Dichloroacetate reverses sepsis-induced hepatic metabolic dysfunction
Figures
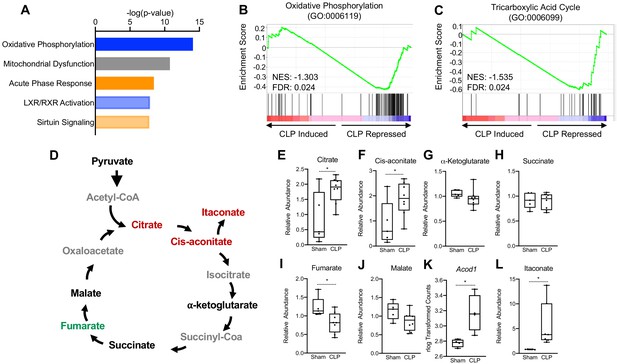
Sepsis Impairs hepatic mitochondrial metabolism.
(A) Top five canonical pathways subject to transcriptional alterations in the liver identified by ingenuity pathway analysis of RNA-seq of sham versus CLP mice (n = 4 mice per group). Blue represents a negative z-score and orange represents a positive z-score. Shading indicates intensity of pathway activation/inhibition. (B) GSEA of the oxidative phosphorylation pathway (GO:0006119) indicating a negative normalized enrichment score (NES = −0.853). (C) GSEA of the TCA cycle pathway (GO:0006099) indicating a negative enrichment score (NES = −1.1956). (D) Schematic representation of hepatic TCA cycle metabolites altered during chronic sepsis. Red denotes a metabolite increased in response to sepsis; green indicates a metabolite decreased in response to sepsis; black indicates a metabolite unchanged in response to sepsis; gray indicates a metabolite not measured in our metabolomic screening. (E–J) Relative metabolite levels measured by ultrahigh-performance liquid chromatography–tandem mass spectroscopy (UPLC–MS/MS) from livers of sham and CLP mice 30 hr post-surgery (n = 5 sham; 8 CLP). (K) rlog transformed counts from RNA-seq of sham and CLP mice 30 hr post-surgery. (L) Relative itaconate levels in livers of sham and CLP mice measured by UPLC–MS/MS (n = 5 sham; 8 CLP). *p<0.05 as determined by Student’s T-test.
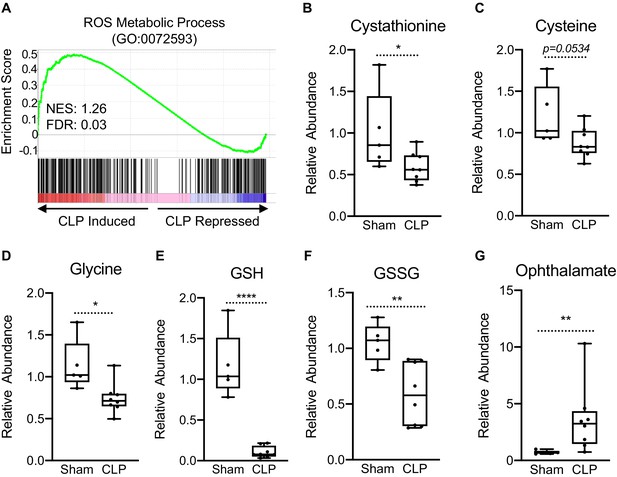
Impaired hepatic redox balance in septic mice.
(A) GSEA of the ROS metabolic pathway (GO:0072593) from RNA-seq from livers comparing sham to CLP showing a positive enrichment score (NES = 1.117) (n = 4 mice per group). (B–F) Relative metabolite levels of metabolites involved in redox balance measured by UPLC–MS/MS (n = 5 sham; 8 CLP). (G) Relative levels of ophthalmate involved in redox balance measured by UPLC–MS/MS. *p<0.05, **p<0.01, ****p<0.0001 as determined by Student’s T-test.
