An estimate of the deepest branches of the tree of life from ancient vertically evolving genes
Figures
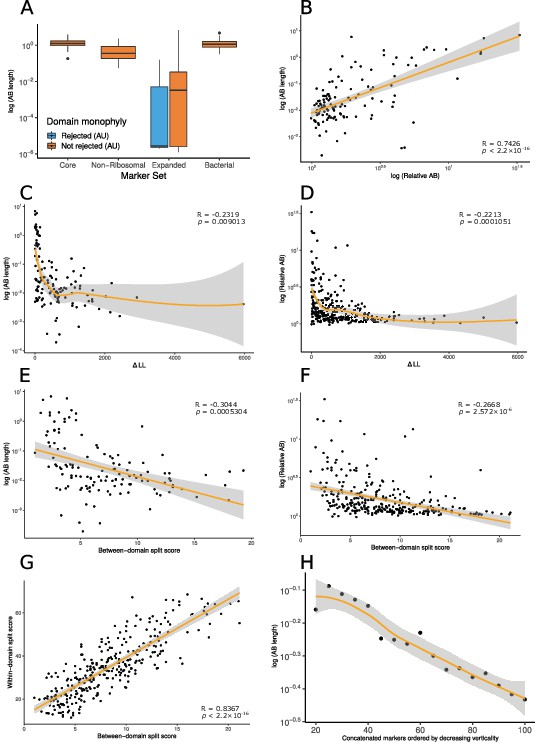
Vertically evolving marker genes support a greater evolutionary distance between Archaea and Bacteria.
(A) Expanded set genes that reject domain monophyly (p<0.05, approximately unbiased [AU] test, with Bonferroni correction [see main text]) support significantly shorter Archaea-Bacteria (AB) branch lengths when constrained to follow a domain monophyletic tree (p=3.653 × 10–6, Wilcoxon rank-sum test). None of the marker genes from several other published analyses significantly reject domain monophyly (Bonferroni-corrected p<0.05, AU test) for all genes tested, consistent with vertical inheritance from the LUCA (last universal common ancestor) to the last common ancestors of Archaea and Bacteria, respectively. (B) Two measures of evolutionary proximity (Zhu et al., 2019), AB branch length and relative AB distance, are positively correlated (R = 0.7426499, p< 2.2 × 10–16). We considered two complementary proxies of marker gene verticality: ∆LL (C: against AB branch length; D: against relative AB length), which reflects the degree to which marker genes reject domain monophyly (C: p=0.009013 and R = –0.2317894; D: p=0.0001051 and R = –0.2213292); and the between-domain split score (E: against AB branch length; F: against relative AB length), which quantifies the extent to which marker genes recover monophyletic Archaea and Bacteria; a higher split score (see Materials and methods) indicates the splitting of domains into multiple gene tree clades due to gene transfer, reciprocal sorting out of paralogs, or lack of phylogenetic resolution (E: p=0.0005304 and R = –0.3043537; F: p=2.572 × 10–6 and R = –0.2667739). We also considered a split score based on within-domain relationships (G); between- and within-domain split scores are positively correlated: R = 0.836679, p<2.2 × 10–16, Pearson’s correlation, indicating that markers that recover Archaea and Bacteria as monophyletic also tend to recover established within-domain relationships. (H) Inferred AB length decreases as marker genes of lower verticality (larger ∆LL) are added to the concatenate. Marker genes were sorted by ∆LL, the difference in log-likelihood between the maximum likelihood gene family tree under a free topology search and the log-likelihood of the best tree constrained to obey domain monophyly. Note that 79/381 expanded set markers had zero or one archaea in the 1000-species subsample and so could not be included in these analyses; of the remaining 302 markers, 176 have AB branch lengths very close to 0 in the constraint tree (as seen in panel A). In these plots, we removed all markers with an AB branch length of <0.00001; see Figure 1—figure supplements 1–13 for all plots. Nonlinear trendlines were estimated using LOESS regression.
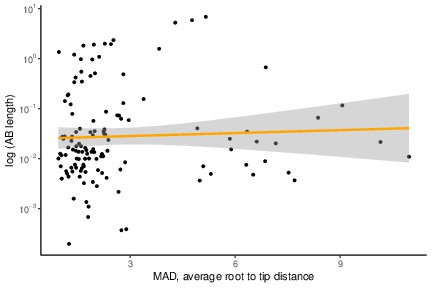
No evidence for a relationship between Archaea-Bacteria (AB) branch length and gene evolutionary rate (average MAD root-to-tip distance).
We did not detect a significant correlation between AB length and rate (average MAD root-to-tip distance) when excluding genes with a nonzero (<0.00001) AB length. p=0.2025, R = 0.1143076, Pearson’s correlation.
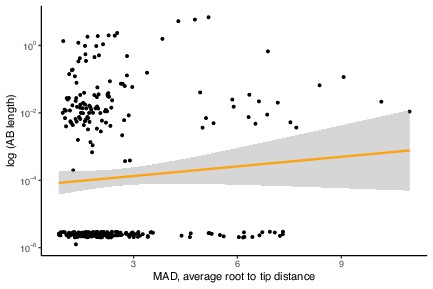
No evidence for a relationship between Archaea-Bacteria (AB) branch length and gene evolutionary rate (average MAD root-to-tip distance).
