Transcriptomic encoding of sensorimotor transformation in the midbrain
Figures
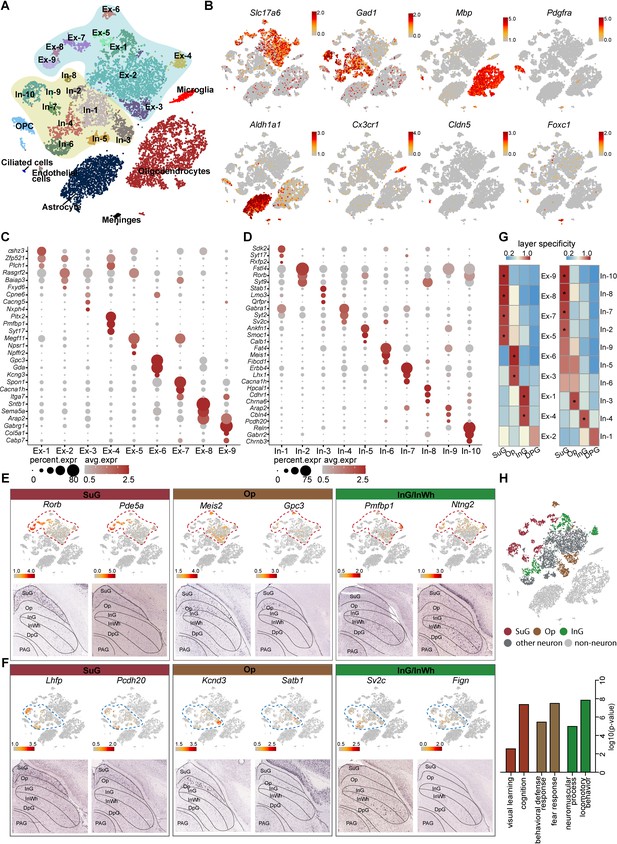
Identification and characterization of cell types and spatial heterogeneity of mouse superior colliculus (SC) neurons.
(A) Unbiased clustering of single-nucleus RNA-sequencing (snRNA-seq) data of mouse SC cells. Each dot represents an individual cell. The cells were grouped into 26 clusters, and the cell types were annotated according to the expression of known marker genes. (B) t-Distributed stochastic neighbor embedding (t-SNE) showing the known markers of major cell types (excitatory neurons, inhibitory neurons, oligodendrocytes, oligodendrocyte progenitor cells (OPCs), astrocytes, microglia, endothelial cells, ciliated cells and meningeal cells) in the mouse SC. The scale bar indicates the relative gene expression level (gray, low; red, high). (C, D) Dot plots showing the differentially expressed genes (DEGs) among 9 excitatory neuron subclusters (C) and 10 inhibitory neuron subclusters (D). (E, F) Spatial expression of the top DEGs of excitatory neuron subclusters (E) and inhibitory neuron subclusters (F). Upper panel: gene expression levels projected onto the two-dimensional t-SNE and colored according to relative gene expression level (gray, low; red, high). Red dashed line, excitatory neuron subclusters; blue dashed line, inhibitory neuron subclusters. Lower panel: in situ hybridization staining of mouse SC for the identified excitatory neuron layer markers (from the Allen Brain Atlas). (G) Heatmap showing the computed layer specificity score for each excitatory neuron subcluster (left) and each inhibitory neuron subcluster (right). Statistical analyses were performed by ANOVA (*p<0.05). (H) SC layer information annotation of neurons. Upper panel: cells colored by layer information as indicated by the legend on the bottom. Lower panel: gene ontology enrichment analysis of layer-annotated SC neurons.
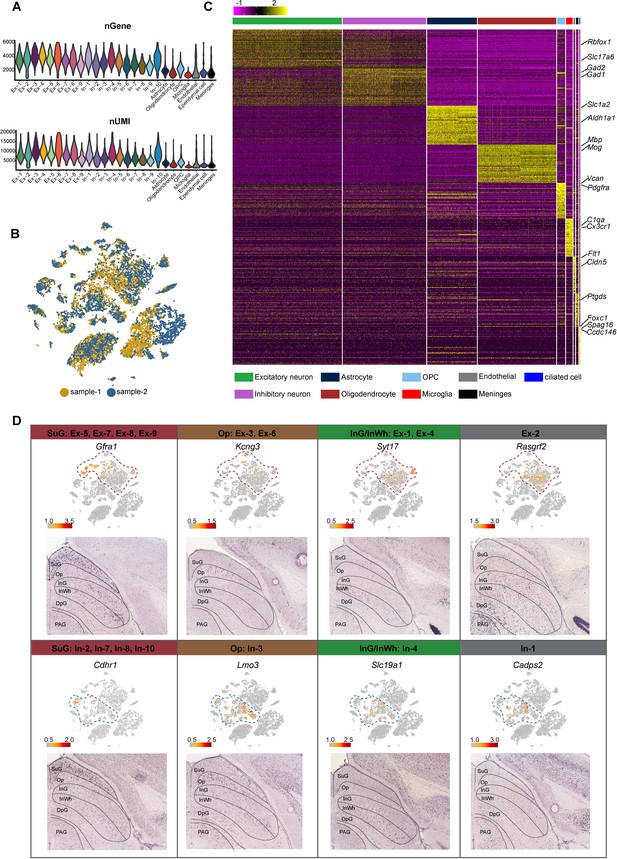
Quality of single-nucleus RNA-sequencing (snRNA-seq) metrics and the spatial distribution of neurons.
