Co-targeting myelin inhibitors and CSPGs markedly enhances regeneration of GDNF-stimulated, but not conditioning-lesioned, sensory axons into the spinal cord
Figures
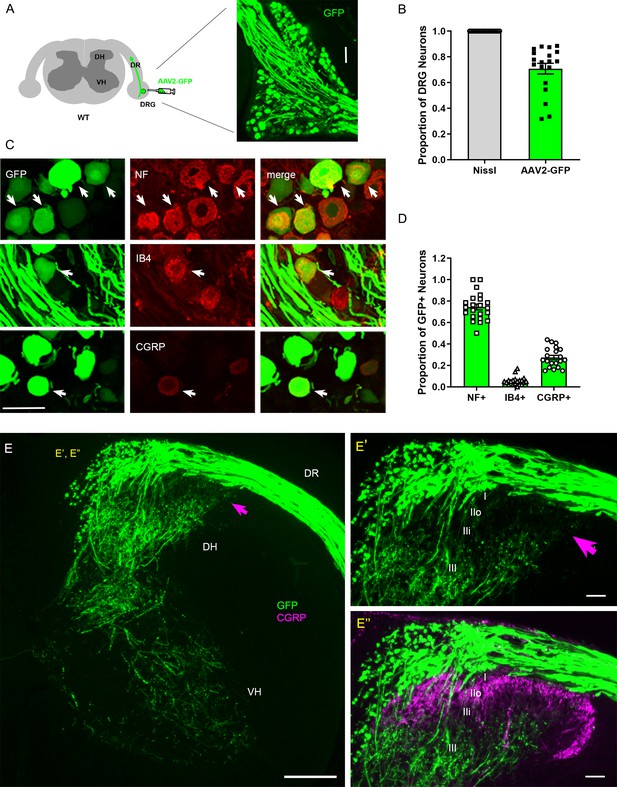
Intraganglionic AAV2-GFP labels proprioceptive and mechanoreceptive axons.
(A) Schematic illustration of intraganglionic injection of scAAV2-eGFP and a representative dorsal root ganglion (DRG) showing infected neurons expressing GFP at 2 weeks post-injection. (B) Mice expressing GFP in >70% Nissl-stained neurons were used in the present study. (C) DRG transverse sections showing GFP+ neurons (arrows) co-expressing neurofilament (NF), IB4, or CGRP. (D) Quantitative comparisons of AAV2-GFP-infected neurons illustrating preferential labeling of large-diameter myelinated NF+ neurons, which mediate proprioception and mechanoreception. n > 20 sections, three mice. (E) A transverse section showing GFP+ axons along the root and within the right side of the spinal cord, projecting into dorsal column, deeper laminae of the dorsal horn and into the ventral horn. An arrow denotes superficial laminae I–IIi lacking GFP fluorescence. (E’, E”) Enlarged views of the superficial dorsal horn, illustrating lack of GFP-fluorescence where CGRP+ nociceptive axons (magenta) innervate. DH: dorsal horn; DR: dorsal root; VH: ventral horn. Scale bars = 50 μm (A, C, E’, E”), 200 μm (E).
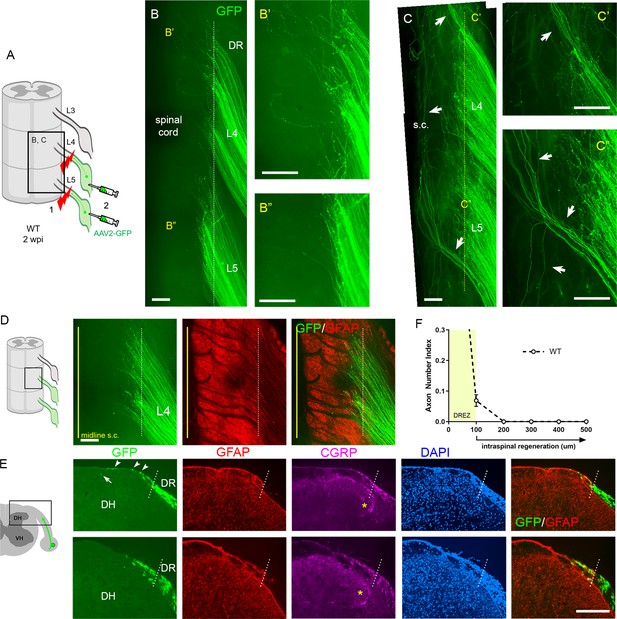
Additional strategies for complete lesions and evaluation of dorsal root (DR) regeneration.
DR regeneration in wildtype (WT) mice assessed in wholemounts (A–D) and transverse sections (E, F) 2 weeks after L4 and L5 DR crush. (A) Schematic illustration of crushing roots prior to intraganglionic AAV2-GFP injections to avoid labeling of degenerating distal stump axons. (B) Wholemount view of completely crushed L4 and L5 DRs illustrating hundreds of GFP+ axons terminated at the entrance of spinal cord. (B’, B”) Enlarged views illustrating most axons terminated near the border. (C–C”) Wholemount views of incompletely crushed DRs showing spared axons with long intraspinal projections. Spared axons are easily detectable in wholemounts and commonly observed in the outermost dorsal rootlets (arrows). (D) Wholemount view of L4 dorsal root entry zone (DREZ) illustrating GFP+ axons that crossed the astrocyte: PNS border (dotted line) and terminated nearby. The astrocytic border is identified by GFAP immunostaining of astrocytes (red). Yellow line denotes spinal cord midline recognized by the midline vein. (E) Four-color immunolabeling of transverse sections illustrating limited penetration of GFP+ or CGRP+ axons through the DREZ. White dotted lines approximate the peripheral boundary of the DREZ (astrocyte: PNS border) by locating peripherally projecting astrocytic processes (red) or by greater abundance of cell nuclei in the PNS (blue). Axons rarely extended >200 μm beyond the border. Arrowheads denote frequently observed axons that grew along the growth-permissive dura. Arrow denotes occasionally observed subdural axons located several hundred microns past the border. (F) Quantitative analysis of DR regeneration on transverse sections (13 sections, three mice). ~90% GFP+ axons terminated within ~100 μm of the border. Axons growing farther than 100 μm are considered as having penetrated the DREZ. DH: dorsal horn; S.C.: spinal cord. Scale bars = 200 μm (B–B”, C–C”, D, E).
