Common host variation drives malaria parasite fitness in healthy human red cells
Figures
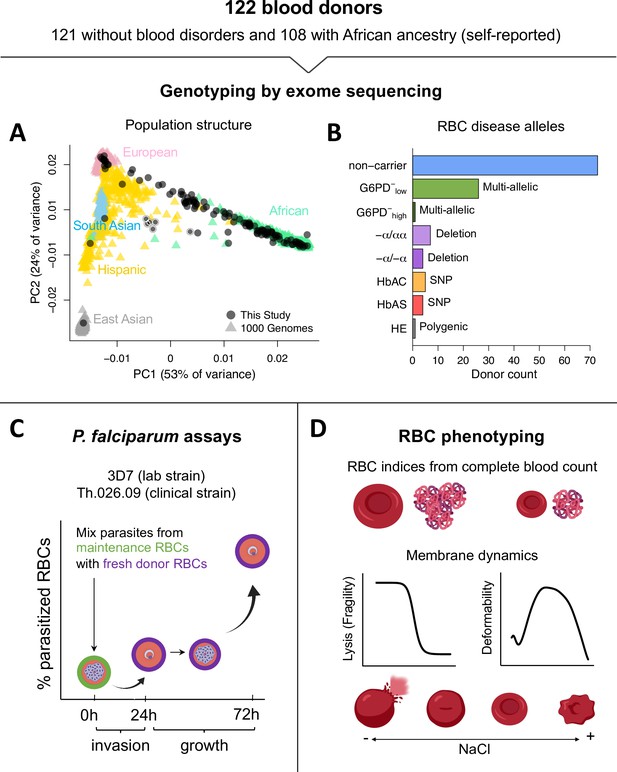
Overview of blood donors and study design.
(A) PCA of genetic variation across 35,759 unlinked exome SNPs. Donors from this study are plotted on coordinate space derived from 1000 Genomes reference populations. Points with white borders represent six related individuals, five of whom were excluded from the study. All exome variants passing quality filters are available in Figure 1—source data 1. (B) Over a third of donors carried alleles for RBC disorders linked to Plasmodium falciparum resistance. Individuals with >1 disease allele were classified by their most severe condition. non-carrier: Donor without any of the following alleles or conditions. G6PD−low: Mild to medium G6PD deficiency (<42% loss of function). G6PD−high: Severe G6PD deficiency (>60% loss of function). −α/αα: heterozygous HBA2 deletion, or alpha thalassemia minima. −α/−α: homozygous HBA2 deletion, or α-thalassemia trait. HbAC: heterozygous HBB:E7K, or hemoglobin C trait. HbAS: heterozygous HBB:E7V, or sickle cell trait. HE: hereditary elliptocytosis. (C) Two components of P. falciparum fitness were measured with flow cytometry at three timepoints. Invasion is the change in parasitemia as schizonts egress from maintenance RBCs (green) and invade fresh acceptor RBCs from the blood donors (purple). Growth is the multiplication rate from a complete parasite cycle in the fresh acceptor RBCs. (D) RBC phenotypes were measured using complete blood counts with RBC indices, osmotic fragility tests, and ektacytometry on fresh samples. This figure was partially created with Biorender.com. RBC, red blood cell; SNP, single-nucleotide polymorphism.
-
Figure 1—source data 1
Individual genotypes, population frequencies, and protein annotations for exome variants passing quality filters (N~160,000).
- https://cdn.elifesciences.org/articles/69808/elife-69808-fig1-data1-v2.zip
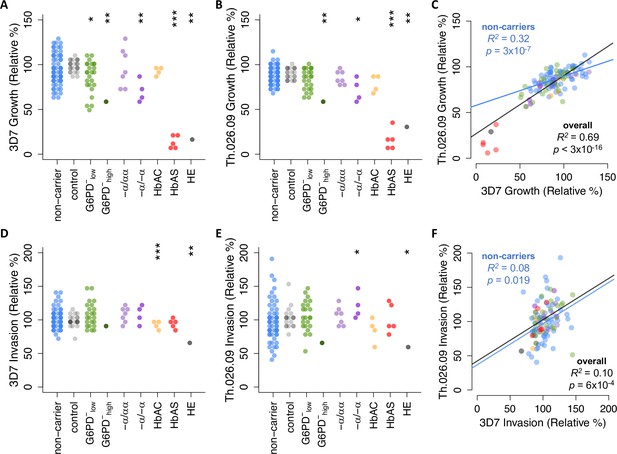
Plasmodium falciparum replication rate varies widely among donor RBCs.
