Single-cell transcriptomes of developing and adult olfactory receptor neurons in Drosophila
Figures
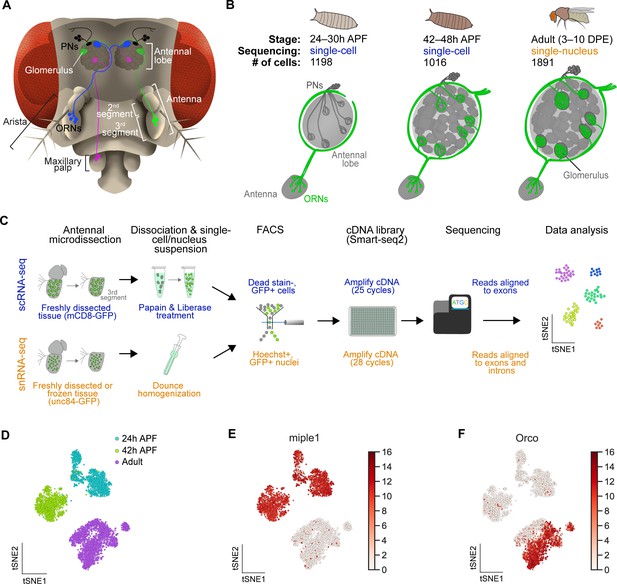
Single-cell transcriptomic profiling of Drosophila olfactory receptor neurons.
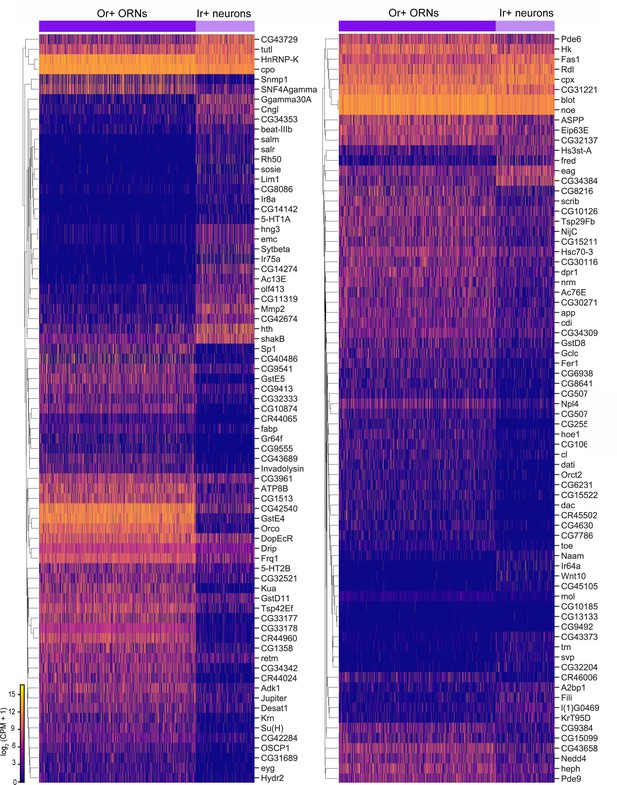
(A) Schematic of the Drosophila olfactory system. Three types of olfactory receptor neurons (ORNs) are portrayed in three different colors (green, magenta, and blue) and their cell bodies are housed in the antennae and maxillary palps. Axons of a given ORN type form synapses with dendrites of a given type of projection neuron (PN; black) in the antennal lobe of the brain in discrete glomeruli. (B) Diagram containing information about the olfactory circuit and sequencing performed at three stages: 24–30 hr APF (h APF; hours after puparium formation), 42–48 hr APF, and adults (3–10 DPE; days post-eclosion). Each ORN type expresses a unique olfactory receptor or a unique combination of receptors and sends its axons to a stereotyped glomerulus. 42 hr APF ORNs were previously sequenced in Li et al., 2020. (C) Diagram highlighting key differences in single-cell (blue) and single-nucleus (orange) RNA sequencing protocols. Cells are labeled with membrane-bound mCD8-GFP, whereas nuclei are marked with the nuclear envelop-bound unc84-GFP. (D) t-distributed stochastic neighbor embedding (t-SNE) plot depicting all sequenced ORNs. Dimensionality reduction was performed using the top 416 differentially expressed (DE) genes from ORNs at all stages. (E–F) Expression of genes found in developing ORNs (E) and a subset of mature ORNs (F). Heat map units, log2(CPM+1) (CPM: counts per million reads).
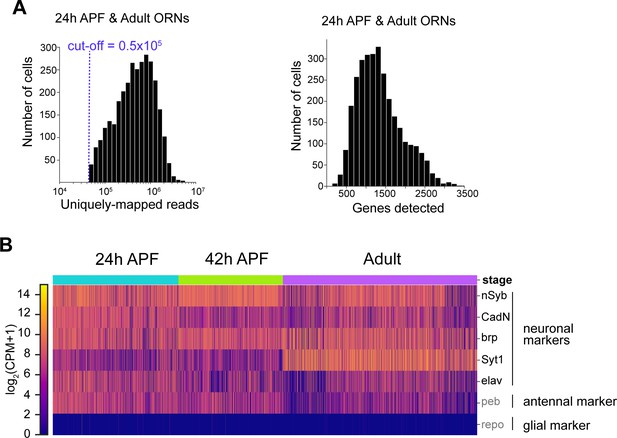
Filtering and quality control of sequenced ORNs.
(A) Distributions of uniquely mapped reads per cell (left) and number of detected genes per cell/nucleus (right). The blue dashed line marks the reads cut-off. (B) Heatmap depicting that ORNs analyzed express five pan-neuronal markers (nSyb, CadN, brp, Syt1, elav), an antennal cell marker (peb), but not a glial marker (repo). We filtered for high-quality ORNs by ensuring that they expressed two out of five neuronal markers (in black) at log2(CPM+1) ≥ 2 (CPM: counts per million reads).
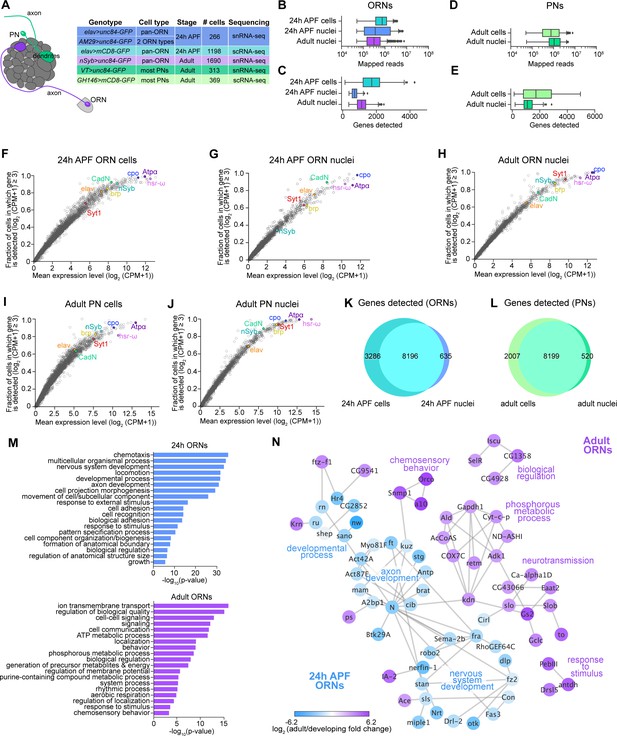
Comparison of single-cell and single-nucleus RNA-seq.
