Molecular basis for functional connectivity between the voltage sensor and the selectivity filter gate in Shaker K+ channels
Figures

Proposed chain of residues involved in the voltage sensor selectivity filter gate (VS-SF) coupling.
(A) Schematic representation of residues involved in the VS-SF coupling. Voltage sensor domain (VSD) is depicted in blue, whereas pore domain (PD) is depicted in light purple. (B) Intramembrane view of the active state. (C) Intramembrane view of S4 in resting state and PD in active state. (D) Extracellular view of the active state S4 segment (blue ribbon) and PD (light purple ribbon) are from an adjacent subunit, based on a computational 3D model of crystallographic structures of KV1.2 (Chen et al., 2010) (Shaker-like channel, PDB: 3LUT). The model shows the VSD in the active/relaxed state for B and D. The S4 in the resting state is based on the consensus model (Vargas et al., 2011; Vargas et al., 2012). Residues proposed to be implicated in the VS-SF coupling mechanism are depicted by their van der Waals volumes (Carvalho-de-Souza and Bezanilla, 2019) (Shaker numbering): L361 (vermilion), L366 (black), L409 (green), S411 (blue), S412 (light purple), F433 (orange), W434 (yellow), and Y445 (sky blue). From the consensus model, it is possible to observe that L361 and L366 are practically in the same horizontal plane with the VS-SF coupling residues from the PD.
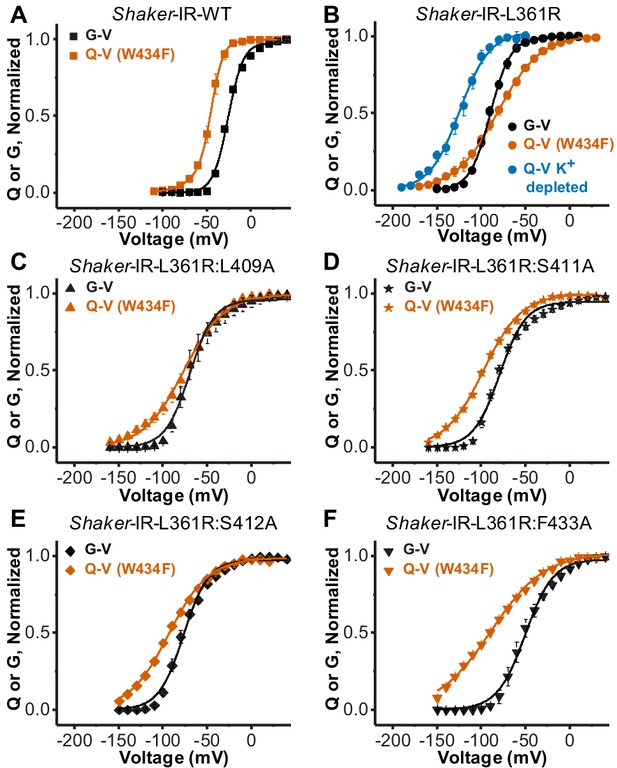
Disrupting the voltage sensor selectivity filter gate (VS-SF) coupling in Shaker-IR-L361R:W434F.
(A) Conductance-voltage (G-V) (black) and charge-voltage (Q-V) (W434F-vermilion) curves from Shaker-IR-WT channels. Note the Q-V curve is displaced to more hyperpolarized potentials with respect to the G-V curve. (B) G-V (black) and the Q-V curves (W434F-vermilion) curves from Shaker-IR-L361R:W434F channels. Note that G-V crosses Q-V (W434F-vermilion), but not the Q-V when K+ is depleted (blue). Adapted from Carvalho-de-Souza and Bezanilla, 2018. (C), (D), (E), and (F) are, respectively, G-V (black) and Q-V (W434F-vermilion) curves for Shaker-IR-L361R:L409A, Shaker-IR-L361R:S411A, Shaker-IR-L361R:S412A, and Shaker-IR-L361R:F433A channels. Note that when any of the residues L409, S411, S412, or F433 are mutated to Ala, the Q-V curve (W434F) does not cross the associated G-V curve, indicating the disruption of the VS-SF coupling. Continuous lines over Q-V and G-V curves are the best fit of Equations 4 and 2 (two-state model), except the Q-V curve for WT channels that was fitted by Equation 3 (three-state model). Best-fitted values are listed in Table 1. Plotted data are mean ± SEM (N = 4-8).
