Pseudohypoxic HIF pathway activation dysregulates collagen structure-function in human lung fibrosis
Figures
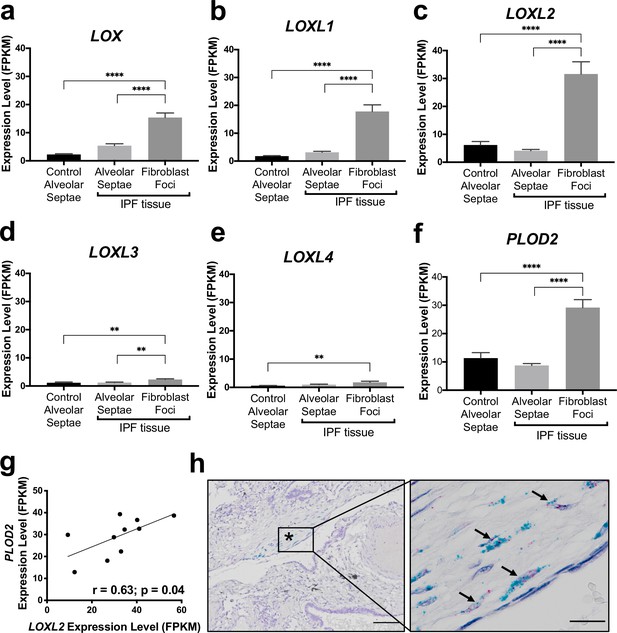
The collagen cross-linking enzymes PLOD2 and LOXL2 are co-expressed at sites of active fibrogenesis in IPF.
(A–F) Expression of LOX, LOXL1, LOXL2, LOXL3, LOXL4, and PLOD2 in healthy alveolar septae, IPF alveolar septae and IPF fibroblast foci (n = 10 individual healthy and IPF donors). Relative expression levels are calculated as Fragments Per Kilobase of transcript per Million mapped reads (FPKM). Bars represent standard geometric means. **p < 0.01; ****p < 0.0001 by Tukey’s multiple comparisons test. (G) Scatterplot of paired fibroblast foci data from (C) and (F) were plotted to compare expression of PLOD2 and LOXL2 (Spearman rank correlation coefficient r = 0.63, p = 0.04). (H) Representative image of mRNA expression of PLOD2 (red chromagen) and LOXL2 (green chromagen) in IPF lung tissue (n = 7 donors) using RNAscope RNA in-situ hybridisation. A fibroblastic focus is identified by * and arrows identify co-expression pattern. Left scale bar 100 μm, right scale bar 20 μm.
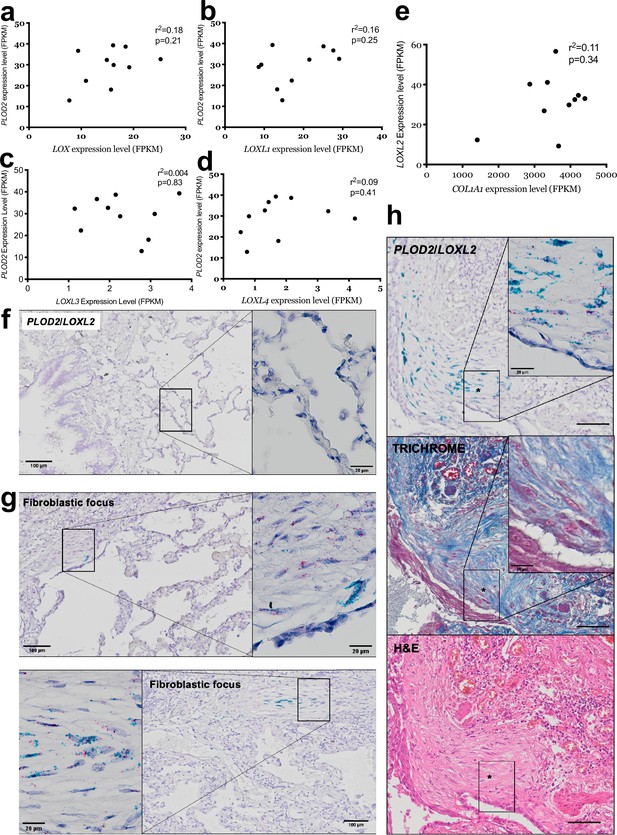
Correlation of PLOD2 with LOXL family members.
(A–D) Scatterplots of paired data from (Figure 1A, B, D-F) comparing gene expression within fibroblast foci (n = 10 donors) of PLOD2 with LOX, LOXL1, LOXL3, and LOXL4. (E) Scatterplot comparing gene expression within fibroblast foci (n = 10 donors) of LOXL2 and COL1A1 expression. Relative expression levels are calculated as Fragments Per Kilobase of transcript per Million mapped reads (FPKM). Strength of correlation calculated using Spearman correlation coefficient. (F and G) Representative image of mRNA expression of PLOD2 (red chromagen) and LOXL2 (green chromagen) in IPF lung tissue (n = 7 donors) using RNAscope RNA in-situ hybridisation. (H) Serial sections of IPF lung tissue were stained for PLOD2 (red chromagen) and LOXL2 (green chromagen) using RNAscope RNA in-situ hybridisation (top panel), with Masson’s trichrome stain (middle panel, collagen shown in blue) or H/E (bottom panel). Scale bars in (F–H) are 100 μm with inserts 20 μm.
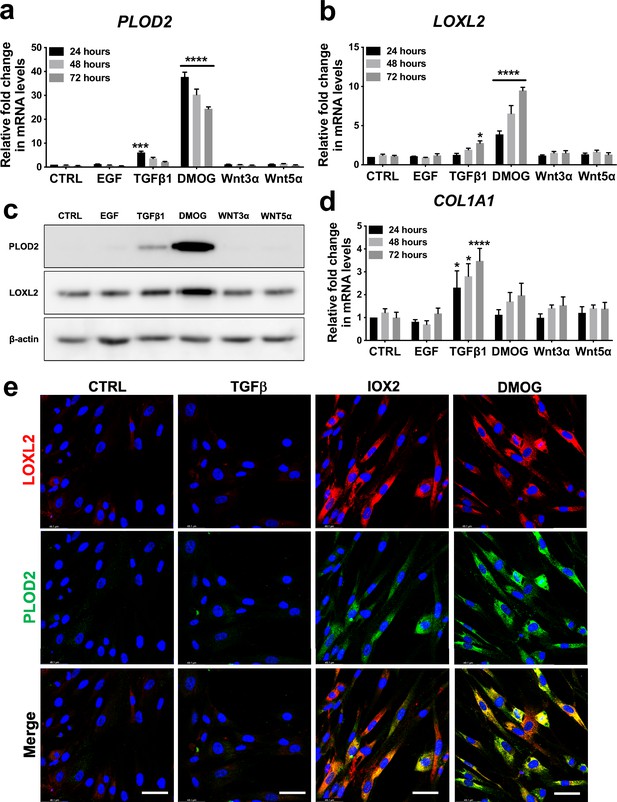
Hypoxia mimetics strongly promote PLOD2 and LOXL2 expression in lung fibroblasts.