(A, B) Growth of P. falciparum lab strain 3D7 (A) or clinical isolate Th.026.09 (B) over a full 48-hr cycle in donor RBCs (see Figure 1C). Growth is presented relative to the average non-carrier rate after correction for batch effects (Figure 2—figure supplement 1; see Materials and methods), including comparison to a repeated RBC control shown in gray. Each carrier group was compared to unrelated non-carriers using Student’s t-test, except in cases where N=1, where asterisks instead indicate the percentile of the non-carrier distribution. Repeated measurements of 11 donors are shown in Figure 2—figure supplement 2. (C) Per-sample growth rates are correlated between the two P. falciparum strains. (D–F) As in (A–C) but for P. falciparum invasion efficiency (see Figure 1C). R2 and p-values are derived from OLS regression. *p<0.1; **p<0.05; ***p<0.01. RBC, red blood cell.
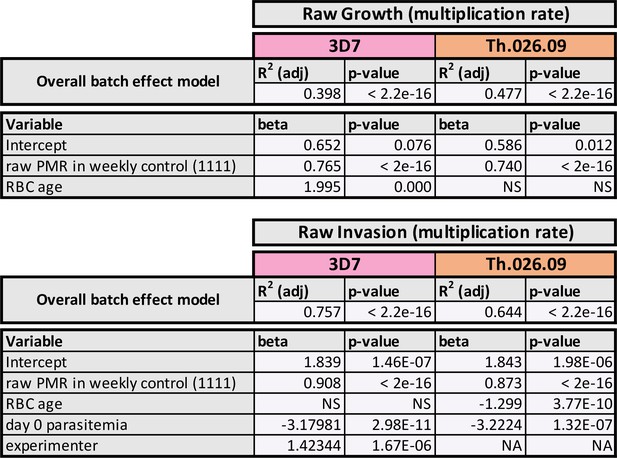
Linear models of batch effects on parasite fitness.
PMR: parasite multiplication rate.
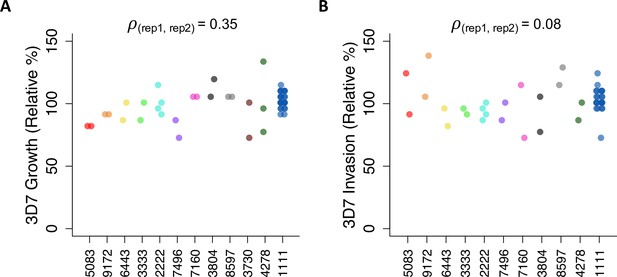
Repeatability of parasite assays in the same donors over time.
Twelve participants (including the weekly control, 1111) donated blood in multiple weeks for independent experiments. Repeated donors were non-carriers except 6443, who carried the HbAC allele; and 7160, 8597, and 4278, who carried alleles for mild G6PD deficiency. Growth and invasion data are shown after standardization and batch correction, as described in Materials and methods. Pearson’s rho is calculated between the first and second assays and can range from –1 to 1.
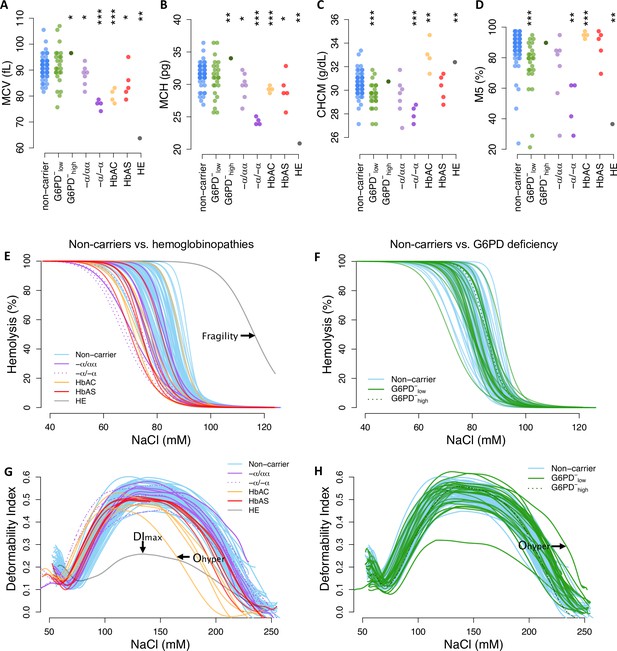
Red cell phenotypes that are abnormal in carriers also vary widely among non-carriers.