(A) Number of genes (nGene) and number of unique molecular identifiers (nUMI) of each cluster. (B) t-Distributed stochastic neighbor embedding (t-SNE) plot of the two replicates of single-nucleus RNA-seqencing (snRNA-seq) experiment. Replicate 1 (sample-1) is colored in yellow and replicate 2 (sample-2) is colored in blue. Each dot represents an individual cell. (C) Heatmap of top 50 differentially expressed genes (DEGs) of each major cell type shown in Figure 1A. The scale bar indicates relative gene expression level (purple, low; yellow, high). (D) Spatial expression of the top DEGs of excitatory neuron subclusters and inhibitory neuron subclusters. Upper panel: gene expression levels projected onto the two-dimensional t-SNE and colored according to relative gene expression level (gray, low; red, high). Red dashed line, excitatory neuron subclusters; blue dashed line, inhibitory neuron subclusters. Lower panel: in situ hybridization staining of mouse superior colliculus for the identified excitatory neuron layer markers (from the Allen Brain Atlas).
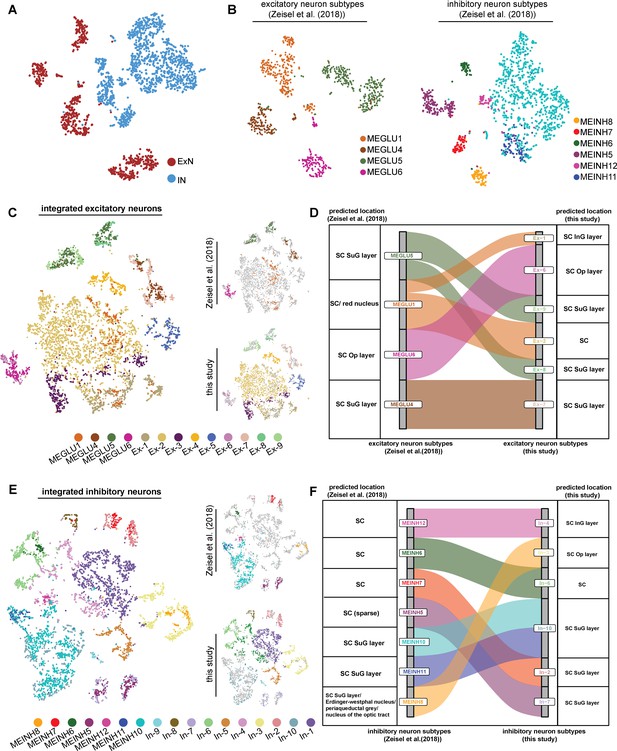
Comparison of neuronal subtypes and spatial mapping results.
(A) t-Distributed stochastic neighbor embedding (t-SNE) plot displaying the distribution of excitatory neurons and inhibitor neurons from Zeisel’s study. (B) t-SNE plots showing the diversity of excitatory neurons and inhibitory neurons from Zeisel’s study. (C) Integration of excitatory neurons from two datasets are visualized by t-SNE plot. (D) Alluvial plot showing the correspondences between excitatory neuron subtypes identified in two studies. (E) Integration of inhibitory neurons from two datasets are visualized by t-SNE plot. (F) Alluvial plot showing the correspondences between inhibitory neuron subtypes identified in two studies.
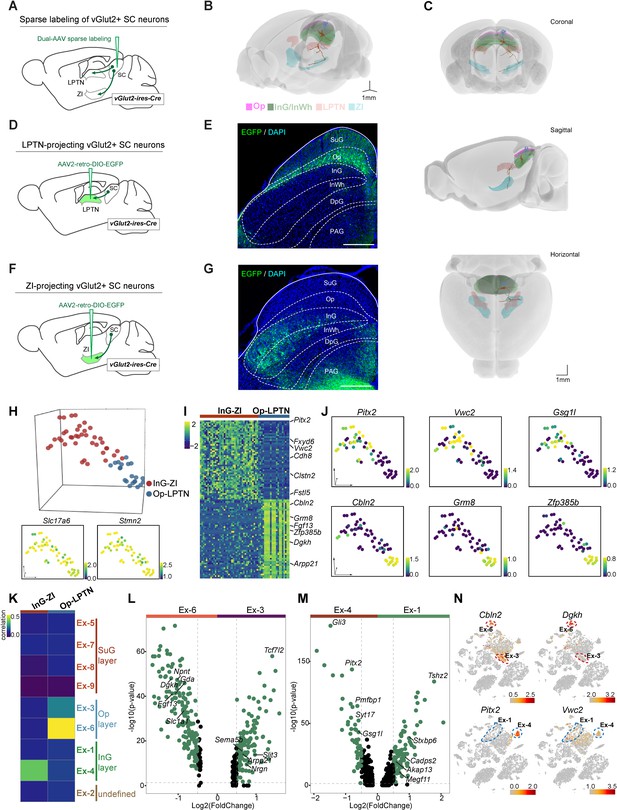
Projection-based analyses of single-cell gene expression profiles.