(A–B, D) Relative gene expression using the ΔΔCt method of PLOD2, LOXL2, and COL1A1 in healthy lung fibroblasts over a 72-hr time course in the presence of EGF, TGFβ1, the hypoxia mimetic DMOG, Wnt3a, Wnt5a, or vehicle control. n = 3 independent experiments. Bars indicate geometric means. *p < 0.05; ***p < 0.001; ****p < 0.0001 by Dunnett’s multiple comparisons test. (C) PLOD2 and LOXL2 protein levels at 72 hr. β-actin loading control. The full blots are shown in Figure 2—source data 1. (E) Representative immunofluorescence images of healthy lung fibroblasts with indicated treatment stained for LOXL2 (red), PLOD2 (green), and DAPI (blue). Scale bar 50 μm.
-
Figure 2—source data 1
Full membrane scans for western blot images for Figure 2c.
- https://cdn.elifesciences.org/articles/69348/elife-69348-fig2-data1-v1.zip
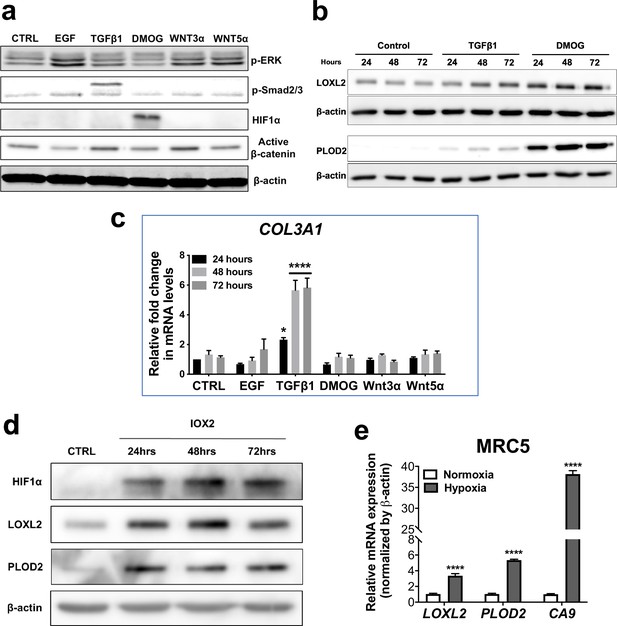
Pro-fibrotic signalling pathways in human lung fibroblasts.
(A–C) Healthy lung fibroblasts exposed to control, EGF, TGFβ1, DMOG, Wnt3α or Wnt5α signalling for 24, 48, or 72 hr. n = 3 independent experiments. (A) Protein expression of phospho-ERK, phospho-SMAD2/3, HIF1α, and active β-catenin at 24 hr of exposure to conditions. β-actin was used as a loading control. The full blots are shown in Figure 2—figure supplement 1—source data 1. (B) LOXL2 and PLOD2 protein levels at 24, 48, or 72 hr of exposure to conditions. β-actin was used as a loading control. The full blots are shown in Figure 2—figure supplement 1—source data 1. (C) Expression of COL3A1 in healthy lung fibroblasts exposed to conditions for 24, 48, or 72 hr using the ΔΔCt method. Bars indicate geometric means. ****p < 0.0001 by Dunnett’s multiple comparisons test. (D) Protein expression of HIF1α, LOXL2, and PLOD2 in IPF fibroblasts exposed to control media or IOX2 for 24, 48, or 72 hr. β-actin was used as a loading control. The full blots are shown in Figure 2—figure supplement 1—source data 1. (E) Fold change in mRNA levels of LOXL2, PLOD2 and the HIF pathway activation marker gene carbonic anhydrase IX/9 (CA9) in MRC5 fibroblasts after incubation in nomoxia (21% O2) or hypoxia (1% O2) for 24 hr. β-actin-normalised mRNA levels under nomoxia were used to set the baseline value at unity. Data are mean ± s.d. n = 3 samples per group. ****p < 0.0001 using unpaired t test.
-
Figure 2—figure supplement 1—source data 1
Full membrane scans for western blot images for Figure 2—figure supplement 1a, b, d.
- https://cdn.elifesciences.org/articles/69348/elife-69348-fig2-figsupp1-data1-v1.zip
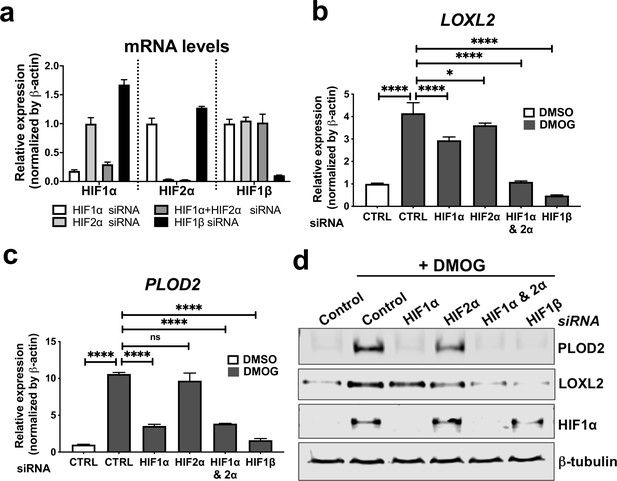
HIF pathway activation regulates PLOD2 and LOXL2 expression in lung fibroblasts from patients with IPF.
(A) Fold changes in mRNA levels of HIF1α (HIF1A), HIF2α (EPAS1), and HIF1β (ARNT) in primary human lung fibroblasts from patients with IPF transfected with indicated siRNA followed by treatment with DMOG. β-actin-normalised mRNA levels in control cells were used to set the baseline value at unity. Data are mean ± s.d. n = 3 samples per group. (B, C) Fold change in mRNA levels of LOXL2 (B) and PLOD2 (C) in IPF fibroblasts transfected with indicated siRNA followed by treatment with DMOG or vehicle control. β-actin-normalised mRNA levels in control cells were used to set the baseline value at unity. Data are mean ± s.d. n = 3 samples per group. ns (not significant, p > 0.05); *p < 0.05; ****p < 0.0001 by Dunnett’s multiple comparisons test. (D) PLOD2, LOXL2 and HIF1α and β-tubulin protein levels in IPF fibroblasts transfected with indicated siRNA followed by treatment of DMSO or DMOG. β-tubulin was used as a loading control. The full blots are shown in Figure 3—source data 1.