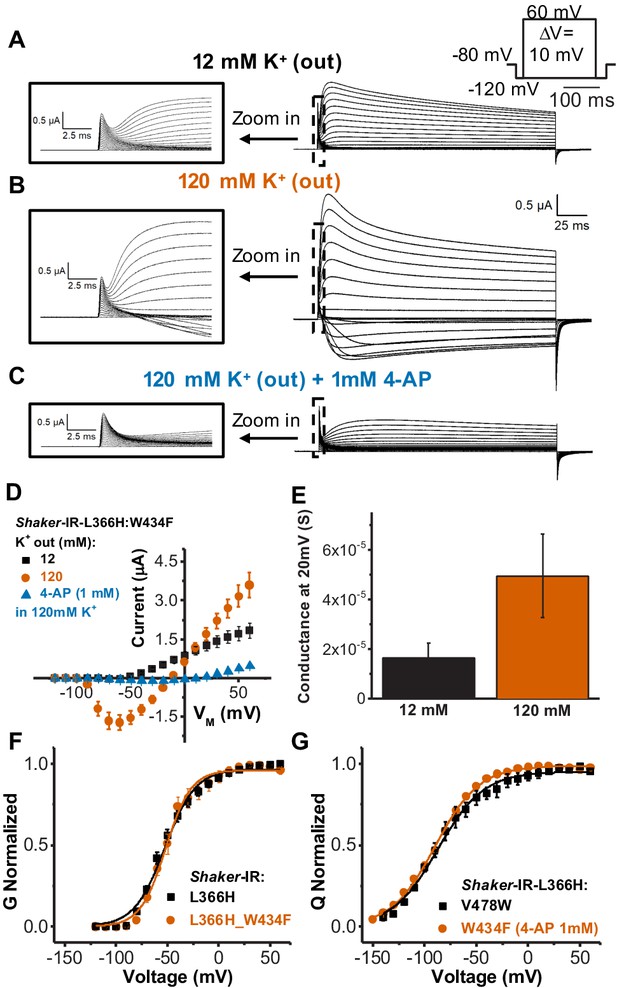
Single amino acid mutation in the voltage sensor partially relieves the inactivation in channels bearing W434F mutation.
(A) Typical currents from Shaker-IR-L366H:W434F channels elicited in 120 mM internal and 12 mM external K+ solution. (B) Currents elicited in symmetric 120 mM K+ solutions. (C) Typical currents in the presence of 1 mM 4-aminopyridine (4-AP) in symmetric 120 mM K+ solutions showing blockage of ionic currents (compared to B). Insets in A, B, and C are a time expansion to better visualize the gating currents. Voltage protocol used to record the currents is inset in A and currents presented in A, B, and C are from the same oocyte. (D) I-V relationships for mutant L366H:W434F in different ionic conditions, as indicated. (E) Conductance at 20 mV for L366H:W434F in 12 and 120 mM external K+ concentration, and 120 mM internal K+ concentration. We used 20 mV for comparison because the conductance-voltage relationship (G-V) at that voltage is already at its maxima. The conductance was calculated using the K+ currents peak as described in 'Materials and methods'. (F) Normalized G-V curves for L366H and L366H:W434F. K+ concentration used was 120 mM internal and 12 mM external. (G) Nearly identical charge-voltage (Q-V) curves measured with L366H:V478W (black squares) and L366H:W434F 1 mM 4-AP (vermilion circles). Continuous lines over G-V and Q-V curves are the best fittings of Equations 2 and 4 (two-state model), respectively. Best-fitted values are listed in Table 1. Data shown as mean ± SEM (N=3).
