Rapid adaptation to malaria facilitated by admixture in the human population of Cabo Verde
Figures
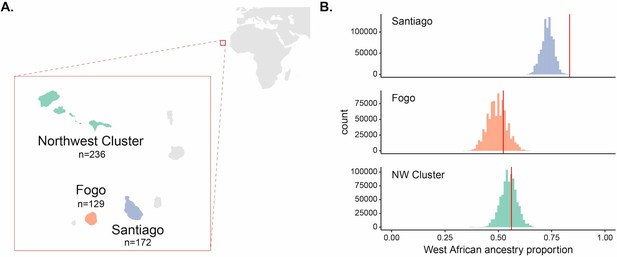
Enrichment of West African ancestry at the DARC locus in Santiago, Cabo Verde.
(A) Map of Cabo Verde islands and sample sizes for number of individuals from each island region. (B) The distribution of West African-related local ancestry proportion across the genome by SNP (n = 881,279) by island, with the DARC locus marked by vertical red lines. Local ancestry was estimated using RFMix (see Materials and methods). The DARC locus is an outlier for high West African-related ancestry in Santiago, but not Fogo or the Northwest Cluster.
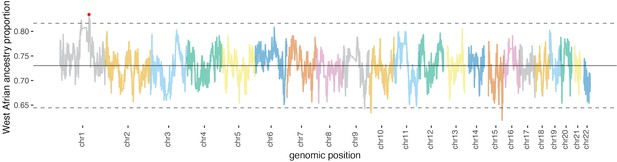
Local ancestry proportion along the genome in Santiago.
The mean is indicated by the solid horizontal line, and dashed horizontal lines represent three standard deviations from the mean. Again, this plot demonstrates Duffy-null (red dot) as the highest value for West African ancestry proportion.
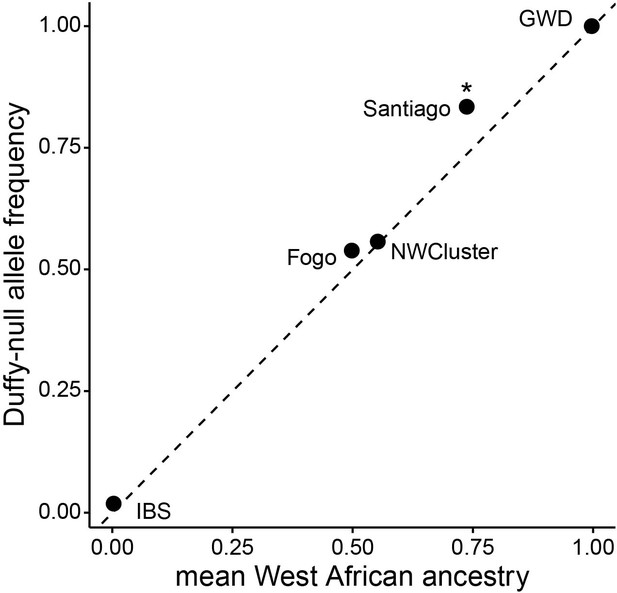
The observed frequency of Duffy-null for each island vs neutral expectation based on mean global ancestry (as estimated by admixture).
*indicates significant p-value <0.001 for binomial test (see Table 1 for sample sizes and further details).
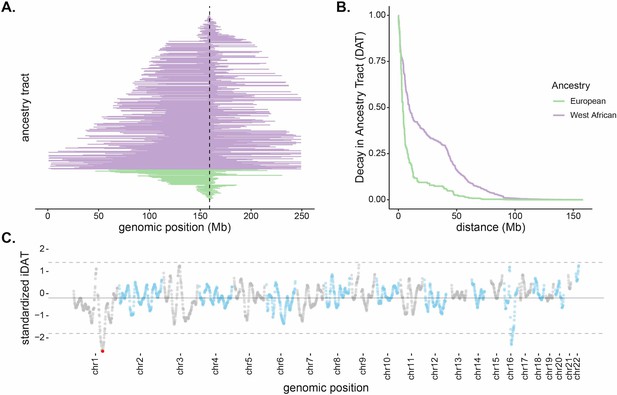
Long, high-frequency West African ancestry tracts span the DARC locus in Santiago.