(A) Schematic depicting ORN axons and PN dendrites in the antennal lobe (left) and summary table of the cell types, cell numbers, genotypes, and stages that were profiled using snRNA-seq and scRNA-seq. Note that 24 hr APF cells were labeled by intersecting elav-GAL4 with ey-FLP and UAS-FRT-STOP-FRT-mCD8-GFP. The reason for fewer adult nuclei here than in Figure 1B is because some adult nuclei were labeled with lam-GFP, but those nuclei were not included in analyses here (see Figure 6—figure supplement 1F). (B–E) Box plots depicting uniquely-mapped reads (B, D) and detected genes (C, E) per cell/nucleus. Only exonic reads are shown for RNAs in cells, whereas exonic and intronic reads are combined for nuclear transcripts. (F–J) Mean expression level and detection rate of all genes in 24 hr APF ORN cells (F), 24 hr APF ORN nuclei (G), adult ORN nuclei (H), adult PN cells (I), and adult PN nuclei (J). Detection failure events can occur due to (1) the gene not being expressed in the cell; (2) gene dropouts resulting from technical artifacts even though mRNA transcripts are present; or (3) gene expression is below detection threshold. Colored circles represent the five neuronal markers [ embryonic lethal abnormal vision (elav); neuronal Synaptobrevin (nSyb); Synaptotagmin 1 (Syt1); Cadherin-N (CadN); bruchpilot (brp)] used for quality control filtering (Figure 1—figure supplement 1B) and three additional genes [couch potato (cpo); Heat shock RNA ω (hsr-ω); Na+ pump α subunit (ATPα)]. (K–L) Venn diagram comparing all genes detected using scRNA-seq and snRNA-seq in 24 hr APF ORNs (K) and in adult PNs (L). Expression was defined as log2(CPM +1) ≥ 2. (M) Gene Ontology (GO) analysis based on the top ~365 genes enriched in 24 hr ORN nuclei compared to adult ORN nuclei (top) and GO analysis of top ~365 genes enriched in adult ORN nuclei compared to 24 hr APF nuclei (bottom). Top 19 GO terms are shown. (N) STRING plot showing gene networks of 24 hr APF and adult ORN nuclei.
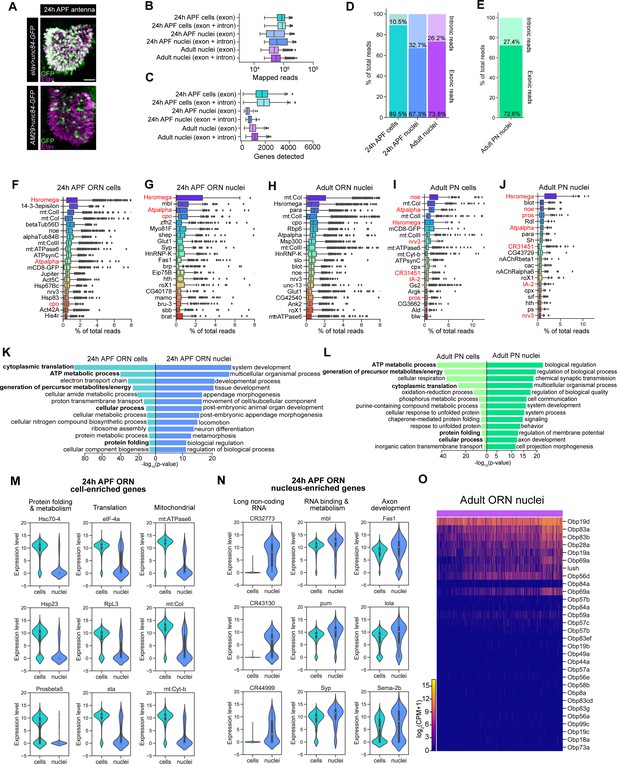
Additional comparisons of scRNA-seq and snRNA-seq.
(A) Confocal images showing expression of the nuclear envelop protein Unc84-GFP in the antenna driven by elav-GAL4 and AM29-GAL4; anti-GFP labeling is in green and anti-Elav (pan-neuronal nuclear marker) labeling is in magenta. Scale bar, 20 µm. (B–C) Box plots showing distribution of uniquely-mapped reads [average reads are, 24 hr APF cells (exon): 8.4 × 105; 24 hr APF cells (exon + intron): 9.4 × 105; 24 hr APF nuclei (exon): 7.4 × 105; 24 hr APF nuclei (exon + intron): 1.1 × 106; adult (exon): 3.6 × 105; adult (exon + intron): 5.0 × 105] (B) and genes detected [average genes detected are, 24 hr APF cells (exon): 1,706; 24 hr APF cells (exon + intron): 1,847; 24 hr APF nuclei (exon): 520; 24 hr APF nuclei (exon + intron): 714; adult (exon): 956; adult (exon + intron): 1,135] (C) per cell/nucleus in ORNs. These plots depict a breakdown of all cells/nuclei with exonic reads only and exonic and intronic reads combined. Analyses in Figure 2 and the rest of the study were performed on cells with exonic reads only and nuclei with combined exonic and intronic reads. (D–E) Stacked bar plots of proportion of exonic and intronic reads in scRNA-seq and snRNA-seq samples where reads were combined in ORNs (D) and PNs (E). (F–J) Top 20 transcripts with highest percentage of total reads in 24 hr APF ORN cells (F), 24 hr APF ORN nuclei (G), adult ORN nuclei (H), adult PN cells (I), and adult PN nuclei (J). Note that top transcripts in adult ORN nuclei contain mitochondrial (mt) transcripts, likely artifacts resulting from dissociation procedures. Transcripts shared between stage-matched cells and nuclei are in red. (K–L) Gene Ontology (GO) analysis based on genes enriched in 24 hr APF ORN cells compared to 24 hr APF ORN nuclei (K) and adult PN cells compared to adult PN nuclei (L). The top 13 significant GO terms are shown. Approximately500 DE genes were used for GO analysis in each category. Bolded terms in the cell categories are shared between ORN and PN cells. (M–N) Violin plots showing cell-enriched (M) and nucleus-enriched (N) transcripts in each category in 24 hr APF ORNs. Note that mitochondrial genes are more enriched in cells, whereas long non-coding RNAs are more enriched in nuclei. (O) Heatmap depicting Obp expression in adult ORN nuclei. The y-axis is arranged according to descending order of Obp expression levels from bulk RNA-sequencing from the antenna (Menuz et al., 2014). Highest detected Obps in Menuz et al. are expressed at levels 10–15 times higher than the highest detected chemosensory co-receptor, Orco. In situ expression analysis suggests that Obps are mostly expressed in supporting cells (Larter et al., 2016); thus, the correlation of Obp expression levels from previous study and the expression patterns shown here suggest that Obp expression seen here is likely due to contamination of highly abundant ambient RNAs associated with neuronal nuclei.
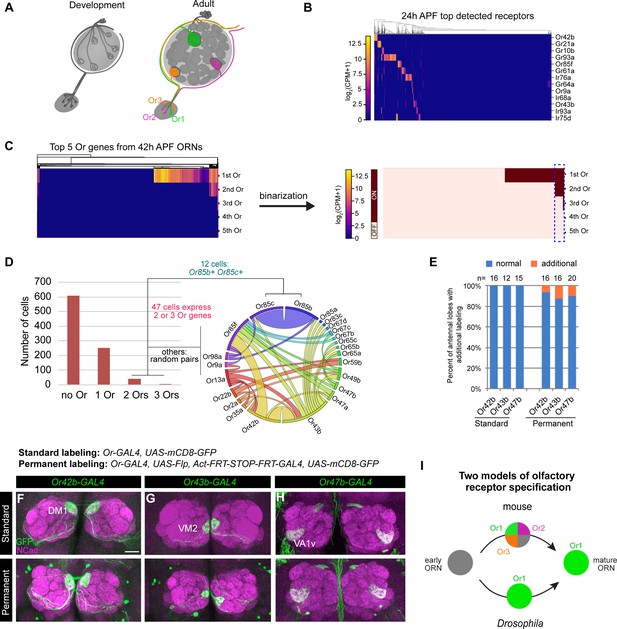
Individual ORNs maintain expression of the same olfactory receptor(s) throughout development.
(A) Schematic depicting ORNs at an early developmental stage (left) and in the mature circuit (right) where each ORN type expresses a distinct olfactory receptor represented by different colors. (B) Heatmap of the top detected [log2(CPM+1) ≥ 3 in > 5 cells] receptor genes in 24 hr APF ORNs. Co-receptors were excluded from this plot. (C) Heatmap (left) and binarization of heatmap (right) of the top five highest expressed Ors in 908 nSybGAL4>mCD8-GFP positive ORNs from 42 hr APF. Olfactory receptors were considered ‘on’ if they were expressed at log2(CPM+1) ≥ 3. Blue dashed box indicates cells that co-express more than one receptor. (D) Quantification of the number of cells expressing zero, one, two, or three Ors in 42 hr APF ORNs (left). Chord plot (right) of co-expressed receptors in the cells that expressed 2 or 3 Ors. Most co-expressed receptors show random patterns of expression. (E) Quantification of the percentage of antennal lobes that displayed labeling of normal (blue) and additional glomeruli (orange) in standard and permanent labeling conditions. In permanently labeled brains, 6.3% of Or42b-GAL4; 12.5% of Or43b-GAL4; and 10% of Or47b-GAL4 antennal lobes displayed additional labeling. (F–H) Confocal images of adult antennal lobes labeled with anti-GFP (green) and anti-NCad (magenta) in standard labeling (top row) or permanent labeling (bottom row) conditions of Or42b-GAL4-positive (F), Or43b-GAL4-positive (G), and Or47b-GAL4-positive (H) ORNs. Scale bar, 20 µm. (I) Schematic depicting olfactory receptor specification in mice and Drosophila. Mouse ORNs express multiple olfactory receptors before one predominates in mature ORNs (top; Hanchate et al., 2015; Tan et al., 2015), whereas our data indicate that Drosophila ORNs maintain expression of the same receptor throughout development (bottom).
