BET family members Bdf1/2 modulate global transcription initiation and elongation in Saccharomyces cerevisiae
Figures
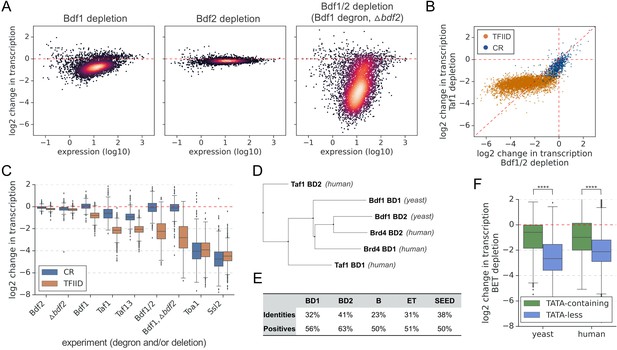
Bdf1/2 are critical for transcription of TFIID-dependent genes and share significant similarities with human bromodomain and extra-terminal domain (BET) factors.
(A) Transcriptional changes caused by degron depletion of Bdf1/2 for 30 min. Scatter plots comparing gene expression level (in log scale) with log2 change in transcription per gene measured by 4-thioU RNA-seq. Mean values from replicate experiments are plotted for 5313 genes with detectable signals in all RNA-seq samples collected in this work. (B) Scatter plot comparing log2 change in transcription after depleting Bdf1/2 (this work) or Taf1 (Donczew et al., 2020). Data for 4883 genes previously classified into TFIID-dependent and coactivator-redundant (CR) categories are shown (Donczew et al., 2020). (C) Boxplot showing log2 change in transcription for 4883 genes measured by 4-thioU RNA-seq after depleting indicated factors. Genes are grouped into TFIID-dependent and CR categories. (D) Phylogenetic analysis of amino acid sequences of individual bromodomains from human Taf1, human Brd4, and yeast Bdf1. (E) BLAST global alignment of amino acid sequences within indicated domains of Bdf1 and Brd4. (F) Boxplot showing log2 change in transcription after depleting BET factors in yeast (this work) or human cells (Winter et al., 2017). Genes are classified depending on the presence of a consensus TATA box (TATAWAW) in their promoter. A list of TATA-containing promoters in human cells was obtained from Eukaryotic Promoter Database (EPD). A single, most representative promoter per gene was used in this analysis. Results of the Welch’s t-test are shown. The asterisks represent p-value with the following cutoff levels: ****0.0001, *** 0.001, ** 0.01, * 0.05, ns = > 0.05. See also Supplementary files 1 and 2.
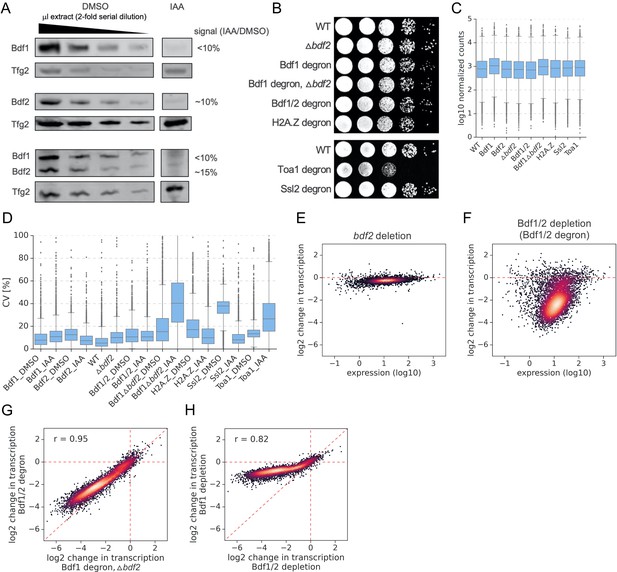
Bdf1/2 are critical for transcription of TFIID-dependent genes.
(A) Western blot analysis of degradation of Bdf1 and/or Bdf2 after 30 min of indole-3-acetic acid (IAA) treatment. Variable amounts of the untreated samples were loaded to allow quantitation of degron efficiency. Blots were probed with α-V5 monoclonal (Bdf1, Bdf2) or control α-Tfg2 polyclonal antibodies. (B) Spot assay comparing the fitness of indicated strains grown at 30°C. A 10-fold serial dilution was used. (C) Boxplot of average gene expression level in indicated experiments following dimethyl sulfoxide (DMSO) treatment. Log10 scale is used for the Y axis. (D) Boxplot showing coefficients of variation (CV) for all 5313 genes with detectable signals in all 4-thioU RNA-seq experiments. Samples were collected in two (Bdf1, Bdf2, Δbdf2, Bdf1Δbdf2, H2A.Z) or three (Bdf1/2, Ssl2, Toa1) biological replicates. CV values were calculated for each gene based on normalized read counts for replicate experiments. (E, F) Scatter plots comparing gene expression level (in log scale) with log2 change in transcription per gene in indicated 4-thioU RNA-seq experiments. Mean values from replicate experiments for 5313 genes with detectable signals in all RNA-seq samples collected in this work are plotted. (G) Scatter plot comparing log2 change in transcription after depleting Bdf1/2 using two different approaches as indicated. Spearman correlation coefficient (r) is shown. (H) Scatter plot comparing log2 change in transcription at 4883 genes after depleting Bdf1/2 or Bdf1. Spearman correlation coefficient (r) is shown.
-
Figure 1—figure supplement 1—source data 1
Original, unedited image of a western blot for the Bdf1 depletion experiment (A).
The blot was scanned at 700/800 nm wavelengths using Li-Cor Odyssey CLx imager. The image shows results obtained for 700 nm channel.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig1-figsupp1-data1-v2.pdf
-
Figure 1—figure supplement 1—source data 2
Original, unedited image of a western blot for the Bdf1 depletion experiment (A).
The blot was scanned at 700/800 nm wavelengths using Li-Cor Odyssey CLx imager. The image shows results obtained for 800 nm channel.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig1-figsupp1-data2-v2.pdf
-
Figure 1—figure supplement 1—source data 3
Original, unedited images of a western blot for the Bdf2 depletion experiment (A).
The blot was scanned at 700/800 nm wavelengths using Li-Cor Odyssey CLx imager. The image shows results obtained for both channels.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig1-figsupp1-data3-v2.pdf
-
Figure 1—figure supplement 1—source data 4
Original, unedited images of a western blot for the Bdf1/2 depletion experiment (A).
The blot was scanned at 700/800 nm wavelengths using Li-Cor Odyssey CLx imager. The image shows results obtained for both channels.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig1-figsupp1-data4-v2.pdf
-
Figure 1—figure supplement 1—source data 5
Uncropped images of western blots shown in (A) with the relevant cropped bands marked with red rectangles.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig1-figsupp1-data5-v2.pdf
-
Figure 1—figure supplement 1—source data 6
Original image of a YPD plate for spot assay analysis (B).
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig1-figsupp1-data6-v2.pdf
-
Figure 1—figure supplement 1—source data 7
Original image of a YPD plate for spot assay analysis (B).
