order and localization of foci represented along the length axis of each half cell. Black half-cells represent those presented, and grey those not presented. (E) DprA-GFP accumulates at the cell poles during competence. 1290 cells and 1128 foci analyzed. (F) Sample immunofluorescence microscopy images of a strain producing wild-type DprA (R1502; wildtype) and a strain lacking DprA (R2018; dprA-) fixed 15 min after competence induction and probed using anti-DprA antibodies. Scale bars, 1 µm.
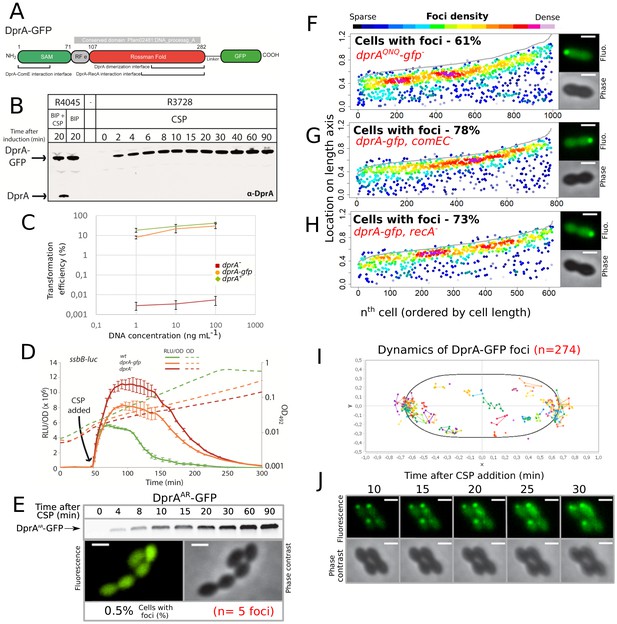
Validation of DprA-GFP activity.
(A) Linear representation of DprA-GFP with the limits of Pfam02481 (in gray) and structural domains indicated. Interfaces of dimerization and interaction with RecA and ComE are indicated and represent the regions within which the majority of point mutations affecting interaction were identified (Mirouze et al., 2013; Quevillon-Cheruel et al., 2012). The GFP protein is separated from DprA by a linker, as described previously (Bergé et al., 2013). (B) Western blot tracking cellular levels of DprA-GFP after induction as shown from strains R4045 and R3728. α-DprA antibody used. Samples at each time point corrected by OD to render direct comparison of cellular levels possible. Full gel shows no degradation of DprA-GFP throughout growth. (C) DprA-GFP is active in transformation. Comparing transformation efficiency of wildtype (R1501), drpA- (R2018) and dprA-gfp (R3728) strains. R304 chromosomal DNA, conferring streptomycin resistance via rpsL41 point mutation (Salles et al., 1992), used to transform at 1, 10, and 100 ng mL−1. Error bars represent triplicate repeats. (D) DprA-GFP is partially active in competence shut-off. Comparing the competence profiles of wildtype (R1502), drpA- (R2018) and dprA-gfp (R3743) strains after CSP addition (100 ng mL−1, t = 40 min). Luminometric and photometric readings of ssbB-luc transcriptional reporter fusion taken every 5 min. Full lines represent competence induction (RLU/OD) and dotted lines represent growth (OD). Error bars represent triplicate repeats. (E) Dimerization of DprA is necessary for accumulation of DprA-GFP at the cell poles. Western blot tracking cellular levels of DprA-GFP after competence induction in strain R4046. α-DprA antibody used. Samples at each time point corrected by OD to render direct comparison of cellular levels possible. In contrast to DprA-GFP (Figure 1B), the levels of DprAAR-GFP continue to increase over the time period of the experiment as competence is not shut-off in this DprAAR strains (Quevillon-Cheruel et al., 2012). Sample fluorescence microscopy images of R4046 strain producing DprAAR-GFP 15 min after competence induction. Scale bars, 1 µm. 1124 cells and five foci analyzed. (F) Disrupting the ability of DprA to interact with RecA (dprAQNQ) does not impact the polar accumulation of DprA. Strain R4047 (comC0, dprAQNQ-gfp) observed 15 min after competence induction. Data presented as in Figure 1E. 767 cells and 612 foci analyzed. Sample microscopy images of strain R4047 15 min after competence induction. Scale bars, 1 µm. (G) Inactivating the comEC transformation pore gene to prevent uptake of exogenous DNA does not impact the polar accumulation of DprA. Strain R4082 (dprA-gfp, comEC-) observed 15 min after competence induction. Data presented as in Figures 1E and 6 32 cells and 778 foci analyzed. Sample microscopy images of strain R4082 15 min after competence induction. Scale bars, 1 µm. (H) Inactivation the recombinase gene recA to prevent homologous recombination does not impact the polar accumulation of DprA. Strain R4061 (dprA-gfp, recA-) observed 15 min after competence induction. Data presented as in Figures 1E and 3 93 cells and 996 foci analyzed. Sample microscopy images of strain R4061 15 min after competence induction. Scale bars, 1 µm. (I) DprA-GFP foci remain stable over time after competence induction. DprA-GFP foci tracked by time-lapse fluorescence microscopy every 5 min from 10 to 30 min after competence induction in strain R3728. Images analyzed by MicrobeJ (Ducret et al., 2016). Each series of linked spots represents a single focus tracked over time, localized in an averaged pneumococcal cell. (J) Sample time-lapse fluorescence microscopy images of strain R3728 used to generate panel H.
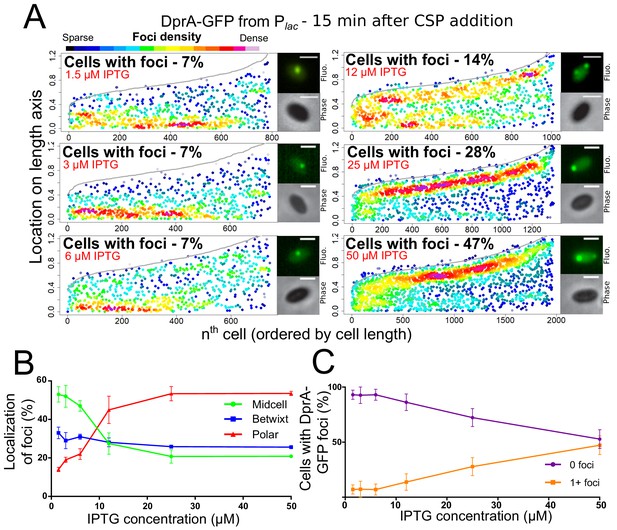
Reducing cellular DprA-GFP levels results in loss of polar accumulation and competence shut-off.