-
Figure 2—source data 1
Source data for quantifying regeneration across the dorsal root entry zone.
- https://cdn.elifesciences.org/articles/63050/elife-63050-fig2-data1-v2.xlsx
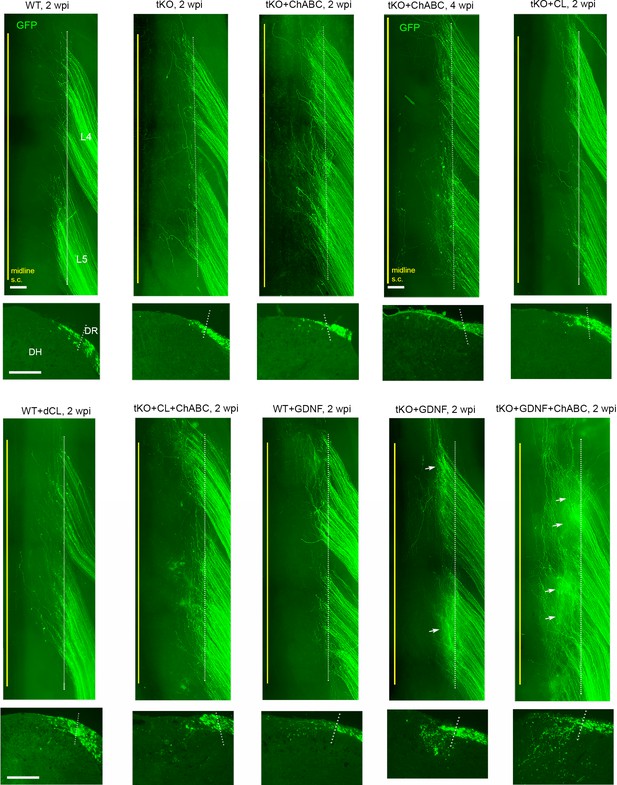
Side-by-side comparison of different mouse groups in wholemounts and transverse sections.
Representative wholemount and transverse sections of wildtype (WT) and Rtn4/Mag/Omg triple knockout (tKO) groups with or without additional removal of chondroitin sulfate proteoglycans (CSPGs) and/or nerve conditioning lesion. Arrows in wholemounts of tKO + glial cell line-derived neurotrophic factor (GDNF) and tKO + GDNF + chondroitinase ABC (ChABC) groups denote intensely fluorescent areas due to abundant subdural GFP+ axons. wpi, weeks post injury; CL: conditioning lesion; dCL: double conditioning lesion. Scale bars = 200 μm.
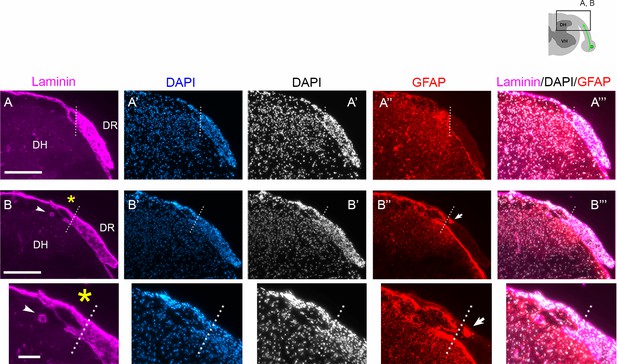
Validation of DAPI as a marker of the CNS:PNS border.
Two sets of wildtype (WT) transverse sections at 2 weeks post injury (wpi) (A–A”’, B–B”’), co-labeled for laminin (A, B), DAPI (A’, B’), and GFAP (A”’, B”’), illustrating that DAPI reveals the astrocyte:PNS border because cell nuclei are much more densely accumulated in the PNS than CNS. The astrocyte:PNS border (dotted line) delineated by laminin, DAPI, and GFAP overlaps closely. Arrowheads in (B) denote radicular arteries also labeled for laminin, which can obscure precise delineation of the boundary. Arrows in (B”) denote astrocytic processes that invaded the PNS after root injury but are readily distinguished from parental astrocytes in CNS territory. Small and large asterisks indicate enlarged views of (B–B”’). Scale bars = 200 μm.
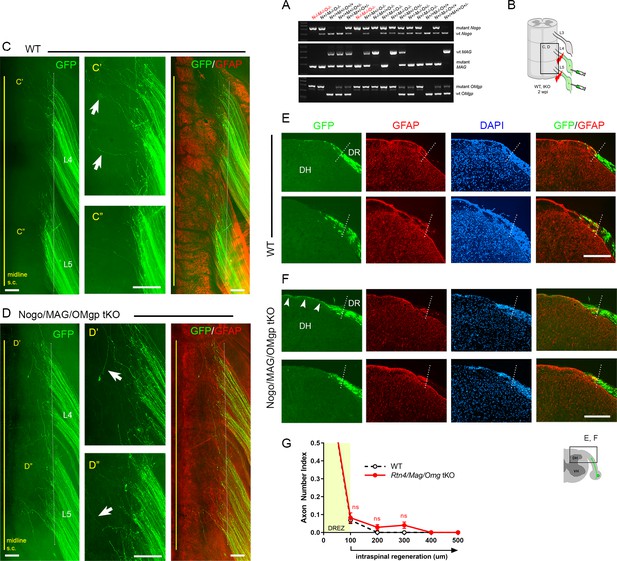
Genetic deletion of Nogo/MAG/OMgp elicits little intraspinal regeneration.