The analysis is the same as in Figure 1—figure supplement 3 but with the inclusion of markers where AB branch length ~0. p=0.0761, R = 0.102226, Pearson’s correlation.
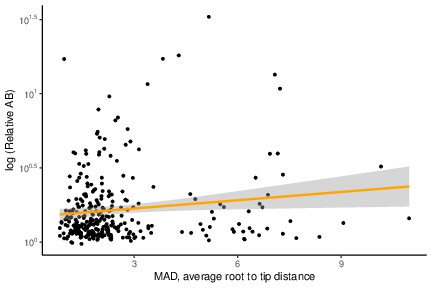
A significant positive relationship between relative Archaea-Bacteria (AB) distance and evolutionary rate (MAD root-to-tip distance).
Genes with a greater relative AB distance have moderately higher evolutionary rates. p=0.007435, R = 0.1537479, Pearson’s correlation.

Two proxies for marker gene verticality, ΔLL and between-domain split score, are highly correlated.
p<2.2 × 10–16, R = 0.836679, Pearson’s correlation.
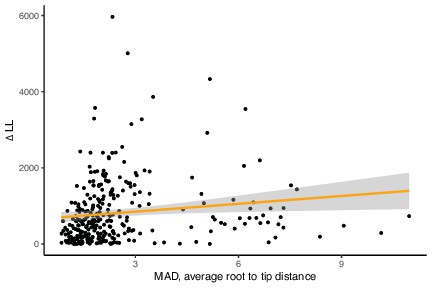
Low-verticality genes (as measured by ΔLL) have a higher evolutionary rate (as measured by the mean root-to-tip distance on MAD-rooted gene trees).
Less vertically evolving marker genes evolve faster, although the effect is moderate (p=0.01506, R = 0.1397803, Pearson’s correlation).

Low-verticality genes (measured by between-domain split score) have a higher evolutionary rate (MAD root-to-tip distance).
There is a positive relationship between between-domain split score (higher split score denotes lower verticality) and gene evolutionary rate (measured as mean MAD root-to-tip distance), p=0.002947, R = 0.1705415, Pearson’s correlation.
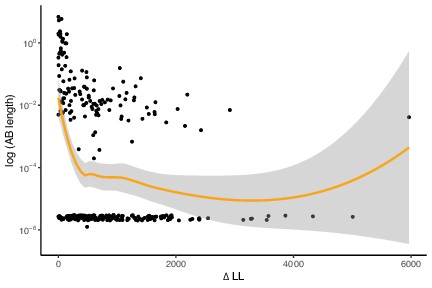
High-verticality marker genes have longer Archaea-Bacteria (AB) branch lengths.
The depicted relationship is negative because high values of ΔLL correspond to lower between-domain verticality. This is the same analysis as in Figure 1C but with marker genes with AB branch length ~0 included. The trendline is estimated using LOESS regression. We used a LOESS regression as the trendline here as the relationship varies across markers of differing verticality, p=0.00145, R = –0.1824596, Pearson’s correlation.
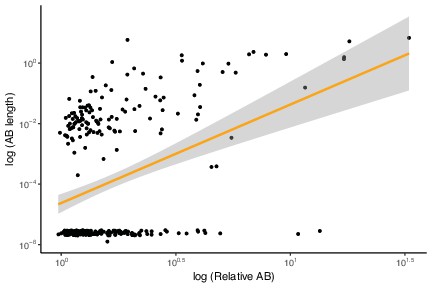
Archaea-Bacteria (AB) branch length and relative AB distance are positively correlated.
This is the same analysis as in Figure 1B but with marker genes with AB branch length ~0 included, p<2.2 × 10–16, R = 0.706099, Pearson’s correlation.
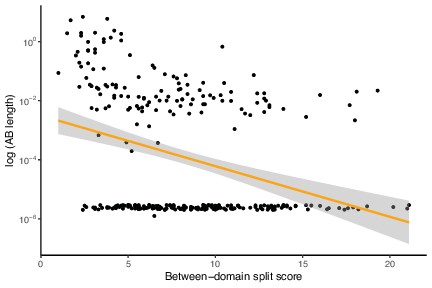
Archaea-Bacteria (AB) branch length is negatively correlated with between-domain split score.
This is the same analysis as Figure 1E but with marker genes with AB branch length ~0 included, p=1.111 × 10–05, R = –0.2498829, Pearson’s correlation.
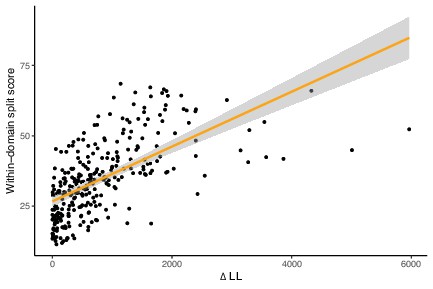
Within-domain split score and ΔLL are strongly correlated, suggesting that both proxies capture a common signal of marker gene verticality.
p<2.2 × 10–16, R = 0.6201967, Pearson’s correlation.