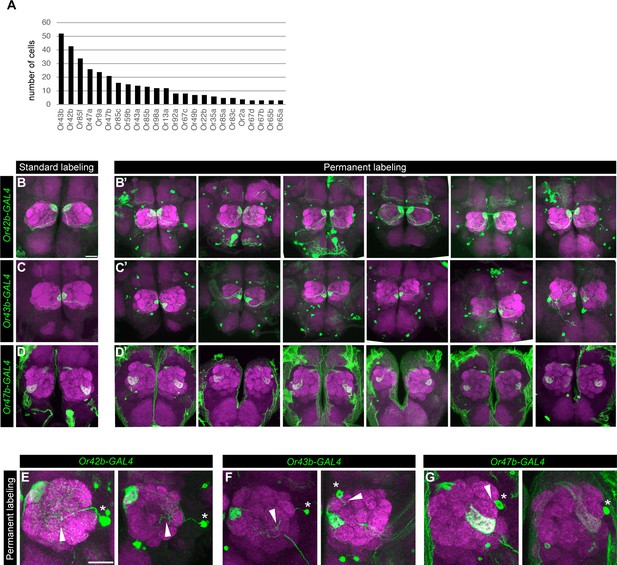
Permanent labeling reveals that Drosophila ORNs express the same receptor across all developmental stages.
(A) Quantification of number of cells expressing each Or in 42 hr APF ORNs. (B–D’) Additional confocal images of adult antennal lobes labeled with anti-GFP (green) and anti-NCad (magenta) in standard (B–D) and permanent labeling (B’–D’) of Or42b-GAL4-positive (B panels), Or43b-GAL4-positive (C panels), Or47b-GAL4-positive (D panels) ORNs. (E–G) Confocal images depicting non-ORN expression of Or42b-GAL4 (E); Or43b-GAL4 (F); Or47b-GAL4 (G) in permanently labeled antennal lobes. Asterisks denote the cell body of antennal lobe neurons and arrowheads point to the neurite(s) of these cells in the antennal lobe. Scale bar, 20 µm.
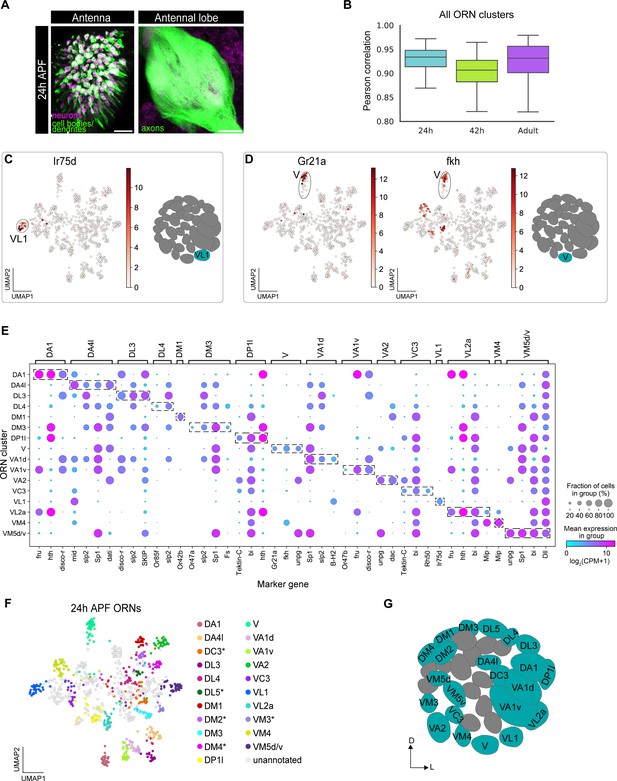
Mapping 24 hr APF transcriptomic clusters to their glomerular types.
(A) Confocal images of 24 hr APF ey-FLP;UAS-FRT-STOP-FRT-mCD8-GFP;elav-GAL4 ORNs in the antenna (left) labeled with anti-GFP (green) and anti-Elav (magenta) and antennal lobe (right) labeled with anti-GFP (green). Scale bar, 20 µm. (B) Quantification of cluster-level transcriptomic similarity of all ORN clusters annotated in this study and in Li et al., 2020 using Pearson’s correlation. 416 differentially expressed genes identified from all three stages were used for this analysis. (C) Uniform manifold approximation and projection (UMAP) plot depicting Ir75d expression in a single cluster (left). Ir75d-positive ORNs target their axons to the VL1 glomerulus as shown in the schematic (right). (D) UMAP plots (left) depicting Gr21a expression in a single cluster and fkh expression in the same cluster. Gr21a-positive ORNs target their axons to the V glomerulus and fkh is a gene expressed in V ORNs at 24 hr and 42 hr APF. Schematic of the glomerular projection of V ORNs (right). (E) Dot plot summarizing the markers used to map 16 transcriptomic clusters at 24 hr APF to their corresponding 42 hr APF clusters. Dashed boxes around dots highlight markers used to map 24 hr APF clusters to 42 hr APF ones. (F) UMAP plot depicting the 21 transcriptomic clusters that were decoded at 24 hr APF. Asterisk next to ORN type indicates that it was annotated using data presented in Figure 6. Note that VM5d/v could not be separated (see Figure 4—figure supplement 2C). (G) Schematic depicting the glomerular targeting patterns of the 22 ORN types mapped in E–F and Figure 6. Heatmap units, log2(CPM+1). For all UMAP plots: dimensionality reduction was performed using differentially expressed genes from 42 hr APF combined with sensory receptors for a total of 335 genes.
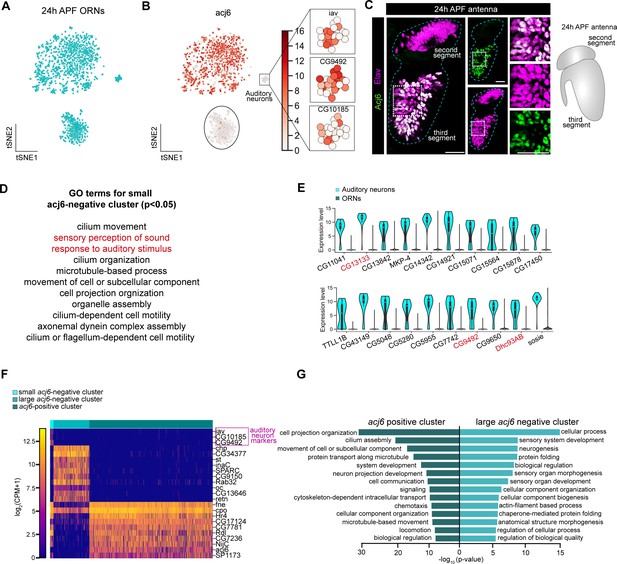
Further analysis of 24 hr APF transcriptomic clusters.