(A) Schematic diagram showing injection of AAV mixture into the superior colliculus (SC) of vGlut2-IRES-Cre mice for sparse labeling of glutamatergic SC neuron projections. (B) Two reconstructed neurons (blue, cells projected from the optic nerve [Op] to the lateral posterior thalamic nucleus [LPTN]; brown, cells projected from the intermediate gray [InG] to the zona incerta [ZI]) were registered to the mouse brain regions (Op, magenta; LPTN, orange; InG/intermediate white (InWh), green; ZI, cyan). (C) Coronal, sagittal, and horizontal views of reconstructed neurons. Scale bar, 1 mm. (D) Schematic diagram showing injection of AAV2-retro-DIO-EGFP into the LPTN of vGlut2-IRES-Cre mice for labeling of LPTN-projecting glutamatergic SC neurons. (E) Sample micrograph showing the distribution of LPTN-projecting glutamatergic SC neurons labeled by EGFP. (F) Schematic diagram showing injection of AAV2-retro-DIO-EGFP into the ZI of vGlut2-IRES-Cre mice for labeling of ZI-projecting glutamatergic SC neurons. (G) Sample micrograph showing the distribution of ZI-projecting glutamatergic SC neurons labeled by EGFP. (H) Three-dimensional (3D) t-distributed stochastic neighbor embedding (t-SNE) plot showing SC cells sequenced by patch-seq. Upper panel: 3D t-SNE displaying the distribution of Op-LPTN and InG-ZI projection neurons. Cells are colored according to their cell projection identities (Op-LPTN, blue; InG-ZI, red). Lower panel: expression profiles of classic markers Slc17a6 and Stmn2 for VGlut-expressing neurons were projected onto the 3D t-SNE. The scale bar indicates the relative gene expression level (blue, low; yellow, high). (I) Heatmap showing the differentially expressed genes of ZI- and LPTN-projecting SC neurons. The scale bar indicates the relative gene expression level. (J) Expression of genes enriched in LPTN-projecting and ZI-projecting neurons visualized as a t-SNE plot (blue, low; yellow, high). (K) Transcriptional correlation between LPTN-projecting and ZI-projecting neurons (patch-seq) and excitatory neuron subtypes (high-throughput single-nucleus RNA-sequencing [snRNA-seq]). The scale bar indicates the correlation coefficient (blue, low; yellow, high). (L, M) Volcano plot showing the differentially expressed genes in excitatory neuron subtypes Ex-3 and Ex-6 (L) and excitatory neuron subtypes Ex-1 and Ex-4 (M). Each dot represents a gene. Significantly upregulated genes are shown in green. (N) t-SNE plot visualizing the expression of differentially expressed genes in LPTN-projecting (top) and ZI-projecting (bottom) neurons in the same layout used in Figure 1A. The scale bar indicates the relative gene expression level (gray, low; red, high).
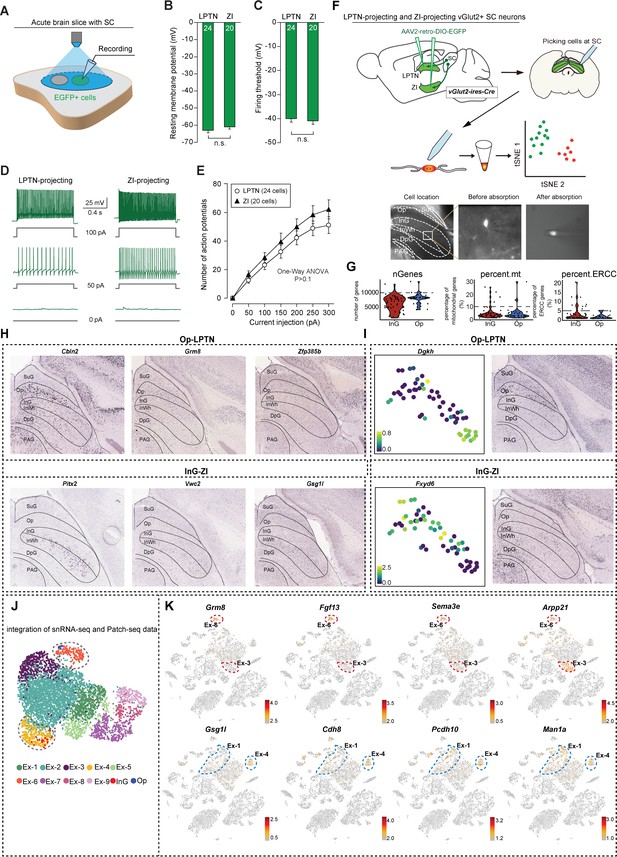
Electrophysiological properties and the expression of differentially expressed genes (DEGs) between lateral posterior thalamic nucleus (LPTN)- and zona incerta (ZI)-projecting superior colliculus (SC) neurons.
(A) Schematic diagram showing current-clamp whole-cell recording from LPTN- or ZI-projecting SC neurons labeled with EGFP. (B, C) Quantitative analyses of resting membrane potential (B) and firing threshold (C) of SC neurons that project to the LPTN (LPTN) and ZI (ZI). (D) Example traces of action potential firing evoked by depolarizing currents injected into EGFP+ LPTN-projecting and ZI-projecting SC neurons. (E) Quantitative analyses of spike number as a function of current intensity. Scale bar is indicated in the graph. (F) Patch-seq strategy of LPTN-projecting neurons from optic nerve (Op) and ZI-projecting neurons from intermediate gray (InG) in SC. A representative example was given to show how the fluorescence labeled cells were picked (bottom). (G) Violin plots showing the quality control metrics of the single-cell RNA-sequencing (scRNA-seq) experiments by patch-seq. Each dot represents an individual cell. Red represents ZI-projecting neurons from the InG layer in SC. Blue represents LPTN-projecting neurons from the Op layer in SC. (H) In situ hybridization visualization (from Allen Brain Atlas) of genes presented in Figure 2J. (I) Examples of gene expression pattern and in situ hybridization visualization (from Allen Brain Atlas) of the DEGs presented in Figure 2I. The scale bar indicates relative gene expression level (blue, low; yellow, high). (J) UMAP showing the integrated single-nucleus RNA-sequencing (snRNA-seq) and patch-seq dataset. Cells were colored according to their cell type identities. (K) t-Distributed stochastic neighbor embedding (t-SNE) plot visualizing the expression of DEGs between LPTN-projecting (upper panel) or ZI-projecting (lower panel) neurons by same layout as Figure 1A. The scale bar indicates the relative gene expression level (gray, low; red, high). Red dash line: Op layer excitatory neuron subclusters.; blue dash line: InG layer excitatory neuron subclusters. Data in (B, C, E) are mean ± SEM. Statistical analyses in (B, C) were performed with Student’s t-test (n.s., p>0.1). Statistical analysis in (E) was performed with one-way ANOVA (n.s., p>0.1). For p-values, see Supplementary file 8.