(A) The distribution of West African (purple) and European (green) ancestry tract lengths spanning the DARC locus (dashed line). Each horizontal line represents a single chromosome in the population (n = 343, one chromosome was excluded due to having unknown ancestry at the DARC locus). (B) Decay in Ancestry Tract (DAT) as function of absolute distance from the Duffy-null allele for West African (purple) and European (green) ancestry tracts. (C) Mean standardized integrated DAT (iDAT) score for 20 Mb sliding windows (step size = 1 Mb), using standardized iDAT for 10,000 random positions across the genome. Horizontal solid gray line indicates mean windowed standardized iDAT score (−0.196), and horizontal dashed gray lines indicate three standard deviations from the mean windowed score. The red dot is the most extreme windowed standardized iDAT score (−2.602), indicative of a larger area under the curve for West African DAT compared to European DAT. This 20 Mb window contains the Duffy-null SNP.
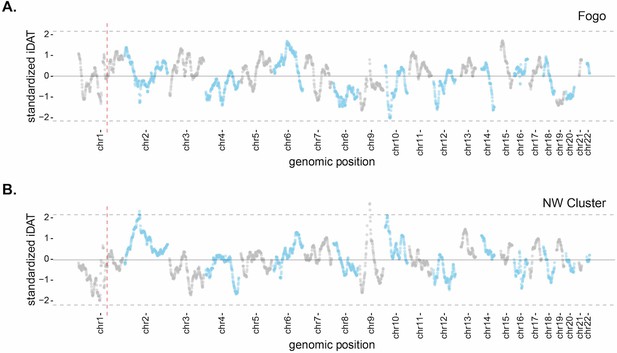
Mean standardized integrated Decay in Ancestry Tract (iDAT) score for 20 Mb sliding windows (step size = 1 Mb), using standardized iDAT for 10,000 random positions across the genome for (A) Fogo and (B) the Northwest Cluster.
Solid gray lines indicate mean windowed standardized iDAT score for each island (Fogo, 0.006; NW Cluster, −0.024) and dashed gray lines indicate three standard deviations from the mean. Vertical dashed red lines indicate the DARC locus, which is not an outlier for either Fogo or the NW Cluster.
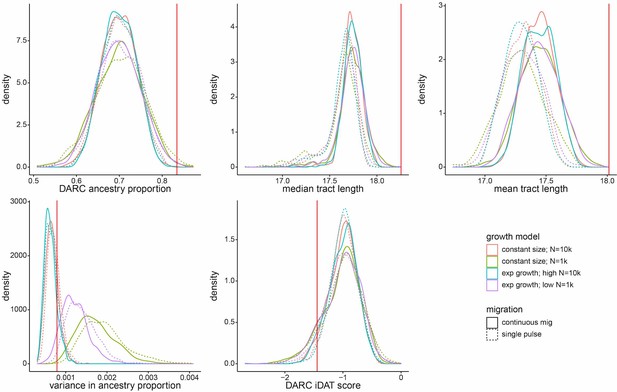
Density distributions for five ancestry-based statistics under eight neutral models.
Summary statistics were calculated from a random sample of 172 individuals from each simulated population, matching the number of individuals from Santiago included in our analyses. High population size models correspond to initial N = 10,000, low population size (high drift) models correspond to initial N = 1000. Exponential growth model corresponds to a rate of 0.05 per generation. Continuous migration refers to 1% total new migrants each generation, at the same proportions as initial admixture contributions for each source population. Vertical red line represents each measure’s observed value for Santiago.
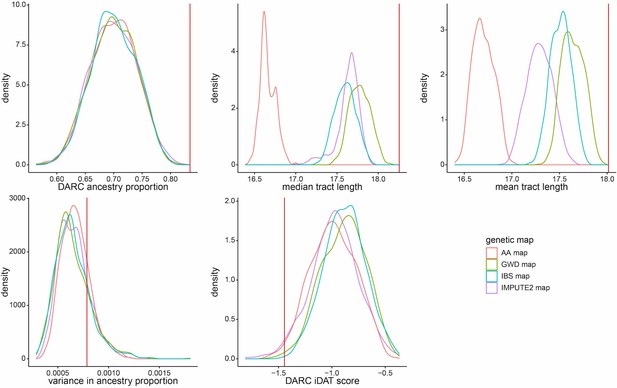
Density distributions for five ancestry-based statistics under simulations using different genetic maps.