(A) t-SNE plot showing that ORNs cluster into two large and one small population(s) of cells when dimensionality reduction was performed using the top 500 over-dispersed genes. (B) t-SNE plot of acj6 in 24 hr APF ORNs (left). Expression of three auditory neuron markers iav, CG9492, CG10185 (right) in the small acj6-negative cluster. These markers are essentially absent from the two larger clusters in this plot. (C) Confocal images (left) of 24 hr APF antenna co-labeled with anti-Acj6 (green) and anti-Elav (magenta). Some cells are Elav-positive but Acj6-negative in the third antennal segment (zoomed-in panels to the right). White squares denote the location where zoomed in panels were obtained. The second antennal segment does not express Acj6. Schematic of 24 hr APF antennal anatomy (far right). Scale bars, 20 µm. (D) List of GO terms enriched in the small acj6-negative cluster derived from the top 110 highest expressed genes in that cluster. Red terms are auditory organ-specific. (E) Violin plots of the top auditory-neuron enriched genes. Genes in red have known expression or function in auditory organs. (F) Heatmap depicting auditory neuron markers (top three genes; marked with magenta); top 10 DE genes between acj6+ and large acj6– cluster (middle 10 genes), and the top 10 highest expressed genes in the acj6+ cluster compared to the large acj6– cluster (last 10 genes). (G) Gene Ontology (GO) analysis based on >500 differentially expressed genes between acj6+ and the large acj6– cluster. Top 14 significant GO terms are shown.
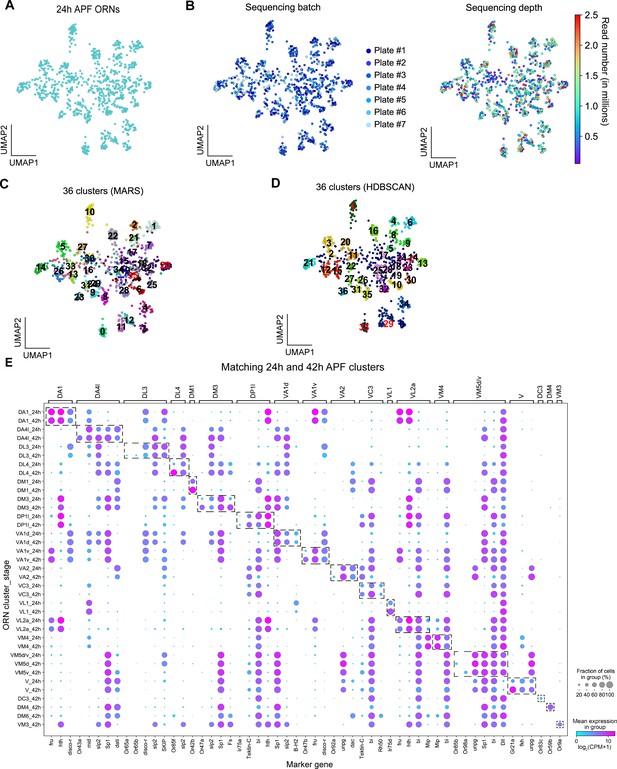
Quality control and additional analysis of 24 hr APF transcriptomes of Acj6-positive ORNs.
(A) UMAP plot depicting 24 hr APF ORNs. (B) UMAP plots depicting 24 hr APF ORNs color-coded by sequencing batch (plate #; left) and colored with a heat map depicting read number (right). (C–D) UMAP plots comparing MARS clustering (C) with HDBSCAN density-based clustering (D) approaches. MARS is better able to separate VA1v and VA1d [clusters # 11 and 12] in (C) and cluster # 29 in (D). (E) Dot plot showing expression of the markers used to match 24 hr APF ORNs to 42 hr APF ORNs. Some 42 hr APF clusters express sensory receptors that are not found in 24 hr APF clusters. Dashed boxes highlight the markers for each cluster. Note this plot also depicts 42 hr APF DC3, DM4, and VM3 clusters as they were decoded in Li et al., 2020 but we were not able to annotate these clusters until Figure 6.
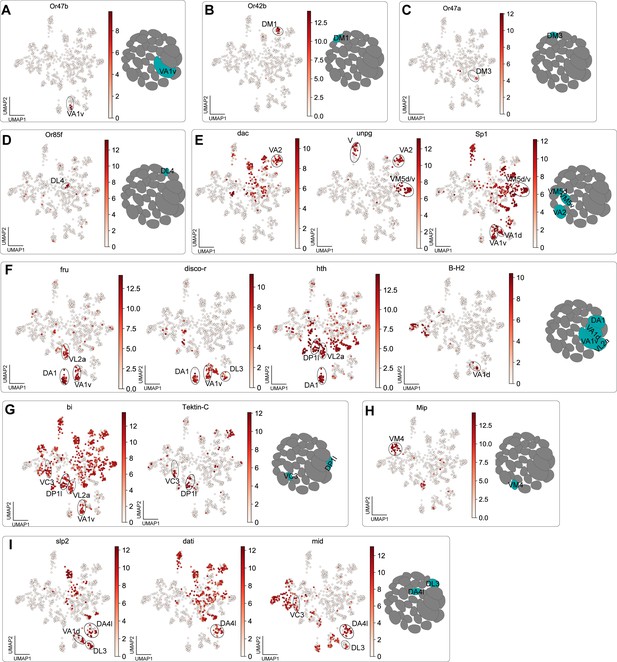
Markers used to match 24 hr APF ORNs to their glomerular type.
(A–I) In each panel, a UMAP plot showing the gene(s) used to annotate clusters is next to a schematic depicting the glomerular target(s) of annotated ORN type(s). Heatmap units, log2(CPM + 1).
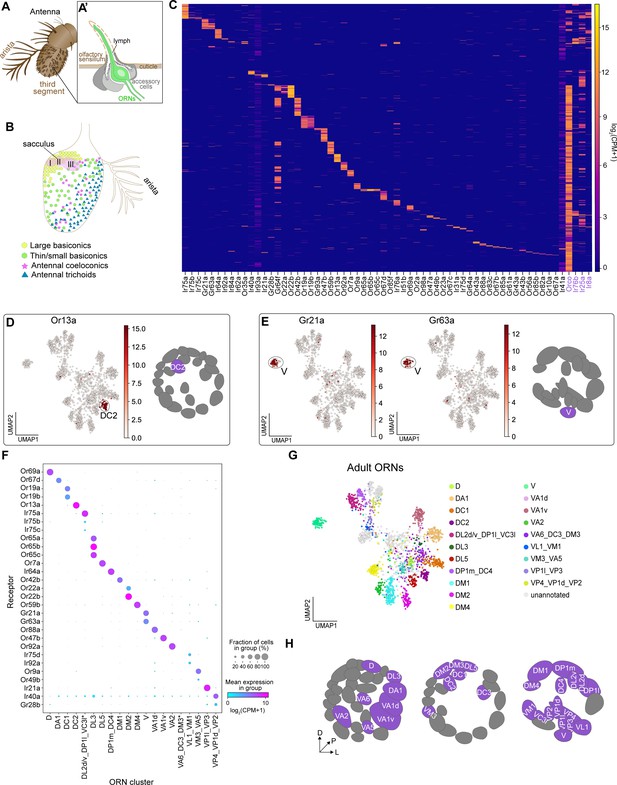
Mapping adult transcriptomic clusters to their glomerular types.
(A–A’) Diagram of the adult antenna (A) and the structure of an individual sensillum (A’) where ORN dendrites and support cells are located. (B) Schematic of the antenna depicting the spatial distribution of sensillar classes and the location of the sacculus and arista. (C) Heatmap showing the expression of all detected sensory receptors belonging to the Or (odorant receptor), Gr (gustatory receptor), and Ir (ionotropic receptor) families. A detected receptor is expressed at a level of log2(CPM+1) ≥3 in > 5 cells. Note low level expression of Ir41a and Ir93a in many cells may be the result of increased read numbers generated by their large genomic regions. Or and Ir co-receptors are in purple. Cells were ordered by hierarchical clustering. (D–E) UMAP plots depicting Or13a expression in a single cluster (left) and glomerular projection of Or13a-positive DC2 ORNs (right) (D) and Gr21a and Gr63a expression in a single cluster (left) and glomerular projection of Gr21a/63a-positive V ORNs (E). (F) Dot plot depicting sensory receptors used to annotate adult neuron clusters. Clusters that contained multiple neuron types are indicated by an underscore separating the names of each type. Asterisk next to VA6_DC3_DM3 and DP1l_DL2d/v_VC3l clusters indicates that some or all of receptors for these cell types are expressed in a small non-overlapping percentage of cells and can be better resolved on a UMAP plot (see Figure 5—figure supplement 2K, M). (G) UMAP plot of 20 adult clusters annotated to 31 neuron types using sensory receptor expression. Note that some clusters contained receptors of multiple ORN types and are labeled as such. (H) Schematic of the glomerular targets of the 31 neuron types mapped in F and G. For all UMAP plots, dimensionality reduction was performed using differentially expressed genes from 42 hr APF combined with sensory receptors for a total of 335 genes. Heatmap units, log2(CPM + 1).