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig1-figsupp1-data7-v2.pdf
-
Figure 1—figure supplement 1—source data 8
Uncropped images of plates shown in (B) with the relevant cropped area marked with a red rectangle.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig1-figsupp1-data8-v2.pdf
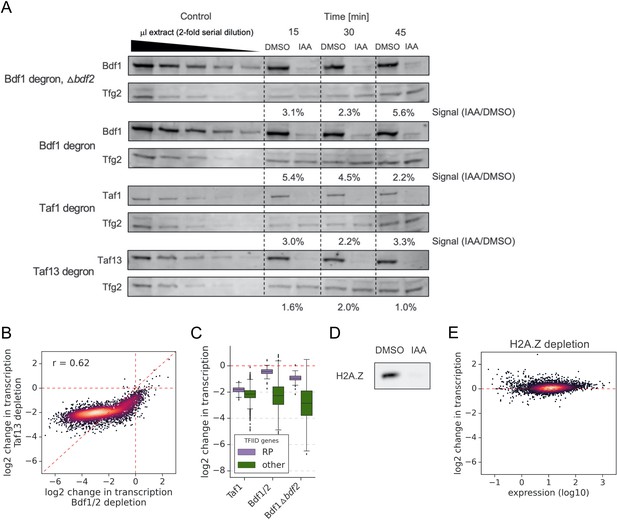
Bdf1 and Tafs are degraded with similar efficiency.
(A) Western blot analysis comparing degradation of Bdf1, Taf1, and Taf13 after 15, 30, and 45 min of indole-3-acetic acid (IAA) treatment. Variable amounts of the sample collected at 0 time point (control) were loaded to allow quantitation. Blots were probed with α-V5 monoclonal (Bdf1, Taf1, and Taf13) or control α-Tfg2 polyclonal antibodies. (B) Scatter plot comparing log2 change in transcription at 4883 genes after depleting Bdf1/2 or Taf13. Spearman correlation coefficient (r) is shown. (C) Boxplot comparing log2 change in transcription at ribosomal protein (RP) genes with the rest of the TFIID-dependent genes in the indicated degron experiments. (D) Western blot analysis of IAA-induced degradation of H2A.Z. Blot was probed with α-V5 monoclonal antibody. (E) Scatter plot comparing expression level (in log scale) with log2 change in transcription per gene after depleting H2A.Z. Data are shown for 5313 genes with detectable signals in all RNA-seq samples collected in this work.
-
Figure 1—figure supplement 2—source data 1
Original, unedited image of a western blot for the experiment comparing the degradation of Bdf1, Taf1, and Taf13 (A).
The blot was scanned at 700/800 nm wavelengths using Li-Cor Odyssey CLx imager. The image shows results obtained for 700 nm channel.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig1-figsupp2-data1-v2.pdf
-
Figure 1—figure supplement 2—source data 2
Original, unedited image of a western blot for the experiment comparing the degradation of Bdf1, Taf1, and Taf13 (A).
The blot was scanned at 700/800 nm wavelengths using Li-Cor Odyssey CLx imager. The image shows results obtained for 800 nm channel.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig1-figsupp2-data2-v2.pdf
-
Figure 1—figure supplement 2—source data 3
Uncropped images of western blots shown in (A) with the relevant cropped bands marked with red rectangles.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig1-figsupp2-data3-v2.pdf
-
Figure 1—figure supplement 2—source data 4
Original, unedited image of a western blot for the H2A.Z depletion experiment (D).
The blot was scanned at 700 nm wavelength using Li-Cor Odyssey CLx imager.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig1-figsupp2-data4-v2.pdf
-
Figure 1—figure supplement 2—source data 5
Uncropped image of a western blot shown in (D) with the relevant cropped bands marked with a red rectangle.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig1-figsupp2-data5-v2.pdf
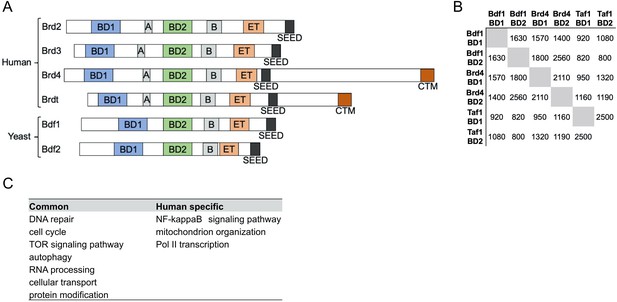
Bdf1/2 share significant similarities with human bromodomain and extra-terminal domain (BET) factors.
(A) Conserved domain organization of human and yeast BET family members. Adapted from Wu and Chiang, 2007. (B) Pairwise alignment scores between amino acid sequences of individual bromodomains of Bdf1, Brd4, and human Taf1 obtained using Clustal Omega (Sievers et al., 2011). (C) Major gene ontology terms enriched among the 25% most BET-sensitive genes in yeast and human cells.
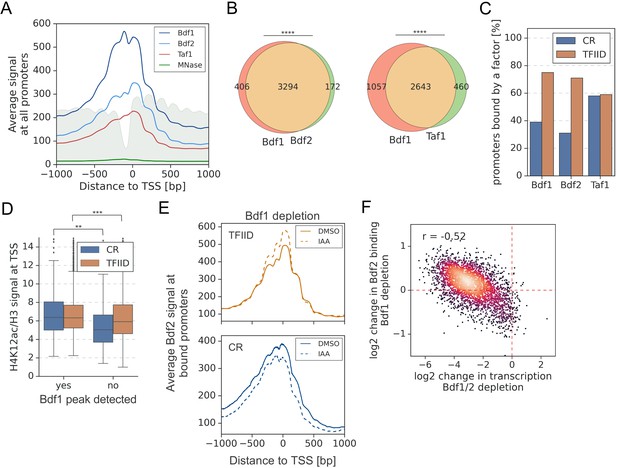
A significant overlap between Bdf1/2 and Taf1 bound promoters.
(A) Average plot of Bdf1, Bdf2, and Taf1 ChEC-seq signals versus free MNase signal at 5888 yeast promoters. Published MNase-seq data are shown as a gray area plot (Oberbeckmann et al., 2019). Mean values from replicate experiments are plotted. (B) Venn diagrams showing the overlap of promoters bound by indicated factors. Results of the hypergeometric test are shown. The asterisks represent p-value with the following cutoff levels: **** 0.0001, ***0.001, ** 0.01, * 0.05, ns = >0.05. (C) Bar plot showing percentage of promoters in each class (TFIID-dependent or coactivator-redundant [CR]) bound by a given factor. (D) Boxplot showing the H4K12ac signal at transcription start site (TSS) normalized to H3 signal. Genes are grouped into TFIID-dependent and CR categories, and data are plotted separately depending on the presence of a significant Bdf1 peak. Signals were calculated in a −100 to +200 bp window relative to TSS. Published H3 ChIP-seq dataset was used for this analysis (Bruzzone et al., 2018). Results of the Welch’s t-test are shown. (E) Average plot comparing Bdf2 ChEC-seq signal before (dimethyl sulfoxide [DMSO], solid line) and after (indole-3-acetic acid [IAA], dashed line) Bdf1 depletion at 3223 promoters bound by Bdf2 and classified into TFIID-dependent and CR categories. (F) Scatter plot comparing log2 change in transcription and log2 change in Bdf2 ChEC-seq occupancy after depleting Bdf1/2 or Bdf1, respectively. Spearman correlation coefficient (r) is shown. Bdf2 signal was calculated in a 200 bp window centered on a dominant peak assigned to 3223 promoters. See also Supplementary file 3.