Dorsal root (DR) regeneration in Rtn4/Mag/Omg triple knockout (tKO) mice assessed in wholemounts (D) or transverse sections (F) 2 weeks after L4 and L5 DR crush. (A) Identification of triple null mutants (red) lacking Nogo (A, B, C), MAG, and OMgp. (B) Schematic illustration of the experimental procedures. (C) Wholemount view of a wildtype (WT) mouse. (C’, C”) Enlarged views of L4 and L5 dorsal root entry zone (DREZ) in (C). Arrows denote axons extending longer processes past the DREZ. (D) Wholemount views of a tKO mouse illustrating termination of hundreds of GFP+ axons near the astrocyte:PNS border (dotted line), as in WT mice. The astrocyte:PNS border is identified by GFAP immunostaining of astrocytes (red). (D’, D”) Enlarged views of L4 and L5 DREZ in (D). Arrows denote axons extending longer processes past the DREZ, which were also frequently observed in WT mice. (E) Representative transverse sections of WT mice. (F) Representative transverse sections of Rtn4/Mag/Omg tKO mice illustrating little if any enhanced regeneration of GFP+ axons across the DREZ. Arrows denote axons that grew dorsally along the pia matter, as also observed in WT mice. (G) Quantitative comparisons illustrating no significant difference in WT and Rtn4/Mag/Omg tKO mice. 100 μm, p=0.9738, df = 162; 200 μm, p=0.5046, df = 162; 300 μm, p=0.1454, df = 162. Two-way repeated-measures ANOVA with Sidak’s multiple comparisons test (WT: 13 sections, three mice; tKO: 16 sections, five mice). S.C.: spinal cord; ns: not significant. Scale bars = 200 μm (C–C”, D–D”, E, F).
-
Figure 3—source data 1
Source data for quantifying regeneration across the dorsal root entry zone.
This Excel file contains raw data used for quantitative analysis shown in Figure 3G.
- https://cdn.elifesciences.org/articles/63050/elife-63050-fig3-data1-v2.xlsx
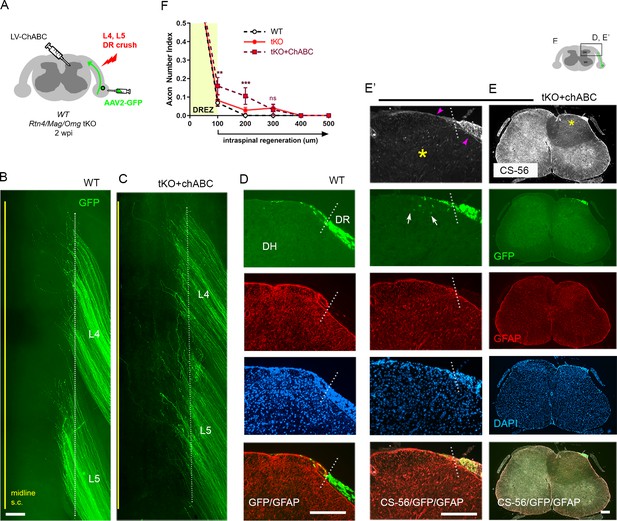
Additional chondroitin sulfate proteoglycan (CSPG) removal slightly increases intraspinal regeneration in triple knockout (tKO) mice.
Dorsal root (DR) regeneration in chondroitinase ABC (ChABC)-expressed Rtn4/Mag/Omg tKO mice assessed in wholemounts (C) and transverse sections (E–E’) 2 weeks after L4 and L5 DR crush. (A) Schematic illustration of the experimental procedures. LV-ChABC was injected into ipsilateral dorsal horn at multiple locations rostrocaudally along the L4–L5 DREZ. (B) Wholemount views of a wildtype (WT) mouse. (C) Wholemount views of a ChABC-expressed tKO showing hundreds of GFP+ axons in L4 and L5 roots terminated near the astrocyte:PNS border (dotted line), as in WT and tKO mice. The astrocyte:PNS border is identified by GFAP immunostaining of astrocytes (red). (D) Representative transverse sections of a WT mouse. (E) Representative transverse sections of a ChABC-expressed tKO illustrating effective degradation of CSPGs and modestly enhanced intraspinal regeneration. CS-56 immunoreactivity is very low in ipsilateral dorsal horn (asterisks), indicating effective removal of inhibitory GAG chains of CSPGs. Arrowheads denote Schwann cell-associated CS-56 immunoreactivity, which is markedly reduced but discernible in ChABC-expressed tKO. (E’) Enlarged views showing a few GFP+ axons that penetrated the DREZ and are located at the top of the dorsal horn (arrows); such axons were not observed in WT or Rtn4/Mag/Omg tKO mice. (F) Quantitative comparisons illustrating modestly improved regeneration in ChABC-expressed Rtn4/Mag/Omg tKO mice: ~15% GFP+ penetrated the dorsal root entry zone (DREZ) and remained within ~200 μm of the border. ChABC-expressed tKO vs. WT: 100 μm, **p=0.0022, df = 186; 200 μm, ***p=0.0003, df = 186; 300 μm, p=0.4818, df = 186. ChABC-expressed tKO vs. tKO: 100 μm, **p=0.0086, df = 186; 200 μm, **p=0.0099, df = 186; 300 μm, p=0.9262, df = 186. Two-way repeated-measures ANOVA with Sidak’s multiple comparisons test (WT: 13 sections, three mice; tKO: 16 sections, five mice; ChABC-tKO: 14 sections, three mice). S.C.: spinal cord; ns: not significant. Scale bars = 200 μm (B, C, D, E–E’).