Simulations shown assumed a single pulse of admixture with exponential growth at a rate of 0.05 per generation and an initial population size of N = 10,000. Initial admixture contributions were drawn from a uniform distribution from 0.65 to 0.75. Summary statistics were calculated from a random sample of 172 individuals from each simulated population, matching the number of individuals from Santiago included in our analyses. Genetic maps correspond to the population-averaged IMPUTE2 map, Iberian Population in Spain (IBS)-specific genetic map, Gambian in Western Division (GWD)-specific genetic map, and African American (AA)-specific genetic map. Vertical red line represents each measure’s observed value for Santiago.
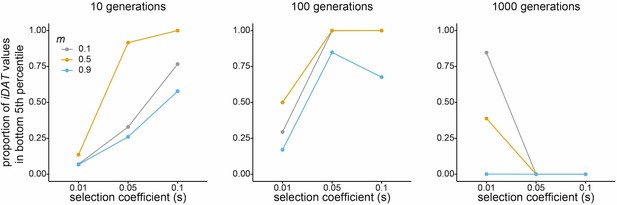
Performance of integrated Decay in Ancestry Tract (iDAT) under various scenarios.
Each plot corresponds to number of generations since admixture (10 – left; 100 – middle; 1000 – right). Line and point colors correspond to source population one admixture contribution at (gray), (yellow), and (blue). Within each plot, the x-axis shows selection strength for the simulated variant at the Duffy-null position, and the y-axis shows the proportion of Duffy-null iDAT values from the selection simulations that are in the bottom fifth percentile of the simulated neutral Duffy-null iDAT distribution. Notably, iDAT cannot be calculated for variants that are fixed in the population, as was the case for many simulations of older admixture (100 or 1000 generations) and high admixture proportion (m = 0.9) and/or stronger selection. This is reflected in the statistic’s performance under these scenarios.
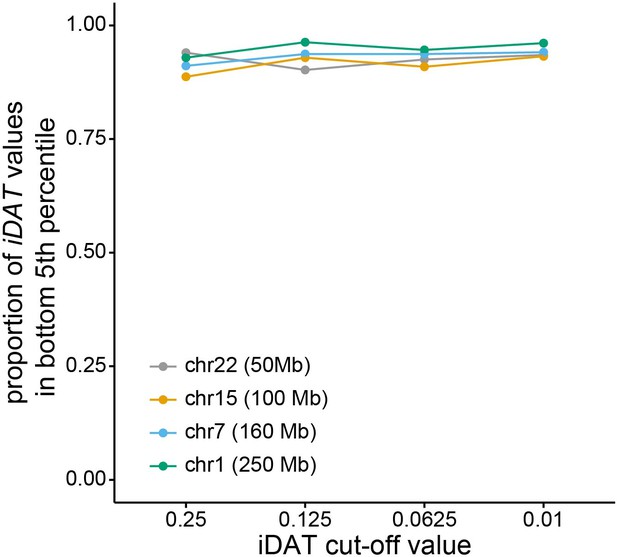
Performance of integrated Decay in Ancestry Tract (iDAT) for various chromosome sizes and cut-off values.
Line and point colors correspond to simulated human chromosome and corresponding size (chr 1 – green; chr 7 – blue; chr 15 – yellow; chr 22 – gray). X-axis shows DAT cut-off values, and y-axis shows proportion of iDAT values at the simulated variant under selection that are in the bottom fifth percentile of simulated neutral iDAT values.