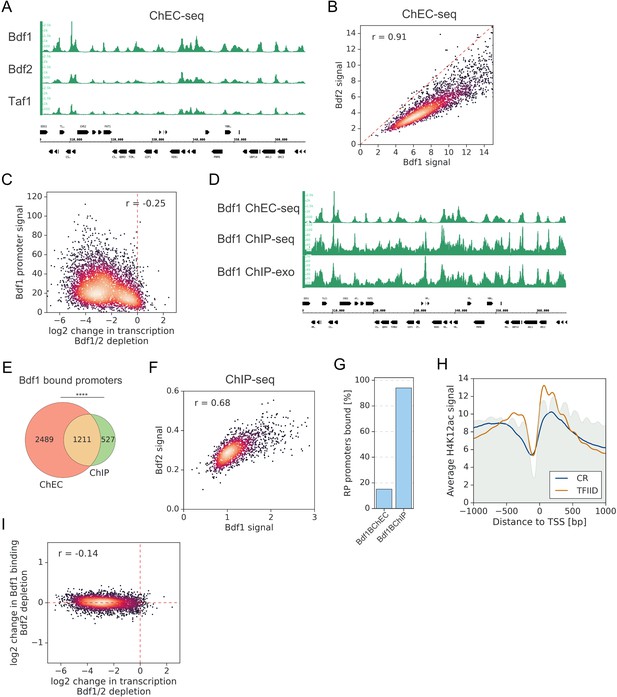
A significant overlap between Bdf1/2 and Taf1 bound promoters.
(A) Genome browser image showing comparison of Bdf1, Bdf2, and Taf1 ChEC-seq signals at a representative genomic location. (B) Scatter plot comparing Bdf1 and Bdf2 ChEC-seq signals at 3294 promoters bound by both factors. Spearman correlation coefficient (r) is shown. (C) Scatter plot comparing log2 change in transcription after depleting Bdf1/2 and the Bdf1 promoter ChEC-seq signal for 4883 genes analyzed with 4-thioU RNA-seq. Spearman correlation coefficient (r) is shown. Bdf1 signal was calculated in a −300 to 200 bp window relative to transcription start site (TSS). (D) Genome browser image showing comparison of Bdf1 ChEC-seq, ChIP-seq (this work), and ChIP-exo signal (Rhee and Pugh, 2012). The same genomic location as in (A) is shown. (E) Venn diagram showing the overlap of promoters bound by Bdf1 as detected by ChEC-seq and ChIP-seq. Results of the hypergeometric test are shown. The asterisks represent p-value with the following cutoff levels: ****0.0001, ***0.001, **0.01, *0.05, ns = > 0.05. (F) Scatter plot comparing Bdf1 and Bdf2 ChIP-seq signals at individual promoters bound by both factors. Spearman correlation coefficient (r) is shown. 1738 Bdf1 bound promoters identified by ChIP-seq were used for this analysis. Signals were calculated in a −100 to 200 bp window relative to TSS. (G) Barplot showing the percentage of ribosomal protein (RP) gene promoters bound by Bdf1 as detected by ChEC-seq or ChIP-seq. (H) Average plot of H4K12ac ChIP-seq signal at 4900 TFIID-dependent or coactivator-redundant (CR) promoters. Published MNase-seq dataset is shown as a gray area plot (Oberbeckmann et al., 2019). (I) Scatter plot comparing log2 change in transcription and log2 change in Bdf1 ChEC-seq occupancy after depleting Bdf1/2 or Bdf2, respectively. Spearman correlation coefficient (r) is shown. Bdf1 signal was calculated in a 200 bp window centered on a dominant peak assigned to 342d6 promoters bound by Bdf1 and classified into TFIID-dependent and CR categories.
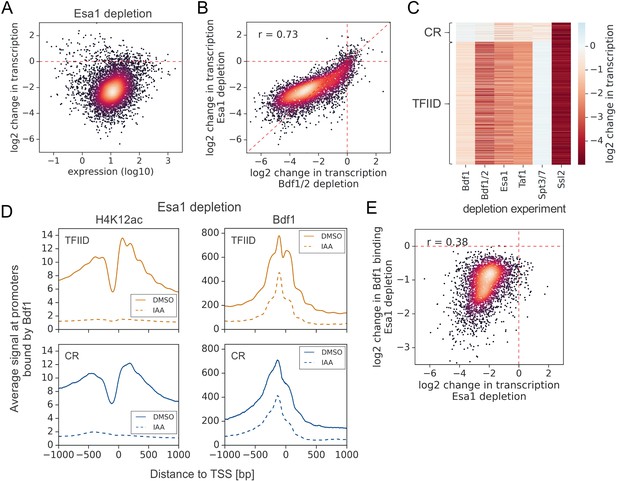
Esa1 regulates similar set of genes as Bdf1/2 but is not essential for Bdf1 targeting to chromatin.
(A) Scatter plot comparing gene expression level (in log scale) with log2 change in transcription per gene after depleting Esa1 for 60 min. Mean values from replicate experiments are plotted for 5313 genes with detectable signals in all RNA-seq samples collected in this work. (B) Scatter plot comparing log2 change in transcription after depleting Bdf1/2 or Esa1. Spearman correlation coefficient (r) is shown. (C) Heatmap comparing results of indicated depletion experiments on transcription of 4883 genes. (D) Average plots comparing H4K12ac ChIP-seq and Bdf1 ChEC-seq signals before (dimethyl sulfoxide [DMSO], solid line) and after (indole-3-acetic acid [IAA], dashed line) Esa1 depletion at 3426 promoters bound by Bdf1 and classified into TFIID-dependent and coactivator-redundant (CR) categories. Mean values from replicate experiments are plotted. (E) Scatter plot comparing log2 change in transcription and log2 change in Bdf1 occupancy after depleting Esa1 at 3426 promoters. Spearman correlation coefficient (r) is shown. Bdf1 signal was calculated in a 200 bp window centered on a dominant peak. See also Supplementary file 3.
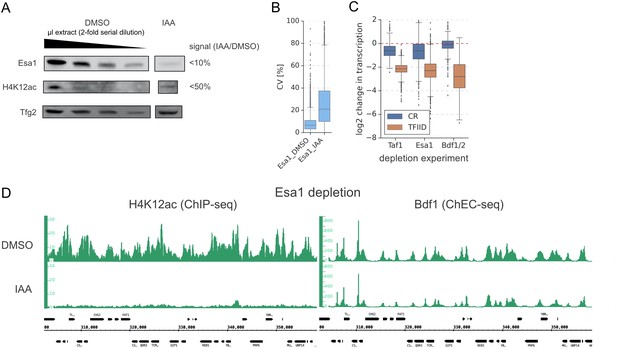
Esa1 regulates similar set of genes as Bdf1/2 but is not essential for Bdf1 targeting to chromatin.