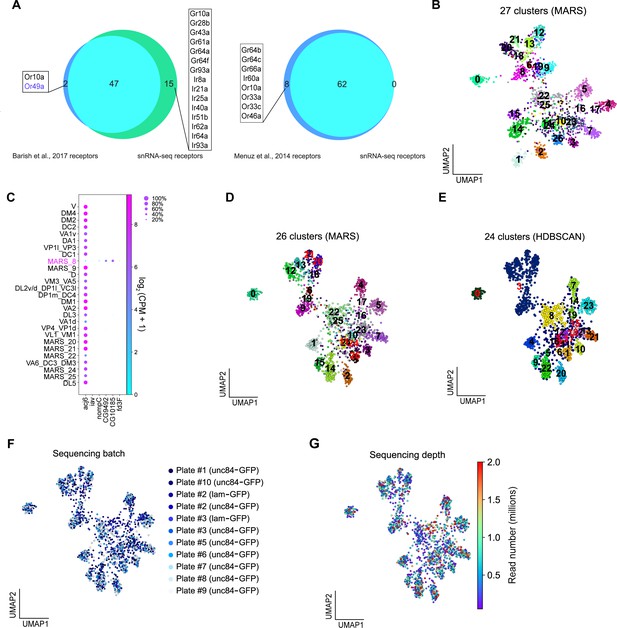
Quality control and additional analysis of adult transcriptomes.
(A) Venn diagrams comparing sensory receptor genes detected in snRNA-seq [log2(CPM+1) ≥ 3 in > 5 cells] and bulk antennal RNA-seq studies in Barish et al., 2017 (left) and Menuz et al., 2014 (right). Or49a is in blue because its detection was low in the bulk RNA-seq study. snRNA-seq and bulk RNA-seq detect largely similar receptors but snRNA-seq missed the expression of Or10a reported in both bulk sequencing studies. (B–C) UMAP plot of 27 clusters of adult ORNs (B); note that MARS_8 was removed from subsequent analysis because it was found to express auditory neuron genes and not the ORN gene, acj6, shown in the dot plot (C). (D–E) UMAP plots comparing MARS (D) with HDBSCAN density-based clustering (E). Both algorithms yield mostly concordant results for Or-expressing clusters; however, MARS predictions are better able to separate Ir-expressing clusters [compare top clusters] in (D) with the same clusters in (E). (F–G) UMAP plots depicting adult transcriptomes color coded by sequencing batch (plate #; F) and sequencing depth (G). Note that we included some adult nuclei labeled with lam-GFP to increase cell type representation in the adult data.
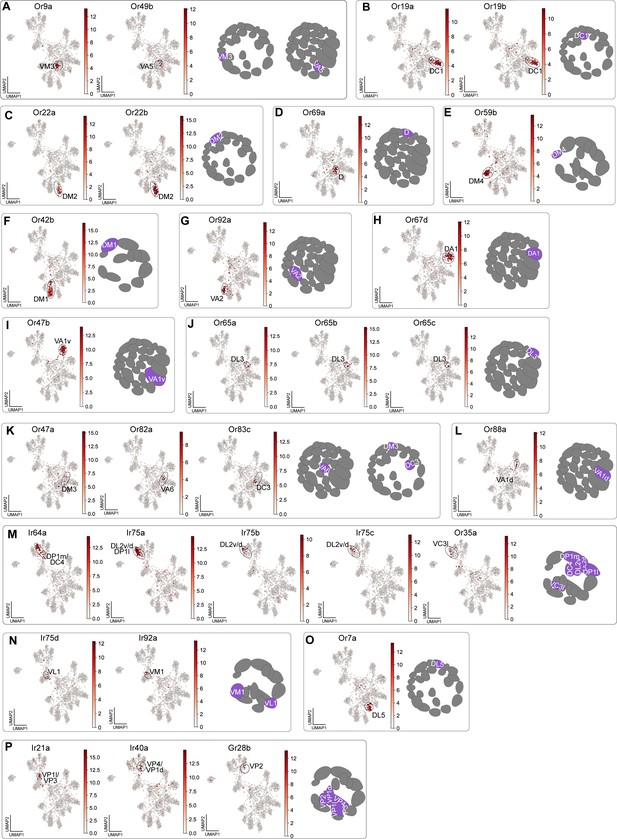
Cluster-specific sensory receptor expression in adult transcriptomes.
(A–P) UMAP plots of sensory receptors used to annotate adult sensory neuron clusters. In each panel, a UMAP plot showing the gene(s) used to annotate clusters is placed next to a schematic showing the glomerular projections of annotated neuron types. Heatmap units, log2(CPM + 1).
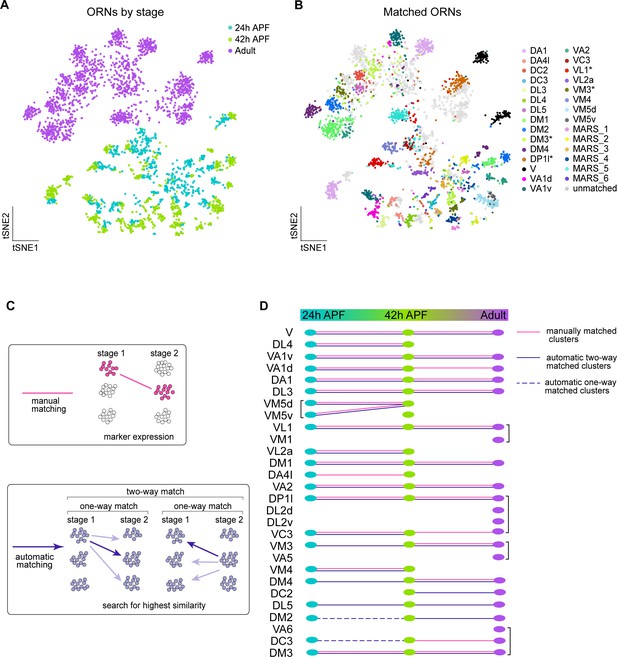
Matching transcriptomic clusters across all stages enabled annotation of additional ORN types.
(A–B) t-SNE plots of ORNs from 24 hr APF, 42 hr APF, and adult stages (A) and matched ORNs across these stages (B). Clusters marked 'MARS_#' are matched but their glomerular identity is unknown. Asterisk next to cluster name indicates that at the adult stage the cluster contained multiple ORN types, but it was annotated with the name of the developmental cluster it mapped to. Dimensionality reduction was performed using differentially expressed genes from 42 hr APF combined with olfactory receptors for a total of 335 genes. (C) Schematic summarizing the two approaches used for matching ORNs from different stages: (1) manual matching using cluster-specific markers/olfactory receptors (top); and (2) automatic matching by calculating the transcriptome similarity between one cluster at a certain stage to all clusters from another stage (bottom). (D) Summary of all matched ORN types from three different stages, the method used to match clusters is represented by the type of line (color, solid, dashed) linking the clusters between timepoints.
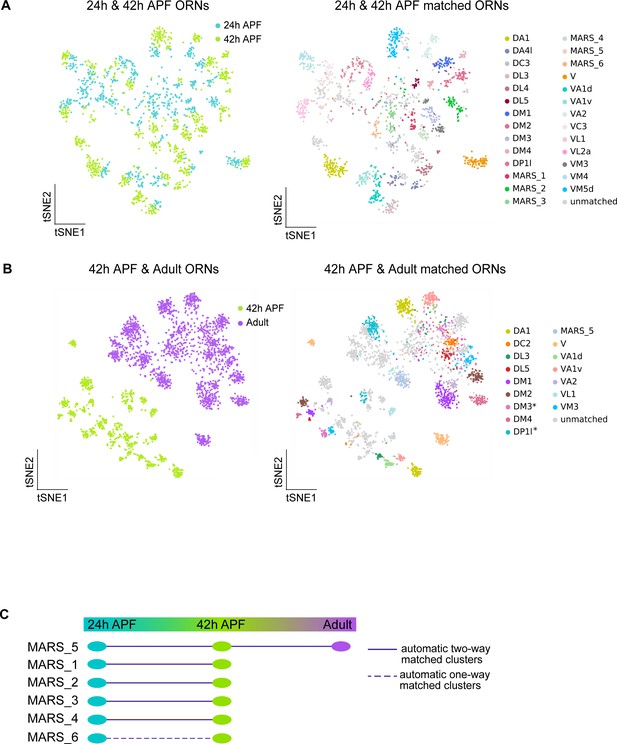
Additional evidence for cluster matching across developmental stages.