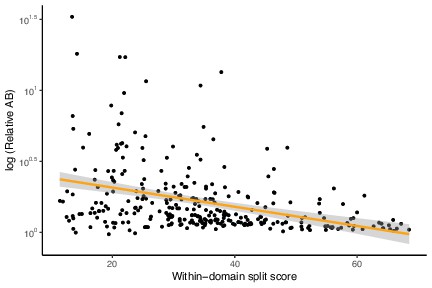
Low-verticality marker genes (measured as within-domain split score) have shorter relative Archaea-Bacteria (AB) distances.
A higher split score denotes lower verticality. p=5.685 × 10–06, R = –0.2577625, Pearson’s correlation.
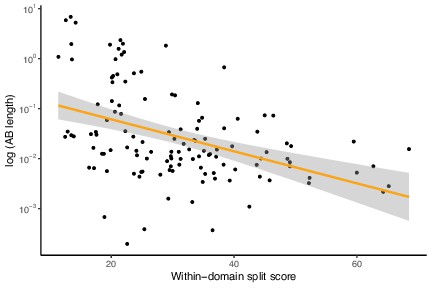
Low-verticality marker genes (measured as within-domain split score) have shorter Archaea-Bacteria (AB) branch lengths.
A higher split score denotes lower verticality. p=0.0001467, R = –0.3318924, Pearson’s correlation.
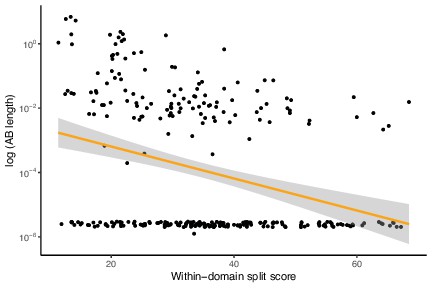
Low-verticality marker genes (measured as within-domain split score) have shorter Archaea-Bacteria (AB) branch lengths.
This is the same analysis as in Figure 1—figure supplement 12 but with marker genes with AB branch length ~0 included. There is a significant but weaker negative correlation between AB branch length and marker gene verticality (within-domain split score) compared to the analysis where this class of genes is excluded. p=7.498 × 10–06, R = –0.2545369, Pearson’s correlation.
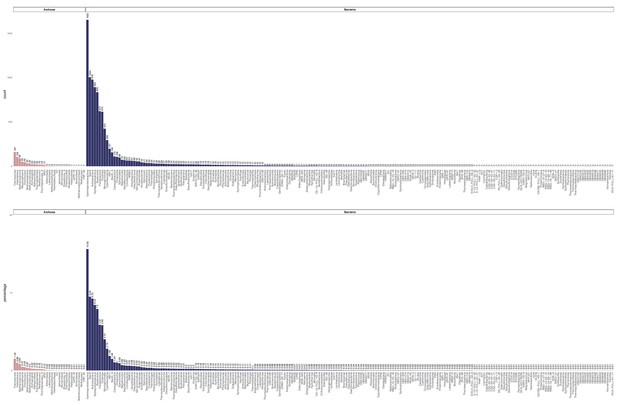
Raw count and percentage distribution of the Genome Taxonomy Database (GTDB)-defined classes for 10,575 archaeal and bacterial genomes in the expanded marker set analysis.
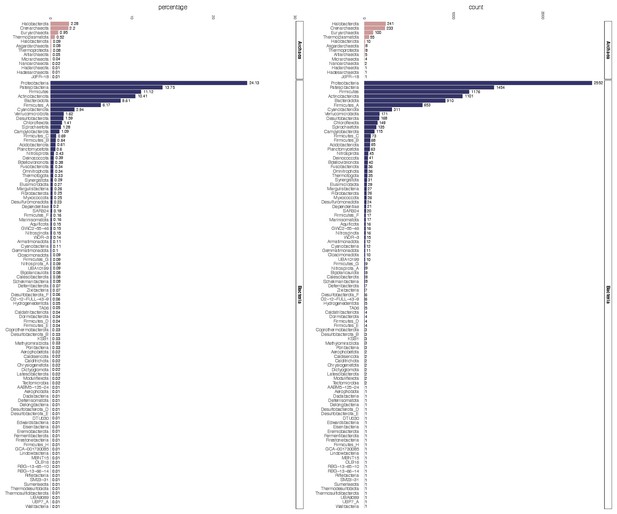
Raw count and percentage distribution of the Genome Taxonomy Database (GTDB)-defined phyla for 10,575 archaeal and bacterial genomes in the expanded marker set analysis.