(A) Western blot analysis of degradation of Esa1 and a corresponding change in H4K12ac levels after 60 min of indole-3-acetic acid (IAA) treatment. Variable amounts of the untreated samples were loaded to allow quantitation. Blots were probed with α-V5 monoclonal (Esa1), α-H4K12ac monoclonal, and control α-Tfg2 polyclonal antibodies. (B) Boxplot showing coefficients of variation (CV) for the Esa1 4-thioU RNA-seq experiment. Two biological replicates were collected. CV values were calculated for each gene based on normalized read counts for replicate experiments. (C) Boxplot showing log2 change in transcription at 4883 genes in indicated 4-thioU RNA-seq experiments. Genes are grouped into TFIID-dependent and coactivator-redundant (CR) categories. (D) Genome browser images showing H4K12ac ChIP-seq and Bdf1 ChEC-seq signals at a representative genomic location before (dimethyl sulfoxide [DMSO]) and after (IAA) Esa1 depletion.
-
Figure 3—figure supplement 1—source data 1
Original, unedited image of a western blot for the Esa1 depletion experiment (A).
The blot was scanned at 700/800 nm wavelengths using Li-Cor Odyssey CLx imager. The image shows results obtained for 700 nm channel.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig3-figsupp1-data1-v2.pdf
-
Figure 3—figure supplement 1—source data 2
Original, unedited image of a western blot for the Esa1 depletion experiment (A).
The blot was scanned at 700/800 nm wavelengths using Li-Cor Odyssey CLx imager. The image shows results obtained for 800 nm channel.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig3-figsupp1-data2-v2.pdf
-
Figure 3—figure supplement 1—source data 3
Uncropped images of western blots shown in (A) with the relevant cropped bands marked with red rectangles.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig3-figsupp1-data3-v2.pdf
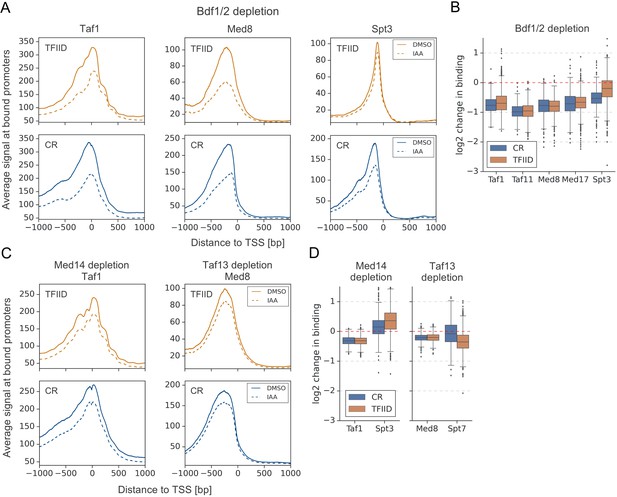
Bdf1/2 participate in recruitment of TFIID and Mediator to chromatin.
(A) Average plots comparing Taf1, Med8, and Spt3 ChEC-seq signals before (dimethyl sulfoxide [DMSO], solid line) and after (indole-3-acetic acid [IAA], dashed line) Bdf1/2 depletion at promoters bound by each factor and classified into TFIID-dependent and coactivator-redundant (CR) categories (2879, 890, and 2526 promoters, respectively). Mean values from replicate experiments are plotted. (B) Boxplot showing log2 change in promoter occupancy of indicated factors after Bdf1/2 depletion. Signals were calculated in a 200 bp window centered on a dominant peak. (C) Average plots comparing Taf1 and Med8 ChEC-seq signals before (DMSO, solid line) and after (IAA, dashed line) Med14 or Taf13 depletion, respectively. Same set of promoters as in (A) was used for this analysis. (D) Boxplot showing log2 change in promoter occupancy of indicated factors after Med14 or Taf13 depletion. Signals were calculated in a 200 bp window centered on a dominant peak. List of Spt3 bound promoters was used to calculate Spt7 occupancy. See also Supplementary file 3.
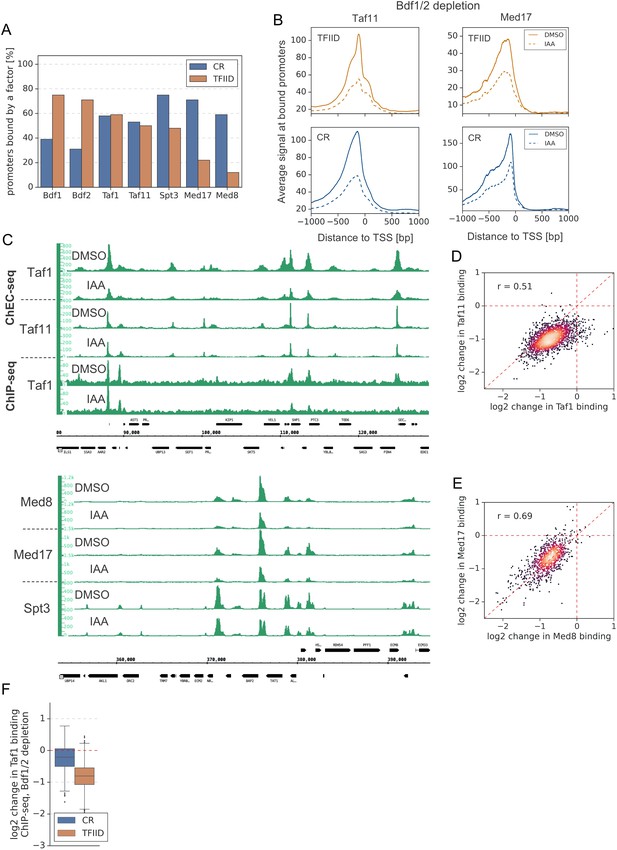
Bdf1/2 participate in recruitment of TFIID and Mediator to chromatin.
(A) Bar plot showing percentage of promoters in each class (TFIID-dependent or coactivator-redundant [CR]) bound by a given factor. (B) Average plots comparing Taf11 and Med17 ChEC-seq signals before (dimethyl sulfoxide [DMSO], solid line) and after (indole-3-acetic acid [IAA], dashed line) Bdf1/2 depletion at promoters bound by each factor and classified into TFIID-dependent and CR categories (2486 and 1414, respectively). Mean values from replicate experiments are plotted. (C) Genome browser images showing Taf1 ChEC, Taf11 ChEC, and Taf1 ChIP (upper panel) or Med8 ChEC, Med17 ChEC, and Spt3 ChEC (lower panel) signals at representative genomic locations before (DMSO) and after (IAA) Bdf1/2 depletion. (D) Scatter plot comparing log2 change in binding of Taf1 and Taf11 after depleting Bdf1/2 at 2070 promoters bound by both Taf1 and Taf11. Spearman correlation coefficient (r) is shown. (E) Scatter plot comparing log2 change in binding of Med8 and Med17 after depleting Bdf1/2 at 853 promoters bound by both Med8 and Med17. Spearman correlation coefficient (r) is shown. (F) Boxplot showing log2 change in ChIP-seq promoter occupancy of Taf1 after Bdf1/2 depletion. Signals were calculated in a −100 to 200 bp window relative to transcription start site (TSS) at 2879 promoters defined as bound by Taf1 in ChEC-seq experiments and classified into TFIID-dependent and CR categories.
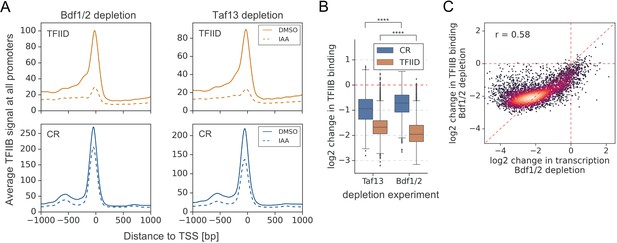
Depletion of Bdf1/2 compromises TFIIB recruitment to promoters.