(A) t-SNE plots depicting 24 hr APF and 42 hr APF ORNs (left) and matched clusters between these two timepoints (right). 'MARS_#' are matched between both timepoints but their glomerular identity is unknown. (B) t-SNE plots depicting 42 hr APF and adult ORNs (left) and matched clusters between these two time points (right). Dimensionality reduction was performed using differentially expressed genes from 42 hr APF combined with olfactory receptors for a total of 335 genes. Asterisk next to cluster name indicates that adult clusters contain multiple ORN types. (C) Summary of matched ORN types where the glomerular identity of the clusters is unknown.
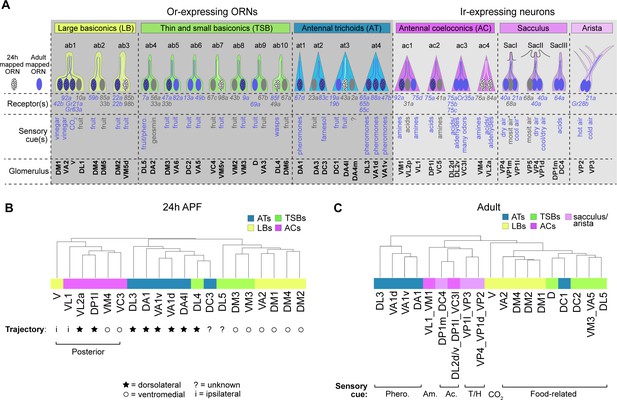
ORN transcriptomes in part reflect axon trajectory at 24 hr APF and sensory modality in adults.
(A) Schematic summary of ORNs, thermo- and hygro-sensory neurons within their respective sensillar class/structure, the sensory receptors each neuron type expresses, and the category of cues each type predominantly responds to. White dotted ORNs were annotated at 24 hr APF, blue ORNs were annotated in adults, and blue dotted ORNs are annotated at both stages. Receptors in blue were detected in either 24 hr APF or adult datasets. Sensory cues marked with asterisks indicate that they are putative stimuli for that neuron type. See references below for information regarding the correspondence between receptor expression and glomerular projection (Couto et al., 2005; Fishilevich and Vosshall, 2005; Silbering et al., 2011; Grabe et al., 2016; Marin et al., 2020) and regarding sensory stimuli detected by specific types (Yao et al., 2005; Hallem and Carlson, 2006; Kurtovic et al., 2007; Kwon et al., 2007; Ray et al., 2007; Semmelhack and Wang, 2009; Gallio et al., 2011; Silbering et al., 2011; Ni et al., 2013; Ronderos et al., 2014 Ebrahim et al., 2015; Lin et al., 2015 ; Enjin et al., 2016; Dweck et al., 2013; Dweck et al., 2018; Budelli et al., 2019). (B–C) Dendrograms of 24 hr APF (B) and adult (C) sensory neuron types that resulted from hierarchical clustering of transcriptomes based on differentially expressed genes at each stage. Axon trajectory/projection information is from Endo et al., 2007; Silbering et al., 2011; Joo et al., 2013; Rybak et al., 2016. The following abbreviations were used to describe the stimuli adult neurons respond to: Phero. = pheromones; Am = amines; Ac = acids; and T/H. = temperature and humidity. Some adult clusters contain multiple neuron types, but these clusters are within the same sensillar class or detect the same cues.
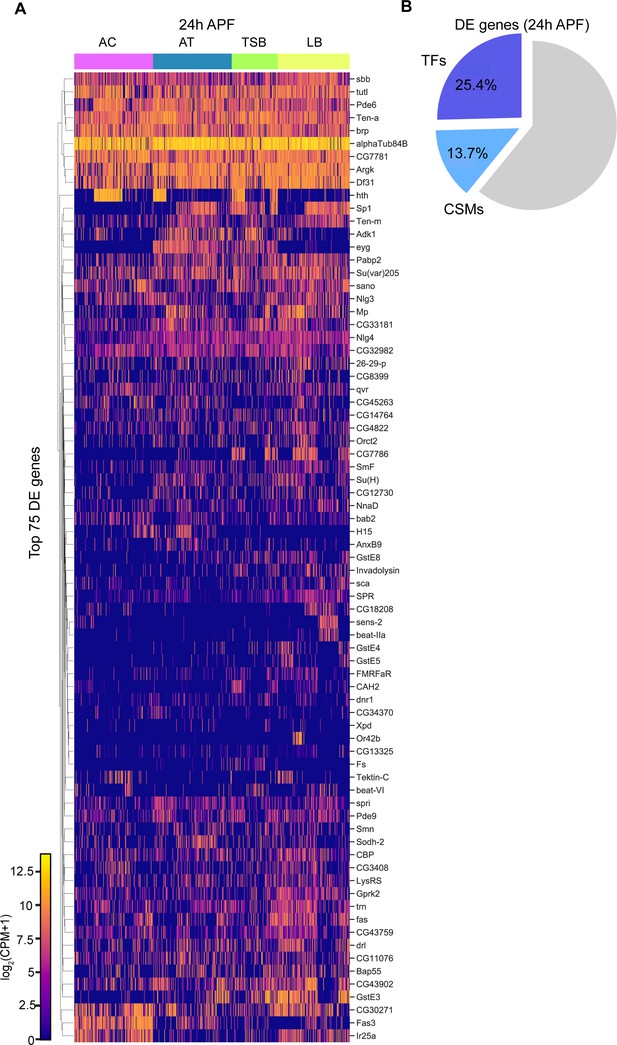
Comparisons of 24 hr APF ORNs by sensillar class.
(A) Heatmap of top 75 differentially expressed genes in 24 hr APF ORNs grouped by sensillar class. (B) Quantification of the percentage of TFs and CSMs in the 437 differentially expressed (DE) genes used to cluster 24 hr APF transcriptomes in Figure 7B.
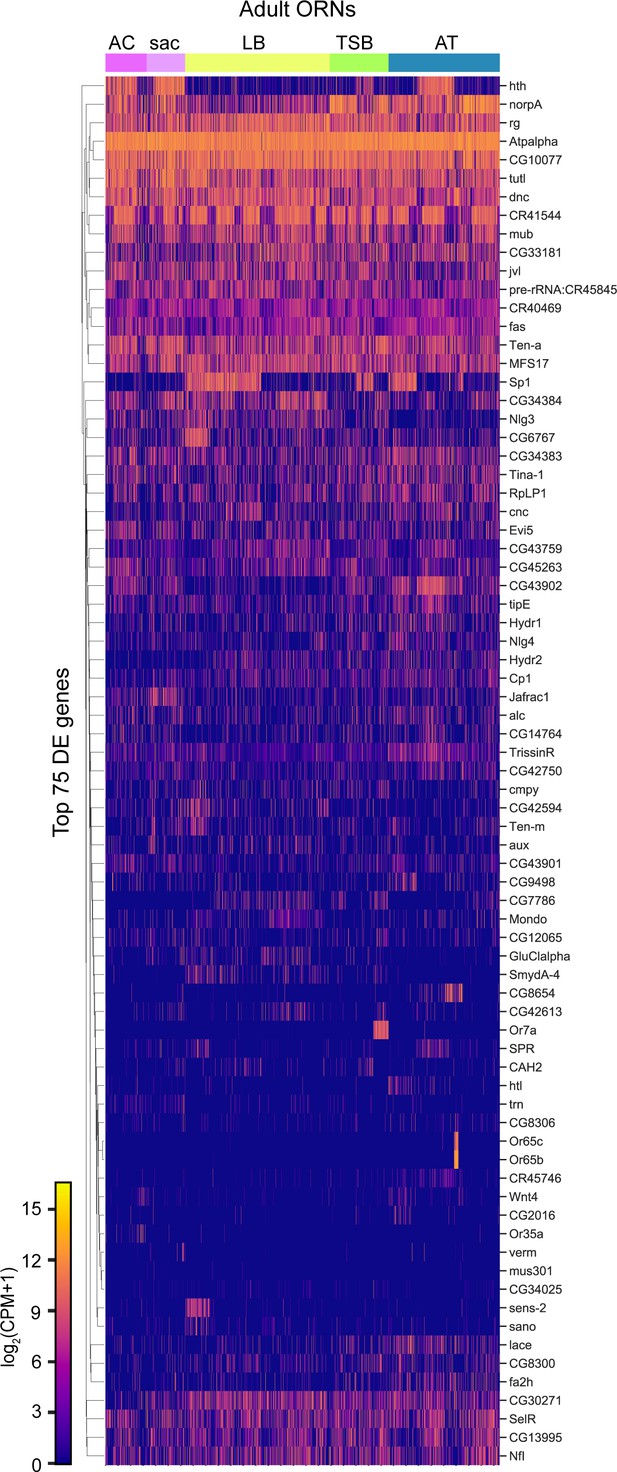
Comparisons of adult ORNs by sensillar class and sensory structure.