(A–D) Red cell indices were measured by an ADVIA hematology analyzer. Additional indices are shown in Figure 3—figure supplement 1. MCV: mean corpuscular (RBC) volume; MCH: mean cellular hemoglobin; CHCM: cellular hemoglobin concentration; M5: fraction of RBCs with normal volume and normal hemoglobin (see Figure 3—figure supplement 2). Statistical tests as in Figure 2. (E, F) Osmotic fragility curves. Fragility is defined as the NaCl concentration at which 50% of RBCs lyse (see Figure 3—figure supplement 4). (G, H) Ektacytometry curves characterize RBC deformability and dehydration under salt stress (Figure 3—figure supplement 5). A heatmap of all phenotypes by carrier status is available in Figure 3—figure supplement 3. RBC, red blood cell.
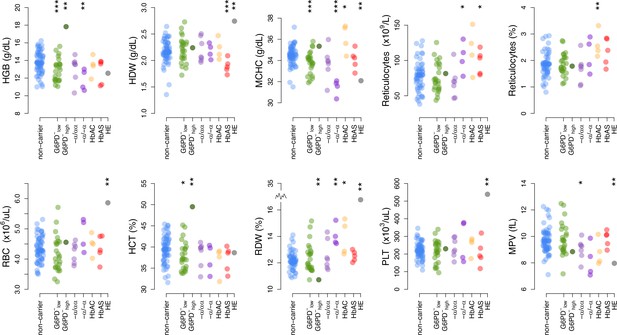
RBC indices data from complete blood counts in donor RBCs.
Donor classifications and statistical tests as in Figure 3. HCT, hematocrit; HDW, hemoglobin distribution width; HGB, hemoglobin; MCHC, mean corpuscular hemoglobin concentration; MPV, mean platelet volume; PLT, platelet count; RBC, red blood cell; RDW, red cell distribution width, .
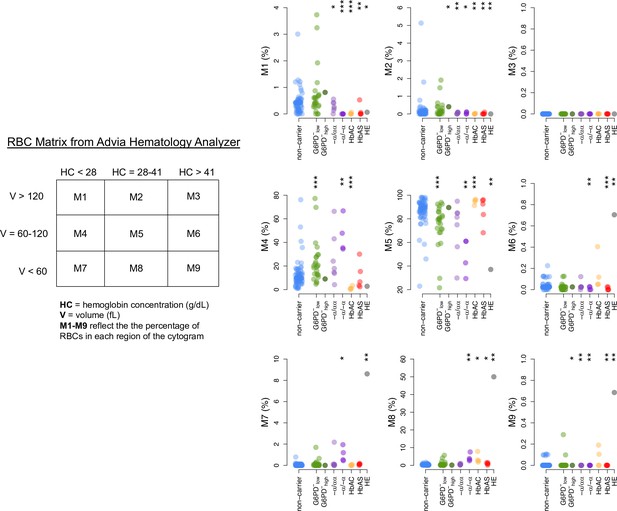
RBC Matrix data from complete blood counts in donor RBCs.
Donor classifications and statistical tests as in Figure 3. RBC, red blood cell.
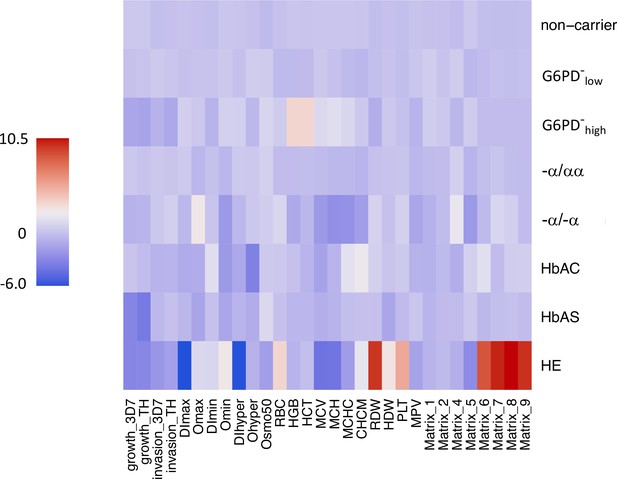
Heatmap of RBC phenotypes by carrier status.