-
Figure 3—source data 1
Full membrane scans for western blot images for Figure 3d.
- https://cdn.elifesciences.org/articles/69348/elife-69348-fig3-data1-v1.zip
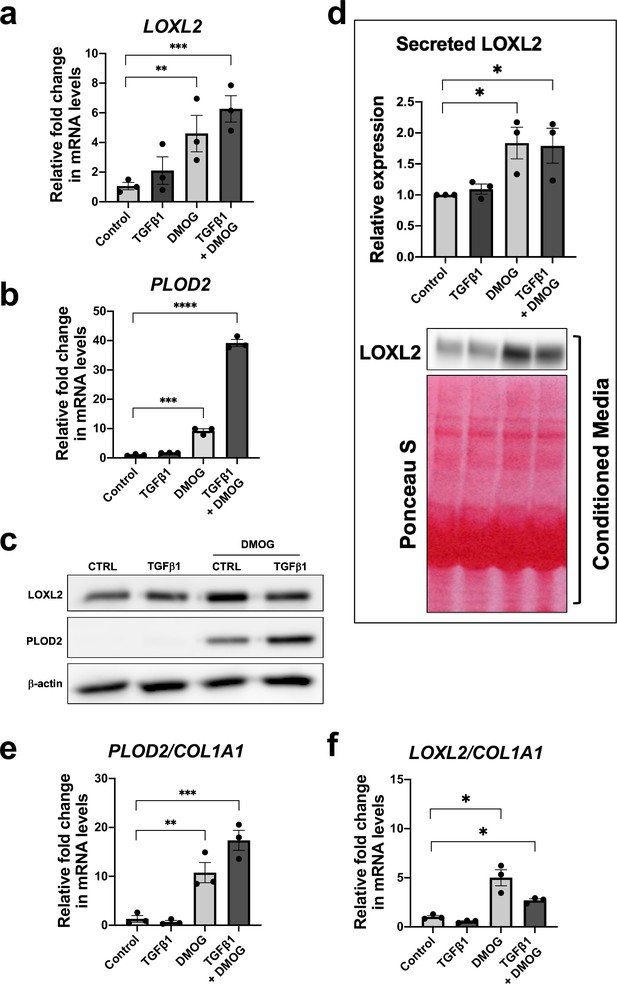
HIF pathway activation promotes PLOD2 and LOXL2 gene expression relative to fibrillar collagen expression.
Lung fibroblasts from IPF donors (n = 3 across two independent experiments) were cultured in the presence or absence TGFβ1, DMOG, combined TGFβ1 and DMOG, or vehicle control for 48 hr. (A, B) Relative gene expression of PLOD2 (A) and LOXL2 (B) using the ΔΔCt method. Bars indicate geometric means. Data are mean ± s.d. **p < 0.01; ***p < 0.001; ****p < 0.0001 by Dunnett’s multiple comparisons test. (C) PLOD2 and LOXL2 protein levels. β-actin was used as a loading control. (D) Protein expression of LOXL2 in conditioned media. Ponceau S staining showing total protein levels. The full blots are shown in Figure 4—source data 1. Bars in graph indicate geometric means. Data are mean ± s.d. **p < 0.01; ***p < 0.001; ****p < 0.0001 by Dunnett’s multiple comparisons test. (E, F) Expression of PLOD2 and LOXL2 from (A and B) was divided by COL1A1 expression (shown in Figure 4—figure supplement 2) to calculate proportionate expression changes of cross-linking enzymes relative to collagen fibrillogenesis gene expression. Bars indicate geometric mean. Grouped analysis was performed using Dunnett’s multiple comparison test. * p < 0.05, ** p < 0.01, *** p < 0.001, ****p < 0.0001.
-
Figure 4—source data 1
Full membrane scans for western blot images for Figure 4a and b.
- https://cdn.elifesciences.org/articles/69348/elife-69348-fig4-data1-v1.zip
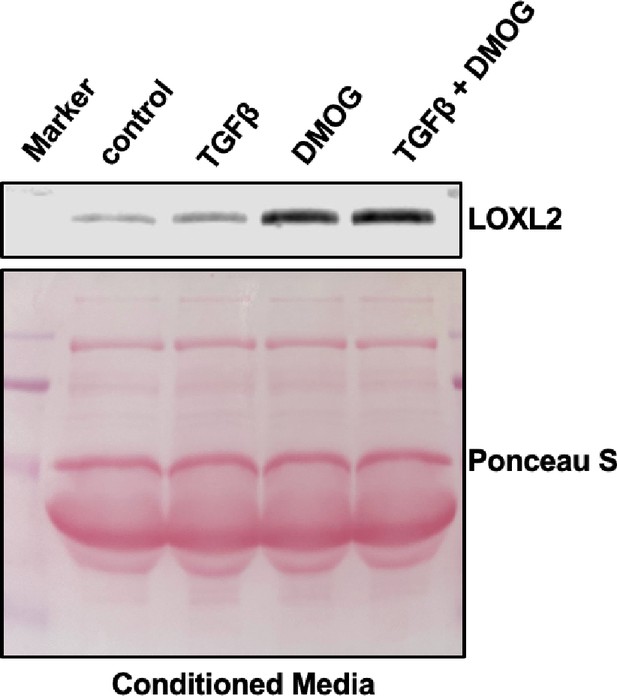
HIF stabilisation increases LOXL2 secretion in control fibroblasts.
Levels of LOXL2 protein in conditioned media from control fibroblasts with the indicated treatment. Ponceau S staining showing total protein levels. The full blots are shown in Figure 4—figure supplement 1—source data 1.
-
Figure 4—figure supplement 1—source data 1
Full membrane scans for western blot images for Figure 4—figure supplement 1.
- https://cdn.elifesciences.org/articles/69348/elife-69348-fig4-figsupp1-data1-v1.zip
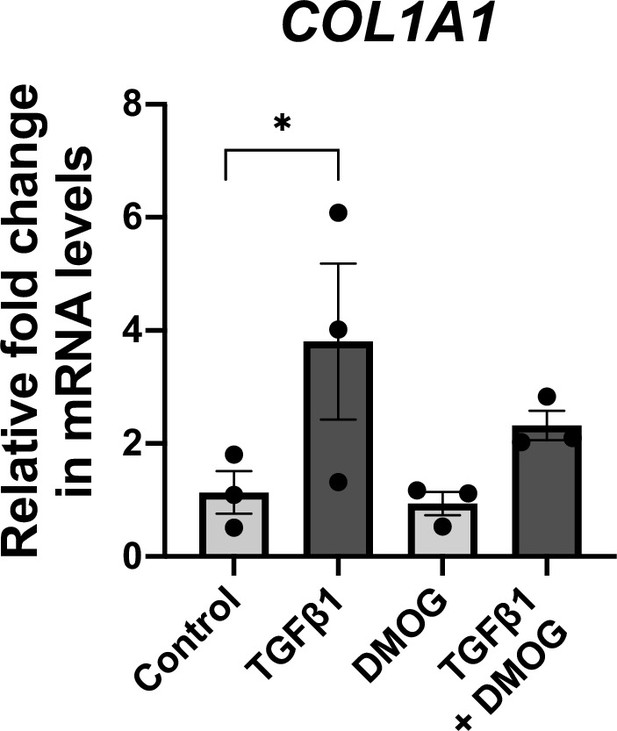
TGFβ1 promotes interstitial collagen gene expression in lung fibroblasts.