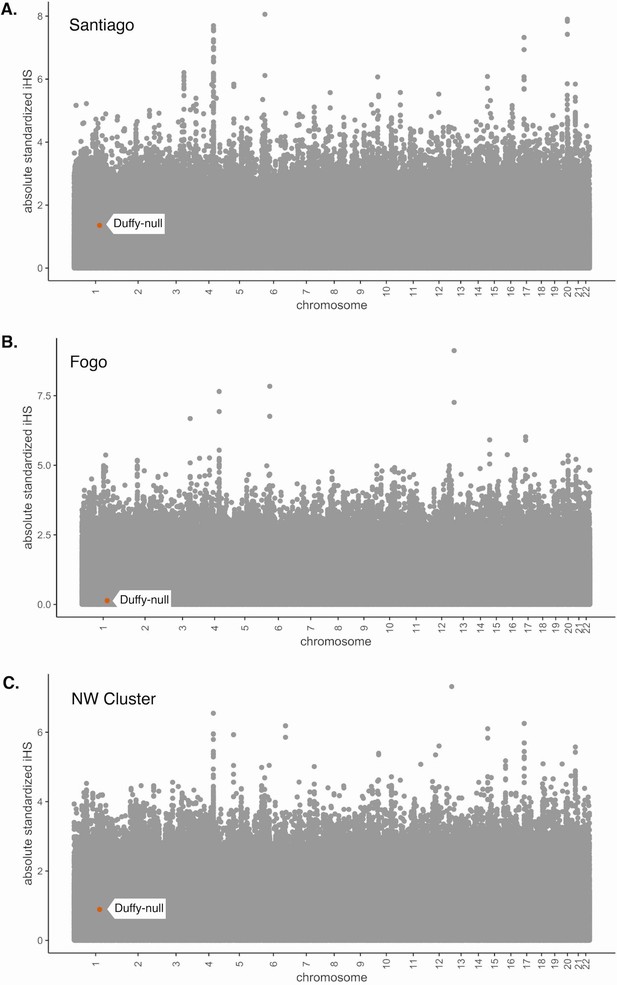
Absolute values of iHS for SNPs in the Cabo Verde data set.
iHS was calculated using the hapbin software and standardized using the default method based on allele frequencies. (A) Santiago, (B) Fogo, and (C) NW Cluster. Value for Duffy-null SNP is indicated by orange dot and white label. Duffy-null iHS value is nonsignificant in all island regions.
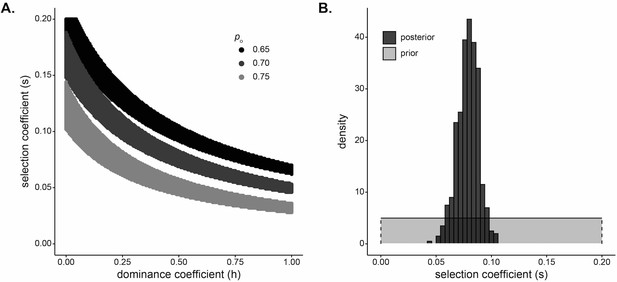
Strong selection inferred at the DARC locus in Santiago.
(A) Pairs of and that result in a small difference in final allele frequency calculated under the model and the allele frequency observed in the Santiago genetic data, under a deterministic population genetic model. Colors indicate the initial Duffy-null frequency: , black; , dark gray; , light gray. (B) Approximate Bayesian computation (ABC) estimates of the selection coefficient for Duffy-null on Santiago. Shaded gray area shows prior distribution of selection coefficient []. Dark gray histogram shows posterior distribution for selection coefficient (median = 0.0795), constructed from regression-adjusted values from accepted simulations.
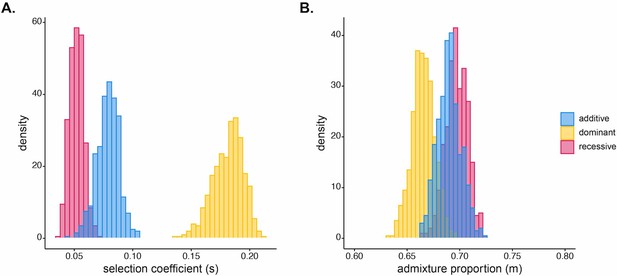
Results of approximate Bayesian computation (ABC) estimation of posterior distributions for (A) selection coefficient for Duffy-null and (B) initial West African ancestry contribution for Santiago.
Duffy-null allele was modeled as additive (blue; ), dominant (yellow; in SLiM), or recessive (pink; in SLiM). Posterior median estimates for selection coefficient: , , ; initial ancestry contribution: , , . Prior distributions were and .
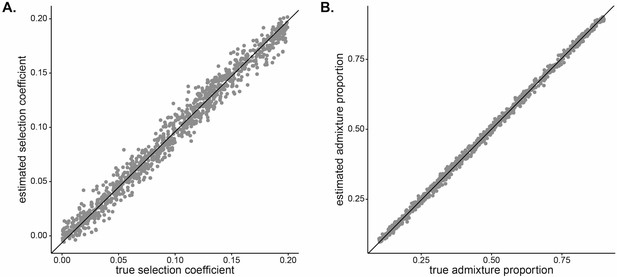
Results of leave-one-out cross-validation of approximate Bayesian computation (ABC) joint estimation.