3D reconstruction of layer-specific neuron projection patterns from SC to downstream brain regions by in vivo sparse-labeling strategy and M-CRITIC, related to Figure 2B–C.
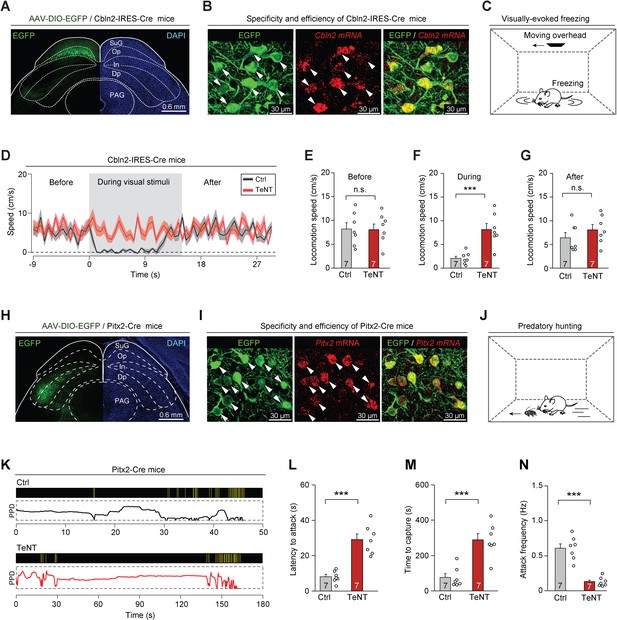
Synaptic inactivation of Cbln2+ and Pitx2+ superior colliculus (SC) neurons.
(A) Sample coronal section showing the restricted distribution of EGFP-expressing neurons in the optic nerve (Op) layer of the SC in Cbln2-IRES-Cre mice. (B) Sample micrographs showing the specificity and efficiency of the Cbln2-IRES-Cre line for labeling of SC neurons expressing Cbln2 mRNA. (C) Schematic diagram showing the behavioral paradigm of the visually evoked freezing response in mice. (D) Time courses of locomotion speed before, during, and after the sweep of an overhead moving visual target in mice without (Ctrl) or with (tetanus neurotoxin [TeNT]) synaptic inactivation of Cbln2+ SC neurons. (E–G) Quantitative analysis of locomotion speed before (E), during (F), and after (G) the sweep of an overhead moving target in mice without (Ctrl) and with (TeNT) synaptic inactivation of Cbln2+ SC neurons. (H) Sample coronal section showing the restricted distribution of EGFP-expressing neurons in the In layer of the SC in Pitx2-Cre mice. (I) Sample micrographs showing the specificity and efficiency of the Pitx2-Cre line for labeling of SC neurons expressing Pitx2 mRNA. (J) Schematic diagram showing the behavioral paradigm of predatory hunting in mice. (K) Behavioral ethograms of predatory hunting in mice without (Ctrl) and with (TeNT) synaptic inactivation of Pitx2+ SC neurons. The yellow vertical lines indicate jaw attacks. The PPD curve shows the time course of prey-predator distance. (L–N) Quantitative analysis of latency to attack (L), time to capture (M), and attack frequency (N) in mice without (Ctrl) and with (TeNT) synaptic inactivation of SC Pitx2+ neurons. The data in (D–G, L–N) are presented as mean ± SEM (error bars). The statistical analyses in (E–G, L–N) were performed using Student’s t-test (n.s. p>0.1; ***p < 0.001). For the p-values, see Supplementary file 8. Scale bars are indicated in the graphs.
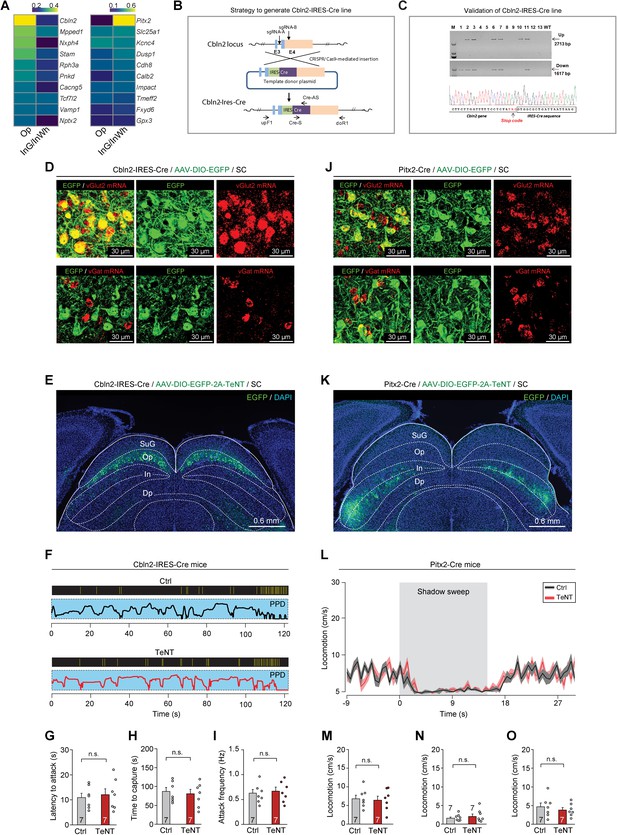
Generation of Cbln2-IRES-Cre mice to test the function of Cbln2+ and Pitx2+ superior colliculus (SC) neurons.