Phenotypes were scaled and centered at 0 with the preProcess function from the caret package in R using all sample data (i.e., not limited to non-carriers). The mean value for each carrier group is shown. RBC, red blood cell.
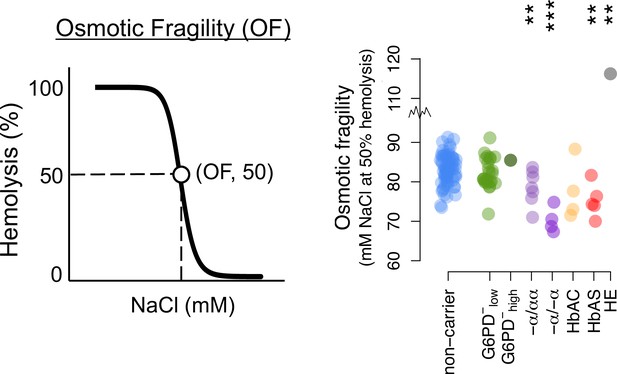
Osmotic fragility diagram and summary data.
Donor classifications and statistical tests as in Figure 3.
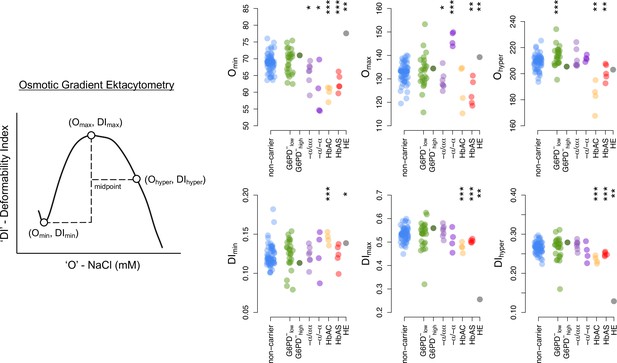
Ektacytometry diagram and summary data.
Donor classifications and statistical tests as in Figure 3.
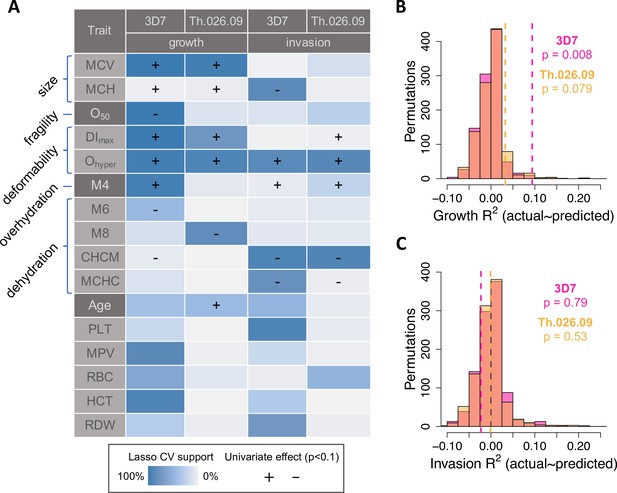
RBC phenotypes predict Plasmodium falciparum fitness in non-carriers.