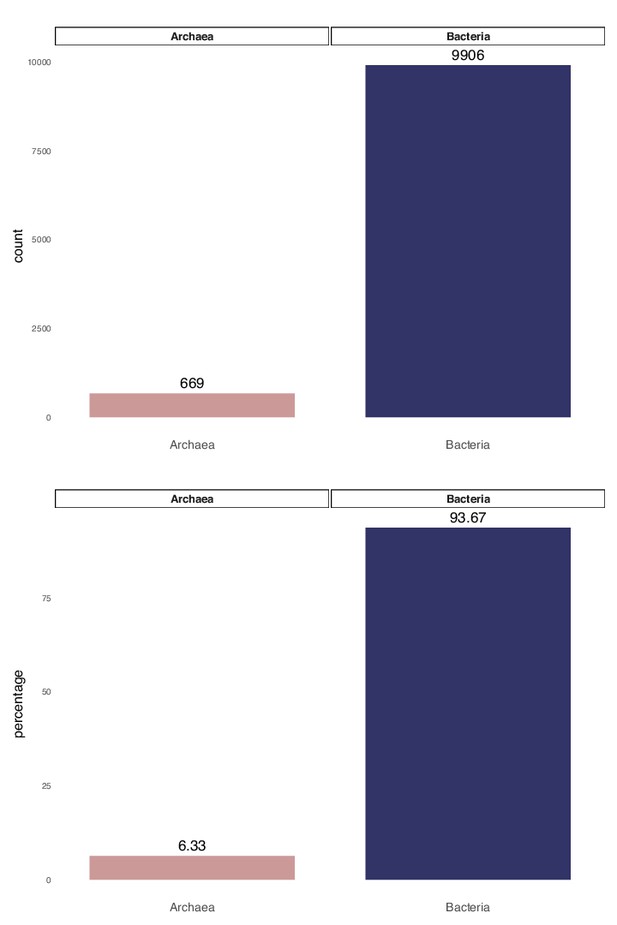
Raw count and percentage distribution of domains for 10,575 archaeal and bacterial genomes in the expanded marker set analysis.
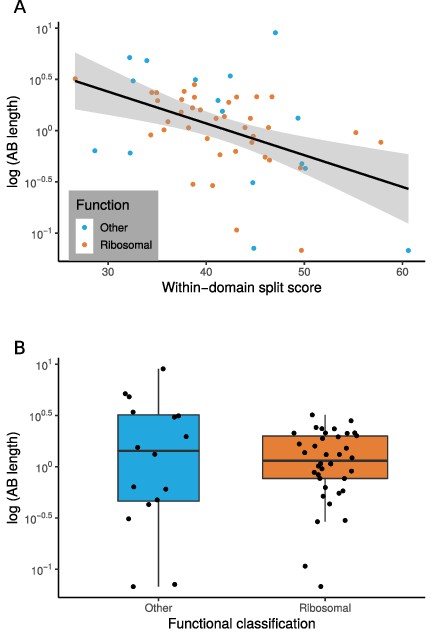
The relationship between marker gene verticality, Archaea-Bacteria (AB) branch length, and functional category.
(A) Vertically evolving phylogenetic markers have longer AB branches. The plot shows the relationship between a proxy for marker gene verticality, within-domain split score (a lower split score denotes better recovery of established within-domain relationships, see Materials and methods), and AB branch length (in expected number of substitutions/site) for the 54 marker genes. Marker genes with higher split scores (that split established monophyletic groups into multiple subclades) have shorter AB branch lengths (p=0.0311, R = 0.294). Split scores of ribosomal and non-ribosomal markers were statistically indistinguishable (p=0.828, Figure 2—figure supplement 1). (B) Among vertically evolving marker genes, ribosomal genes do not have a longer AB branch length. The plot shows functional classification of markers against AB branch length using 54 vertically evolving markers. We did not obtain a significant difference between AB branch lengths for ribosomal and non-ribosomal genes (p=0.6191, Wilcoxon rank-sum test).
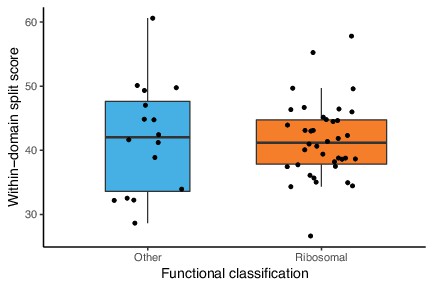
Among vertically evolving marker genes, the split scores of ribosomal and non-ribosomal proteins are statistically indistinguishable.
The plot shows functional classification of markers (ribosomal markers or other) against the split score (a higher split score denotes greater disagreement with established within-domain relationships) using 54 markers from the new analysis. After removing genes that do not appear to have been vertically inherited since the divergence of Archaea and Bacteria, split scores of ribosomal and non-ribosomal markers were statistically indistinguishable (p=0.8275, Wilcoxon rank-sum test).
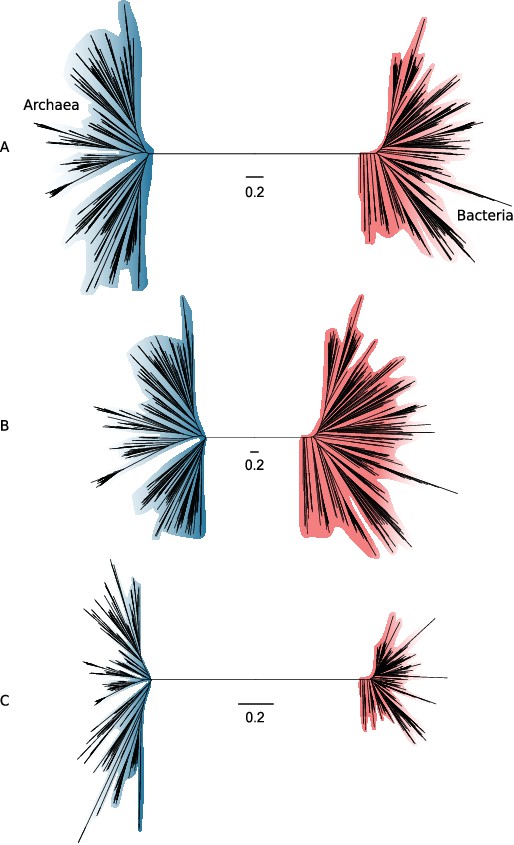
Slow- and fast-evolving sites support different shapes for the universal tree.