(A) Focus density maps of DprA-GFP foci at different cellular levels during competence. CEPlac-dprA-gfp from strain R4262. Cellular DprA-GFP levels were controlled by growing cells in a gradient of IPTG. 1.5 µM IPTG, 11267 cells and 791 foci analyzed; 3 µM IPTG, 10623 cells and 748 foci analyzed; 6 µM IPTG, 10603 cells and 743 foci analyzed; 12.5 µM IPTG, 6985 cells and 1010 foci analyzed; 25 µM IPTG, 2945 cells and 1345 foci analyzed; 50 µM IPTG, 3678 cells and 1964 foci analyzed. Sample microscopy images of strain R4262 in varying IPTG concentrations. Scale bars, 1 µm. (B) Reducing cellular levels of DprA-GFP reduces the number of cells with foci. Error bars represent triplicate repeats. (C) Reducing cellular levels of DprA-GFP results specifically in loss of polar foci. Error bars represent triplicate repeats.
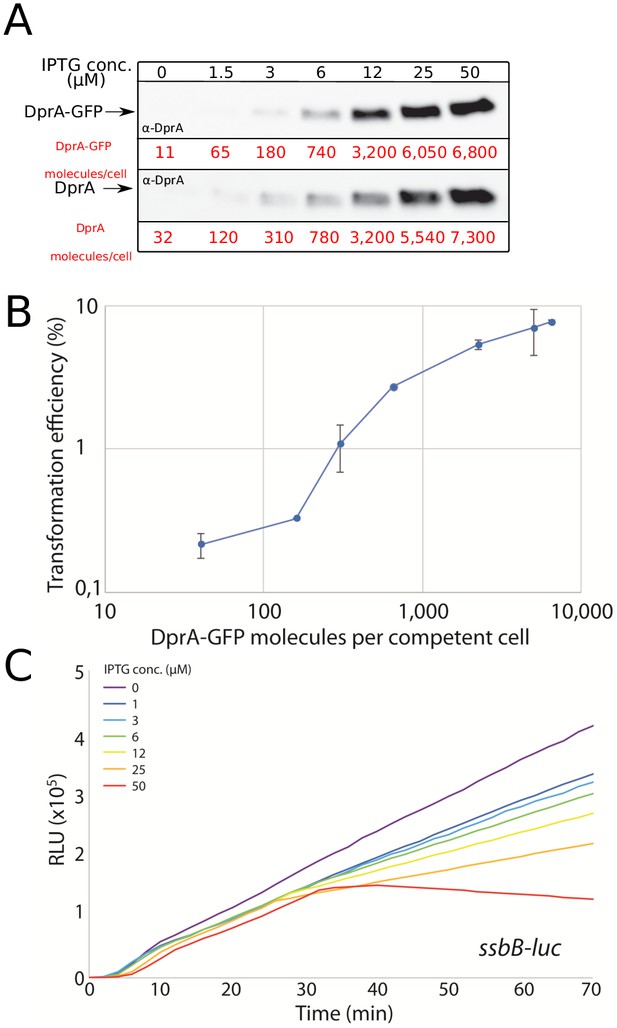
Validation of CEPlac-dprA-gfp activity.
(A) Western blots comparing cellular levels of DprA (R3833) and DprA-GFP (R4262) produced from the CEPlac platform in cells lacking native dprA in varying concentrations of IPTG. α-DprA antibodies used. Samples at each IPTG concentration corrected by OD to render direct comparison of cellular levels possible. Cellular levels of DprA-GFP estimated from Western blots compared to purified DprA as previously described (Johnston et al., 2018). (B) Transformation efficiency of a CEPlac-dprA-gfp, dprA- strain (R4262) in an IPTG concentration gradient. R304 chromosomal DNA (50 ng mL−1), conferring streptomycin resistance via rpsL41 point mutation (Salles et al., 1992), used to transform. Error bars represent triplicate repeats. (C) Competence profiles of a R4262 strain possessing CEPlac-dprA-gfp and ssbB-luc in an IPTG concentration gradient. Appropriate IPTG concentration present from the beginning of the culture. Competence induced by addition of CSP at t = 0, when OD = ~0.05. Plots representative of triplicate repeats.
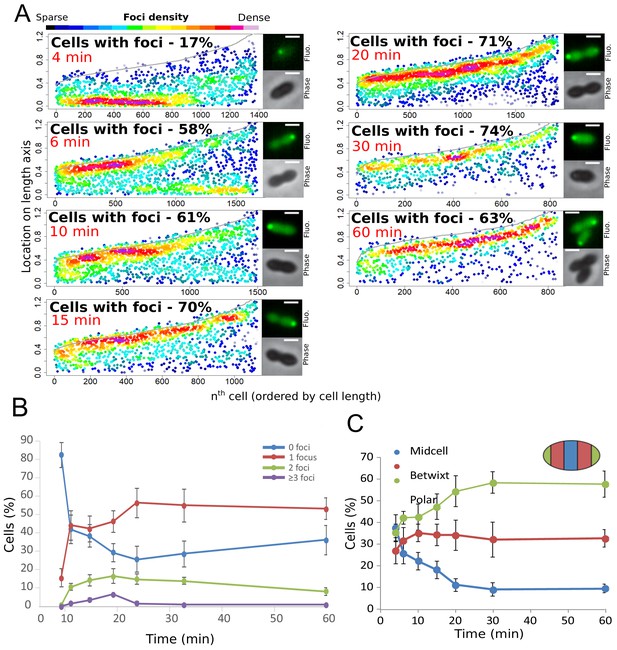
Analysis of the cellular localization of DprA-GFP.
(A) DprA-GFP foci persist at the cell pole after competence shut-off. Data at different time points after CSP addition represented as in Figure 1E. 4 min, 8336 cells and 1383 foci analyzed; 6 min, 2871 cells and 1674 foci analyzed; 10 min, 2614 cells and 1502 foci analyzed; 15 min, 1290 cells and 1128 foci analyzed; 20 min, 2110 cells and 1900 foci analyzed; 30 min, 789 cells and 831 foci analyzed; 60 min, 842 cells and 839 foci analyzed. Sample microscopy images of strain R3728 at varying times after competence induction. Scale bars, 1 µm. (B) Most competent cells possess a single polar focus of DprA-GFP. Error bars represent triplicate repeats. (C) Most cells possess polar DprA-GFP foci. Along the length of a cell of arbitrary length 1, polar foci are found between positions 0–0.15 and 0.85–1, midcell foci are found between 0.35 and 0.65, and anything in between is localized as betwixt. Error bars represent triplicate repeats.