Relative gene expression of COL1A1 in lung fibroblasts from IPF donors (n = 3) exposed to 48 hr of control media, TGFβ1, DMOG or combined TGFβ1 and DMOG conditions using the ΔΔCt method. Bars indicate geometric means. *p < 0.05 by Dunnett’s multiple comparisons test.
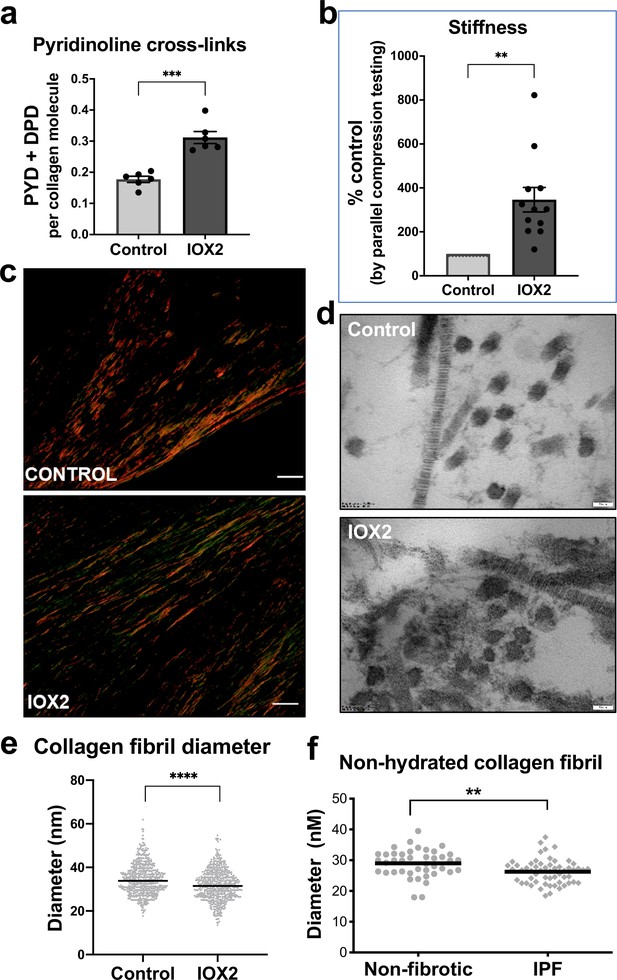
HIF pathway activation promotes pyridinoline cross-linking, alters collagen nano-architecture, and increases tissue stiffness.
Lung fibroblasts from IPF patients (n = 3 donors, two experiments per donor) were used in the 3D model of fibrosis in the presence of IOX2 or vehicle control. Bars indicate geometric mean + s.e.m. Analysis was performed using a Mann-Whitney t-test (two-tailed) **p < 0.01; ***p < 0.001; ****p < 0.0001. (A) Total mature trivalent (PYD+ DPD) collagen cross-links determined by ELISA. n = 6 samples from three IPF donors. (B) Tissue stiffness measured from parallel-plate compression testing (n = 12 samples from three IPF donors) determined by Young’s modulus and represented as proportion of control. (C) Representative images of histological sections of samples stained with picrosirius red and imaged under polarised light. Scale bar 20 μm. (D) Representative electron microscopy images of collagen fibrils within the 3D model of fibrosis. Scale bar 50 nm. (E) Collagen fibril diameter within the 3D model of fibrosis measured in transverse section (300 fibrils for each condition from two IPF donors, measured by a blinded investigator). (F) Atomic force microscopy indentation modulus of collagen fibrils (3–7 fibrils per donor) from control (n = 42 fibrils from eight donors) or IPF lung tissue (n = 57 fibrils from 10 donors) under non-hydrated conditions; each data point represents the mean of 30–50 force-displacement curves per fibril.
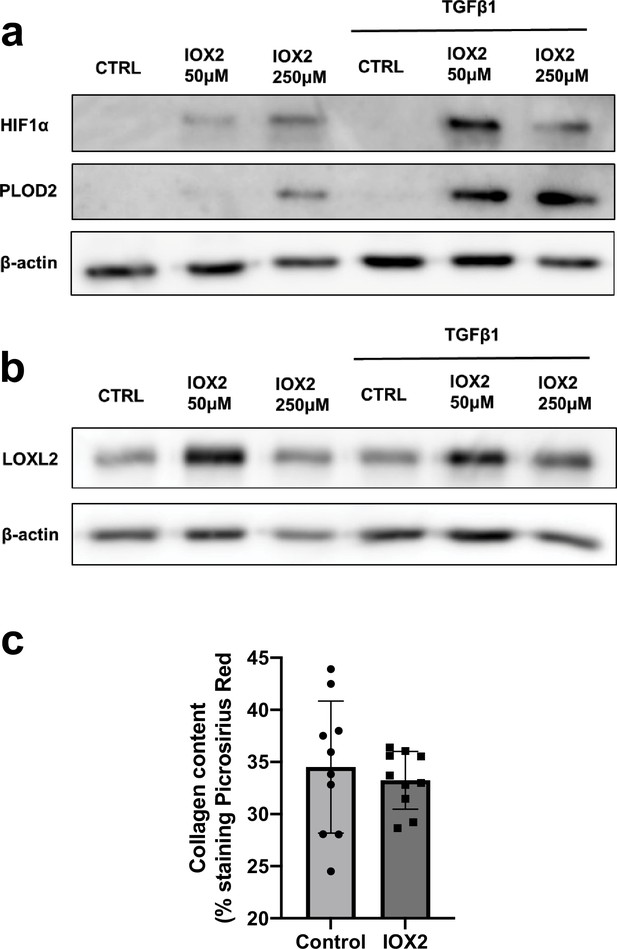
IOX2-mediated HIF pathway activation promotes PLOD2 and LOXL2 expression in the 3D in vitro model of fibrosis.