(A) Tree of Archaea (blue) and Bacteria (red) inferred from a concatenation of 27 core genes using the best-fitting model (LG + C60 + G4 + F). (B) Tree inferred from the fastest-evolving sites. (C) Tree inferred from the slowest-evolving sites. To facilitate comparison of relative diversity, scale bars are provided separately for each panel; for a version of this figure with a common scale bar for all three panels, see Figure 3—figure supplement 1. Slow-evolving sites support a relatively long inter-domain branch and less diversity within the domains (i.e., shorter between-taxa branch lengths within domains). This suggests that substitution saturation (overwriting of earlier changes) may reduce the relative length of the AB branch at fast-evolving sites and genes.
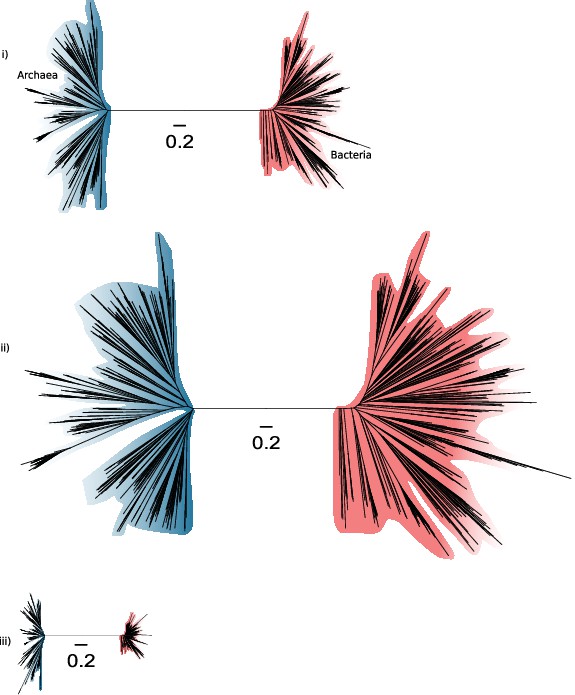
Slow- and fast-evolving sites support different shapes for the universal tree.
(i) Tree of Archaea (blue) and Bacteria (red) inferred from a concatenation of 27 core genes using the best-fitting model (LG + C60 + G4 + F). (ii) Tree inferred from the fastest-evolving sites. (iii) Tree inferred from the slowest-evolving sites. Identical scale bars are provided for comparison.
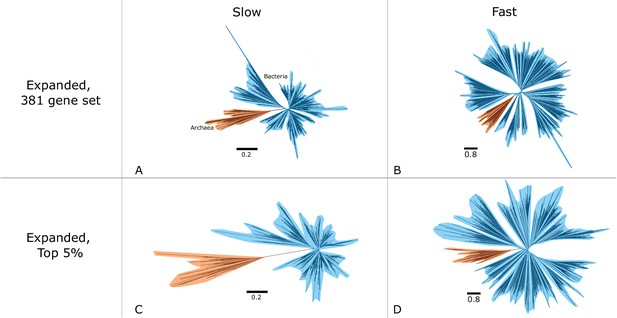
Vertically evolving genes and slow-evolving sites support a longer relative Archaea-Bacteria (AB) branch length.
We estimated site-specific evolutionary rates for all marker genes in the expanded dataset (A, B), as well as for the 20 genes with the smallest ΔLL (top 5%) in that dataset (C, D). Concatenations based on the slowest sites (A, C) and on the top 5% vertical genes (C, D) support a longer AB branch. This suggests that the inference of a short AB branch is impacted by both substitutional saturation and unmodeled inter-domain transfer of marker genes. Phylogenies were inferred under the LG + G4 + F model in IQ-TREE 2. Branch lengths are the expected number of substitutions per site, as indicated by the scale bars. Alignment lengths in amino acids: (A) 36797, (B) 67274, (C) 2736, and (D) 3,884.
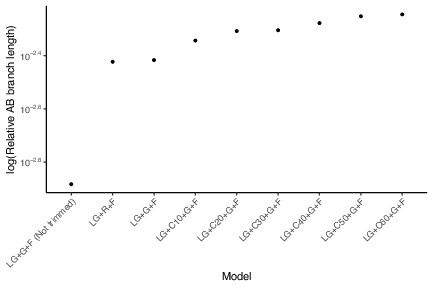
The effect of modeling site compositional heterogeneity on Archaea-Bacteria (AB) branch length.