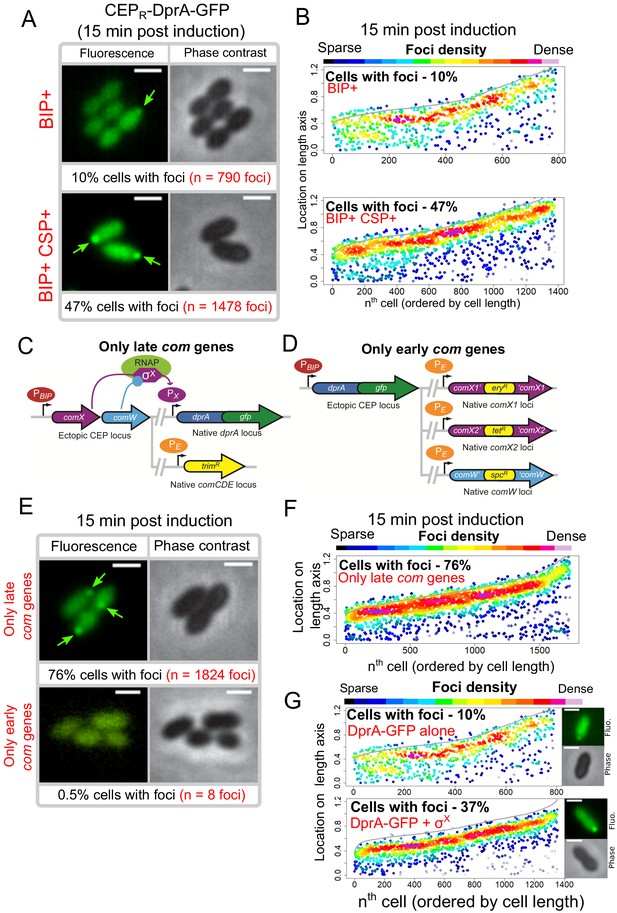
Polar accumulation of DprA-GFP appears to depend on late com regulon expression.
(A) Sample fluorescence microscopy images of strain R4060 producing DprA-GFP 15 min after induction with BIP or BIP and CSP. Scale bars, 1 µm. (B) Competence induction is required for optimal accumulation of DprA-GFP at the cell poles. Focus density maps as in Figure 1E. BIP+, 7845 cells and 790 foci analyzed; BIP+ CSP+, 2707 cells and 1478 foci analyzed. (C) Genetic context strain R4107 expressing dprA-gfp and only the late com regulon. PBIP and PX as in panel A, PE represents early com promoter controlled by ComE. Light blue circle, ComW; light green oval, RNA polymerase; purple hexagon, σX. (D) Genetic context of strain R4140 expressing CEPR-dprA-gfp and only the early com regulon. PBIP as in panel A, PE as in panel D. (E) Sample fluorescence microscopy images of strains producing DprA-GFP with only late (R4107) or only early (R4140) com operons 15 min after competence induction. Scale bars, 2 µm. (F) Induction of the late com regulon is required for accumulation of DprA-GFP at the cell poles. Focus density maps as in Figure 1E. 1988 cells and 1824 foci analyzed. (G) Focus density maps produced as in Figure 1E from images where DprA-GFP was produced outside of competence in presence or absence of σX. DprA-GFP alone, 7845 cells and 790 foci analyzed; DprA-GPF + σX, 3545 cells and 1355 foci analyzed. Strains used: DprA-GFP alone, R4060; DprA-GFP and σX, R4489.
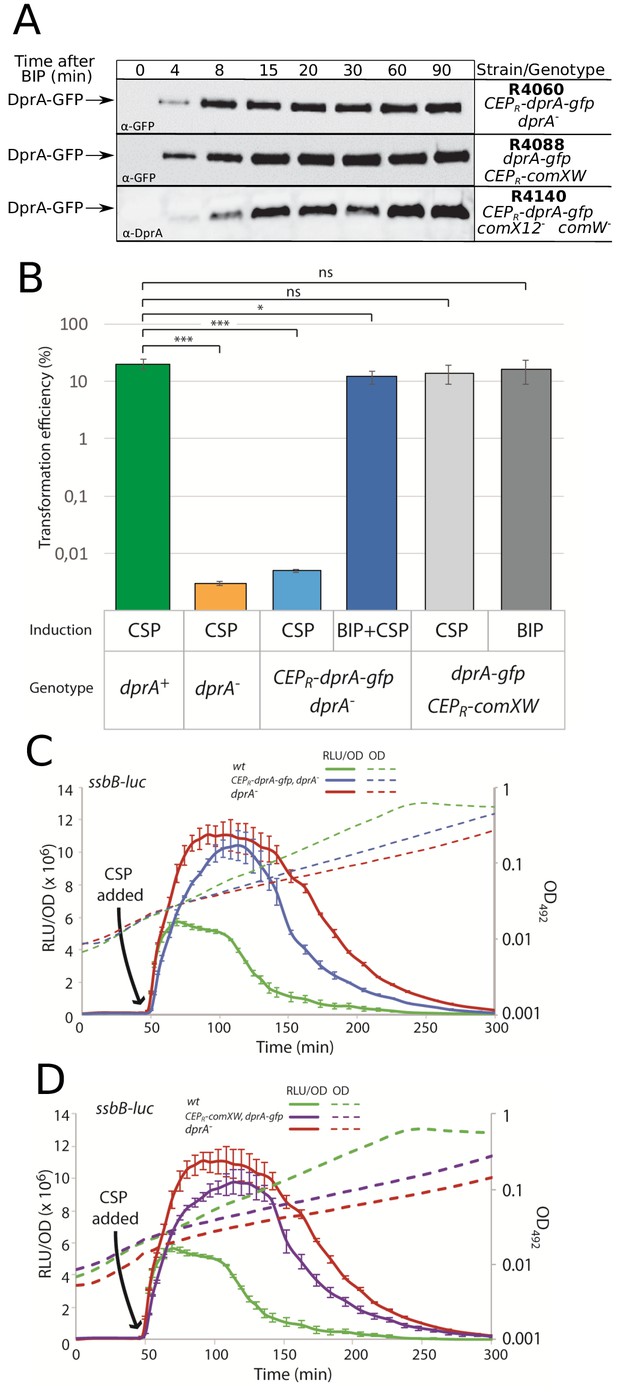
Validation of strains expressing CEPR-dprA-gfp and the late com regulon alone.