-
Figure 4—source data 1
Source data for quantifying regeneration across the dorsal root entry zone.
This Excel file contains raw data used for quantitative analysis shown in Figure 4F.
- https://cdn.elifesciences.org/articles/63050/elife-63050-fig4-data1-v2.xlsx
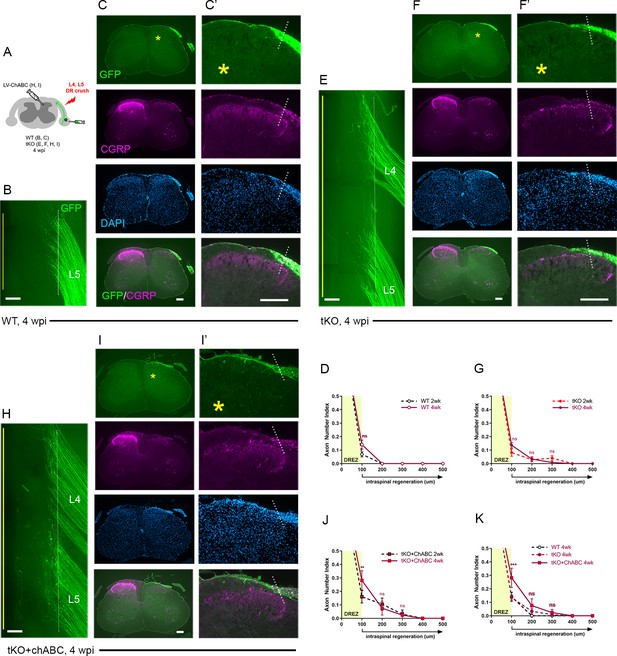
Chronic regeneration failure at the dorsal root entry zone (DREZ) lacking Nogo/MAG/OMgp and chondroitin sulfate proteoglycans (CSPGs).
Dorsal root (DR) regeneration in wildtype (WT) (A–D), Rtn4/Mag/Omg triple knockout (tKO) (E–G), and chondroitinase ABC (ChABC)-expressed Rtn4/Mag/Omg tKO mice (H–K) analyzed 4 weeks after L4 and L5 DR crush. (A) Schematic illustration of the experimental procedures. (B) Wholemount view of L5 DREZ in a WT mouse showing no noticeably enhanced regeneration into the spinal cord. (C, C’) Transverse sections showing no improved penetration of GFP+ (green) and CGRP+ axons (magenta) through the DREZ. (D) Quantitative comparisons of WT mice at 2 weeks post injury (wpi) and 4 wpi illustrating no significant difference. 100 μm, p=0.5292, df = 102. Two-way repeated-measured ANOVA with Sidak’s multiple comparisons test (WT-2 wpi: 13 sections, three mice; WT-4 wpi: 14 sections, four mice). (E) Wholemount view of L4–L5 DREZ in a tKO showing no marked increase in intraspinal regeneration. (F, F’) Transverse sections of a tKO mouse showing GFP+ (green) and CGRP+ axons (magenta) remaining at the DREZ at 4 wpi. (G) Quantitative comparisons of tKO mice at 2 wpi and 4 wpi illustrating no significant difference. 100 μm, p=0.5067, df = 168; 200 μm, p>0.9999, df = 168; 300 μm, p>0.9999, df = 168. Two-way repeated-measures ANOVA with Sidak’s multiple comparisons test (tKO-2 wpi: 16 sections, five mice; tKO-4 wpi: 14 sections, five mice). (H) Wholemount view of L4–L5 DREZ in a ChABC-expressed tKO showing no noticeably enhanced intraspinal regeneration. (I, I’) Transverse sections of a ChABC-expressed tKO showing GFP+ (green) and CGRP+ axons (magenta) remaining at the DREZ at 4 wpi. (J) Quantitative comparisons of ChABC-expressed tKO mice at 2 wpi and 4 wpi illustrating no significant increase in GFP+ axons that penetrated the DREZ. 100 μm, p=0.0027, df = 60; 200 μm, p=0.936, df = 60; 300 μm, p>0.9999, df = 60. Two-way repeated-measures ANOVA with Sidak’s multiple comparisons test (ChABC-tKO-2 wpi: 14 sections, three mice; ChABC-tKO-4 wpi: 12 sections, three mice). (K) Quantitative comparisons of WT, tKO, and ChABC-expressed tKO at 4 wpi showing no significant difference in GFP+ axons crossing the DREZ. ChABC-expressed tKO vs. WT: 100 μm, ***p=0.0001, df = 144; 200 μm, p=0.668, df = 144; 300 μm, p=0.7582, df = 144. Two-way repeated-measures ANOVA with Sidak’s multiple comparisons test. Scale bars = 200 μm (B, C–C’, E–F’, H–I’).
-
Figure 5—source data 1
Source data for quantifying regeneration across the dorsal root entry zone.
This Excel file contains raw data used for quantitative analysis shown in Figure 4D, G, J, K.