(A) Selection coefficient (, ) and (B) initial West African admixture contribution (, ).
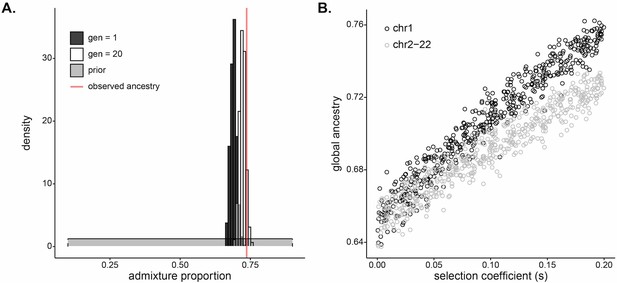
Selection at a single locus impacts genome-wide ancestry proportion.
(A) Inferred (dark gray), simulated (white), and observed (red) mean of global ancestry in Santiago over time. The dark gray histogram plots the posterior distribution for initial West African ancestry contribution inferred using approximate Bayesian computation (ABC) (median, 0.690); the prior distribution [] is in light gray. The red line plots the mean global ancestry estimated by admixture from modern genetic data from Santiago, 0.737. The observed global ancestry is higher than most values of the initial contributions inferred in dark gray. The white histogram plots the distribution of West African global ancestry proportion calculated after 20 generations in populations simulated with selection coefficients and initial ancestries drawn from the ABC-inferred values (median, 0.723). The global ancestry calculated after 20 generations of simulated selection (white) more closely matches that observed from Santiago genetic data (red line). (B) West African mean global ancestry proportion calculated for 500 simulated populations after 20 generations under varying single-locus selection coefficients, . We simulated whole autosomes, setting the initial West African ancestry contribution to 0.65. Black circles indicate mean ancestry on chromosome 1 alone. Gray circles indicate mean ancestry on the other autosomes (2–22). The increase in ancestry with selection for gray circles demonstrates that selection impacts global ancestry beyond the local effects of the chromosome under selection.
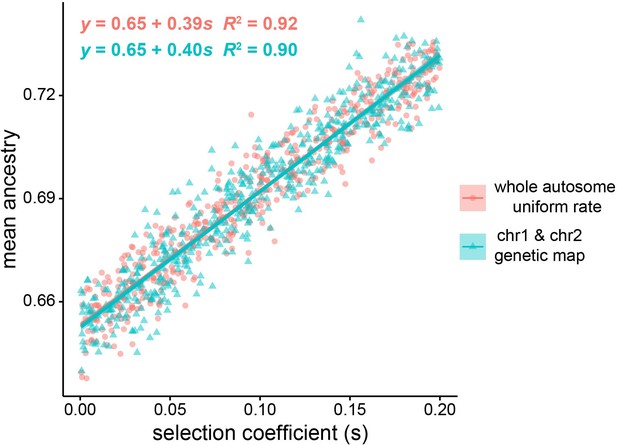
Effect of selection on global ancestry across simulation methods.
Pink circles indicate West African mean global ancestry after 20 generations versus selection coefficient for whole autosome (22 chromosome) simulations, using a uniform recombination rate within each chromosome. Green triangles represent mean weighted ancestry for chromosome 1 and chromosome 2, with chromosome 2 representing the 92% of the genome that segregates independently from chromosome 1, using a human genetic map for recombination rates. We performed 500 simulations for each model, and all simulations started with West African ancestry contribution of 0.65. The estimated slope and intercept for the two methods of simulating global ancestry are highly similar. ANCOVA results suggest there is a significant effect of selection coefficient on global ancestry: F(1,997) = 1.0519 × 104, p < 2 × 10−6, but there is no significant effect of recombination rate and simulation model on global ancestry estimate after controlling for selection coefficient: F(1,997) = 1.6350 × 10−1, p = 0.686.