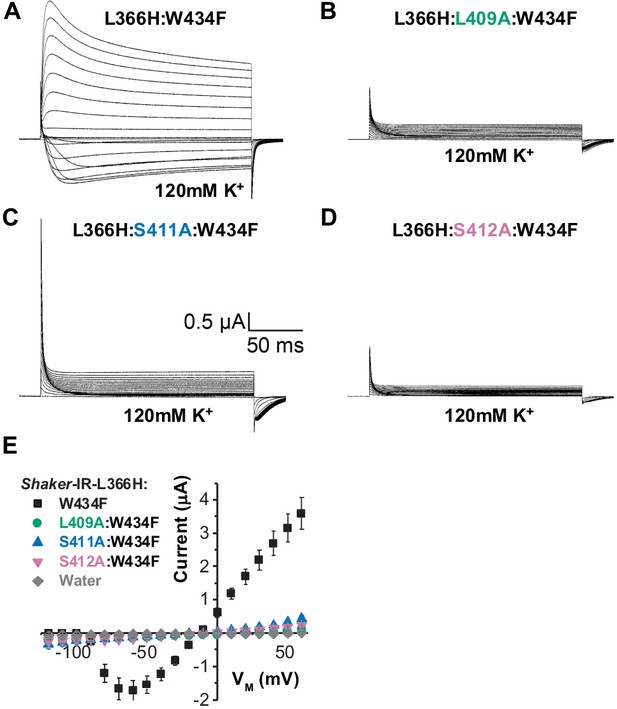
Disrupting the voltage sensor selectivity filter gate (VS-SF) coupling in Shaker-IR-L366H:W434F.
(A) Typical currents from L366H:W434F channels elicited in symmetric 120 mM K+ solutions as indicated. (B), (C), and (D) are currents also elicited in symmetric K+ solutions for L409, S411, or S412 mutated to alanine in L366H:W434F channels, respectively. Voltage protocol used is the same shown in Figure 3a. Note that K+ currents are not present in recording shown in B, C, and D, indicating the disruption of the VS-SF coupling. (E) I-V relationship taken from the peaks of K+ currents show that when L409, S411, and S412 are mutated to Ala, the K+ currents are dramatically diminished. The small remaining current is indistinguishable from leak. Data shown as mean ± SEM (N=4).
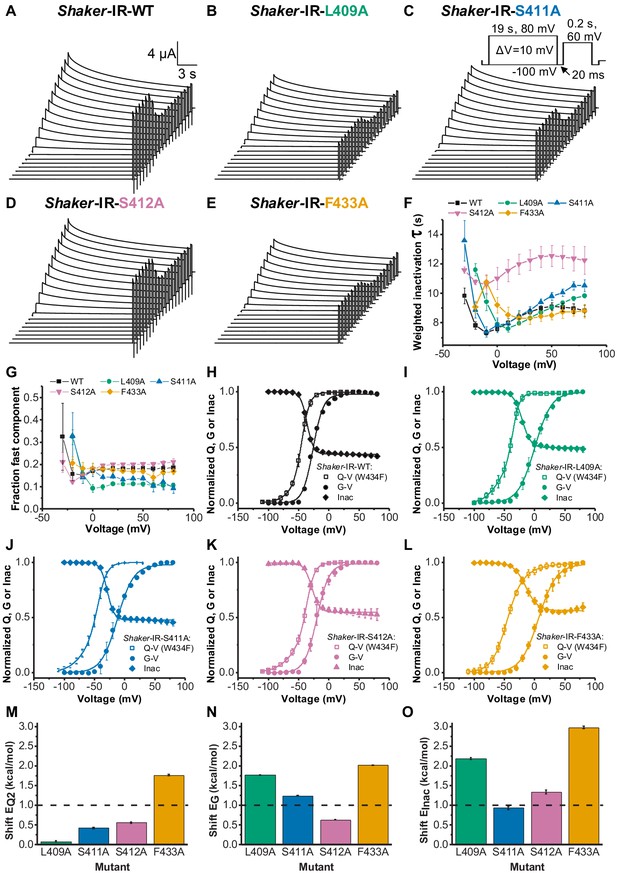
Voltage sensor domain (VSD) to pore opening coupling and C-type inactivation are affected by L409A, S411A, S412A, and F433A.