(A) Heatmap showing the computed layer specificity score of 10 differentially expressed genes (DEGs) of lateral posterior thalamic nucleus (LPTN)-projecting and zona incerta (ZI)-projecting neurons. (B) Schematic diagram of CRISPR/Cas9-mediated knock-in of a Cre coding cassette into downstream of Cbln2 gene of mice. The Cre expression was driven by IRES. (C) Top, PCR amplification of Cre cassette knock-in at the endogenous Cbln2 gene locus. Arrows indicate PCR amplification of the genome joint with the left homologous arm (up, 2713 bp) or the right homologous arm (down, 1617 bp), and part of the Cre report cassette. The primers used for PCR amplification are indicated in (B) and shown in Key resources table. PCR amplicons were cloned and sequenced for insertion analysis. Samples #2, #3, #6, #7, and #11 showed correct recombination. The insertion information was furthered confirmed by Sanger sequencing. M indicates marker DL2000 (Takara). Bottom, the chromatographs from the sequence files showed that CRISPR/Cas9-mediated precise insertion at the target locus. (D) Example micrographs showing, in Cbln2-IRES-Cre mice, the Cbln2+ SC neurons labeled by EGFP were predominantly positive for vGlut2 mRNA (first row), while they were mostly negative for vGat mRNA (second row). (E) Example coronal section of Cbln2-IRES-Cre mice showing expression of EGFP and tetanus neurotoxin (TeNT) in Cbln2+ SC neurons that were predominantly distributed in the SC optic nerve (Op) layer. (F) Behavioral ethograms of predatory hunting in example mice without (Ctrl) and with (TeNT) inactivation of Cbln2+ SC neurons. (G–I) Quantitative analyses of latency to attack (G), time to capture (H), and attack frequency (I) of Cbln2-IRES-Cre mice without (Ctrl) and with (TeNT) inactivation of Cbln2+ SC neurons. (J) Example micrographs showing, in Pitx2-Cre mice, the Pitx2+ SC neurons labeled by EGFP were predominantly positive for vGlut2 mRNA (first row), while they were mostly negative for vGat mRNA (second row). (K) Example coronal section of Pitx2-Cre mice showing expression of EGFP and TeNT in Pitx2+ neurons that were predominantly distributed in the SC intermediate layers (In). (L) Time courses of locomotion speed of Pitx2-Cre mice without (Ctrl) and with (TeNT) inactivation of Pitx2+ SC neurons before, during, and after the sweep of overhead moving visual target. (M–O) Quantitative analyses of locomotion speed of Pitx2-Cre mice without (Ctrl) and with (TeNT) inactivation of Pitx2+ SC neurons before (M), during (N), and after (O) the sweep of overhead moving visual target. Data in (G–I, M–O) are mean ± SEM. Statistical analyses in (G–I, M–O) were performed with Student’s t-test (n.s., p>0.1). For p-values, see Supplementary file 8. Scale bars are indicated in the graph.
An example video showing that synaptic inactivation of Cbln2+ SC neurons by TeNT impaired visually-evoked freezing responses.
Related to Figure 3D.
An example video showing that synaptic inactivation of Pitx2+ SC neurons by TeNT impaired prey capture behavior in the arena.
Related to Figure 3K.
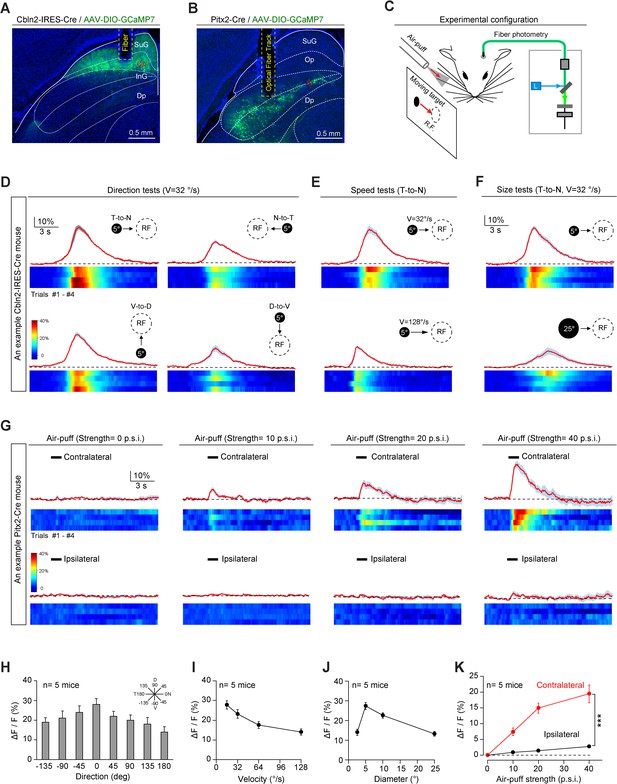
Sensory responses of Cbln2+ and Pitx2+ superior colliculus (SC) neurons.