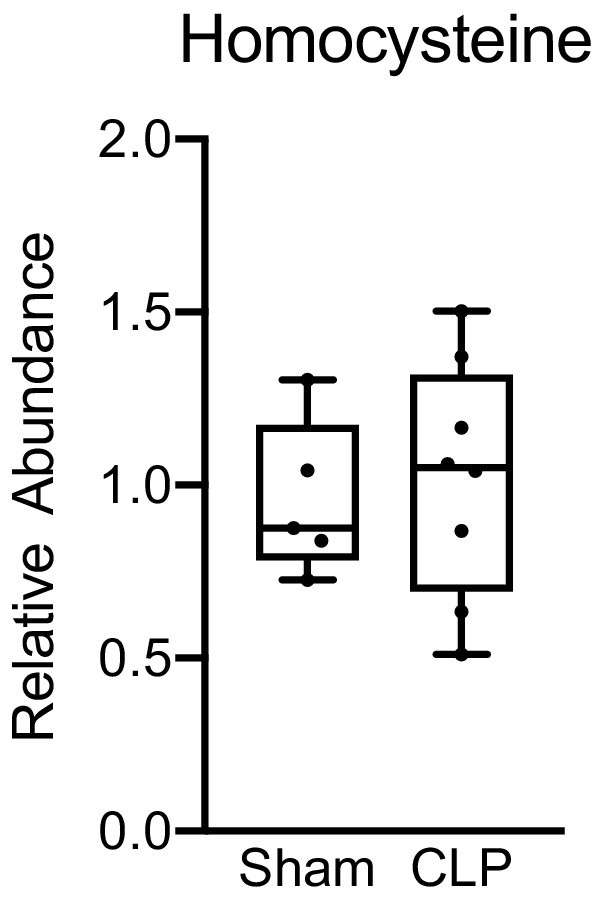
Hepatic homocysteine levels in sham and CLP mice.
Ultrahigh-performance liquid chromatography–tandem mass spectroscopy measurement of homocysteine from isolated hepatocytes 30 hr post-surgery. n = 5 sham, n = 8 CLP.
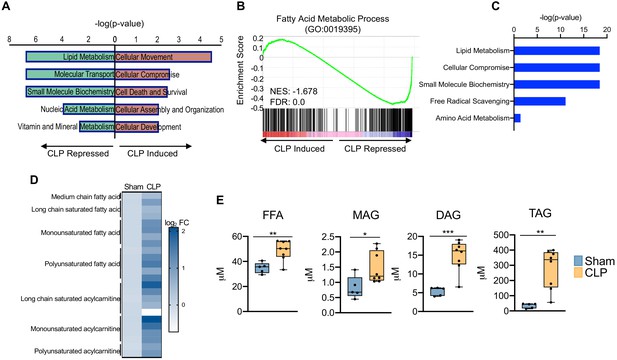
Sepsis promotes hepatic steatosis.
(A) Top five induced and repressed physiological pathways in the liver of sham versus CLP mice identified by IPA of RNA-seq. (B) GSEA of the fatty acid metabolic pathway (GO:0019395) from RNA-seq from livers comparing sham to CLP showing a negative enrichment score (NES = −1.018) (n = 4 mice per group). (C) IPA of top five metabolic pathways significantly altered in the liver in response to sepsis identified by global metabolomic screening. (D) Heatmap representation of log2 fold change of different lipid species in sham and CLP mice measured by UPLC–MS/MS. (E) Quantification of lipids by UPLC–MS/MS in hepatocytes isolated from sham and CLP mice 30 hr post-surgery (n = 5 sham; n = 8 CLP). *p<0.05, **p<0.01, ***p<0.001 as determined by Student’s T-test.
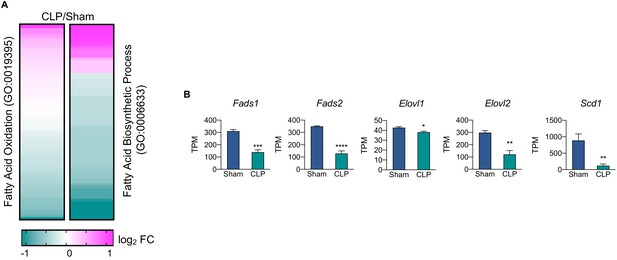
Fatty acid oxidation and biosynthetic pathways in septic livers.