A, B, C, D, and E are, respectively, typical K+ currents elicited by the voltage protocol inset in C (not to scale), for Shaker-IR-WT, Shaker-IR-L409A, Shaker-IR-S411A, Shaker-IR-S412A, and Shaker-IR-F433A channels. (F) Weighted inactivation time constants calculated from a double exponential fit to currents from WT, L409A, S411A, S412A, and F433A channels. (G) Fraction of the fast component of inactivation for channels tested as indicated in the graph. (H, I, J, K, and L) are, respectively, charge-voltage (Q-V) (open square), conductance-voltage (G-V) (solid circle), and Inac-voltage (Inac-V) curves (solid diamond) for WT (black), L409A (green), S411A (blue), S412A (light purple), and F433A (orange) channels. Currents used to build Inac-V curves were taken from the peak currents elicited by the 60 mV testing pulse after the 19 s inactivating period. M, N, and O are, respectively, shifts (from WT values) in the energy involved in the second component of Q-V, G-V, and Inact-V curves, as caused by L409A, S411A, S412A, and F433A mutations. Continuous lines over G-V, Q-V, and Inac-V curves are the best fits to Equations 2, 3, and 7, respectively. Q-V from F433A:W434F was fitted using Equation 4. The best-fitted values are listed in Table 1. Data shown as mean ± SEM for F to L; and mean ± SE for M to O (N=5-8).

Interaction between residues S411 and F433 does not depend on W434F mutation.
(A) Q-V for Shaker-IR-V478W (black), Shaker-IR-S411A:V478W (blue), Shaker-IR-F433A:V478W (orange), and Shaker-IR-S411A:F433A:V478W (purple). (B) Diagram showing the thermodynamic mutant cycle analysis used to estimate the interaction between residues S411 and F433. The energies were calculated using the generalized interaction-energy analysis (GIA). The VMedian and energy calculated for each channel are shown in Table 2. Data shown as mean ± SEM (N=4-8).
Tables
Best-fitted values for all the mutants studied.
| Mutant | Q-V | G-V | Inac-V | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| V0 | z0 | V1 | z1 | V1/2G | zG | V1/2Inac | zInac | |||||||||||||||||
| WT | −26.0 | ± | 0.6 | 2.9 | ± | 0.2 | −36 | ± | 0 | 4.6 | ± | 0.3 | ||||||||||||
| W434F | −51.9 | ± | 1.9 | 1.7 | ± | 0.2 | −46.5 | ± | 1.1 | 3.4 | ± | 0.1 | ||||||||||||
| L361R* | −124.3 | ± | 1.1 | 1.5 | ± | 0.1 | −89.7 | ± | 0.4 | 2.2 | ± | 0.1 | ||||||||||||
| L361R:W434F | −81.5 | ± | 0.3 | 1.0 | ± | 0.0 | ||||||||||||||||||
| L361R:L409A | −69.4 | ± | 1.1 | 1.8 | ± | 0.1 | ||||||||||||||||||