(A, B) Sample micrographs showing the optical fiber tracks above GCaMP7-positive SC neurons in Cbln2-IRES-Cre (A) and Pitx2-Cre (B) mice. (C) Schematic diagram of the experimental configuration showing vibrissal tactile stimulation (air puff) and visual stimulation; the latter was presented as a black circle moving across the receptive field (RF) on a tangent screen. (D) Normalized GCaMP fluorescence changes (ΔF/F) and heatmaps of Cbln2+ SC neurons in an example mouse in response to a stimulus consisting of a black circle (5°) moving at 32°/s in various directions (T-to-N, N-to-T, V-to-D, D-to-V). N, D, T, and V indicate nasal, dorsal, temporal, and ventral, respectively. (E) Normalized GCaMP fluorescence changes (ΔF/F) and heatmaps of Cbln2+ SC neurons in an example mouse in response to a black circle (5°) moving (T-to-N) at different velocities (32°/s and 128°/s). (F) Normalized GCaMP fluorescence changes (ΔF/F) and heatmaps of Cbln2+ SC neurons in an example mouse in response to black circles of different sizes (5° and 25°) moving in a T-to-N direction at 32°/s. (G) Normalized GCaMP fluorescence changes (ΔF/F) and heatmaps of Pitx2+ SC neurons in an example mouse in response to air puffs of different strengths (0, 10, 20, and, 40 p.s.i.) directed toward the contralateral or ipsilateral vibrissal area. (H) Quantitative analysis of peak GCaMP responses of Cbln2+ SC neurons to black circles moving in eight directions. Inset, eight directions spaced by 45°. (I) Quantitative analysis of the peak GCaMP responses of Cbln2+ SC neurons to black circles moving at different velocities. (J) Quantitative analysis of the peak GCaMP responses of Cbln2+ SC neurons to moving black circles with different diameters. (K) Quantitative analysis of the peak GCaMP responses of Pitx2+ SC neurons to air puffs of different strengths directed toward the contralateral or ipsilateral vibrissal areas. The data in (D–K) are presented as mean ± SEM (error bars). The statistical analyses in (K) were performed by one-way ANOVA (***p < 0.001). For the p-values, see Supplementary file 8. Scale bars are indicated in the graphs.
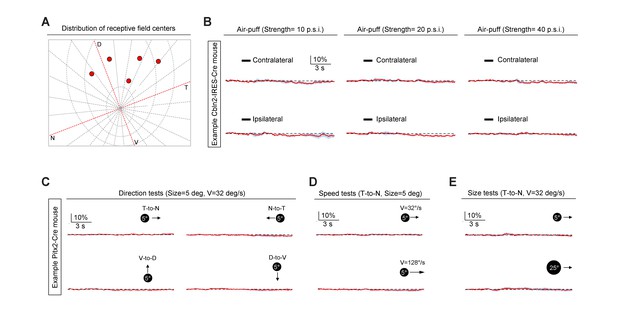
Sensory response properties of Cbln2+ and Pitx2+ superior colliculus (SC) neurons.
(A) Distribution of receptive field centers (red spots) of recorded Cbln2+ SC neurons in five Cbln2-IRES-Cre mice in the meridian plot of the mouse retina on the tangent screen. The coordinate origin of the meridian plot was the visual axis. Concentric circles were spaced by 15° visual angles. Note the receptive fields of Cbln2+ SC neurons were localized within the dorsal quadrant, consistent with their role in detecting aerial predator. (B) Normalized GCaMP fluorescence changes (ΔF/F) of Cbln2+ SC neurons in response to air puffs directed toward contralateral or ipsilateral vibrissal area with different strengths (10, 20, 30 p.s.i.). (C) Normalized GCaMP fluorescence changes (ΔF/F) of Pitx2+ SC neurons in response to a black circle (5°) moving (32°/s) in different directions (T-to-N, N-to-T, V-to-D, D-to-V). N, D, T, and V represent nasal, dorsal, temporal, and ventral, respectively. (D) Normalized GCaMP fluorescence changes (ΔF/F) of Pitx2+ SC neurons in response to a black circle (5°) moving (T-to-N) with different velocities (32°/s and 128°/s). (E) Normalized GCaMP fluorescence changes (ΔF/F) of Pitx2+ SC neurons in response to a moving black circle with different diameters (5° and 25°) (T-to-N, 32°/s).
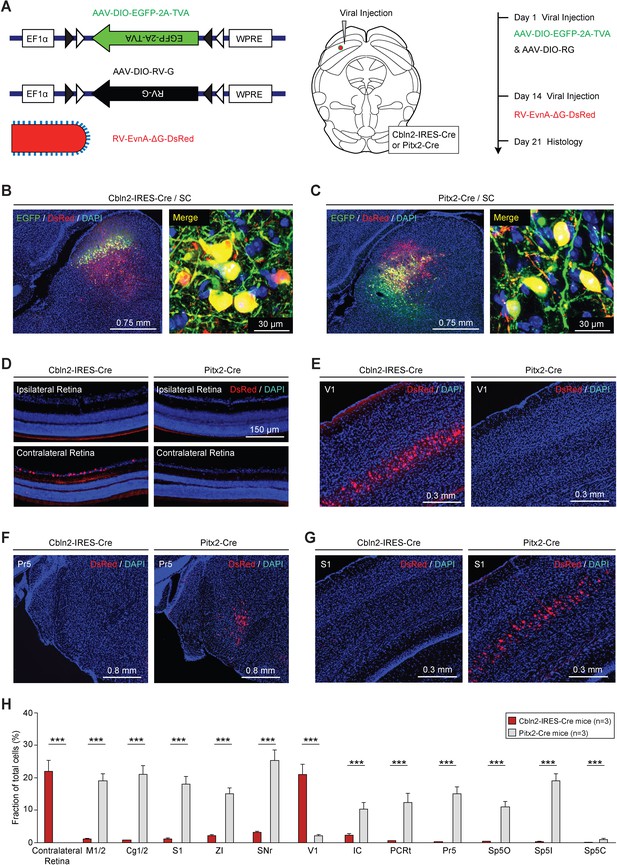
Retrograde tracing of Cbln2+ and Pitx2+ superior colliculus (SC) neurons using rabies virus (RV).