(A) Western blots tracking cellular levels of DprA-GFP in R4060 (CEPR-dprA-gfp dprA-), R4088 (dprA-gfp CEPR-comXW) and R4140 (CEPR-dprA-gfp comX12- comW-) strains after induction with BIP. Either α-GFP or α-DprA antibodies used. Samples at each timepoint corrected by OD to render direct comparison of cellular levels possible. (B) Comparing transformation efficiencies of isogenic strains impaired for competence auto-induction. R1501 (‘wildtype’), R2018 (dprA-), R4060 (CEPR-dprA-gfp dprA-), and R4088 (dprA-gfp CEPR-comXW) strains after induction with either CSP, BIP or both, as shown. rpsL41 PCR fragment, conferring streptomycin via point mutation (Salles et al., 1992), used to transform at 50 ng mL−1. Error bars represent triplicate repeats. Statistical differences between wildtype and mutants determined by Student’s T-test using GraphPad Prism. p values as followed compared to wildtype. dprA-, p=0.0002; CEPR-dprA-gfp dprA- CSP+, p=0.0002; CEPR-dprA-gfp dprA- CSP+ BIP+, p=0.0109; dprA-gfp CEPR-comXW CSP+, p=0.286; dprA-gfp CEPR-comXW BIP+, p=0.218. (C) Exploring the competence profile of a strain ectopically expressing DprA-GFP under the control of BIP. R1501 (wildtype), R2018 (dprA-) and R4060 (CEPR-dprA-gfp dprA-) strains. Experiment carried out as as in Figure 1—figure supplement 1D. Error bars represent triplicate repeats. (D) Exploring the competence profile of a strain ectopically expressing σX and ComW under the control of BIP and DprA-GFP from its native promoter. R1501 (wildtype), R2018 (dprA-), and R4088 (CEPR-comXW, dprA-gfp, cbpD::cat) strains. Experiment carried out as in Figure 1—figure supplement 1D. Error bars represent triplicate repeats.
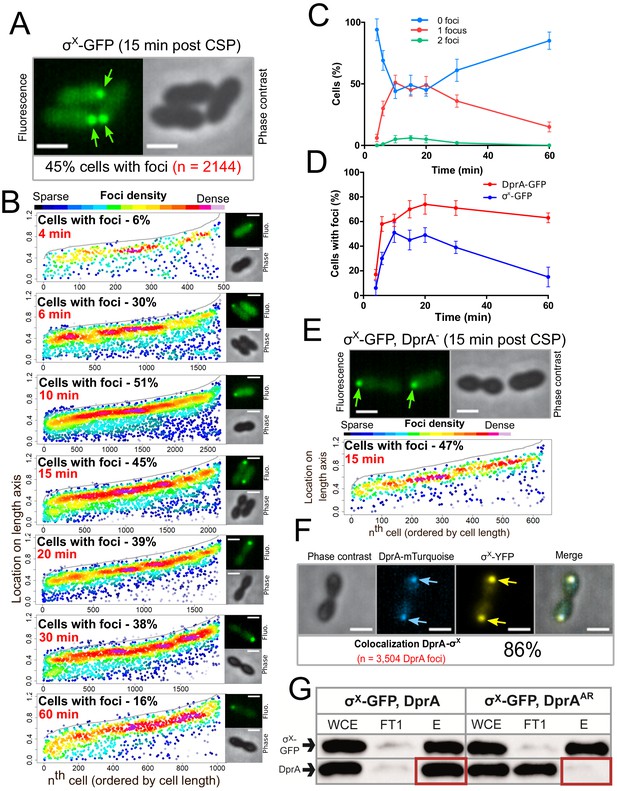
σX-GFP interacts directly with DprA at the cell pole during competence.
(A) Sample fluorescence microscopy images of strain R4451 producing σX-GFP from comX1 and wild-type σX from comX2 15 min after competence induction. Scale bars, 1 µm. (B) σX-GFP accumulates at the cell pole during competence. Focus density maps presented as in Figure 1E. 4 min, 7544 cells and 489 foci analyzed; 6 min, 5442 cells and 1711 foci analyzed; 10 min, 4358 cells and 2691 foci analyzed; 15 min, 3746 cells and 2144 foci analyzed; 20 min, 4211 cells and 1754 foci analyzed; 30 min, 4695 cells and 1920 foci analyzed; 60 min, 5713 cells and 1016 foci analyzed. (C) Most cells have a single σX-GFP focus. Data from the time-course experiment presented in panel B showing the number of foci per cell at each time point. Error bars represent triplicate repeats. (D) DprA-GFP foci persist in cells longer than σX-GFP foci. Comparison of cells with foci at different timepoints from timecourse experiments. DprA-GFP from Figure 3A, σX-GFP from panel B. Error bars represent triplicate repeats. (E) Accumulation of σX at the cell poles does not depend on DprA. Sample microscopy images of a comX1-gfp, dprA- strain (R4469). Focus density maps generated from cells visualized 15 min after competence induction presented as in Figure 1E. 1104 cells and 638 foci analyzed. (F) σX and DprA colocalize at the cell pole. Colocalization of σX-YFP and DprA-mTurquoise in R4473 cells visualized 15 min after competence induction. 7460 cells and 3504 DprA-mTurquoise foci analyzed. Scale bars, 1 µm. (G) DprA is copurified with σX-GFP while DprAAR is not. Western blot of pull-down experiment carried out on strains producing σX-GFP and either DprA (R4451) or DprAAR (R4514) 10 min after competence induction. WCE, whole cell extract; FT1, flow through; E, eluate.