(A) Average plots comparing TFIIB ChIP-seq signals before (dimethyl sulfoxide [DMSO], solid line) and after (indole-3-acetic acid [IAA], dashed line) Bdf1/2 or Taf13 degradation at 4900 promoters classified into TFIID-dependent and coactivator-redundant (CR) categories. Mean values from replicate experiments are plotted. (B) Boxplot showing log2 change in promoter occupancy of TFIIB after Taf13 or Bdf1/2 degradation. Signals were calculated in a −200 to 100 bp window relative to transcription start site (TSS). Results of the Welch’s t-test are shown. (C) Scatter plot comparing log2 change in transcription and log2 change in TFIIB occupancy after Bdf1/2 degradation at 4883 promoters/genes analyzed in RNA-seq experiments. Spearman correlation coefficient (r) is shown. See also Supplementary file 3.
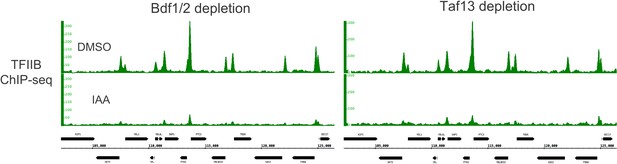
Genome browser images showing TFIIB ChIP-seq signals at a representative genomic location before (dimethyl sulfoxide [DMSO]) and after (indole-3-acetic acid [IAA]) Bdf1/2 or Taf13 degradation.
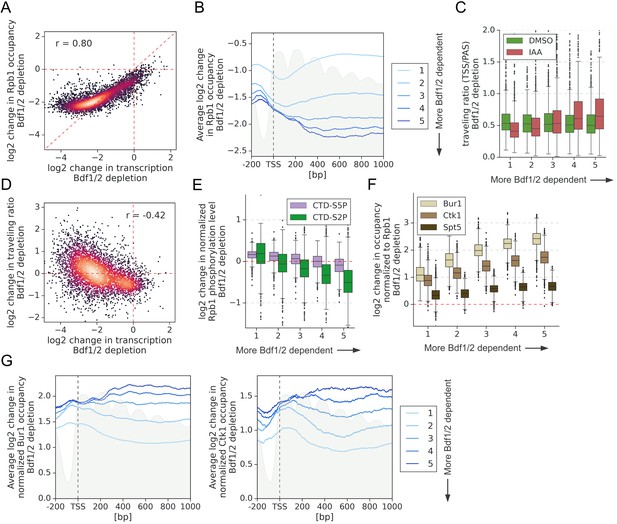
Bdf1/2 are involved in transcription elongation.
(A) Scatter plot comparing log2 change in transcription and occupancy of the largest Pol II subunit Rpb1 after Bdf1/2 degradation. Rpb1 occupancy was calculated along the whole transcribed region for 4615 genes longer than 300 bp, and with annotated transcription start site (TSS) and polyadenylation site (PAS) locations (Park et al., 2014). Spearman correlation coefficient (r) is shown. Mean values from replicate experiments are plotted. (B) Average plot showing log2 change in Rpb1 occupancy from 200 bp upstream to 1000 bp downstream of TSS after depleting Bdf1/2. Data in this and following panels are divided into five groups based on gene dependence on Bdf1/2 as measured by 4-thioU RNA-seq. Data for 3438 genes longer than 1 kb and with annotated TSS and PAS locations (Park et al., 2014) are plotted. Published MNase-seq dataset is shown as a gray area plot (Oberbeckmann et al., 2019). (C) Boxplot comparing Pol II traveling ratio (TR) in dimethyl sulfoxide (DMSO) or indole-3-acetic acid (IAA)-treated samples in the Bdf1/2 degron experiment. TR represents a ratio of Rpb1 occupancy calculated in 100 bp windows at the beginning and end of a transcribed region. The same set of 4615 genes as in (A) was used. (D) Scatter plot comparing log2 change in transcription and log2 change in Pol II TR after depleting Bdf1/2. Spearman correlation coefficient (r) is shown. (E) Boxplot comparing log2 change in Rpb1 CTD phosphorylation status at Ser2 and Ser5 residues after depleting Bdf1/2. Data were calculated along the whole transcribed region for the same set of 4615 genes as in (A) and signals were normalized to the total Rpb1 signal. (F) Boxplot comparing log2 change in Bur1, Ctk1, and Spt5 occupancy after depleting Bdf1/2. Data for DMSO and IAA-treated samples were calculated along the whole transcribed region for the same set of 4615 genes as in (A) and signals were normalized to the total Rpb1 signal. See also Supplementary file 4. (G) Average plots showing log2 changes in Bur1 and Ctk1 occupancy from 200 bp upstream to 1000 bp downstream of TSS after depleting Bdf1/2. Data for 3438 genes longer than 1 kb and with annotated TSS and PAS locations (Park et al., 2014) are plotted. Published MNase-seq dataset is shown as a gray area plot (Oberbeckmann et al., 2019).
-
Figure 6—source data 1
Processed data used to prepare average line plot in (B) showing log2 change in Rpb1 occupancy following Bdf1/2 depletion.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig6-data1-v2.csv.zip
-
Figure 6—source data 2
Processed data used to prepare average line plot in (G) showing log2 change in Bur1 occupancy following Taf1 depletion.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig6-data2-v2.csv.zip
-
Figure 6—source data 3
Processed data used to prepare average line plot in (G) showing log2 change in Ctk1 occupancy following Taf1 depletion.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig6-data3-v2.csv.zip
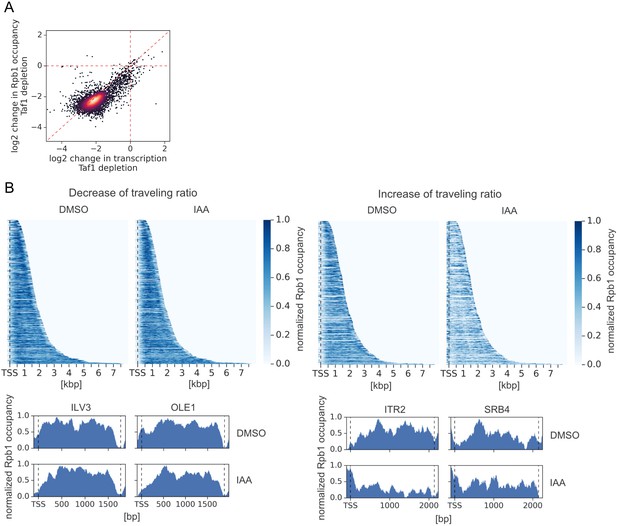
Bdf1/2 are involved in transcription elongation.