Increasing the number of protein mixture profiles, as well as trimming poorly aligned positions, is associated with a change in AB branch length on the expanded marker set. All analyses used LG exchangeabilities, four rate categories (gamma-distributed or freely estimated), and included a general composition vector containing the empirical amino acid frequencies (+F). Modeling of site heterogeneity with the C10-C60 models increases the inferred AB branch length approximately twofold. Trimming poorly aligned sites slightly increases the AB branch estimation, whereas relaxing the gamma rate categories slightly decreases estimation of AB branch length. LG, LG substitution matrix; G, four gamma rate categories; F, empirical site frequencies estimated from the data; C10–60, number of protein mixture profiles used; R, four free rate categories that relax the assumption of a gamma distribution for rates; BMGE, trimming using Block Mapping and Gathering with Entropy (Criscuolo and Gribaldo, 2010).

A phylogeny of Archaea and Bacteria inferred from a concatenation of 27 marker genes.
Consistent with some recent studies (Dombrowski et al., 2020; Guy and Ettema, 2011; Raymann et al., 2015; Williams et al., 2017), we recovered the DPANN, TACK, and Asgard Archaea as monophyletic groups. Although the deep branches within Bacteria are poorly resolved, we recovered a sister group relationship between candidate phyla radiation (CPR) and Chloroflexota, consistent with recent reports (Taib et al., 2020; Coleman et al., 2021). The tree was inferred using the best-fitting LG + C60 + G4 + F model in IQ-TREE 2 (Minh et al., 2020). Branch lengths are proportional to the expected number of substitutions per site. Support values are ultrafast (UFBoot2) bootstraps (Hoang et al., 2018). Numbers in parenthesis refer to the number of taxa within each collapsed clade. Please note that the collapsed taxa in the Archaea and Bacteria roughly correspond to order- and phylum-level lineages, respectively.
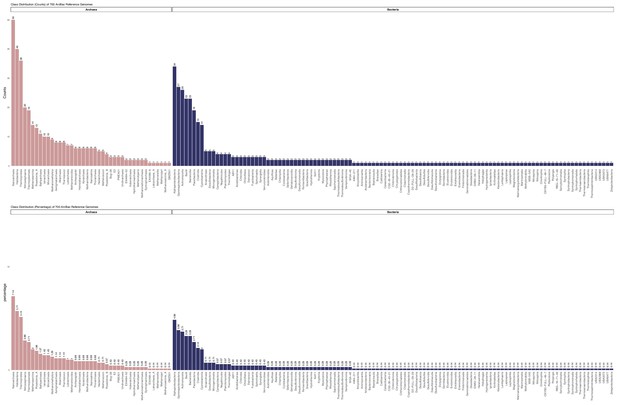
Raw count and percentage distribution of the Genome Taxonomy Database (GTDB)-defined classes of 350 archaea and 350 bacteria used in the 27 marker gene set analysis.
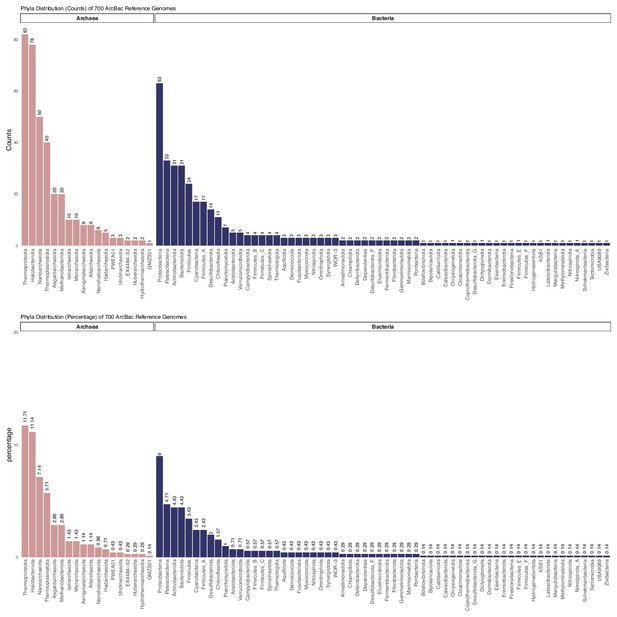
Raw count and percentage distribution of the Genome Taxonomy Database (GTDB)-defined phyla of 350 archaea and 350 bacteria used in the 27 marker gene set analysis.
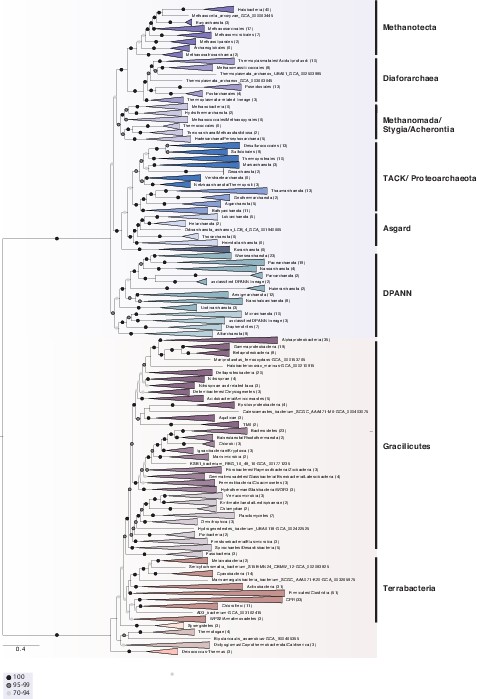
A phylogeny of Archaea and Bacteria inferred from a concatenation of 25 marker genes.