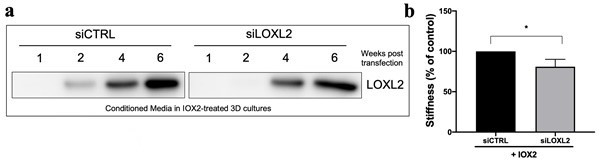
Lung fibroblasts from IPF patients were used in the 3D model of fibrosis in the presence of IOX2 or vehicle control as indicated. Protein expression of (A) HIF1α, PLOD2, and (B) LOXL2 following 2 weeks of culture in the presence or absence of TGFβ1 with or without IOX2 (50 μM or 250 μM) or vehicle control. β-actin loading control. Blots representative of experiments from two separate IPF donors. The full blots are shown in Figure 5—figure supplement 1—source data 1.
-
Figure 5—figure supplement 1—source data 1
Full membrane scans for western blot images for Figure 5—figure supplement 1a, b.
- https://cdn.elifesciences.org/articles/69348/elife-69348-fig5-figsupp1-data1-v1.zip
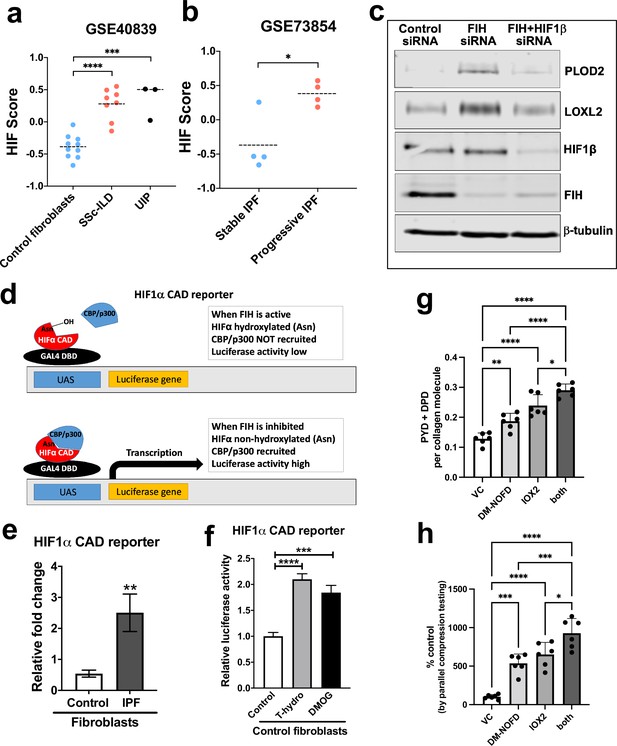
Pseudohypoxia and loss of FIH activity promotes HIF pathway signalling in IPF fibroblasts and increases tissue stiffness.
(A) HIF GSVA scores calculated in human lung fibroblasts derived from control or patients with interstitial lung disease (scleroderma lung or a usual interstitial pneumonia / IPF pattern) (GSE40839). Data are mean ± s.d. ***p < 0.001; ****p < 0.0001 by Dunnett’s multiple comparisons test. (B) HIF GSVA scores calculated in human bronchoalveolar lavage derived mesenchymal stromal cells from patients with stable and progressive IPF (GSE73854). Data are mean ± s.d. *p < 0.05 by the unpaired t test. (C) PLOD2, LOXL2, HIF1β, FIH, and β-tubulin protein levels in lung fibroblasts from patients with IPF transfected with indicated siRNA. β-tubulin was used as a loading control. The full blots are shown in Figure 6—source data 1. (D) Diagram explaining the HIF1α CAD reporter assay in E and F. In brief, the FIH asparaginyl hydroxylase hydroxylates HIF1α CAD, inhibiting its binding with CBP/p300 and decreasing luciferase activity. When FIH is inhibited, the non-hydroxylated HIF1α CAD can bind with CBP/p300 increasing luciferase activity. (E) HIF1α CAD reporter assays in normal human lung fibroblasts (control fibroblasts) or IPF lung fibroblasts (IPF fibroblasts). Values represent the relative fold increase of firefly luciferase in relation to Renilla luciferase, normalised against control (1.0). Data are mean ± s.d. n = 3 samples per group. **p < 0.01 by unpaired t test. (F) HIF1α CAD reporter assays in control fibroblasts with indicated treatment (hydrogen peroxide (T-hydro), DMOG, or vehicle control). Values represent relative fold of firefly luciferase in relation to Renilla luciferase, normalised against control (1.0). Data are mean ± s.d. n = 3 samples per group. (G and H) Control lung fibroblasts (n = 3 donors, two experiments per donor) were used in the 3D model of fibrosis in the presence of IOX2 and/or DM-NOFD or vehicle control as indicated. (G) Total mature trivalent (PYD+ DPD) collagen cross-links determined by ELISA. n = 6 samples from three donors. (H) Tissue stiffness measured from parallel-plate compression testing (n = 6 samples from three donors) determined by Young’s modulus and represented as proportion of control. * p < 0.05, ** p < 0.01, *** p < 0.001, ****p < 0.0001 by Dunnett’s multiple comparisons test.
-
Figure 6—source data 1
Full membrane scans for western blot images for Figure 6.
- https://cdn.elifesciences.org/articles/69348/elife-69348-fig6-data1-v1.zip
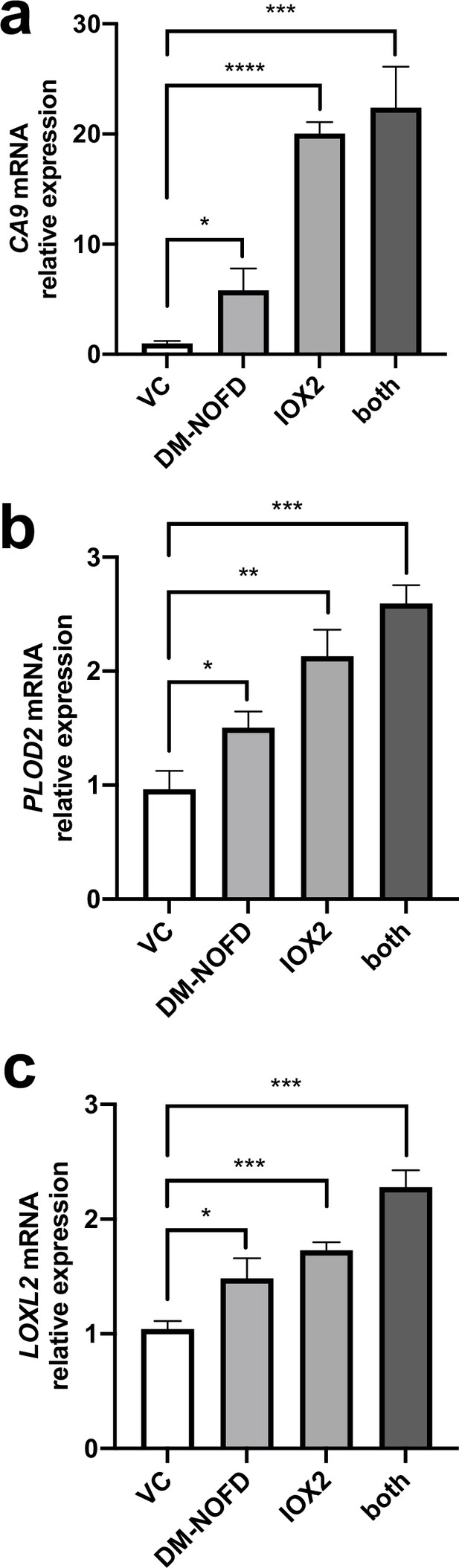
Pseudohypoxia and loss of FIH activity promotes HIF pathway signalling and increases LOXL2 and PLOD2 expression.