(A) Scatter plot comparing log2 change in transcription and occupancy of the largest Pol II subunit Rpb1 after Taf1 degradation. Rpb1 occupancy was calculated along the whole transcribed region for 4615 genes longer than 300 bp and with annotated transcription start site (TSS) and polyadenylation site (PAS) locations (Park et al., 2014). Mean values from replicate experiments are plotted. (B) Upper panel: heatmap showing Rpb1 occupancy at 10% (462) genes with the biggest decrease (left) and 10% (462) genes with the biggest increase (right) in Pol II traveling ratio after depleting Bdf1/2. Data were normalized individually for each gene. TSS position is marked with a dashed line. Separate plots for dimethyl sulfoxide (DMSO) and indole-3-acetic acid (IAA)-treated samples are shown. Lower panel: examples of Rpb1 distribution at representative genes with the biggest decrease (left) and the biggest increase (right) in Pol II traveling ratio after depleting Bdf1/2. TSS and PAS positions are marked with dashed lines. Data were normalized individually for each gene. Separate plots for DMSO and IAA-treated samples are shown.
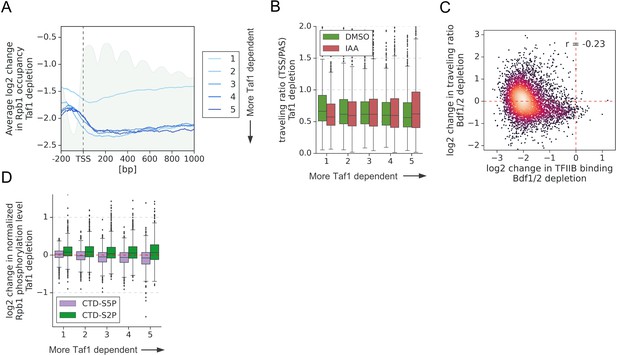
TFIID depletion does not affect transcription elongation.
(A) Average plot showing log2 change in Rpb1 occupancy from 200 bp upstream to 1000 bp downstream of transcription start site (TSS) after depleting Taf1. Data in this and following panels are divided into five groups based on gene dependence on Taf1 as measured by 4-thioU RNA-seq. Data for 3438 genes longer than 1 kb and with annotated TSS and PAS locations (Park et al., 2014) are plotted. Published MNase-seq dataset is shown as a gray area plot (Oberbeckmann et al., 2019). (B) Boxplot comparing Pol II traveling ratio (TR) in dimethyl sulfoxide (DMSO) or indole-3-acetic acid (IAA)-treated samples in Taf1 degron experiment. TR represents a ratio of Rpb1 occupancy along a 100 bp window at the beginning and end of a transcribed region. Data for 4615 genes longer than 300 bp and with annotated TSS and PAS locations (Park et al., 2014) are plotted. (C) Scatter plot comparing log2 change in TR and log2 change in TFIIB promoter signal after depleting Bdf1/2. Spearman correlation coefficient (r) is shown. Data for the same set of 4615 genes as in (B) are shown. (D) Boxplot comparing log2 change in Rpb1 CTD phosphorylation status at Ser2 and Ser5 residues after depleting Taf1. Data were calculated along the whole transcribed region for the same set of 4615 genes as in (B).
-
Figure 6—figure supplement 2—source data 1
Processed data used to prepare average line plot in (A) showing log2 change in Rpb1 occupancy following Taf1 depletion.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig6-figsupp2-data1-v2.csv.zip
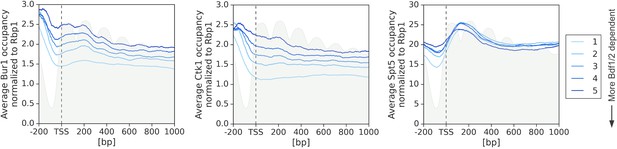
Average plots showing Bur1, Ctk1, and Spt5 occupancy from 200 bp upstream to 1000 bp downstream of transcription start site (TSS).
Data are divided into five groups based on gene dependence on Bdf1/2 as measured by 4-thioU RNA-seq. Data for 3438 genes longer than 1 kb and with annotated TSS and PAS locations (Park et al., 2014) are plotted. Published MNase-seq dataset is shown as a gray area plot (Oberbeckmann et al., 2019).
-
Figure 6—figure supplement 3—source data 1
Processed data used to prepare average line plot showing Bur1 occupancy.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig6-figsupp3-data1-v2.csv.zip
-
Figure 6—figure supplement 3—source data 2
Processed data used to prepare average line plot showing Ctk1 occupancy.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig6-figsupp3-data2-v2.csv.zip
-
Figure 6—figure supplement 3—source data 3
Processed data used to prepare average line plot showing Spt5 occupancy.
- https://cdn.elifesciences.org/articles/69619/elife-69619-fig6-figsupp3-data3-v2.csv.zip
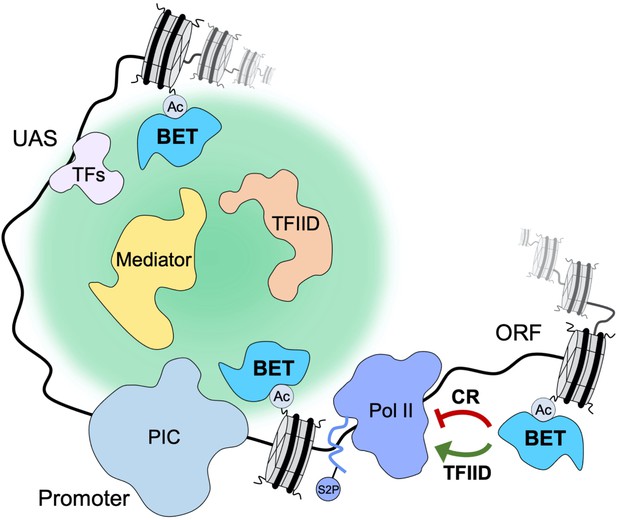
Yeast bromodomain and extra-terminal domain (BET) proteins regulate transcription initiation and elongation.