- https://cdn.elifesciences.org/articles/63050/elife-63050-fig5-data1-v2.xlsx
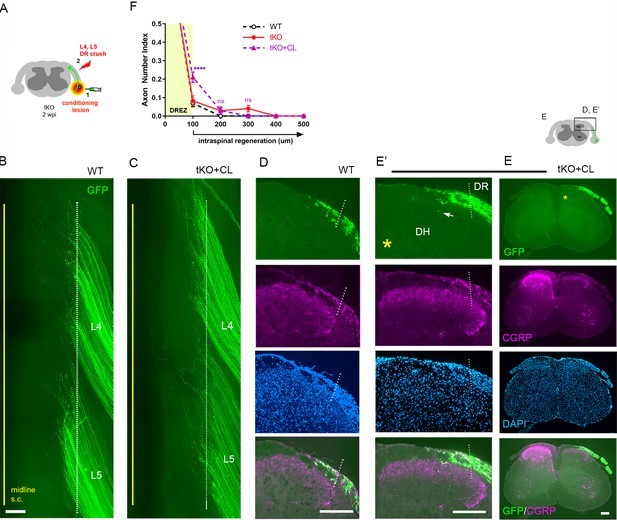
Nerve conditioning lesion does not promote regeneration across the dorsal root entry zone (DREZ) in triple knockout (tKO) mice.
(A) Schematic illustration of the experimental procedures. Rtn4/Mag/Omg tKO mice received a nerve conditioning lesion 10 days before L4 and L5 dorsal root (DR) crush and were assessed at 2 weeks post injury (wpi). (B) Wholemount view of a wildtype (WT) mouse. (C) Wholemount view of a conditioned tKO showing hundreds of GFP+ axons terminated near the astrocyte:PNS border (dotted line), as in WT and tKO mice. (D) Transverse sections of a WT mouse. (E, E’) Transverse sections of a conditioned tKO illustrating little if any enhanced regeneration of GFP+ (green) or CGRP+ axons (magenta) across the DREZ. An arrow denotes occasionally observed GFP+ axons that reached dorsolateral gray matter. (F) Quantitative comparisons illustrating no significant difference in WT, tKO, and conditioned tKO mice. tKO vs. conditioned tKO: 100 μm, ****p<0.0001, df = 220; 200 μm, p=0.9991, df = 220; 300 μm, p>0.9999, df = 220. Two-way repeated-measures ANOVA with Sidak’s multiple comparisons test (WT: 13 sections, three mice; tKO: 16 sections, five mice; conditioned-tKO: 11 sections, three mice). ns: not significant. Scale bars = 200 μm (B, C, D, E, E’).
-
Figure 6—source data 1
Source data for quantifying regeneration across the dorsal root entry zone.
- https://cdn.elifesciences.org/articles/63050/elife-63050-fig6-data1-v2.xlsx
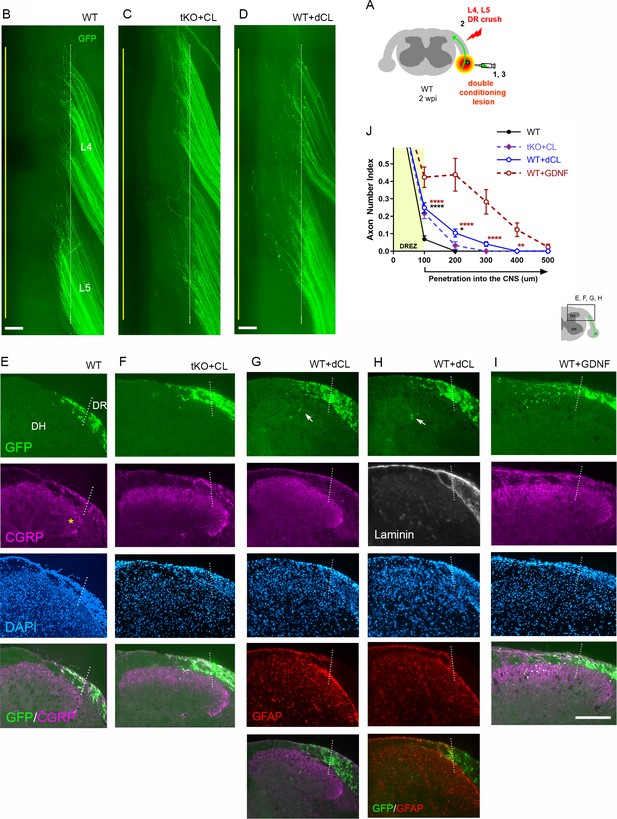
Double conditioning lesion modestly enhances regeneration across the dorsal root entry zone (DREZ) in wildtype (WT) mice.
(A) Schematic illustration of the experimental procedures. WT mice received two conditioning lesions (WT + dCL [double conditioning lesion]); the ipsilateral sciatic nerve was transected 3 days before and 7 days after L4 and L5 dorsal root (DR) crush. Mice were assessed at 2 weeks post injury (wpi). Wholemount and transverse section views of double lesioned WT mice (D, G, H) are presented side-by-side with those of WT (B, E), single lesioned triple knockout (tKO) (C, F), and glial cell line-derived neurotrophic factor (GDNF)-expressed WT mice (I). (B) Wholemount view of a WT that received no conditioning lesion. (C) Wholemount view of a Rtn4/Mag/Omg tKO mouse that received a single nerve crush conditioning lesion. (D) Wholemount view of a WT mouse that received two conditioning lesions showing hundreds of GFP+ axons terminated near the astrocyte:PNS border (dotted line), as in WT and single lesioned tKO mice. (E) Transverse sections of a WT that received no conditioning lesion. (F) Transverse sections of a Rtn4/Mag/Omg tKO mouse that received a single conditioning lesion. (G, H) Transverse sections of a WT mouse that received two conditioning lesions. Arrows indicate occasional GFP+ axons that enter dorsal gray matter. (I) Transverse sections of a GDNF-expressed WT showing numerous GFP+ axons that enter gray matter. See also Figure 8. (J) Quantitative comparisons illustrating modest regeneration of double conditioned DR axons through the DREZ: ~10% GFP+ axons extended ~100 μm past the DREZ. 100 μm, ****p<0.0001, df = 426 (WT vs. WT + dCL, WT + dCL vs. WT + GDNF), p=0.8369, df = 426 (WT + dCL vs. tKO + CL); 200 μm, *p=0.0239, df = 426 (WT vs. WT + dCL), ****p<0.0001, df = 426, p=0.2695, df = 426 (WT + dCL vs. tKO + CL); 300 μm, p=0.6437; df = 426 (WT vs. WT + dCL), ****p<0.0001, df = 426 (WT + dCL vs. WT + GDNF); p=0.7119, df = 426 (WT + dCL vs. tKO + CL); 400 μm, **p=0.0030, df = 426 (WT + dCL vs. WT + GDNF). Two-way repeated-measures ANOVA with Sidak’s multiple comparisons test (WT: 13 sections, three mice; tKO+CL: 11 sections, three mice; WT + dCL: 37 sections, four mice; WT + GDNF: 15 sections, three mice). Scale bars = 200 μm (B–I).