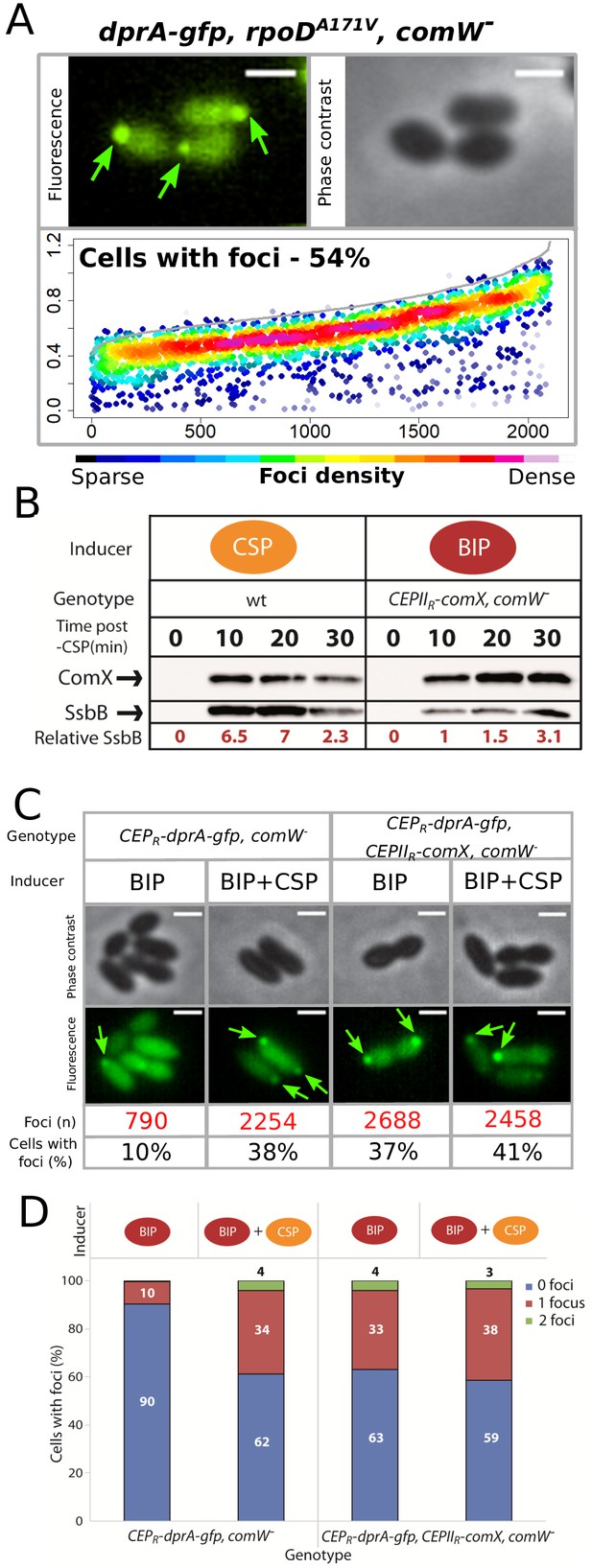
σX is necessary and sufficient to mediate accumulation of DprA at the cell poles.
(A) DprA-GFP accumulates at the cell poles in the absence of ComW. Localization of DprA-GFP in R4168 strain (dprA-gfp, rpoDA171V, comW-) 15 min after competence induction. Sample fluorescence microscopy images shown. Scale bars, 1 µm. Focus density maps presented as in Figure 3B. 3592 cells and 2101 foci analyzed. (B) Production of σX outside of competence promotes accumulation of DprA-GFP at the cell poles. Sample fluorescence microscopy images of cells producing DprA-GFP (R4060) or DprA-GFP and σX (R4489) outside of competence. Focus density maps presented as in Figure 3B. DprA-GPF + σX, 3545 cells and 1355 foci analyzed. CEPR-dprA-gfp induced by BIP, 7845 cells and 790 foci analyzed; CEPR-dprA-gfp induced by BIP and CSP, 5889 cells and 2254 foci analyzed; CEPR-dprA-gfp, CEPIIR-comX induced by BIP, 3545 cells and 1355 foci analyzed; CEPR-dprA-gfp, CEPIIR-comX induced by BIP and CSP, 6085 cells and 2458 foci analyzed. Scale bars, 1 µm. (C) Data from analysis of images in panel A split into cells with 0, 1 or two foci. (D) Western blot comparing expression of late com regulon by tracking cellular levels of SsbB in R1501 (wildtype) and R4509 (CEPIIR-comX, comW-) strains. R1501 induced by CSP at t = 0 and R4509 induced by BIP. Samples at each time point corrected by OD to render direct comparison of cellular levels possible. Relative SsbB levels measured by quantifying SsbB bands and normalizing to the band of SsbB expression 10 min after BIP addition to R4509 as 1. α-σX and α-SsbB antibodies used as shown.
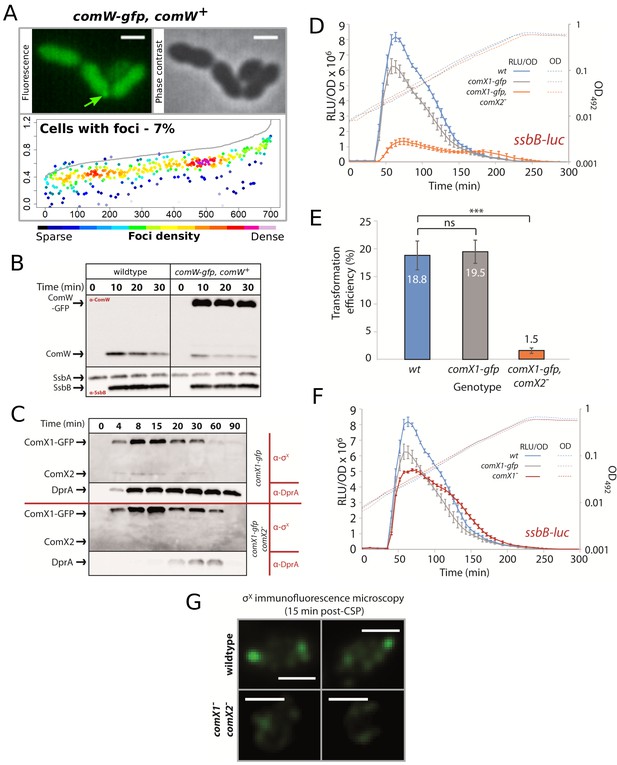
Validation of comW and comX fluorescent fusions.