(A) Heatmap representation of average log2 fold change of genes involved in fatty acid oxidation (GO:0019395) and fatty acid biosynthetic process (GO:0006633) in CLP versus sham operated mice (n = 5 sham; n = 8 CLP). (B) Transcripts per million for lipid elongation and desaturation genes from RNA-seq. n = 4 samples per group. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001 as determined by Student’s T-test.
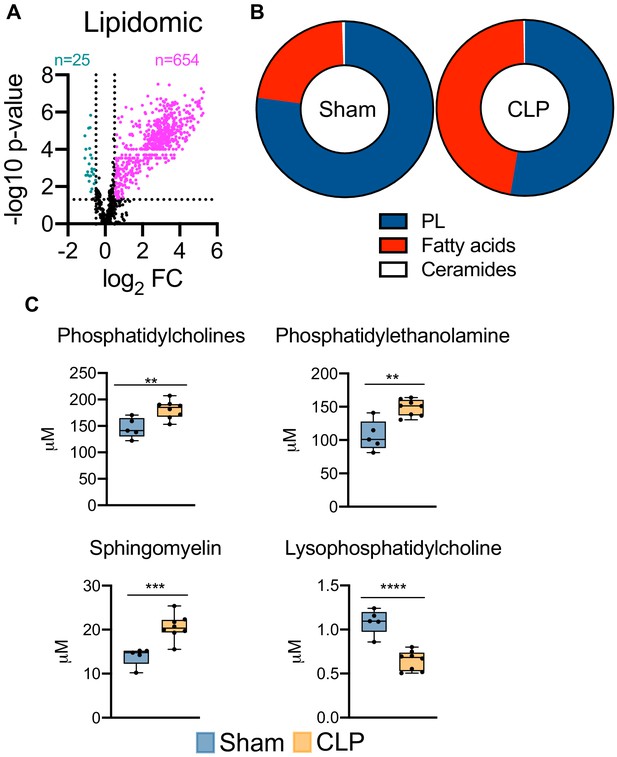
Sepsis reprograms the hepatic lipidome.
(A) Volcano plot of significantly altered lipids from sham and CLP hepatocytes measured by UPLC–MS/MS (n = 5 sham; n = 8 CLP). (B) Pie chart of lipid compositions from sham and CLP mice. (C) Quantification of lipids by UPLC–MS/MS in hepatocytes isolated from sham and CLP mice 30 hr post-surgery (n = 5 sham; n = 8 CLP). **p<0.01, ***p<0.001, ****p<0.0001 as determined by Student’s T-test.
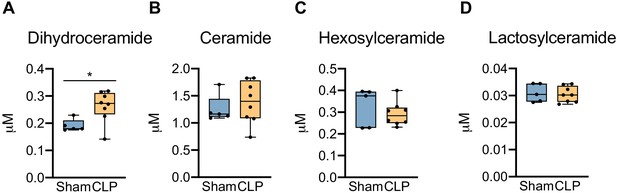
Hepatic ceramide levels in sham and CLP mice.
Ultrahigh-performance liquid chromatography–tandem mass spectroscopy measurement of (A) dihydroceramide, (B) ceramide, (C) hexosylceramide, and (D) lactosylceramide from isolated hepatocytes 30 hr post-surgery. n = 5 sham, n = 8 CLP. *p<0.05 determined by Student’s T-test.
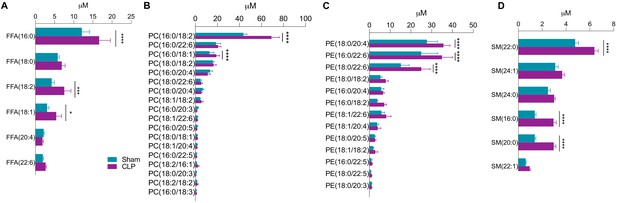
Phosphatidylcholine, phosphatidylethanolamine, and sphingomyelin quantifications.
Ultrahigh-performance liquid chromatography–tandem mass spectroscopy measurement of (A) free fatty acid species, (B) phosphatidylcholine species, (C) phosphatidylethanolamine species, and (D) sphingomyelin species from isolated hepatocytes 30 hr post-surgery. n = 5 sham, n = 8 CLP. ****p<0.0001 determined by two-way ANOVA followed by multiple comparisons test.