Control lung fibroblasts (n = 3 donors, two experiments per donor) were used in the 3D model of fibrosis in the presence of IOX2 and/or DM-NOFD or vehicle control as indicated. Fold change in mRNA levels of CA9 (A), LOXL2 (B) and PLOD2 (C) in the 3D model of fibrosis with indicated treatment. Bars indicate geometric means. Data are mean ± s.d. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001 by Dunnett’s multiple comparisons test.
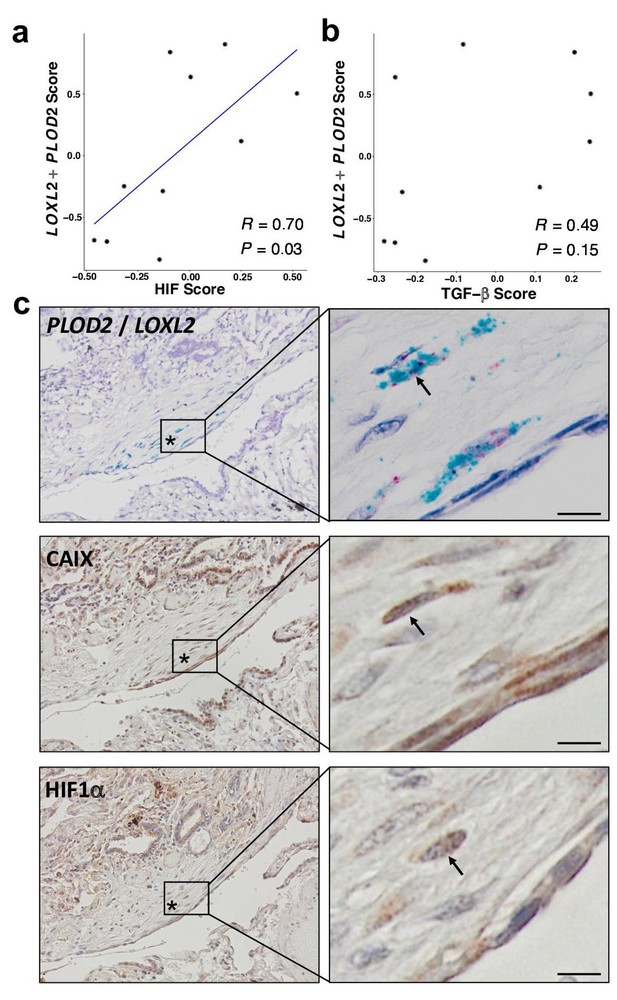
HIF pathway activation localises in areas of active fibrogenesis to cells co-expressing LOXL2 and PLOD2.
(A–B) Scatterplots showing correlations between LOXL2/PLOD2 expression and HIF scores (A) or TGFβ scores (B) in IPF fibroblast foci (n = 10) using the Spearman rank correlation coefficient. (C) Representative images of serial sections of lung tissue from patients with IPF (n = 3). mRNA expression of PLOD2 (red chromagen) and LOXL2 (green chromagen) using RNAscope RNA in-situ hybridisation with immunohistochemical staining for Carbonic anhydrase IX (CA-IX) and HIF1α using DAB (brown). A fibroblastic focus is identified by *. Scale bar 20 μm.
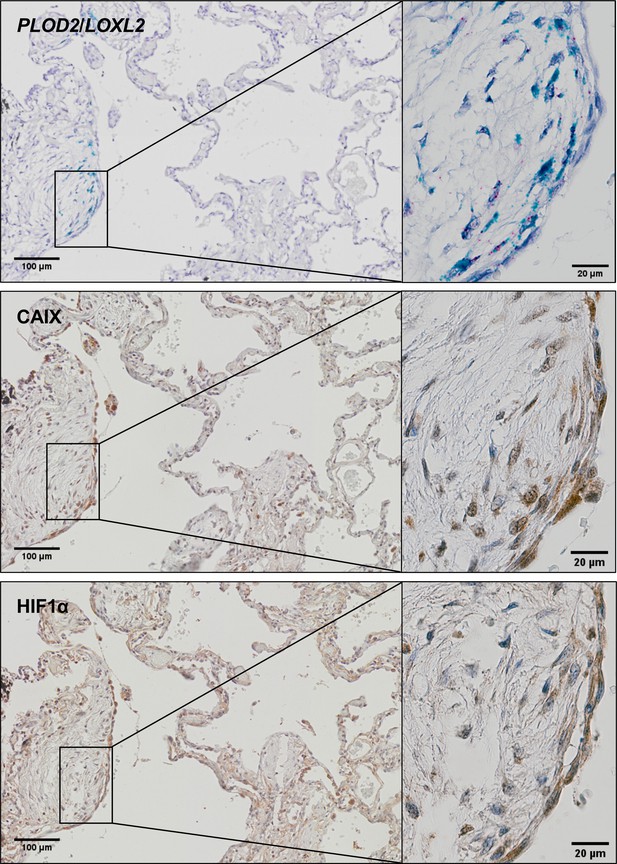
HIF pathway activation localises in areas of active fibrogenesis to cells co-expressing LOXL2 and PLOD2.
Representative images of serial sections of lung tissue from patients with IPF (n = 3). mRNA expression of PLOD2 (red chromagen) and LOXL2 (green chromagen) using RNAscope RNA in-situ hybridisation with immunohistochemical staining for Carbonic anhydrase IX (CA-IX) and HIF1α using DAB (brown). Scale bars are 100 μm with inserts 20 μm.