(A) Phenotypes selected by LASSO in at least 40% of train data sets (blue shading; see Materials and methods) in at least one of four models of parasite replication (columns). Each model was trained on ~90% of the data (B, C) and tested on the remaining 10% (B, C). (+/−) shows the direction of effect if the phenotype was significantly correlated (p<0.1) with the parasite fitness component in a separate, univariate linear model (Figure 4—figure supplement 1; ). MCV: mean RBC volume (fl).MCH: mean corpuscular hemoglobin (pg/RBC). O50: Osmotic fragility (mM NaCl; see Figure 3—figure supplement 4). DImax: Maximum membrane deformability (arbitrary units; see Figure 3—figure supplement 5). Ohyper: Tendency to resist osmotic dehydration and loss of deformability. M4: fraction of RBCs with normal volume and low hemoglobin (see Figure 3—figure supplement 2). M6: fraction of RBCs with normal volume and high hemoglobin. M8: fraction of RBCs with low volume and normal hemoglobin. CHCM: cellular hemoglobin concentration mean (g/dl). MCHC: mean corpuscular hemoglobin concentration (g/dl). PLT: platelet number (×103/µl). MPV: mean platelet volume (fl). RBC: red cell number (×106/µl). HCT: hematocrit, or the fraction of blood volume composed of RBCs. RDW: red cell distribution width (%). (B, C) Variance in parasite fitness explained by RBC phenotypes in LASSO models. Dashed lines indicate average R2 for the measured test data. Each histogram shows the same procedure on 1000 permutations of the measured test data. RBC, red blood cell.
-
Figure 4—source data 1
Association statistics for individual phenotypic predictors with non-zero LASSO support.
- https://cdn.elifesciences.org/articles/69808/elife-69808-fig4-data1-v2.xlsx
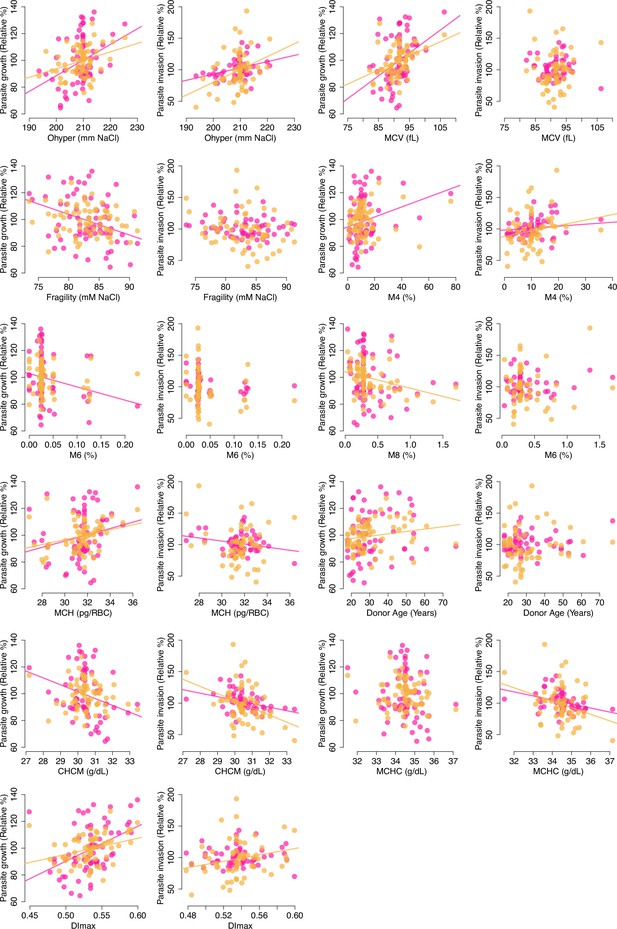
Scatterplots of RBC phenotypes against parasite fitness in non-carriers.
Phenotypes from Figure 4A are shown if at least one OLS test had p<0.1. Lines of best fit are shown if p<0.1. All phenotypes except Fragility and Donor Age are clustered around the median as a result of normalization, which equalized the median value across weeks (see Materials and methods). Pink: 3D7; Orange: Th.026.09. RBC, red blood cell.
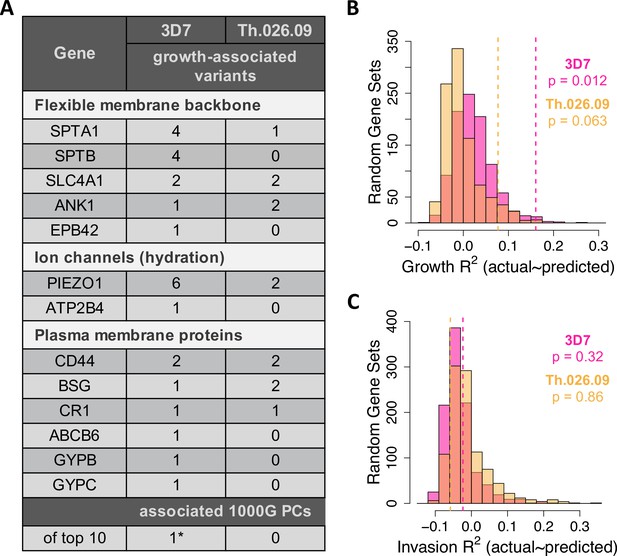
Common variation in malaria-associated genes predicts Plasmodium falciparum fitness in non-carrier RBCs.