Heatmap of the top 75 differentially expressed genes in adult ORNs grouped by sensory structure.
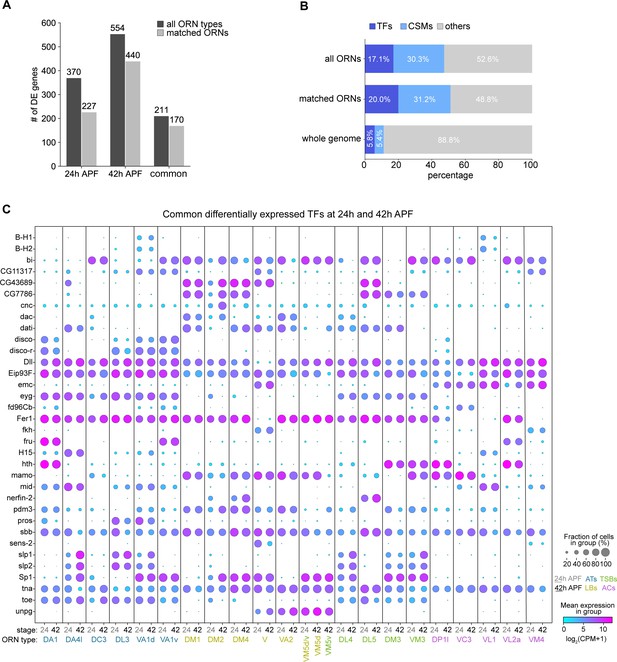
Cell-type-specific expression of transcription factors (TFs) and cell-surface molecules (CSMs) in developing ORNs.
(A) Quantification of differentially expressed (DE) genes identified in all ORN clusters (dark gray) or in clusters matched between 24 and 42 hr APF (light gray) in each pupal stage. (B) Percentage of TFs or CSMs from the common list of DE genes from 24 and 42 hr APF ORNs compared to the genome-wide percentage. (C) Dot plot depicting expression of the 34 common DE TFs from both 24 hr APF and 42 hr APF stages.

Common differentially expressed CSMs in 24 hr and 42 hr APF ORNs.
Dot plot depicting the expression of the 53 differentially expressed CSMs that are shared between 24 and 42 hr APF ORNs.

All differentially expressed CSMs in 24 hr and 42 hr APF ORNs, Part I.
Dot plot depicting the expression of the first half (58) of all of the CSMs DE from either 24 hr or 42 hr APF.

All differentially expressed CSMs in 24 hr and 42 hr APF ORNs, Part II.
Dot plot depicting the expression of the second half (57) of all of the DE CSMs from either 24 hr or 42 hr APF.

All differentially expressed TFs in 24 hr or 42 hr APF ORNs.
Dot plot depicting the expression of all 58 DE TFs from either 24 hr or 42 hr APF.
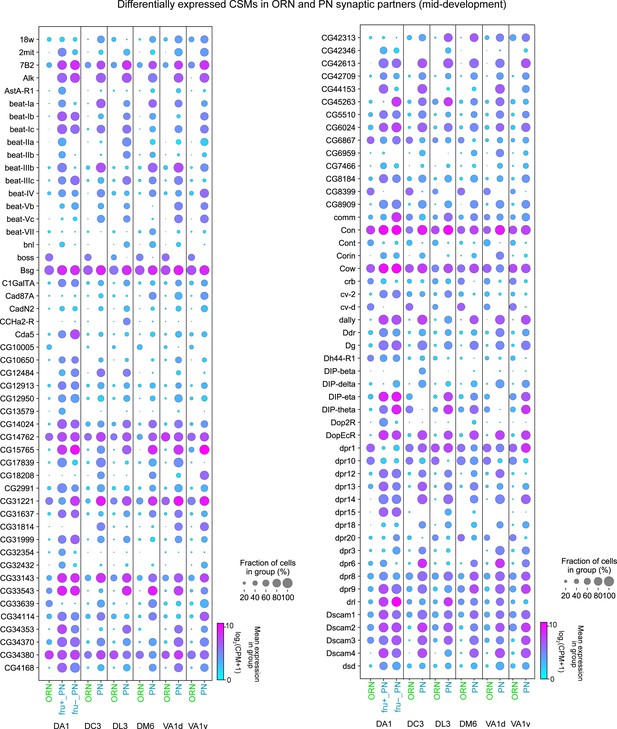
CSM expression in synaptic partner ORNs and PNs, Part I.
Dot plots depicting the expression of the first half (101) of the DE CSMs from 42 hr APF ORN and 48 hr APF PN synaptic partners.

CSM expression in synaptic partner ORNs and PNs, Part II.
Dot plots depicting the expression of the second half (101) of the DE CSMs from 42 hr APF ORN and 48 hr APF PN synaptic partners.
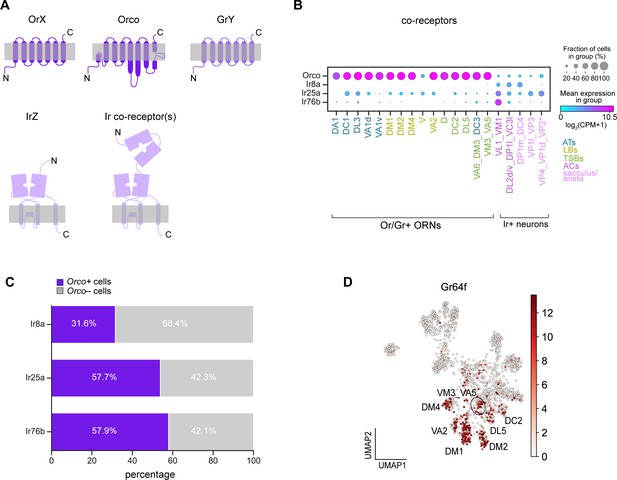
Broad co-receptor expression in adult ORNs.
(A) Schematic of three classes of Drosophila sensory receptors. OrX (odorant receptors) and their co-receptor Orco are seven-pass transmembrane receptors, as are GrY (gustatory receptors) (top). IrZ (ionotropic receptors) and their co-receptors are relatives of ionotropic glutamate receptors. Orco diagram is based on Butterwick et al., 2018 and Ir/Ir co-receptor diagram is based on Abuin et al., 2019. (B) Dot plot depicting Or and Ir co-receptor expression in adult clusters. Clusters containing multiple neuron types are noted using an underscore to separate names of each type. Asterisk next to VP2 ORNs indicates that these cells express a Gr and not an Ir. We observe Orco co-expression in many of the same ORN types as Task et al., 2020; however, although Orco expression is substantially lower than that of Obps, we could not fully rule out the possibility that Orco expression in a subset of cells was due to ambient contamination as was likely the case of Obps (Figure 2—figure supplement 1). (C) Stacked bar graph of the percentage of cells that express each Ir co-receptor that also express Orco, with a stringent expression cut-off of log2(CPM+1)≥5 for all co-receptors (based on Figure 9—figure supplement 1F–H). (D) UMAP plot of Gr64f expression in adult ORNs.