(A) Series of schematic diagrams showing the strategy for monosynaptic retrograde tracing of Cbln2+ and Pitx2+ SC neurons using a combination of AAV and RV. Left, AAV helpers and RV used for injection. Middle, injection into the SC of Cbln2-IRES-Cre and Pitx2-Cre mice. Right, timing of AAV and RV injections. (B, C) Sample micrographs showing the expression of EGFP (green) and DsRed (red) in Cbln2+ and Pitx2+ neurons in the SC of Cbln2-IRES-Cre (B) and Pitx2-Cre mice (C). The dually labeled cells indicate starter cells. For single-channel images, see Figure 5—figure supplement 1. (D–G) Sample micrographs showing DsRed+ cells in various brain regions, including the contralateral and ipsilateral retina (D), the primary visual cortex (V1) (E), the contralateral principal trigeminal nucleus (Pr5) (F), and the ipsilateral primary somatosensory cortex (S1) (G), of Cbln2-IRES-Cre and Pitx2-Cre mice. (H) Fractional distribution of total DsRed-labeled cells in various brain regions that monosynaptically project to Cbln2+ and Pitx2+ SC neurons. The scale bars are labeled in the graphs. The number of mice (H) is indicated in each graph. The data in (H) are presented as mean ± SEM. The statistical analyses in (H) were performed using Student’s t-test (***p < 0.001). For the p-values, see Supplementary file 8.
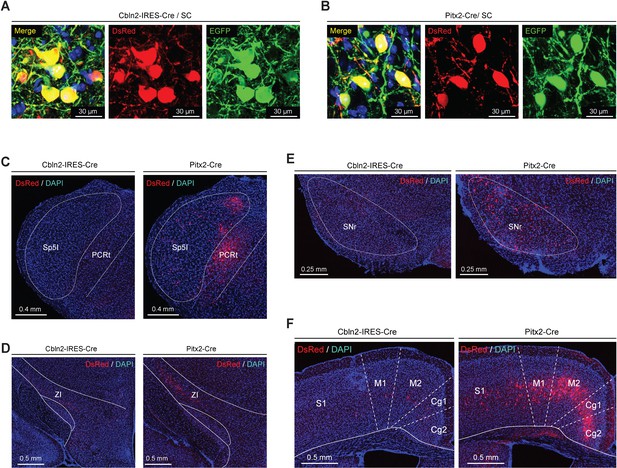
Rabies virus (RV) tracing of Cbln2+ and Pitx2+ superior colliculus (SC) neurons.
(A, B) Example micrographs showing the expression of EGFP (green) and DsRed (red) in the Cbln2+ and Pitx2+ neurons in the SC of Cbln2-IRES-Cre (A) and Pitx2-Cre mice (B), respectively. Note the dually labeled cells indicate starter cells. (C–F) Example micrographs showing DsRed+ cells in different brain regions, including contralateral spinal trigeminal nucleus interpolar part (Sp5I) and the PCRt (C), the ipsilateral zona incerta (ZI) (D), the substantia nigra reticular part (SNr) (E), and the ipsilateral M1/M2 and Cg1/2 (F) of Cbln2-IRES-Cre and Pitx2-Cre mice.
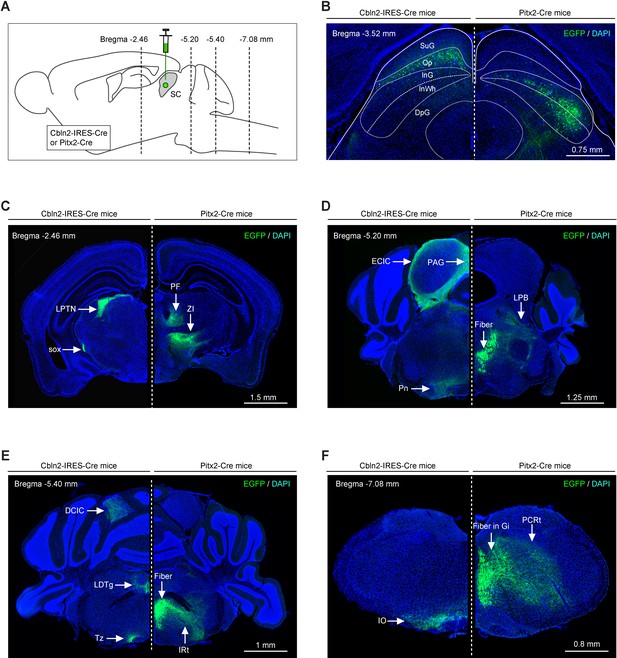
Efferent projections of Cbln2+ and Pitx2+ superior colliculus (SC) neurons are segregated.
(A) Schematic diagram showing the strategy to map the efferent projections of Cbln2+ and Pitx2+ SC neurons. (B) Example coronal sections of Cbln2-IRES-Cre and Pitx2-Cre mice showing the distribution of infected neurons in the SC. (C–F) Example micrographs showing EGFP+ axons of Cbln2+ and Pitx2+ SC neurons in the target brain regions at the level of thalamus (C), midbrain (D), pons (E), and medulla (F). Abbreviations: LPTN, lateral posterior thalamic nucleus; PF, parafascicular nucleus; ZI, zona incerta; Pn, pontine nucleus; LPB, lateral parabrachial nucleus; IO, inferior olive; PCRt, parvicellular reticular nucleus; LDTg, laterodorsal tegmental nucleus; IRt, intermediate reticular nucleus; PAG, periaqueductal gray; ECIC, external cortex of the inferior colliculus; Tz, nucleus of the trapezoid body. Scale bars are labeled in the graphs.