(A) Variants in 23 malaria-related genes (Figure 5—source data 1) and genetic PCs selected by LASSO in at least >40% of train data sets. Each model was trained on ~90% of the measured data (B C) and tested on the remaining 10% (B C). The following genes had no associated variants in non-carriers: CD55, EPB41, FPN, G6PD, GYPA, GYPE, HBA1/2, HBB, and HP. *The only significant PC association was driven by a single East Asian donor (Figure 5—figure supplement 5). (B, C) Variance in parasite fitness explained by LASSO models including 23 malaria-related genes, the top 10 PCs, and RBC phenotypes. Dashed lines indicate average R2 for models using the measured test data. Each histogram shows R2 for models including variants from 23 random genes in the RBC proteome (Figure 5—source data 2) instead of malaria-related genes. All predictors with non-zero LASSO support are shown in Figure 5—source data 3. Additional histograms from permuted data are shown in Figure 5—figure supplement 1. The variance explained by variants undiscovered by previous GWAS is shown in Figure 5—figure supplement 4. GWAS, genome-wide association studies; PC, principal component; RBC, red blood cell.
-
Figure 5—source data 1
Twenty-three RBC genes with strong links to malaria in the literature.
- https://cdn.elifesciences.org/articles/69808/elife-69808-fig5-data1-v2.xlsx
-
Figure 5—source data 2
Proteins present in mature RBCs.
This list was derived from the Red Blood Cell Collection database (rbcc.hegelab.org) using a medium-confidence filter.
- https://cdn.elifesciences.org/articles/69808/elife-69808-fig5-data2-v2.csv
-
Figure 5—source data 3
All genetic and phenotypic predictors with non-zero LASSO support.
Growth predictors selected in at least 40% of train data sets are indicated in bold. Genetic predictors are summarized in Figure 5A. NA indicates predictors that were only present as singletons in the smaller invasion data set.
- https://cdn.elifesciences.org/articles/69808/elife-69808-fig5-data3-v2.xlsx
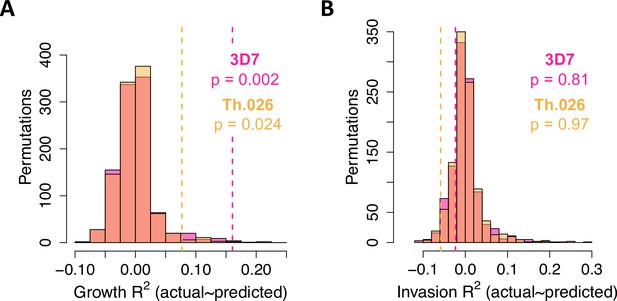
Variance in parasite fitness explained by permuted data in LASSO models.
Each model was trained on ~90% of the measured data and tested on the remaining 10%. Dashed lines indicate average R2 for the measured test data. Each histogram shows the same procedure on 1,000 permutations of the measured test data.
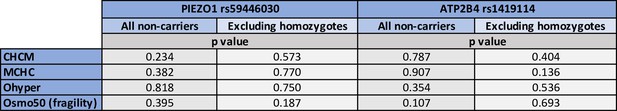
Lack of association between RBC dehydration phenotypes and PIEZO1 rs59446030 or ATP2B4 rs1419114.
Each cell shows the p-value from a linear model between the genetic variant and trait in non-carriers. RBC, red blood cell.
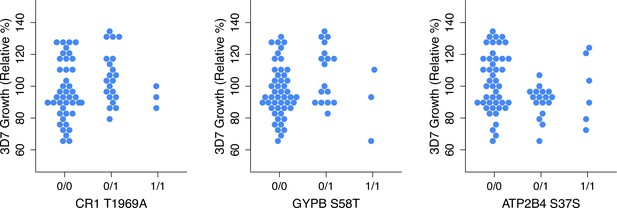
Three non-carrier variants with potentially overdominant effects on 3D7 growth.