-
Figure 6—figure supplement 1—source data 1
Source data for quantifying regeneration across the dorsal root entry zone.
This Excel file contains raw data used for quantitative analysis shown in Figure 6—figure supplement 1J.
- https://cdn.elifesciences.org/articles/63050/elife-63050-fig6-figsupp1-data1-v2.xlsx
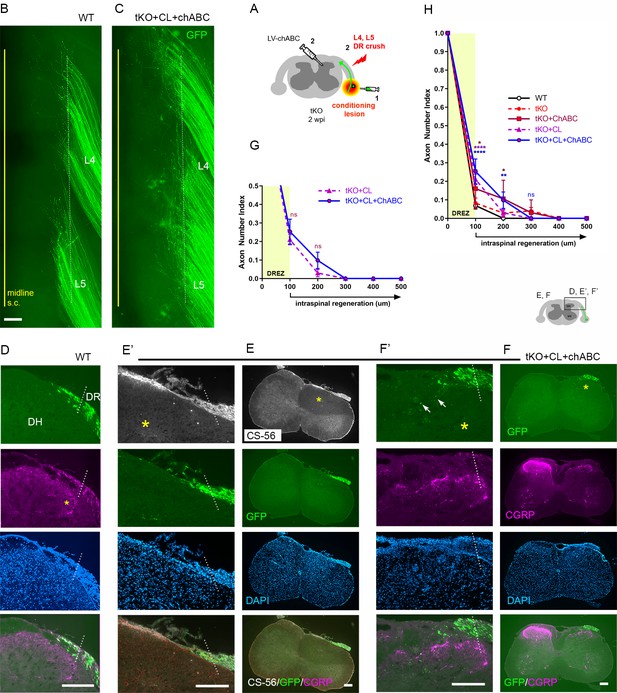
Additional chondroitin sulfate proteoglycan (CSPG) removal minimally enhances regeneration of conditioning lesioned axons in triple knockout (tKO) mice.
(A) Schematic illustration of the experimental procedures. LV-chondroitinase ABC (LV-ChABC) was injected into Rtn4/Mag/Omg tKO mice that received a conditioning lesion 10 days before L4 and L5 dorsal root (DR) crush. (B) Wholemount view of a wildtype (WT) mouse. (C) Wholemount view of a ChABC/conditioned tKO showing hundreds of GFP+ axons that remain near the border (dotted line). (D) Transverse sections of a WT mouse. (E–E’) Transverse sections of a ChABC/conditioned tKO illustrating effective CSPG degradation confirmed by the lack of CS-56 immunoreactivity (asterisks) and little if any intraspinal regeneration of GFP+ axons. (F–F’) Additional transverse sections of a ChABC/conditioned tKO illustrating limited intraspinal regeneration of GFP+ or CGRP+ axons (magenta). Arrows denote occasionally observed GFP+ axons that enter dorsal gray matter. (G) Quantitative comparisons illustrating no significant difference in ChABC/conditioned tKO and conditioned tKO mice. 100 μm, p=0.7629, df = 114; 200 μm, p=0. 2671, df = 114. Two-way repeated-measures ANOVA with Sidak’s multiple comparisons test (conditioned tKO: 11 sections, three mice; ChABC/conditioned tKO: 10 sections, four mice). (H) Quantitative summary illustrating minimal intraspinal regeneration of even conditioned axons after concurrent removal of myelin inhibitors and CSPGs; only ~10% GFP+ axons extended ~100 μm past the dorsal root entry zone (DREZ). 100 μm, *p=0.0488, df = 300 (WT vs. ChABC expressed tKO), ****p<0.0001, df = 300 (WT vs. conditioned tKO, WT vs. ChABC/conditioned tKO); 200 μm, *p=0.014, df = 300 (WT vs. ChABC expressed tKO), **p=0.0024, df = 300 (WT vs. ChABC/conditioned tKO); 300 μm, p>0.9999; df = 300. Two-way repeated-measures ANOVA with Sidak’s multiple comparisons test. Scale bars = 200 μm (B, C, D, E–E’, F–F’).
-
Figure 7—source data 1
Source data for quantifying regeneration across the dorsal root entry zone.
This Excel file contains raw data used for quantitative analysis shown in Figure 7G, H.