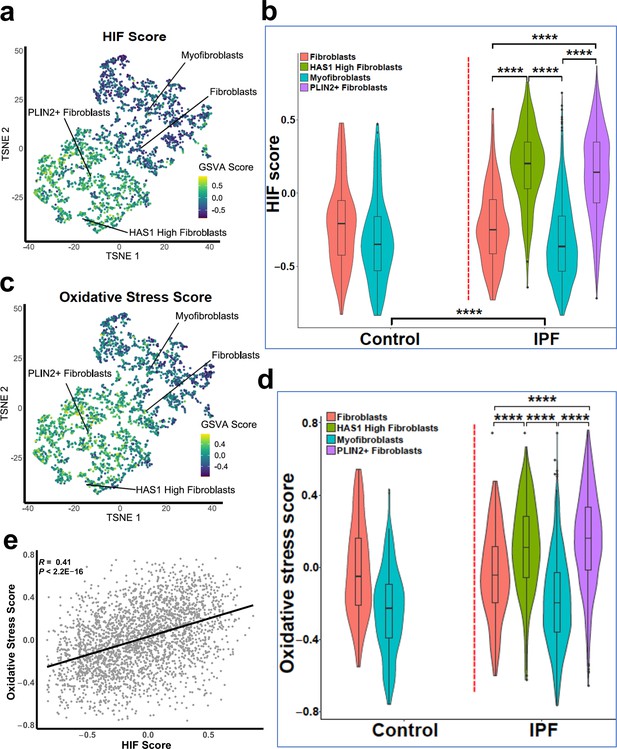
Gene set variance analysis of single-cell RNAseq fibroblast populations identifies co-enrichment of HIF score and oxidative stress genes.
(A) HIF score GSVA in control and IPF fibroblasts sequenced by single-cell RNAseq (GSE135893). Colours correspond to calculated GSVA score for each cell. (B) Plot of mean HIF GSVA scores for each fibroblast type in control and IPF fibroblast cell populations and compared using Dunnett’s multiple comparison test, ****p < 0.0001. (C) GSVA scores for genes upregulated in IPF in this dataset associated with the Gene Set: HALLMARK_REACTIVE_OXYGEN_SPECIES_PATHWAY (M5938). (D) Plot of upregulated oxidative stress GSVA scores for each fibroblast type in control and IPF cells. (E) Correlation plot of HIF score vs upregulated oxidative stress GSVA score for single cell RNAseq data. Correlation coefficient is Pearson’s product-moment coefficient.
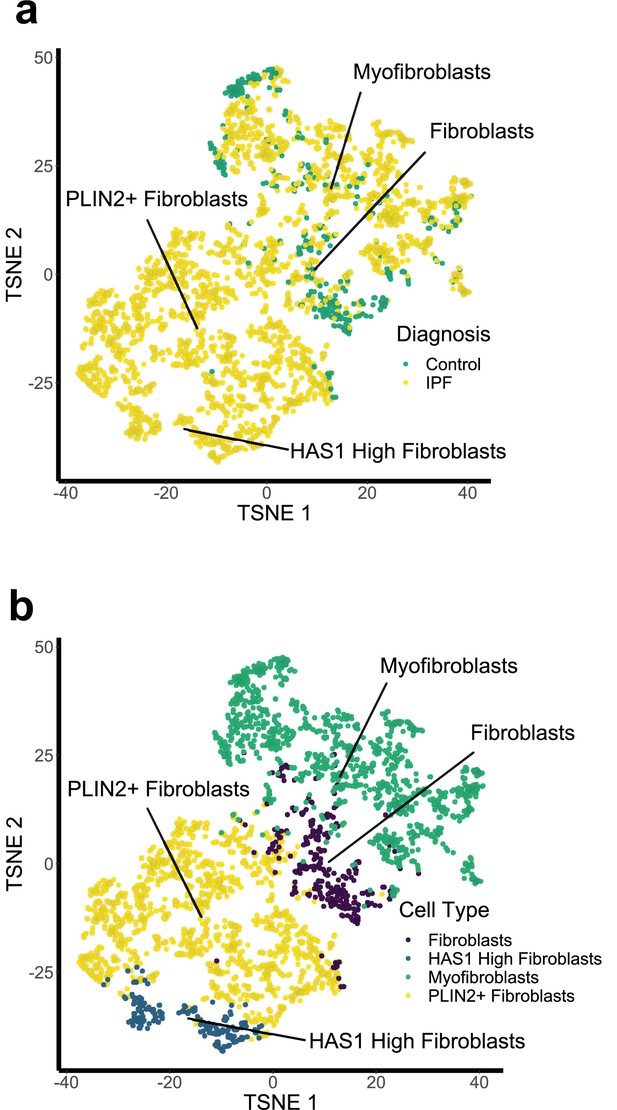
Fibroblast populations identified within a single-cell RNA sequencing dataset.
(A) t-stochastic nearest neighbour embedding (t-SNE) of single cell sequencing data (GSE135893) showing clustering of different lung fibroblast types. (B) t-SNE plot of single cell fibroblast data showing diagnosis of the patients of origin for each fibroblast.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| transfected construct (human) | GAL4DBD-HIF1αCAD (residues 652–826) for HIF1α CAD reporter assay | Ratcliffe lab (University of Oxford) Coleman et al., 2007 | ||
| transfected construct (Homo-sapiens) | UAS-luc reporter for HIF1α CAD reporter assay | Ratcliffe lab (University of Oxford) Coleman et al., 2007 | ||
| transfected construct (Homo-sapiens) | Plasmid for Dual-Luciferase Reporter Assay | Promega | phRL-CMV | |
| transfected construct (Homo-sapiens) | siRNA to human HIF1AN (FIH) | Dharmacon/ Thermo Fisher Scientific | MU-004073-02-0002 | |
| transfected construct (Homo-sapiens) | siRNA to human HIF1A (HIF1α) | Dharmacon/ Thermo Fisher Scientific | MU-00401805-05-0002 | |
| transfected construct (Homo-sapiens) | siRNA to human EPAS1(HIF2α) | Dharmacon/ Thermo Fisher Scientific | MU-004814-01-0002 | |
| transfected construct (Homo-sapiens) | siRNA to human ARNT (HIF1β) | Dharmacon/ Thermo Fisher Scientific | MU-007207-01-0002 | |
| transfected construct (Homo-sapiens) | siGENOME RISC-Free | Dharmacon/ Thermo Fisher Scientific | D-001220-01-05 | |
| Antibody | Anti-CAIX (Rabbit polyclonal) | Novus Biologicals | Cat. #: NB100-417 | IHC 1:500 |
| Antibody | Anti-HIF1A (Rabbit polyclonal) | Cayman Chemical | Cat. #: 10006421 | IHC 1:500 |
| antibody | anti-human HIF1α (Mouse polyclonal IgG1k) | BD Biosciences | Cat #:610,958 | WB (1:1000) |
| antibody | Anti-HIF1β (Rabbit polyclonal) | Cell Signaling Technology | Cat #:5,537 | WB (1:1000) |
| antibody | Anti-phospho- Smad2 (Rabbit polyclonal) | Cell Signaling Technology | Cat #: 3,104 | WB (1:1000) |
| antibody | anti-β-tubulin (Mouse polyclonal) | Cell Signaling Technology | Cat #: 86,298 | WB (1:1000) |
| antibody | anti-PLOD2 (Mouse monoclonal IgG2B) | R&D Systems | Cat #: MAB4445 | WB (1:500) |
| antibody | Anti- human LOXL2 (Goat polyclonal) | R&D Systems | Cat #: AF2639 | WB (1:1000)IF (1:100) |
| antibody | anti-human FIH (Mouse monoclonal 162 C) | Ratcliffe lab (University of Oxford) Stolze et al., 2004 | WB (1:200) | |
| Antibody | Anti-P-ERK (polyclonal rabbit Thr202/Tyr204) | Cell Signalling Technology | Cat #: 9,101 | WB (1:1000) |
| Antibody | Anti-P-SMAD2/3 (Rabbit polyclonal Ser465/467) | Cell Signalling Technology | Cat #: 8,828 | WB (1:1000) |
| Antibody | IRDye 800CW Donkey anti-Goat IgG Secondary Antibody | LI-COR Biosciences | Cat #: 926–32214 | WB (1:5000) |
| Antibody | IRDye 800CW Goat anti-Rabbit IgG Secondary Antibody | LI-COR Biosciences | Cat #: 926–32211 | WB (1:5000) |
| Antibody | IRDye 680LT Goat anti-Mouse IgG Secondary Antibody | LI-COR Biosciences | Cat #: 926–68020 | WB (1:5000) |