Homozygotes for the minor allele were ignored when estimating effect sizes for these alleles with OLS for Figure 6E. Effect size estimates that include all homozygotes are shown in Figure 5—source data 3.

Variants undiscovered by previous GWAS drive most of the association signal between parasite replication rate and the 23 malaria-related genes.
‘Variance explained’ is the R2 of a linear model in non-carriers (excluding using only these variants as predictors. Details on the variants and previous GWAS traits are provided in Figure 5—source data 3). GWAS, genome-wide association studies.
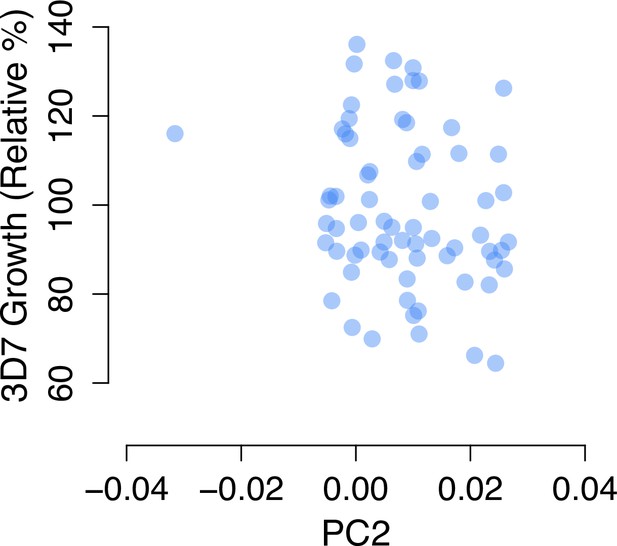
An outlier individual for PC2 drives an apparent association between PC2 and 3D7 growth.
See also Figure 1A.
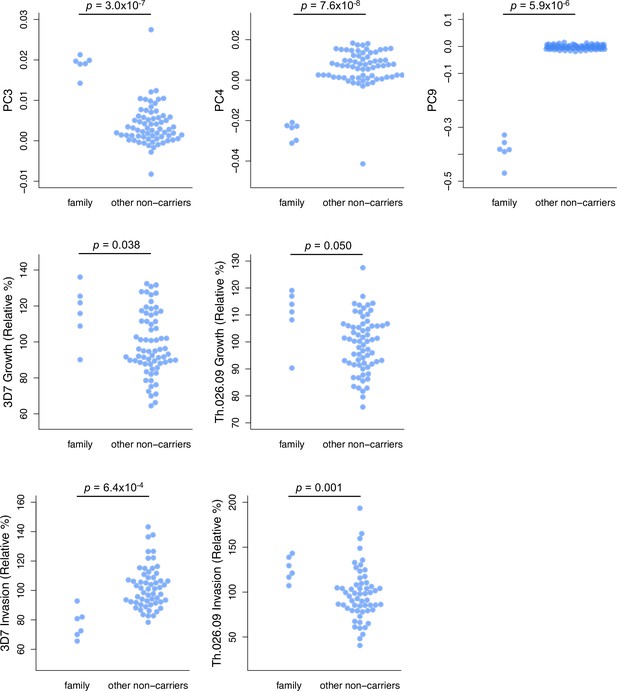
A six-member family has unique ancestry and parasite susceptibility compared to other non-carrier donors.
Only PCs that distinguish the family from other non-carriers are shown. P-values are derived from t-tests. PC, principal component.
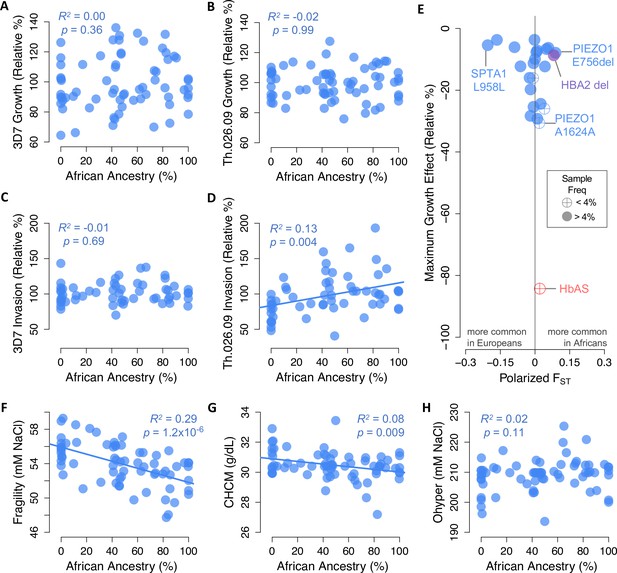
Little evidence of widespread selection in Africa for slower Plasmodium falciparum replication, protective alleles, or protective phenotypes in non-carriers.