(A) ComW-GFP is diffuse within the cell cytoplasm. Localization of ComW-GFP in R4513 strain (comW-gfp, comW+) 15 min after competence induction. Sample fluorescence microscopy images shown. Scale bars, 2 µM. Focus density maps presented as in Figure 1E. 9705 cells and 701 foci analyzed. (B) Western blots showing expression of ComW, ComW-GFP, SsbA and SsbB in response to CSP addition to either R1501 (wildtype) or R4513 (comW-gfp, comW+) cells. α-ComW or α-SsbB antibodies used as shown. α-SsbB antibodies recognize both SsbA and SsbB. Samples at each time point corrected by OD to render direct comparison of cellular levels possible. (C) Western blots tracking cellular levels of σX, σX-GFP and DprA after competence induction in R4451 (comX1-gfp) and R4461 (comX1-gfp, comX2-) strains. α-σX and α-DprA antibodies used as shown. Samples at each time point corrected by OD to render direct comparison of cellular levels possible. (D) σX-GFP is partially active in competence induction. Comparing the competence profiles of R1502 (ssbB-luc), R4471 (comX1-gfp, ssbB-luc), and R4465 (comX1-gfp, comX2-, ssbB-luc) strains. Data presented as described in Figure 1—figure supplement 1D. Error bars represent triplicate repeats. (E) Comparing transformation efficiencies of R1501 (wildtype), R4451 (comX1-gfp), and R4461 (comX1-gfp, comX2-) strains. rpsL41 PCR fragment, conferring streptomycin via point mutation (Salles et al., 1992), used to transform at 50 ng mL−1. Error bars represent triplicate repeats. Statistical differences between wildtype and mutants determined by Student’s T-test using GraphPad Prism. p Values as followed compared to wildtype. comX1-gfp, 0.81; comX1-gfp comX2-, 0.0009. (F) σX-GFP is active in the shut-off of competence. Comparison of competence profiles of R1502 (ssbB-luc), R4471 (comX1-gfp, ssbB-luc), and R4466 (comX1- ssbB-luc) strains as described in Figure 1—figure supplement 1D. Comparison of data from panel E and Figure 7B. Error bars represent triplicate repeats. (G) Sample immunofluorescence microscopy images of a strain producing wildtype σX (R1501; wildtype) and a strain lacking σX (R4072; comX1-, comX2-) fixed 15 min after competence induction and probed using anti- σX antibodies. Scale bars, 1 µm.
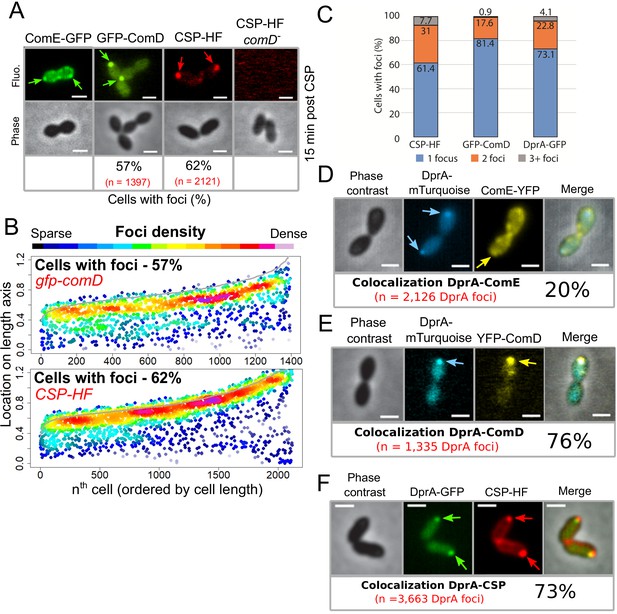
The main actors of competence induction and shut-off colocalize at the cell pole.
(A) Sample fluorescence microscopy images of cells producing ComE-GFP (R4010), GFP-ComD (R3914) or wildtype cells (R1501) and comD- cells (R1745) exposed to CSP-HF. Scale bars, 1 µm. GFP-ComD, 1933 cells and 1397 foci; CSP-HF, 2105 cells and 2121 foci. (B) Focus density maps of cells producing GFP-ComD (R3914) and wild-type (R1501) cells exposed to CSP-HF 15 min after competence induction. Data presented as in Figure 1E. GFP-ComD, 1933 cells and 1397 foci analyzed; CSP-HF, 2105 cells and 2121 foci analyzed. (C) Number of foci present in cells possessing foci of different fluorescent fusions 15 min after competence induction. Data taken from panel B except for DprA-GFP, taken from Figure 3. (D) DprA-mTurquoise and ComE-YFP colocalization in competent R4176 cells visualized by fluorescence microscopy 15 min after competence induction. 3143 cells and 2126 DprA-mTurquoise foci analyzed. Scale bars, 1 µm. (E) DprA-mTurquoise and YFP-ComD colocalization in competent R4111 cells visualized by fluorescence microscopy 15 min after competence induction. 2857 cells and 1335 DprA-mTurquoise foci analyzed. Scale bars, 1 µm. (F) DprA-GFP and CSP-HF colocalization in competent R4062 cells visualized by fluorescence microscopy 15 min after competence induction. 7588 cells and 3663 DprA-mTurquoise foci analyzed. Scale bars, 1 µm.