| Antibody | Anti-non-phospho (active) β-catenin (Rabbit monoclonal IgG) | Cell Signalling Technology | Cat. #: 8,814 S | WB (1:1000) |
| Antibody | Anti-mouse IgG HRP-linked whole antibody | Life Sciences | Cat. #: NXA931 | WB (1:1000) |
| Antibody | Anti-goat Immunoglobulins/ HRP (affinity isolated) | Dako | Cat. #: P0449 | WB (1:1000) |
| Antibody | Anti-goat IgG H&L (Alexa Fluor 647) | Abcam | Cat. #: Ab150131 | ICC 1:100 |
| sequence-based reagent | Human HIF1A (HIF1α) | Qiagen | QuantiTect PCR primersCat #: QT00083664 | |
| sequence-based reagent | Human EPAS1 (HIF2α) | Qiagen | QuantiTect PCR primersCat #: QT00069587 | |
| sequence-based reagent | Human ARNT (HIF1β) | Qiagen | QuantiTect PCR primersCat #: QT00023177 | |
| sequence-based reagent | Human ACTB(β-actin) | Qiagen | QuantiTect PCR primersCat #: QT01680476 | |
| Sequence-based reagent | LOXL2 | Primer Design | ||
| Sequence-based reagent | PLOD2 | Primer Design | ||
| Sequence-based reagent | COL1A1 | Primer Design | ||
| Sequence-based reagent | COL3A1 | Primer Design | ||
| Peptide, recombinant protein | Recombinant Human TGF- beta 1 Protein | R&D Systems | Cat. #: 240-B-010 | |
| Peptide, recombinant protein | Recombinant Human EGF GMP Protein | R&D Systems | Cat. #: 236-GMP-200 | |
| Commercial assay or kit | RNAscope 2.5 HD Duplex Assay | Advanced Cell Diagnostics | Cat. #: 322,430 | |
| Commercial assay or kit | RNAscope probe- Hs-LOXL2-C1 | Advanced Cell Diagnostics | Cat. #: 311,341 | |
| Commercial assay or kit | RNAscope probe- Hs-PLOD2-C2 | Advanced Cell Diagnostics | Cat. #: 547761-C2 | |
| Commercial assay or kit | MicroVue Bone PYD EIA | Quidel | Cat. #: 8,010 | |
| Commercial assay or kit | Hydroxyproline Assay Kit | Merck | Cat. #: MAK008 | |
| Commercial assay or kit | Total Protein Assay | QuickZyme Biosciences | Cat. #: QZBtotprot | |
| Commercial assay or kit | Picro Sirius Red Stain Kit (Connective Tissue Stain) | Abcam | Cat. #: Ab150681 | |
| commercial assay or kit | Lipofectamine 3,000 | Thermo Fisher Scientific | Cat. #: L3000008 | |
| commercial assay or kit | Lipofectamine RNAiMAX | Thermo Fisher Scientific | Cat. #: 13778–075 | |
| commercial assay or kit | Dual-Luciferase Reporter Assay System | Promega | Cat. #: E1910 | |
| commercial assay or kit | QuantiNova SYBR Green RT-PCR kits | Qiagen | Cat. #: 208052 | |
| chemical compound, drug | Dimethyloxaloylglycine (DMOG) | Sigma Aldrich | Cat #: D3695CAS: 89464-63-1 | |
| chemical compound, drug | N-[[1,2-Dihydro-4-hydroxy-2-oxo-1-(phenylmethyl)–3-quinolinyl]carbonyl]-glycine (IOX2) | Selleck Chemicals | Cat #: S2919CAS: 931398-72-0 | |
| chemical compound, drug | Dimethyl N-oxalyl-D-phenylalanine (DM-NOFD) | Schofield lab (University of Oxford) McDonough et al., 2005 | ||
| chemical compound, drug | DMSO | Sigma Aldrich | Cat #: 276,855CAS: 67-68-5 | |
| chemical compound, drug | T-hydro (tert-butyl hydroperoxide) | Sigma Aldrich | Cat #: 19,999CAS: 75-91-2 | 20 µM (fresh prepared) |
Additional files
-
Supplementary file 1
Semiquantitative analysis of mRNA expression & Fibroblast demographic details.
(a) Semiquantitative analysis of LOX2 and PLOD2 mRNA expression identified by RNAscope in situ hybridization in cell subtypes in IPF lung tissue (n = 7 donors). FF, fibroblast focus. (b) Fibroblast donor demographic details.
- https://cdn.elifesciences.org/articles/69348/elife-69348-supp1-v1.docx
-
Supplementary file 2
Short interfering RNA (siRNA) oligo sequences.
- https://cdn.elifesciences.org/articles/69348/elife-69348-supp2-v1.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/69348/elife-69348-transrepform1-v1.pdf
-
Source code 1
Source code for RNAseq analyses for Figure 6a, b.
- https://cdn.elifesciences.org/articles/69348/elife-69348-code1-v1.zip
-
Source code 2
Source code for RNAseq analyses for Figure 7a, b.
- https://cdn.elifesciences.org/articles/69348/elife-69348-code2-v1.zip
-
Source code 3
Source code for RNAseq analyses for Figure 8 and Figure 8—figure supplement 1.
- https://cdn.elifesciences.org/articles/69348/elife-69348-code3-v1.zip