(A–D) Parasite replication versus the exome-wide fraction of African ancestry in non-carriers, determined with ADMIXTURE by comparison to 1000 Genomes reference populations. R2 and p-values are shown for OLS regression. (E) Alleles in 23 malaria-related genes that predict slower P. falciparum growth in non-carriers (Figure 5A) are not enriched for higher frequencies in Africa versus Europe. Effect sizes are shown for one allele copy for 3D7 or Th.026.09 growth, whichever was greater. Effect sizes were determined from additive models except for three alleles that appeared overdominant (Figure 5—figure supplement 3). FST was calculated from African and European samples in gnomAD (see Materials and methods). HbAS and the HBA2 deletion are shown for comparison. (F–H) RBC phenotypes associated with P. falciparum growth versus the exome-wide fraction of African ancestry in non-carriers. Slower P. falciparum growth in RBCs is predicted by greater fragility (F), greater dehydration (G), and lower Ohyper (H) (Figure 4A). Additional phenotypes are shown in Figure 6—figure supplement 1.
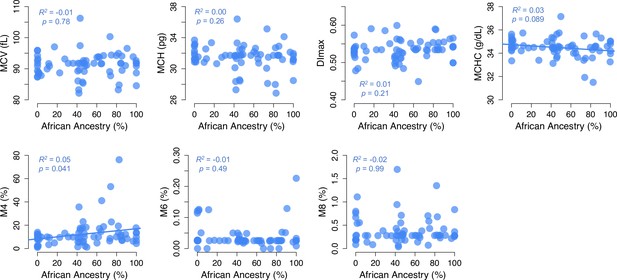
Scatterplots of RBC phenotypes versus African ancestry in non-carriers.
Phenotypes from Figure 4A are shown. R2 and p-values were estimated from OLS regression. RBC, red blood cell.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Biological sample (Homo sapiens) | Primary whole blood samples | This paper | Freshly drawn from de-identified human subjects into CPDA tubes (IRB #40479) | |
| Strain, strain background (Plasmodium falciparum) | 3D7 | PMID:3299700; Obtained from Walter and Eliza Hall Institute, Melbourne, Australia | ||
| Strain, strain background (P. falciparum) | Th026.09 | PMID:22430961; Gift from Daouda Ndiaye and Sarah Volkman, Senegal | ||
| Commercial assay or kit | DNeasy Blood and Tissue Kit | QIAGEN | ||
| Commercial assay or kit | KAPA Hyperplus Kit | Roche | ||
| Commercial assay or kit | SeqCap EZ Prime Exome Kit | Roche | ||
| Sequence-based reagent | Primers amplifying PIEZO1 exon 17 | PMID:32265284 | ||
| Software, algorithm | bwa mem | http://arxiv.org/abs/1303.3997 | 0.7.17-r1188 | |
| Software, algorithm | GATK | https://gatk.broadinstitute.org/hc/en-us | 4.0.0.0 | |
| Software, algorithm | vcftools | doi:10.1093/bioinformatics/btr330 | 0.1.15 | |
| Software, algorithm | ANNOVAR | PMID:20601685 | 2018-04-16 | |
| Software, algorithm | PLINK | PMID:17701901 | v1.90b6.8 64-bit | |
| Software, algorithm | ADMIXTURE | PMID:21682921 | 1.3.0 | |
| Software, algorithm | R | https://www.R-project.org/ | 3.5.1 | |
| Other | SYBR Green I nucleic acid stain | Invitrogen | S7563 | |
| Other | Drabkin’s Reagent | Ricca Chemical | 2660–32 |