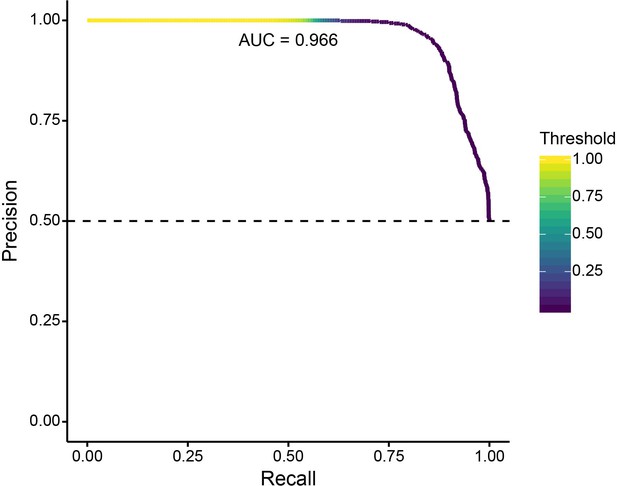
Precision-recall curve for validation of SWIF(r) classification of neutral and positively selected variants, using 1000 neutral and 1000 positive selection simulations.
With our ancestry-based measures, SWIF(r) achieved an area under the curve (AUC) of 0.966, where an AUC of 1 represents a classifier with perfect skill. Horizontal dashed line indicates the no-skill classifier for this data set.
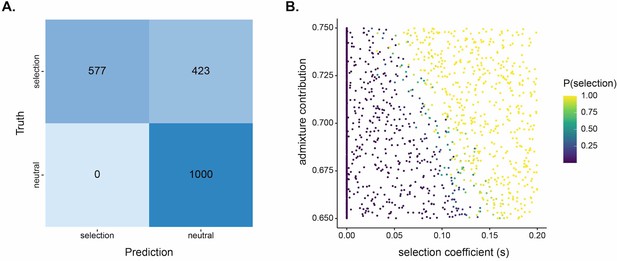
SWIF(r) classification results for 1000 neutral and 1000 positive selection simulations used for the test set based on Santiago’s demographic history.
(A) Confusion matrix with threshold P(selection)>0.5. There are no false positives in test set and a high rate of false negatives. (B) Scatterplot of initial admixture contribution vs selection coefficient. Points colored by P(selection) as estimated by SWIF(r). The majority of false-negative classifications (i.e. classifying selection scenarios as neutral) occurred with low starting admixture proportion or low selection strength. If we consider approximate Bayesian computation (ABC) estimates for admixture proportion (~0.7) and selection strength (~0.08), Duffy-null on Santiago would sit around the transition from high to low rate of false negatives. SWIF(r) returned high probability for Duffy-null, P(selection)>0.999.
Tables
Expected and observed Duffy-null allele frequencies for each island and source population.
Expected Duffy-null frequencies are approximated by mean West African global ancestry proportion for each island, calculated using the admixture software.
| Population | n (sampled individuals) | Expected frequency | Observed frequency | Binomial test p-value |
|---|---|---|---|---|
| Santiago | 172 | 0.737 | 0.834 | 2.193 ×10−5 |
| Fogo | 129 | 0.498 | 0.539 | 0.192 |
| NW Cluster | 236 | 0.552 | 0.557 | 0.817 |
| GWD | 107 | 0.997 | 1.000 | - |
| IBS | 107 | 0.002 | 0.019 | - |
Demographic models used for single-chromosome neutral simulations relevant to Cabo Verde demographic history.
| Initial population size (N) | Population growth model | Population growth rate (per generation) | Admixture type | Proportion of new migrants (per generation) | Scenario number |
|---|---|---|---|---|---|
| 1000 | Constant size | - | Single-pulse | - | 1 |
| Continuous | 0.01 | 2 | |||
| Exponential | 0.05 | Single-pulse | - | 3 | |
| Continuous | 0.01 | 4 | |||
| 10,000 | Constant size | - | Single-pulse | - | 5 |
| Continuous | 0.01 | 6 | |||
| Exponential | 0.05 | Single-pulse | - | 7 | |
| Continuous | 0.01 | 8 |
Additional files
-
Supplementary file 1
Chromosome 16:46582888–60359576 GO terms.
File containing ENSEMBL gene IDs and associated GO terms for the 10 genes that overlap with region showing extreme iDAT signatures.
- https://cdn.elifesciences.org/articles/63177/elife-63177-supp1-v2.zip
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/63177/elife-63177-transrepform-v2.docx