In addition to our focal analysis, we inferred a tree from a 25-gene subset of the 27 highest-ranked marker genes. This analysis excluded two genes (COG0480 and COG5257) that, while scoring well by split score, had either low representation in Bacteria or had previously been suggested to have a complex evolutionary history in Archaea (Narrowe et al., 2018). The tree topologies and Archaea-Bacteria (AB) branch lengths (25-marker supermatrix: 2.3738 substitutions/site, 27-marker supermatrix: 2.5178 substitutions/site) were closely similar between the 27- and 25-gene analyses, with the exception of several lineages that proved difficult to place. In the 25-gene analysis, the Korarchaeota branched outside the TACK + Asgard clade with moderate (92%) bootstrap support, and the Hadesarchaea, Persephonarchaea, Theionarchaea, Methanofastidiosa, and Thermococcales formed a clade sister to Methanomada (98% bootstrap support). The tree was inferred using the best-fitting LG + C60 + G4 + F model in IQ-TREE 2 (Minh et al., 2020). Branch lengths are proportional to the expected number of substitutions per site. Support values are ultrafast (UFBoot2) bootstraps (Hoang et al., 2018). Numbers in parenthesis refer to the number of taxa within each collapsed clade. Please note that the collapsed taxa in the Archaea and Bacteria roughly correspond to order- and phylum-level lineages, respectively.
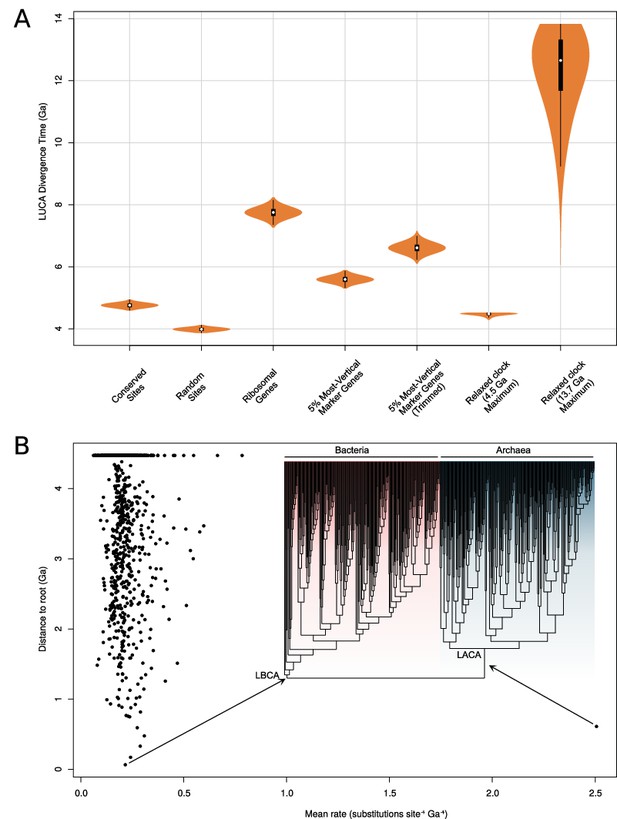
Molecular clock estimates of the LUCA and LACA age are uncertain due to a lack of deep calibrations and maximum ages for microbial clades.
(A) Posterior node age estimates from Bayesian molecular clock analyses of (1) conserved sites as estimated previously (Zhu et al., 2019); (2) random sites (Zhu et al., 2019); (3) ribosomal genes (Zhu et al., 2019); (4) the top 5% of marker gene families according to their ∆LL score (including only one ribosomal protein); and (5) the same top 5% of marker genes trimmed using BMGE (Criscuolo and Gribaldo, 2010) to remove poorly aligned sites. In each case, a strict molecular clock was applied, with the age of the Cyanobacteria-Melainabacteria split constrained between 2.5 and 2.6 Ga. In (6) all annotations for the top 20 genes and (7) an expanded set of fossil calibrations was implemented with a relaxed (lognormal) molecular clock. In (6), a soft maximum age of 4.520 Ga was applied, representing the age of the moon-forming impact (Kleine et al., 2005). In (7), a soft maximum age corresponding to the estimated age of the universe (Aghanim, 2018) was applied. (B) Inferred rates of molecular evolution along the phylogeny in a relaxed clock analysis where the maximum age was set to 4.520 Ga. We plotted per-branch rates (site–1 Ga–1) against distance to the root. The rate of evolution along the archaea stem lineage was a clear outlier (mean = 2.51, 95% HPD = 1.6–3.5subs. site–1 Ga–1). The phylogeny used was that depicted in Figure 4.