Yeast BET proteins bind to regions of the genome highly acetylated at histone H4. Together with sequence-specific transcription factors (TFs), they provide a nucleation center for dynamic recruitment of TFIID and Mediator to the promoter-proximal nuclear territory. In turn, TFIID and Mediator create a platform that facilitates preinitiation complex (PIC) assembly and stimulates transcription initiation. Yeast BET proteins also affect early elongation serving as negative regulatory factors at coactivator-redundant (CR) genes and positive regulatory factors at TFIID-dependent genes.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Saccharomyces cerevisiae) | BY4705 | PMID:9483801 | ATCC: 200869 | Laboratory of Dan Gottschling; WT |
| Genetic reagent (Schizosaccharomyces pombe) | SHY1058 | PMID:28918900 | h- RPB3-3x Flag::NatMX | WT |
| Genetic reagent (S. pombe) | SHY1110 | PMID:18216783 | h + leu1-32 ade6-216 OtrR1:ura4 Abp1-3xFlag::KanMX Cbh1-13xMyc::NatMX | Laboratory of Toshio Tsukiyama; WT |
| Genetic reagent (S. cerevisiae) | SHY1036 | PMID:28918900 | RPB3-3x Flag::NatMX, pGPD1-OSTIR::HIS3 | WT-degron tagging (BY4705 derivative) |
| Genetic reagent (S. cerevisiae) | SHY1235 | Laboratory of Steven Hahn | pGPD1-OSTIR::HIS3 | WT-degron tagging (BY4705 derivative) |
| Genetic reagent (S. cerevisiae) | SHY1039 | PMID:31913117 | RPB3-3x Flag::NatMX, pGPD1-OSTIR::HIS3, TAF1-3xV5 IAA7::KanMX | Taf1 degron |
| Genetic reagent (S. cerevisiae) | SHY1059 | Laboratory of Steven Hahn | RPB3-3x Flag::NatMX, pGPD1-OSTIR::HIS3, bdf2△::HPH | bdf2△ |
| Genetic reagent (S. cerevisiae) | RDY27 | Laboratory of Steven Hahn | RPB3-3x Flag::NatMX, pGPD1-OSTIR::HIS3, BDF1-3xV5 IAA7::KanMX | Bdf1 degron |
| Genetic reagent (S. cerevisiae) | RDY28 | Laboratory of Steven Hahn | RPB3-3x Flag::NatMX, pGPD1-OSTIR::HIS3, BDF2-3xV5 IAA7::KanMX | Bdf2 degron |
| Genetic reagent (S. cerevisiae) | RDY70 | Laboratory of Steven Hahn | pGPD1-OSTIR::HIS3, BDF2-3xV5 IAA7::KanMX | Bdf2 degron |
| Genetic reagent (S. cerevisiae) | RDY73 | Laboratory of Steven Hahn | pGPD1-OSTIR::HIS3, BDF2-3xV5 IAA7::KanMX, BDF1-3xV5 IAA7::URA3 | Bdf1/2 degron |
| Genetic reagent (S. cerevisiae) | RDY1 | Laboratory of Steven Hahn | RPB3-3x Flag::NatMX, pGPD1-OSTIR::HIS3, bdf2△::HPH, BDF1-3xV5 IAA7::KanMX | bdf2△, Bdf1 degron |
| Genetic reagent (S. cerevisiae) | SHY1137 | Laboratory of Steven Hahn | RPB3-3x Flag::NatMX, pGPD1-OSTIR::HIS3, TOA1-3xV5 IAA7 | Toa1 degron |
| Genetic reagent (S. cerevisiae) | SHY1213 | Laboratory of Steven Hahn | pGPD1-OSTIR::HIS3, ssl2△::KanMX/pSH1947 (LEU2 SSL2-3xV5 IAA7) | Ssl2 degron |
| Genetic reagent (S. cerevisiae) | RDY29 | Laboratory of Steven Hahn | RPB3-3x Flag::NatMX, pGPD1-OSTIR::HIS3, HTZ1-3xV5 IAA7::KanMX | H2A.Z degron |
| Genetic reagent (S. cerevisiae) | RDY57 | Laboratory of Steven Hahn | RPB3-3x Flag::NatMX, pGPD1-OSTIR::HIS3, esa1△::KanMX/pRD8 (LEU2 ESA1-3xV5 IAA7) | Esa1 degron |
| Genetic reagent (S. cerevisiae) | RDY44 | Laboratory of Steven Hahn | pBDF2-MNase::TRP1 | Free MNase (BDF2 promoter) |
| Genetic reagent (S. cerevisiae) | RDY34 | Laboratory of Steven Hahn | RPB3-3x Flag::NatMX, pGPD1-OSTIR::HIS3, BDF1-3xV5 IAA7, BDF2-MNase::TRP1 | Bdf1 degron, Bdf2-MNase |
| Genetic reagent (S. cerevisiae) | RDY33 | Laboratory of Steven Hahn | RPB3-3x Flag::NatMX, pGPD1-OSTIR::HIS3, BDF2-3xV5 IAA7, BDF1-MNase::TRP1 | Bdf2 degron, Bdf1-MNase |
| Genetic reagent (S. cerevisiae) | RDY59 | Laboratory of Steven Hahn | RPB3-3x Flag::NatMX, pGPD1-OSTIR::HIS3, esa1△::KanMX, BDF1-MNase::TRP1/pRD8 (LEU2 ESA1-3xV5 IAA7) | Esa1 degron, Bdf1-MNase |
| Genetic reagent (S. cerevisiae) | RDY85 | Laboratory of Steven Hahn | pGPD1-OSTIR::HIS3, BDF1-3xFlag::NatMX | Bdf1-3xFlag |
| Genetic reagent (S. cerevisiae) | RDY86 | Laboratory of Steven Hahn | pGPD1-OSTIR::HIS3, BDF2-3xFlag::NatMX | Bdf2-3xFlag |
| Genetic reagent (S. cerevisiae) | RDY30 | Laboratory of Steven Hahn | RPB3-3x Flag::NatMX, pGPD1-OSTIR::HIS3, bdf2△::HygMX, BDF1-3xV5 IAA7::KanMX, TAF1-MNase::TRP1 | bdf2△, Bdf1 degron, Taf1-MNase |
| Genetic reagent (S. cerevisiae) | RDY21 | Laboratory of Steven Hahn | RPB3-3x Flag::NatMX, pGPD1-OSTIR::HIS3, bdf2△::HygMX, BDF1-3xV5 IAA7::KanMX, TAF11-MNase::TRP1 | bdf2△, Bdf1 degron, Taf11-MNase |
| Genetic reagent (S. cerevisiae) | RDY26 | Laboratory of Steven Hahn | RPB3-3x Flag::NatMX, pGPD1-OSTIR::HIS3, bdf2△::HygMX, BDF1-3xV5 IAA7::KanMX, MED8-MNase::TRP1 | bdf2△, Bdf1 degron, Med8-MNase |
| Genetic reagent (S. cerevisiae) | RDY50 | Laboratory of Steven Hahn | RPB3-3x Flag::NatMX, pGPD1-OSTIR::HIS3, bdf2△::HygMX, BDF1-3xV5 IAA7::KanMX, MED17-MNase::TRP1 | bdf2△, Bdf1 degron, Med17-MNase |
| Genetic reagent (S. cerevisiae) | RDY51 | Laboratory of Steven Hahn | RPB3-3x Flag::NatMX, pGPD1-OSTIR::HIS3, bdf2△::HygMX, BDF1-3xV5 IAA7::KanMX, SPT3-MNase::TRP1 | bdf2△, Bdf1 degron, Spt3-MNase |
| Genetic reagent (S. cerevisiae) | SHY1172 | Laboratory of Steven Hahn | pGPD1-OSTIR::HIS3, MED14-3xV5 IAA7::KanMX, SPT3-MNase::TRP1 | Med14 degron, Spt3-MNase |
| Genetic reagent (S. cerevisiae) | SHY1301 | Laboratory of Steven Hahn | pGPD1-OSTIR::HIS3, MED14-3xV5 IAA7::KanMX, TAF1-MNase::TRP1 | Med14 degron, Taf1-MNase |
| Genetic reagent (S. cerevisiae) | SHY1327 | Laboratory of Steven Hahn | pGPD1-OSTIR::HIS3, TAF13-3xV5 IAA7::KanMX, MED8-MNase::TRP1 | Taf13 degron, Med8-MNase |
| Genetic reagent (S. cerevisiae) | SHY1331 | Laboratory of Steven Hahn | pGPD1-OSTIR::HIS3, TAF13-3xV5 IAA7::KanMX, SPT7-MNase::TRP1 | Taf13 degron, Spt7-MNase |