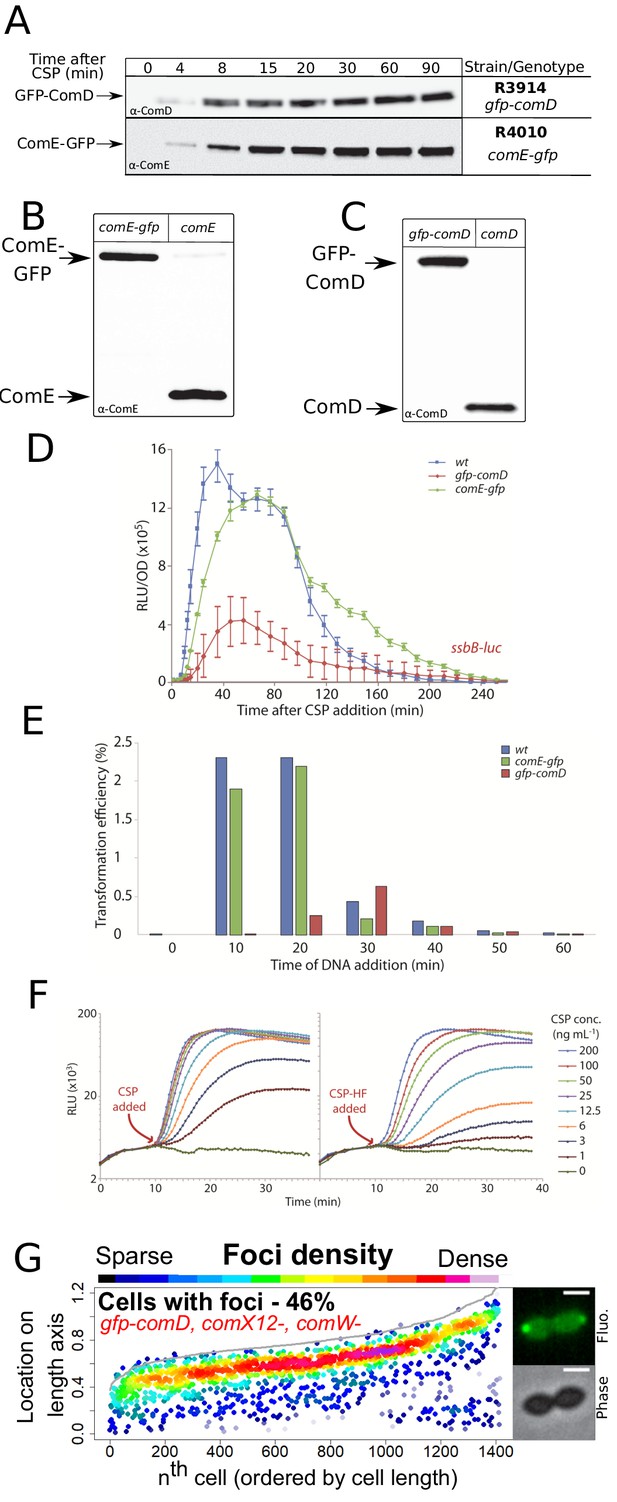
Validation of ComE-GFP, GFP-ComD, and CSP-HF.
(A) ComE-GFP and GFP-ComD are produced stably after induction of competence. Western blots tracking cellular levels of ComE-GFP (R4010) and GFP-ComD (R3914) after induction of competence. α-ComD and α-ComE antibodies used. Samples corrected by OD to render direct comparison of cellular levels possible. (B) Western blot comparing ComE-GFP (R4010) and ComE (R1501) 15 min after competence induction, showing that ComE-GFP is not degraded. (C) Western blot comparing GFP-ComD (R3914) and ComD (R1501) 15 min after competence induction, showing that GFP-ComD is not degraded. (D) Comparison of competence induction profiles of R1502 (ssbB-luc), R4087 (comE-gfp, ssbB-luc) and R3915 (gfp-comD, ssbB-luc) strains as described in Figure 1—figure supplement 1D. Error bars represent triplicate repeats. (E) Comparison of transformation efficiencies at varying time points after competence induction in R1501 (wild-type), R4010 (comE-gfp) and R3914 (gfp-comD) strains. R304 chromosomal DNA (50 ng mL−1), conferring streptomycin resistance via rpsL41 point mutation (Salles et al., 1992), used to transform at indicated time points. (F) Comparison of competence induction profiles of concentration gradients of CSP and CSP-HF. Strain used, R1501 (ssbB-luc) Luminometric readings taken every 30 s during the experiment. (G) ComD localizes to the cell pole in the absence of the late com regulon. Focus density maps of strain R4916 (gfp-comD, comX1-, comX2-, comW-) 15 min after competence induction presented as in Figure 1E. Sample microscopy images of strain R4196 15 min after competence induction. Scale bars, 1 µm.
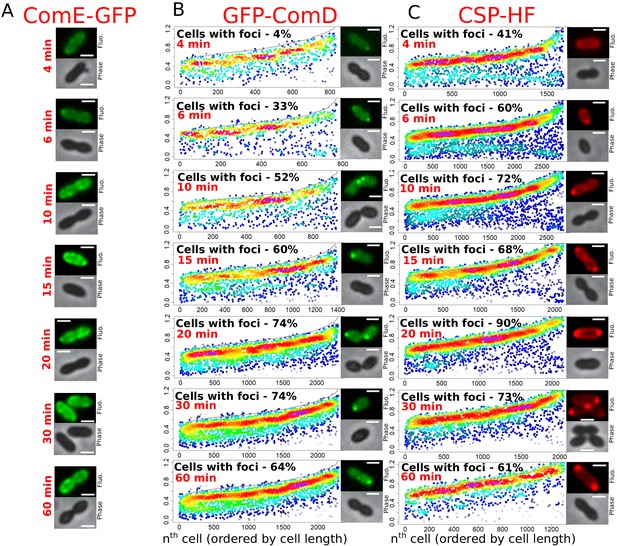
Time-course experiments tracking the localization of ComE-GFP, GFP-ComD, and CSP-HF after competence induction.
(A) ComE-GFP forms patches around the cell membrane in competent cells. Sample microscopy images of strain R4010 at varying time points after competence induction. Scale bars, 1 µm. (B) GFP-ComD accumulates at the cell poles in competent cells. Focus density maps of GFP-ComD (R3914) in time-course experiment after competence induction. Data presented as in Figure 1E. 4 min, 18772 cells and 832 foci analyzed; 6 min, 2115 cells and 770 foci analyzed; 10 min, 1581 cells and 944 foci analyzed; 15 min, 1933 cells and 1397 foci analyzed; 20 min, 2549 cells and 2311 foci analyzed; 30 min, 2526 cells and 2299 foci analyzed; 60 min, 3358 cells and 2428 foci analyzed. Sample microscopy images of strain R3914 at varying time points after competence induction. Scale bars, 1 µm. (C) CSP-HF accumulates at the cell poles in competent cells. Focus density maps of CSP-HF in time-course experiment after competence induction of R1501 strain. Data presented as in Figure 1E. 4 min, 2928 cells and 1671 foci analyzed; 6 min, 2581 cells and 2283 foci analyzed; 10 min, 2391 cells and 2795 foci analyzed; 15 min, 2105 cells and 2121 foci analyzed; 20 min, 1279 cells and 2515 foci analyzed; 30 min, 1997 cells and 2052 foci analyzed; 60 min, 993 cells and 735 foci analyzed. Sample microscopy images of strain R1501 at varying time points after competence induction. Scale bars, 1 µm.