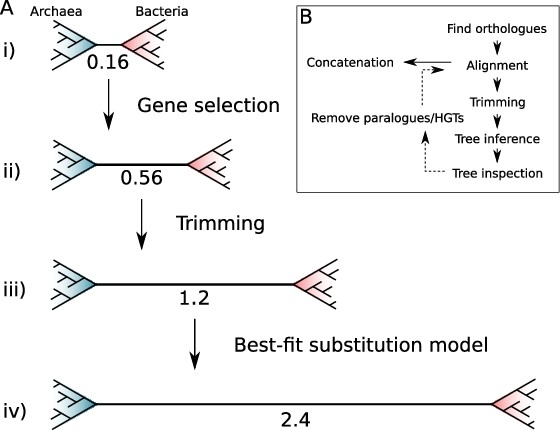
The impact of marker gene choice, phylogenetic congruence, alignment trimming, and substitution model fit on estimates of the Archaea-Bacteria (AB) branch length.
(A) Analysis using a site-homogeneous model (LG + G4 + F) on the complete 381-gene expanded set (i) results in an AB branch substantially shorter than previous estimates. Removing the genes most seriously affected by inter-domain gene transfer (ii), trimming poorly aligned sites (iii) using BMGE (Criscuolo and Gribaldo, 2010) in the original alignments (see below), and using the best-fitting site-heterogeneous model (iv) (LG + C60 + G4 + F) substantially increase the estimated AB length, such that it is comparable with published estimates from the ‘core’ set: 3.3 (Williams et al., 2020) and the consensus set of 27 markers identified in the present study: 2.5. Branch lengths measured in expected number of substitutions/site. (B) Workflow for iterative manual curation of marker gene families for concatenation analysis. After inference and inspection of initial ortholog trees, several rounds of manual inspection and removal of HGTs and distant paralogs were carried out. These sequences were removed from the initial set of orthologs before alignment and trimming. For a detailed discussion of some of these issues, and practical guidelines on phylogenomic analysis of multi-gene datasets, see Kapli et al., 2020 for a useful review.
Tables
Archaea-Bacteria (AB) branch lengths and AB branch lengths as a proportion of total tree length inferred from ribosomal and non-ribosomal concatenates are similar.
The data do not support a faster evolutionary rate for ribosomal proteins on the AB branch compared to other kinds of ancient proteins.
| AB branch length | Total tree length | AB branch length as a proportion of total tree length | ||||
|---|---|---|---|---|---|---|
| Ribosomal | Non-ribosomal | Ribosomal | Non-ribosomal | Ribosomal | Non-ribosomal | |
| 27 marker set | 1.9541 | 3.7723 | 250.7255 | 239.8203 | 0.0078 | 0.0157 |
| 54 marker set | 1.8647 | 2.5414 | 271.3327 | 288.8470 | 0.0069 | 0.0088 |
The inferred Archaea-Bacteria (AB) branch length from a concatenation of the top 27 markers using a simple model compared to models that account for site compositional heterogeneity.
Models that account for across-site compositional heterogeneity fit the data better (as assessed by lower Bayesian information criterion [BIC] scores) and infer a longer AB branch length.
| Substitution model | BIC (∆BIC) | AB branch length |
|---|---|---|
| LG + G4 + F | 5935950.053 | 1.4491 |
| LG + C20 + G4 + F | (152046.1) | 2.1394 |
| LG + C40 + G4 + F | (179126.7) | 2.4697 |
| LG + C60 + G4 + F | (189063.8) | 2.5178 |
Additional files
-
Supplementary file 1
Marker metadata, KO and Pfam annotations and descriptions, and manual inspection notes for reciprocal monophyly and presence of paralogs for 381 marker genes used in Zhu et al., 2019 (Materials and methods).
- https://cdn.elifesciences.org/articles/66695/elife-66695-supp1-v2.xlsx
-
Supplementary file 2
Marker metadata, KO and Pfam annotations and descriptions, and manual inspection notes for 95 markers in the core, bacterial, and non-ribosomal marker gene sets (Materials and methods).
- https://cdn.elifesciences.org/articles/66695/elife-66695-supp2-v2.xlsx
-
Supplementary file 3
NCBI taxonomic information for 350 archaeal and 350 bacterial genomes sampled in the new analyses.
- https://cdn.elifesciences.org/articles/66695/elife-66695-supp3-v2.xlsx
-
Supplementary file 4
Clade definitions for quantifying taxonomic splits and split score statistical summaries for ranking of the core, bacterial, non-ribosomal marker genes, and 381 marker genes (Materials and methods).
- https://cdn.elifesciences.org/articles/66695/elife-66695-supp4-v2.xlsx
-
Supplementary file 5
Annotations of the top 20 genes from the expanded set, and a list of fossil calibrations.
(a) Functional annotations for the top 20 genes used and in Figure 6 and referred to in Figure 6A. (b) A list of fossil calibrations employed in relaxed molecular clock analyses. All calibrations were modeled as uniform distributions between a hard minimum and a soft maximum. The probability that the maximum could be exceeded was modeled as a 2.5% probability tail.
- https://cdn.elifesciences.org/articles/66695/elife-66695-supp5-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/66695/elife-66695-transrepform1-v2.docx