- https://cdn.elifesciences.org/articles/63050/elife-63050-fig7-data1-v2.xlsx
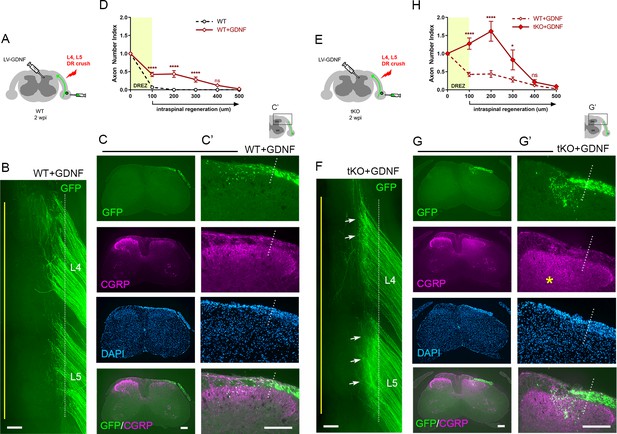
Nogo/MAG/OMgp removal markedly enhances regeneration of glial cell line-derived neurotrophic factor (GDNF)-stimulated dorsal root (DR) axons.
GDNF-induced intraspinal regeneration analyzed in wildtype (WT) (A–D) and Rtn4/Mag/Omg triple knockout (tKO) mice (E–H) 2 weeks after L4 and L5 DR crush. (A) Schematic illustration showing intraspinal injections of LV-GDNF at the time of root crush and AAV2-GFP injections in WT mice. (B) Wholemount view of a GDNF-expressed WT illustrating hundreds of GFP+ axons largely remaining near the border. (C–C’) Transverse sections of a GDNF-expressed WT showing a number of GFP+ (green) or CGRP+ axons (magenta) that cross the DREZ and extend further into the dorsal funiculus and gray matter. (D) Quantitative comparisons illustrating significantly enhanced penetration of GFP+ axons across the DREZ. 100 μm, 200 μm, and 300 μm, ****p<0.0001, df = 156; 400 μm, p=0.2446, df = 156. Two-way repeated-measures ANOVA with Sidak’s multiple comparisons test (WT: 13 sections, three mice; GDNF-expressed WT: 15 sections, three mice). (E) Schematic illustration of the experimental procedures in Rtn4/Mag/Omg tKO mice. (F) Wholemount view of a GDNF-expressed tKO mouse revealing intensely fluorescent area of the L4 and L5 DREZ (arrows), likely due to densely accumulated subdural GFP+ axons. (G–G’) Transverse sections of a GDNF-expressed tKO mouse displaying numerous GFP+ axons regenerating deep into dorsal horn. Asterisks denote CGRP immunoreactivity in deep dorsal laminae (magenta), presumably indicating enhanced regeneration of CGRP+ axons in GDNF-expressed tKO, as compared to that in GDNF-expressed WT. (H) Quantitative comparisons illustrating markedly greater intraspinal growth of GFAP-labeled axons in GDNF-expressed tKO than in GDNF-expressed WT mice. 100 μm and 200 μm, ****p<0.0001, df = 174; 300 μm, *p=0.028, df = 174; 400 μm, p=0.9975, df = 174. Two-way repeated-measures ANOVA with Sidak’s multiple comparisons test (GDNF-expressed WT: 15 sections, three mice; GDNF-expressed tKO: 16 sections, five mice). ns: not significant. Scale bars = 200 μm (B, C, C’, F, G, G’).
-
Figure 8—source data 1
Source data for quantifying regeneration across the dorsal root entry zone.
This Excel file contains raw data used for quantitative analysis shown in Figure 8D, H.
- https://cdn.elifesciences.org/articles/63050/elife-63050-fig8-data1-v2.xlsx
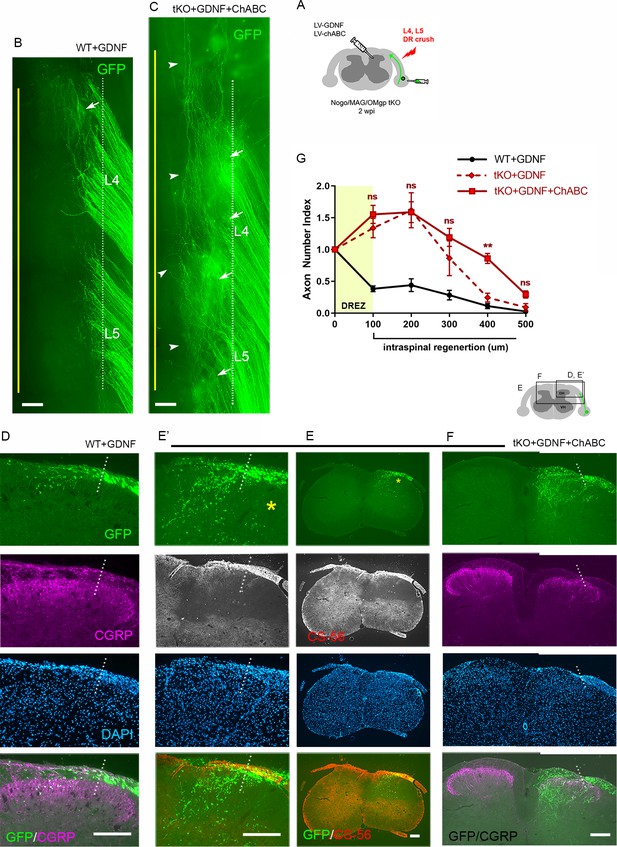
Chondroitin sulfate proteoglycan (CSPG) removal further enhances regeneration of glial cell line-derived neurotrophic factor (GDNF)-stimulated axons in triple knockout (tKO) mice.