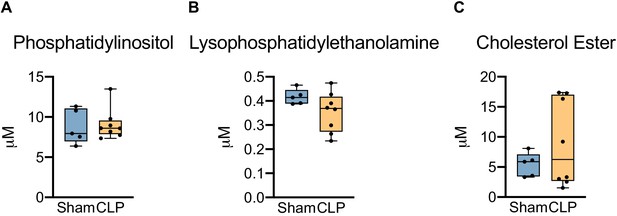
PI, LPE, and CE levels in septic livers.
Ultrahigh-performance liquid chromatography–tandem mass spectroscopy measurement of (A) phosphatidylinositol, (B) lysophosphatidylethanolamine species, and (C) cholesterol ester from isolated hepatocytes 30 hr post-surgery. n = 5 sham, n = 8 CLP. ****p<0.0001 determined by two-way ANOVA followed by multiple comparisons test.
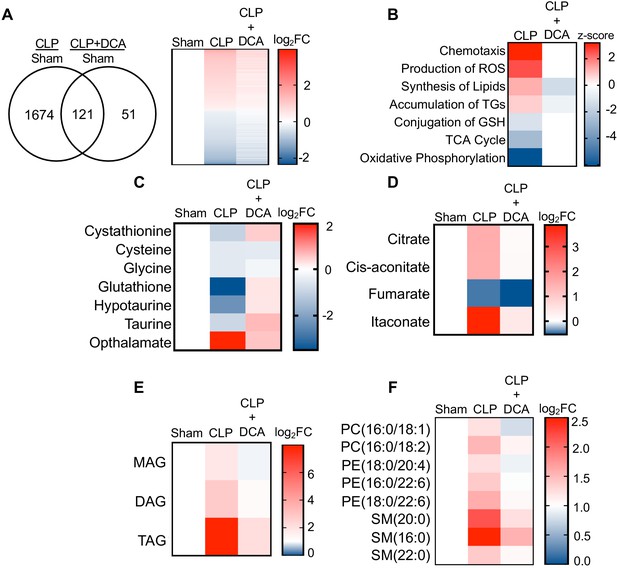
PDK inhibition restores hepatic metabolism in septic mice.
(A) Venn diagram of differentially expressed genes (DEGs) assessed by RNA-seq in CLP versus sham compared to CLP+DCA versus sham 30 hr post-surgery (left). Heatmap of average log2 fold change of DEGs in sham, CLP, and CLP+DCA (right) (n = 4 mice per group). (B) Heatmap depiction of z-scores of top canonical pathways identified by IPA in CLP versus sham and CLP+DCA versus sham. (C) Heatmap depiction of average log2 fold change in metabolite levels involved in redox balance in sham, CLP, and CLP+DCA 24 hr post-surgery measured by UPLC–MS/MS (n = 5 sham; n = 8 CLP and CLP+DCA). (D) Heatmap depiction of average log2 fold change in metabolite levels involved in TCA cycle in sham, CLP, and CLP+DCA 30 hr post-surgery measured by UPLC–MS/MS (n = 5 sham; n = 8 CLP and CLP+DCA). (E) Heatmap depiction of average log2 fold change in MAG, DAG, and TAG levels in sham, CLP, and CLP+DCA 30 hr post-surgery measured by UPLC–MS/MS (n = 5 sham; n = 8 CLP and CLP+DCA). (F) Heatmap depiction of average log2 fold change in phospholipids in sham, CLP and CLP+DCA 30 hr post-surgery measured by UPLC–MS/MS (n = 5 sham; n = 8 CLP and CLP+DCA).
Additional files
-
Source data 1
Lipidomic analysis.
- https://cdn.elifesciences.org/articles/64611/elife-64611-data1-v2.xlsx
-
Source data 2
Metabolomic analysis.
- https://cdn.elifesciences.org/articles/64611/elife-64611-data2-v2.xlsx
-
Source data 3
Transcriptomic analysis.
- https://cdn.elifesciences.org/articles/64611/elife-64611-data3-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/64611/elife-64611-transrepform-v2.docx