| L361R:L409A:W434F | −76.2 | ± | 1.1 | 1.1 | ± | 0.0 | ||||||||||||||||||
| L361R:S411A | −79.2 | ± | 1.2 | 1.7 | ± | 0.1 | ||||||||||||||||||
| L361R:S411A:W434F | −97.4 | ± | 0.5 | 1.2 | ± | 0.0 | ||||||||||||||||||
| L361R:S412A | −76.4 | ± | 1.0 | 1.8 | ± | 0.1 | ||||||||||||||||||
| L361R:S412A:W434F | −93.2 | ± | 0.5 | 1.1 | ± | 0.1 | ± | ± | ||||||||||||||||
| L361R:F433A | −50.1 | ± | 1.0 | 1.6 | ± | 0.1 | ||||||||||||||||||
| L361R:F433A:W434F | −89.3 | ± | 1.7 | 0.9 | ± | 0.1 | ||||||||||||||||||
| L366H | −52.5 | ± | 1.2 | 1.6 | ± | 0.1 | ||||||||||||||||||
| L366H:W434F | −87.2 | ± | 1.2 | 1.1 | ± | 0.1 | −52 | ± | 1 | 2.0 | ± | 0.1 | ||||||||||||
| L366H:V478W | −89.8 | ± | 0.5 | 1.1 | ± | 0.1 | ||||||||||||||||||
| L409A | 0.2 | ± | 0.6 | 1.8 | ± | 0.1 | −20.6 | ± | 0.5 | 3.5 | ± | 0.2 | ||||||||||||
| L409A:W434F | −62.4 | ± | 0.6 | 2.2 | ± | 0.1 | −36.3 | ± | 0.2 | 4.2 | ± | 0.1 | ||||||||||||
| S411A | −11.9 | ± | 1.0 | 1.9 | ± | 0.1 | −28.7 | ± | 0.4 | 4.4 | ± | 0.3 | ||||||||||||
| S411A:W434F | −67.2 | ± | 0.6 | 1.7 | ± | 0.1 | −44.4 | ± | 0.4 | 3.1 | ± | 0.1 | ||||||||||||
| S412A | −20.0 | ± | 0.6 | 2.5 | ± | 0.1 | −28.0 | ± | 0.6 | 3.9 | ± | 0.3 | ||||||||||||
| S412A:W434F | −60.9 | ± | 0.6 | 1.7 | ± | 0.1 | −37.2 | ± | 0.3 | 3.5 | ± | 0.1 | ||||||||||||
| F433A | 6.6 | ± | 0.7 | 1.7 | ± | 0.1 | −14.6 | ± | 0.8 | 2.6 | ± | 0.2 | ||||||||||||
| F433A:W434F | −44.0 | ± | 0.3 | 1.8 | ± | 0.1 | ||||||||||||||||||
-
*Q-V calculated by K+ depletion. Please note that L361R, L361R:W434F, L361R:S412A:W434F, L361R:F433A:W434F, L366H:W434F, L366H:V478W, L366H:S412A:W434F, and F433A:W434F, a two-state model Boltzmann equation, Equation 4, was used to fit the Q-V curves.
VMedian and the free energy for the interacting residues S411 and F433.
| Mutant | VMedian (mV) | G (kcal/mol) | ||||
|---|---|---|---|---|---|---|
| V478W | −44.29 | ± | 1.24 | −13.90 | ± | 0.39 |
| S411A:V478W | −48.54 | ± | 1.13 | −15.24 | ± | 0.35 |
| F433A:V478W | −54.34 | ± | 1.38 | −17.06 | ± | 0.43 |
| S411A:F433A:V478W | −52.21 | ± | 1.11 | −16.39 | ± | 0.35 |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene Drosophila melanogaster | Shaker zH4 K+ channel with Inactivation removed | Hoshi et al., 1990 doi:10.1126/science.2122519 | ||
| Biological sample (Xenopus laevis, female) | Oocytes | Nasco | #LM00531 | Protocol #71475 (IACUC) |
| Chemical compound, drug | 4-Aminopyridine | Sigma-Aldrich | 275875–1G | 1 mM in dissolved in external solution |