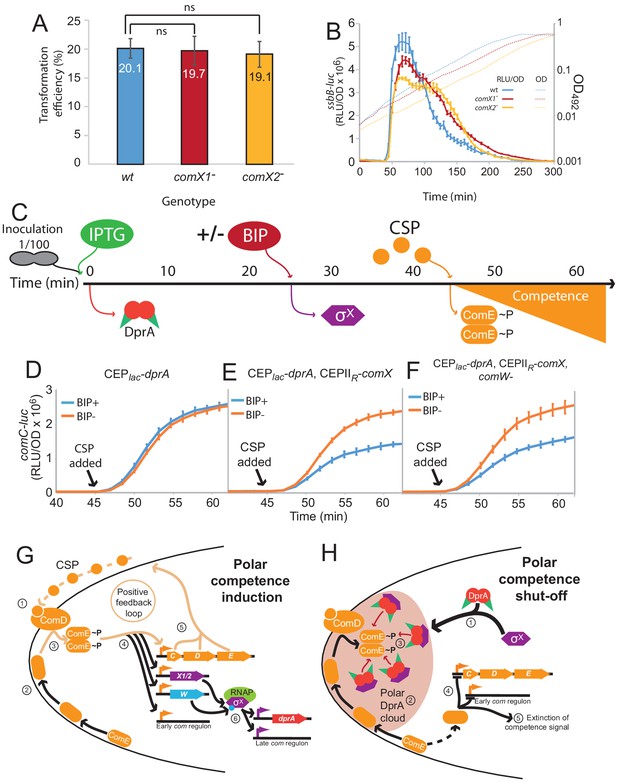
Pre-competence production of DprA and σX antagonizes competence induction.
(A) Inactivation of comX1 or comX2 does not impact transformation efficiency. Error bars represent triplicate repeats. (B) Inactivation of comX1 or comX2 delays the shut-off of competence. Data is plotted as Relative Light Units, corrected by optical density (RLU/OD). Error bars represent triplicate repeats. Differences in transformation efficiency of mutant strains determined as non-significant compared to wild-type using GraphPad Prism software (p=0.954 for comX1-, p=0.983 for comX2-). (C) Visual representation of experiment exploring the impact of pre-competence production of DprA and σX on induction. (D) BIP induction of cells lacking CEPIIR-comX does not alter competence induction. Cells possessing comC-luc and CEPlac-dprA (R4511) were treated as described in Panel C. Error bars represent triplicate repeats. (E) Production of σX and DprA prior to competence induction antagonizes competence. Cells possessing comC-luc and CEPlac-dprA and CEPIIR-comX (R4500) were treated as in panel C. Error bars represent triplicate repeats. (F) Inactivation of comW does not alter the antagonization of competence induction mediated by early DprA and σX production. Cells possessing comC-luc, CEPlac-dprA, CEPIIR-comX and ΔcomW::trim (R4509) were treated as in panel C. Error bars represent triplicate repeats. (G) Polar competence induction in Streptococcus pneumoniae. (1) Extracellular CSP interacts with ComD at the cell poles, prompting ComD autophosphorylation. (2) Patches of ComE navigate around the cell membrane. (3) Polar ComD phosphorylates ComE. (4) Active ComE~P dimers leave the cell poles to interact with genomic targets, inducting the early com regulon and (5) launching an autocatalytic feedback loop. (6) Among the early com genes, comX1, comX2, and comW produce σX and its activator ComW, which induce the late com regulon, including DprA. Orange arrows, early com promoters; purple arrows, late com promoters. C, comC; D, comD; E, comE; X1/2, comX1/comX2; W, comW; RNAP, RNA polymerase. (H) Polar shut-off of pneumococcal competence. (1) σX interacts directly with DprA, promoting accumulation of DprA at the cell pole, generating a polar DprA ‘cloud’ in competent cells, near ComD (2). (3) Polar DprA interacts directly with neosynthesized ComE~P, preventing the regulator from accessing its genomic targets. (4) Unphosphorylated ComE interacts with early com promoters, acting as a repressor to prevent induction (Martin et al., 2013), promoting extinction of the competence signal, resulting in competence shut-off (5).
Videos
Timelapse move of DprA-GFP after competence induction by CSP addition.
Images taken every 5 min. Left panel, fluorescence; right panel, phase contrast; middle panel, overlay.
Timelapse move of ComE-GFP after competence induction by CSP addition.
Images taken every 2 min. Left panel, fluorescence; right panel, phase contrast; middle panel, overlay.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Streptococcus pneumoniae) | Various | This paper | NBCI Taxon: 1313 | See Supplementary file 3 |
| Sequence-based reagent | Various oligonucleotides | This paper (Eurofins-MWG) | Primers for cloning | See Supplementary file 3 |
| Software, algorithm | Fiji | doi:10.1038/nmeth.2019 | RRID:SCR_002285 | |
| Software, algorithm | MicrobeJ | doi: 10.1038/nmicrobiol.2016.77 | www.microbeJ.com | |
| Software, algorithm | R project for statistical computing | http://www.r-project.org/ | RRID:SCR_001905 |
Additional files
-
Supplementary file 1
Validation of cin box mutant inactivations.
Successful cin box inactivation was confirmed in three individual cin box mutants with known transformation deficits, and each strain displayed the expected transformation deficit, showing that as expected, full cin box mutation successfully abrogated σX-mediated expression, and allowing extrapolation to the 18 cin box mutants generated.
- https://cdn.elifesciences.org/articles/62907/elife-62907-supp1-v2.docx
-
Supplementary file 2
Effect of cin box inactivations on the presence of polar DprA-mKate2 foci.
- https://cdn.elifesciences.org/articles/62907/elife-62907-supp2-v2.docx
-
Supplementary file 3
Strains, plasmids and primers used in this study.
a Antibiotic resistances: SpcR, spectinomycin; KanR, kanamycin; TrimR, trimethoprim; SmR, streptomycin; CmR, chloramphenicol; RifR, rifampicin; NovR, novobiocin; EryR, erythromycin; TetR, tetracycline; AmpR, ampicillin. b Underlined, italic bases represent restriction sites used for cloning. References cited in this file: Campbell et al., 1998; Burghout et al., 2007; Desai and Morrison, 2006; Attaiech et al., 2008; Morrison et al., 2007; Martin et al., 1995; Sanchez-Puelles et al., 1986; Laurenceau et al., 2013; Laurenceau et al., 2015; Diallo et al., 2017; Lefevre et al., 1979; Caymaris et al., 2010; Mortier-Barrière et al., 2019; Akerley et al., 1998; van Raaphorst et al., 2017; Guiral et al., 2006.
- https://cdn.elifesciences.org/articles/62907/elife-62907-supp3-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/62907/elife-62907-transrepform-v2.docx