| Genetic reagent (S. cerevisiae) | RDY82 | Laboratory of Steven Hahn | pGPD1-OSTIR::HIS3, BDF2-3xV5 IAA7::KanMX, BDF1-3xV5 IAA7::URA3, TAF1-3xFlag::NatMX | Bdf1/2 degron, Taf1-3xFlag |
| Genetic reagent (S. cerevisiae) | SHY1089 | Laboratory of Steven Hahn | RPB3-3x Flag::NatMX, pGPD1-OSTIR::HIS3, TAF13-3xV5 IAA7::KanMX, SUA7-13xMyc::HPH | Taf13 degron, TFIIB-13xMyc |
| Genetic reagent (S. cerevisiae) | RDY92 | Laboratory of Steven Hahn | pGPD1-OSTIR::HIS3, BDF2-3xV5 IAA7::KanMX, BDF1-3xV5 IAA7::URA3, SUA7-3xFlag::NatMX | Bdf1/2 degron, TFIIB-3xFlag |
| Genetic reagent (S. cerevisiae) | RDY79 | Laboratory of Steven Hahn | pGPD1-OSTIR::HIS3, BDF2-3xV5 IAA7::KanMX, BDF1-3xV5 IAA7::URA3, BUR1-3xFlag::NatMX | Bdf1/2 degron, Bur1-3xFlag |
| Genetic reagent (S. cerevisiae) | RDY81 | Laboratory of Steven Hahn | pGPD1-OSTIR::HIS3, BDF2-3xV5 IAA7::KanMX, BDF1-3xV5 IAA7::URA3, CTK1-3xFlag::NatMX | Bdf1/2 degron, Ctk1-3xFlag |
| Genetic reagent (S. cerevisiae) | RDY94 | Laboratory of Steven Hahn | pGPD1-OSTIR::HIS3, BDF2-3xV5 IAA7::KanMX, BDF1-3xV5 IAA7::URA3, SPT5-3xFlag::NatMX | Bdf1/2 degron, Spt5-3xFlag |
| Antibody | Rabbit polyclonal anti-Tfg2 | Laboratory of Steven Hahn | Rabbit #1260K | WB (1:2000) |
| Antibody | Mouse monoclonal anti-V5 | Invitrogen | Cat. #R960-25 | WB (1:1000) |
| Antibody | Rabbit monoclonal anti-H4K12ac | Cell Signaling Technology | Cat. #13944 | WB (1:1000) ChIP-seq (1:100) |
| Antibody | Mouse monoclonal anti-Flag | Sigma-Aldrich | Cat. #F3165 | ChIP-seq (1:100) |
| Antibody | Mouse monoclonal anti-Myc | Cell Signaling Technology | Cat. #2276 | ChIP-seq (1:100) |
| Antibody | Mouse monoclonal anti-Rpb1 CTD | Cell Signaling Technology | Cat. #2629 | ChIP-seq (1:50) |
| Antibody | Rabbit monoclonal anti-Phospho-Rpb1 CTD (Ser2) | Cell Signaling Technology | Cat. #13499 | ChIP-seq (1:50) |
| Antibody | Rabbit monoclonal anti-Phospho-Rpb1 CTD (Ser5) | Cell Signaling Technology | Cat. #13523 | ChIP-seq (1:50) |
| Recombinant DNA reagent | pFA6a-3V5-IAA7-KanMX6 | PMID:30192227 | Degron tagging vector | |
| Recombinant DNA reagent | pSH1855 | Laboratory of Steven Hahn | Degron tagging vector | |
| Recombinant DNA reagent | pGZ110 | PMID:26490019 | MNase tagging vector | |
| Recombinant DNA reagent | p2L-3Flag-NAT | Laboratory of Toshio Tsukiyama | 3xFlag tagging vector | |
| Recombinant DNA reagent | pFA6a-13Myc-Hyg | Laboratory of Toshio Tsukiyama | 13xMyc tagging vector | |
| Recombinant DNA reagent | pRS303 | PMID:2659436 | ATTC: 77138 | |
| Recombinant DNA reagent | pRD15 | Laboratory of Steven Hahn | pRS303 derivative-free MNase under BDF2 promoter | |
| Recombinant DNA reagent | pRS315 | PMID:2659436 | ATTC: 77144 | |
| Recombinant DNA reagent | pRS316 | PMID:2659436 | ATTC: 77145 | |
| Recombinant DNA reagent | pRD5 | Laboratory of Steven Hahn | pRS315 derivative-ESA1 gene with native promoter | |
| Recombinant DNA reagent | pRD21 | Laboratory of Steven Hahn | pRS316 derivative-ESA1 gene with native promoter and degron tag | |
| Commercial assay or kit | RiboPure yeast kit | Thermo Fisher Scientific | Cat. #AM1926 | |
| Commercial assay or kit | MyOne Streptavidin C1 Dynabeads | Thermo Fisher Scientific | Cat. #65002 | |
| Commercial assay or kit | Protein G Dynabeads | Thermo Fisher Scientific | Cat. #10003D | |
| Commercial assay or kit | RNeasy Kit | Qiagen | Cat. #74104 | |
| Commercial assay or kit | Qubit HS RNA assay | Thermo Fisher Scientific | Cat. #Q32852 | |
| Commercial assay or kit | Qubit HS DNA assay | Thermo Fisher Scientific | Cat. #Q32851 | |
| Commercial assay or kit | Ovation Universal RNA-seq Library Preparation Kit | Tecan | Cat. #0364-A01 | Including S. cerevisiae AnyDeplete reagent |
| Chemical compound, drug | MTSEA biotin-XX | Biotium | Cat. #90066-1 | |
| Chemical compound, drug | 4-thiouracil | Sigma-Aldrich | Cat. #440736 | |
| Chemical compound, drug | Indole-3-acetic acid | Sigma-Aldrich | Cat. #I3750 | |
| Software, algorithm | HTseq | PMID:25260700 | ||
| Software, algorithm | Bowtie 2 | PMID:22388286 | ||
| Software, algorithm | HOMER | PMID:20513432 | ||
| Software, algorithm | Clustal Omega | PMID:21988835 | ||
| Software, algorithm | Jalview v 2 | PMID:19151095 | ||
| Software, algorithm | PANTHER v 16.0 | PMID:33290554 |
Additional files
-
Supplementary file 1
Summary of 4-thioU RNA-seq experiments.
- https://cdn.elifesciences.org/articles/69619/elife-69619-supp1-v2.xlsx
-
Supplementary file 2
Results of gene ontology analysis on 25% most bromodomain and extra-terminal domain (BET)-sensitive genes in yeast (this work) and human (Winter et al., 2017) system.
Related to Figure 1.
- https://cdn.elifesciences.org/articles/69619/elife-69619-supp2-v2.xlsx
-
Supplementary file 3
Summary of ChEC-seq and ChIP-seq experiments.
- https://cdn.elifesciences.org/articles/69619/elife-69619-supp3-v2.xlsx
-
Supplementary file 4
Summary of ChIP-seq experiments on Rpb1, Ctk1, Bur1, and Spt5.
Related to Figure 6.
- https://cdn.elifesciences.org/articles/69619/elife-69619-supp4-v2.xlsx
-
Supplementary file 5
Yeast strains used in this study.
- https://cdn.elifesciences.org/articles/69619/elife-69619-supp5-v2.xlsx
-
Supplementary file 6
Number of biological replicates collected for all experiments in this study.
- https://cdn.elifesciences.org/articles/69619/elife-69619-supp6-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/69619/elife-69619-transrepform-v2.docx