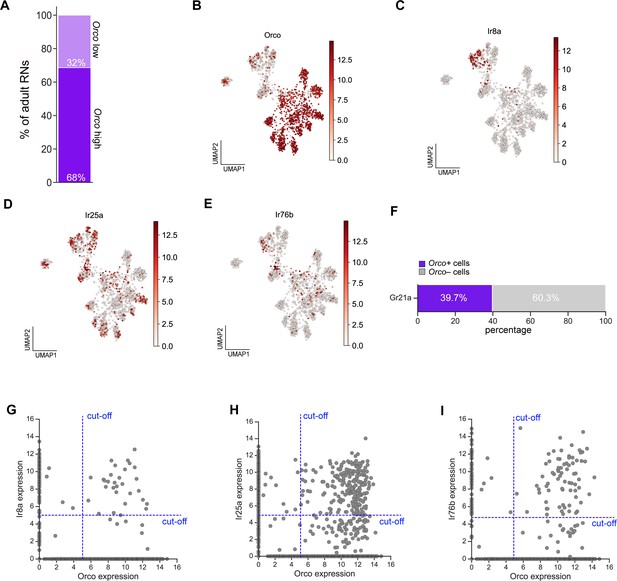
Expression of Orco and Ir co-receptors in adult ORNs.
(A) Stacked bar plot quantifying the percentage of adult antennal ORNs that are Orco high (e.g. express Orco at log2(CPM+1)≥5) and Orco low. (B–E) UMAP plots of Orco (B), Ir8a (C), Ir25a (D), and Ir76b (E) expression in adult ORNs. (F) Stacked bar graph of the percentage of Gr21a-positive cells that also express Orco. (G–I) Scatter plots depicting cells that co-express Orco and Ir8a (G), Ir25a (H), and Ir76b (I) and the reads cut-off used to generate graph in Figure 9C.
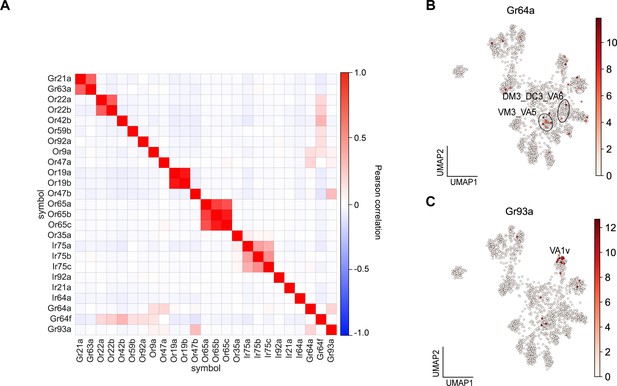
Expression of some gustatory receptors is correlated with the expression of a subset of olfactory receptors.
(A) Correlation matrix of the subset sensory receptors used to annotate transcriptomic clusters. (B–C) UMAP plots depicting expression of Gr64a (B) and Gr93a (C) in adult ORNs.
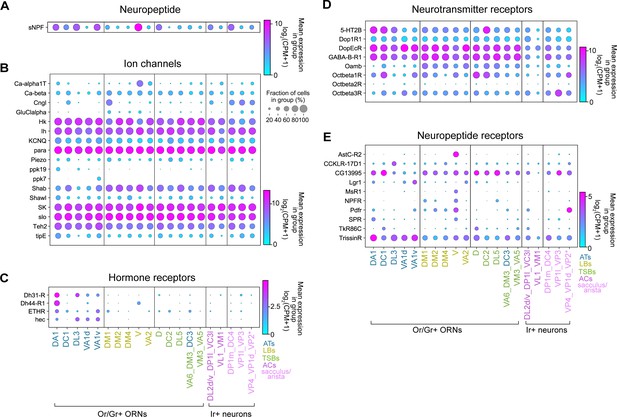
Differential expression of genes involved in neuronal communication in distinct ORN types.
(A–E) Dot plots depicting differential expression of a neuropeptide (A), ion channels (B), hormone receptors (C), small-molecule neurotransmitter receptors (D), and neuropeptide receptors (E) in annotated adult ORN clusters. Asterisk next to VP2 ORNs indicates that these cells express a Gr and not an Ir.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (D. melanogaster) | nSyb-GAL4 | n/a | RRID:BDSC_51635 | n/a |
| Genetic reagent (D. melanogaster) | AM29-GAL4 | Endo et al., 2007 | n/a | n/a |
| Genetic reagent (D. melanogaster) | elav-GAL4 | Luo et al., 1994 | n/a | n/a |
| Genetic reagent (D. melanogaster) | VT033006-GAL4 | Tirian and Dickson, 2017 | RRID:BDSC_73333 | n/a |
| Genetic reagent (D. melanogaster) | UAS-FRTSTOP-FRTmCD8GFP | Potter et al., 2010 | RRID:BDSC_30125 | n/a |
| Genetic reagent (D. melanogaster) | GH146-GAL4 | Stocker et al., 1997 | RRID:BDSC_30026 | n/a |
| Genetic reagent (D. melanogaster) | Act-FRT-STOPFRT-GAL4 | Pignoni and Zipursky, 1997 | n/a | n/a |
| Genetic reagent (D. melanogaster) | UAS-Flp | Duffy et al., 1998 | n/a | n/a |
| Genetic reagent (D. melanogaster) | ey-FLP | Chotard et al., 2005 | n/a | n/a |
| Genetic reagent (D. melanogaster) | Or42b-GAL4 | n/a | RRID:BDSC_9972 | n/a |
| Genetic reagent (D. melanogaster) | Or43b-GAL4 | n/a | RRID:BDSC_23895 | n/a |
| Genetic reagent (D. melanogaster) | Or47b-GAL4 | n/a | RRID:BDSC_9984 | n/a |
| Genetic reagent (D. melanogaster) | UAS-lam-GFP | n/a | BDSC_97378 | n/a |
| Genetic reagent (D. melanogaster) | UAS-unc84-GFP | Henry et al., 2012 | n/a | n/a |
| Antibody | Rat anti-Ncad (rat monoclonal) | Developmental Studies Hybridoma Bank | RRID:AB_528121 | (1:40 in 5% normal goat serum) |
| Antibody | Chicken anti-GFP (Chicken polyclonal) | Aves Labs | RRID:AB_10000240 | (1:1000 in 5% normal goat serum) |
| Antibody | Mouse anti-Acj6 (mouse monoclonal) | Developmental Studies Hybridoma Bank | RRID:AB_528067 | (1:40 in 5% normal goat serum) |
| Antibody | Rat anti-Elav (rat monoclonal) | Developmental Studies Hybridoma Bank | RRID:AB_528218 | (1:100 in 5% normal goat serum) |
| Antibody | Mouse anti-Elav (mouse monoclonal) | Developmental Studies Hybridoma Bank | RRID:AB_528217 | (1:100 in 5% normal goat serum) |
| Software, algorithm | ZEN | Carl Zeiss | RRID:SCR_013672 | n/a |
| Software, algorithm | Fiji | National Institutes of Health | RRID:SCR_002285 | n/a |
| Software, algorithm | Illustrator | Adobe | RRID:SCR_010279 | n/a |
| Software, algorithm | STAR 2.5.4 | Dobin et al., 2013 | RRID:SCR_015899 | https://github.com/alexdobin/STAR |
| Software, algorithm | Htseq 0.11.2 | Anders et al., 2015 | RRID:SCR_005514 | https://github.com/htseq/htseq |
| Software, algorithm | MARS | Brbić et al., 2020 | n/a | https://github.com/snap-stanford/mars |
| Software, algorithm | Scanpy | Wolf et al., 2018 | RRID:SCR_018139 | https://scanpy.readthedocs.io/en/stable |
| Software, algorithm | HDBSCAN | McInnes et al., 2018 | n/a | https://hdbscan.readthedocs.io/en/latest/how_hdbscan_works.html |
| Software, algorithm | Flymine | n/a | RRID:SCR_002694 | http://www.flymine.org |
| Software, algorithm | Revigo | n/a | RRID:SCR_005825 | http://revigo.irb.hr |
| Software, algorithm | SciPy | n/a | RRID:SCR_008058 | http://www.scipy.org |
| Software, algorithm | NumPy | n/a | RRID:SCR_008633 | http://www.numpy.org |
| Software, algorithm | Seaborn | n/a | RRID:SCR_018132 | https://seaborn.pydata.org |
| Software, algorithm | Pandas | n/a | RRID:SCR_018214 | https://pandas.pydata.org |
| Software, algorithm | Scikit learn | n/a | n/a | https://scikit-learn.org/stable |
| Software, algorithm | STRING | n/a | RRID:SCR_005223 | http://string.embl.de |
| Software, algorithm | Cytoscape v3.7.1 | n/a | RRID:SCR_003032 | http://cytoscape.org |