(A) Schematic illustration of the experimental procedures. LV-chondroitinase ABC (LV-ChABC) and LV-GDNF were injected into dorsal horn along the L4–L5 dorsal root entry zone (DREZ) in Rtn4/Mag/Omg tKO mice. (B) Wholemount view of a GDNF-expressed wildtype (WT) mouse. (C) Wholemount view of a ChABC/GDNF-expressed tKO showing broader areas of the DREZ and the CNS with densely accumulated GFP+ axons (arrows). Arrowheads denote numerous axons extending rostrocaudally close to the midline. (D) Transverse sections of a GDNF-expressed WT mouse. (E–E’) Transverse sections of a ChABC/GDNF-expressed tKO showing effective degradation of CSPGs confirmed by CS-56 immunoreactivity and many GFP+ axons densely filling broad and deep areas of the dorsal horn. (F) Transverse sections of a ChABC/GDNF-expressed tKO showing enhanced intraspinal regeneration of GFP+ axons and CGRP+ axons (magenta). CGRP+ immunoreactivity is bright, dense, and remarkably restricted to the superficial laminae. (G) Quantitative comparisons illustrating significantly more GFP+ axons in deeper portions of the dorsal horn in ChABC/GDNF-expressed tKO. 100 μm, p=0.5389, df = 210; 200 μm, p=0.9891, df = 210; 300 μm, p=0.2358, df = 210; 400 μm, p=0.0074, df = 210; 500 μm, p=0.5805, df = 210. Two-way repeated-measures ANOVA with Sidak’s multiple comparisons test (ChABC/GDNF-expressed tKO: 19 sections, eight mice; GDNF-expressed tKO: 16 sections, five mice). ns: not significant. Scale bars = 200 μm (B, C, D, E–E’, F).
-
Figure 9—source data 1
Source data for quantifying regeneration across the dorsal root entry zone.
This Excel file contains raw data used for quantitative analysis shown in Figure 9G.
- https://cdn.elifesciences.org/articles/63050/elife-63050-fig9-data1-v2.xlsx
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Mus musculus) | Rtn4/Omg double knockout | Dr. Binhai Zheng (UCSD) Lee et al., 2010 | RRID:MGI:3624445 RRID:MGI:3821705 | |
| Strain, strain background (Mus musculus) | Mag knockout | Dr. Jae K. Lee (U.Miami) | Stock # 006865; RRID:IMSR JAX:006865 | |
| Strain, strain background (Mus musculus) | C57BL/6J | The Jackson Laboratory | Stock # 000664; RRID:IMSR JAX:000664 | |
| Other | Vectashield | Vector Laboratories, Burlingame | H-100 RRID:AB2336789 | Mounting medium |
| Transfected construct (Mus musculus) | scAAV2-eGFP | Dr. George M. Smith (Temple University) | AAV construct to transfect and express eGFP (Liu et al., 2014) | |
| Transfected construct (Mus musculus) | LV-chABC | Dr. George M. Smith (Temple University) | Lentiviral construct to transfect and express chABC (Curinga et al., 2007) | |
| Transfected construct (Mus musculus) | LV-GDNF | Dr. George M. Smith (Temple University) | Lentiviral construct to transfect and express GDNF (Zhang et al., 2013) | |
| Antibody | Anti-NF200 (rabbit polyclonal) | Sigma | #N4142 RRID:AB477272 | IHC (1:500) |
| Chemical compound, drug | Isolectin B4, biotin conjugate | Sigma | #L2140 RRID:AB2313663 | IHC (1:200) |
| Antibody | Anti-CGRP (rabbit polyclonal) | Peninsula Labs | #T4032 RRID:AB2307330 | IHC (1:2000) |
| Antibody | Anti-GFP (mouse monoclonal) | Avés Labs Inc | #GFP-1020 RRID:AB10000240 | IHC (1:500) |
| Antibody | Anti-GFAP (rabbit polyclonal) | Agilent | #N1506 RRID:AB10013482 | IHC (1:500) |
| Antibody | Alexa Fluor 647-goat anti-rabbit IgG secondary antibody | Invitrogen | #31573 RRID:AB2536183 | IHC (1:400) |
| Antibody | Fluorescein (FITC)-conjugated goat anti-rabbit IgG secondary antibody | Millipore | #AP307F RRID:AB92652 | IHC (1:400) |
| Antibody | Alexa Fluor 568-conjugated goat anti-mouse IgG1 secondary antibody | Invitrogen | #A21124 RRID:AB141611 | IHC (1:400) |
| Antibody | Rhodamine (TRITC)-conjugated streptavidin secondary antibody | Jackson ImmunoResearch Labs Inc | #016-020-084 RRID:AB2337237 | IHC (1:400) |
| Antibody | Alexa-Fluor 568-conjugated goat anti-rabbit secondary antibody | Invitrogen | #A11011 RRID:AB143157 | IHC (1:400) |
| Antibody | Alexa Fluor 488-donkey anti-chicken IgG secondary antibody | Jackson ImmunoResearch Labs Inc | #703-545-155 RRID:AB2340375 | IHC (1:400) |
| Other | DAPI stain | Thermo Fisher Scientific | #D1306 RRID:AB2629482 | IHC (1:1000) |
| Other | Nissl substance stain | Thermo Fisher Scientific | #N21482 RRID:AB2620170 | IHC (1:200) |
| Software, algorithm | MetaMorph Image Analysis Software | Molecular Devices | RRID:SCR002368 | |
| Software, algorithm | AxioVision Imaging System | Zeiss | RRID:SCR002677 | |
| Software, algorithm | Axio Imager | Zeiss | RRID:SCR018876 | |
| Software, algorithm | Imaris | Bitplane | RRID:SCR007370 | |
| Software, algorithm | Adobe Photoshop | Adobe Inc | RRID:SCR014199 | |
| Software, algorithm | PRISM 8.0 | GraphPad | RRID:SCR002798 |