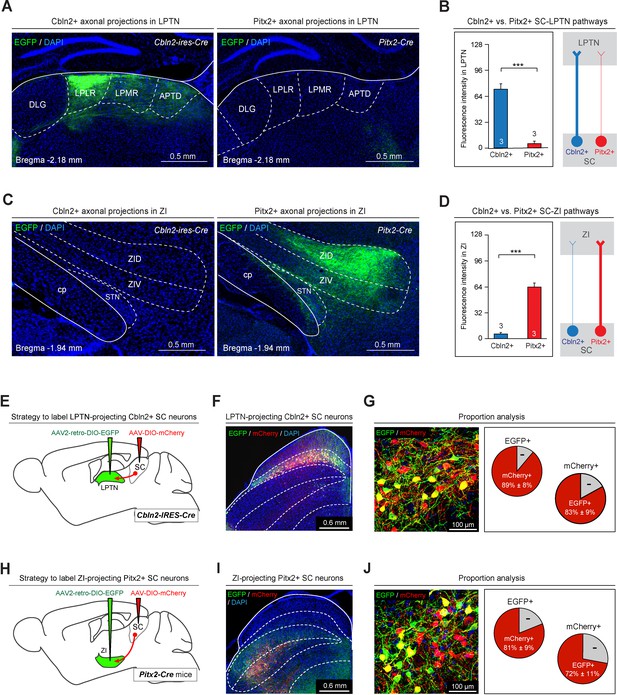
Anterograde and retrograde mapping of Cbln2+ superior colliculus (SC)-lateral posterior thalamic nucleus (LPTN) and Pitx2+ SC-zona incerta (ZI) pathways.
(A) Sample micrographs showing the distribution of EGFP-positive axons in the LPLR and LPMR (collectively the LPTN) of Cbln2-IRES-Cre (left) and Pitx2-Cre (right) mice. (B) Quantitative analysis of fluorescence signals from EGFP+ axons in the LPTN of Cbln2-IRES-Cre and Pitx2-Cre mice. (C) Sample micrographs showing the distribution of EGFP-positive axons in the ZI of Cbln2-IRES-Cre (left) and Pitx2-IRES-Cre (right) mice. (D) Quantitative analysis of fluorescence signals from EGFP+ axons in the ZI of Cbln2-IRES-Cre and Pitx2-Cre mice. (E) Schematic diagram showing the viral injection strategy used to label LPTN-projecting Cbln2+ SC neurons. (F) Coronal section from a Cbln2-IRES-Cre mouse showing the distribution of Cbln2+ SC neurons labeled by AAV2-retro-DIO-EGFP and AAV-DIO-mCherry. (G) Sample micrograph (left) and quantitative analysis (right) showing the number of LPTN-projecting Cbln2+ SC neurons (EGFP+) relative to total Cbln2+ SC neurons (mCherry+). (H) Schematic diagram showing the viral injection strategy used to label ZI-projecting Pitx2+ SC neurons. (I) Coronal section from a Pitx2-Cre mouse showing the distribution of Pitx2+ SC neurons labeled by AAV2-retro-DIO-EGFP and AAV-DIO-mCherry. (J) Sample micrograph (left) and quantitative analysis (right) showing the number of ZI-projecting Pitx2+ SC neurons (EGFP+) relative to total Pitx2+ SC neurons (mCherry+). The data in A, B, D, F, I, and L are presented as mean ± SEM (error bars). The statistical analyses in D and F were performed using Student’s t-test (***p < 0.001). For the p-values, see Supplementary file 8. Scale bars are indicated in the graphs.
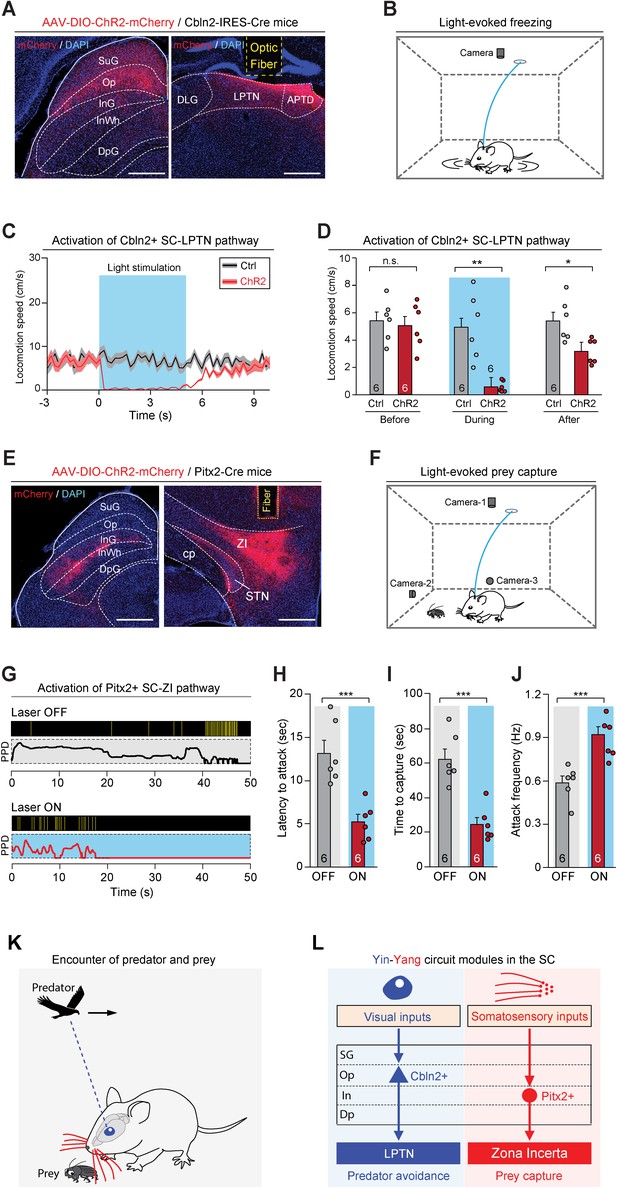
Activation of the Cbln2+ superior colliculus (SC)-lateral posterior thalamic nucleus (LPTN) and Pitx2+ SC-zona incerta (ZI) pathways.
(A) Sample micrographs showing the expression of ChR2-mCherry in the Cbln2+ SC neurons of Cbln2-IRES-Cre mice (left) and the optical fiber track above the ChR2-mCherry+ axons in the LPTN (right). (B) Schematic diagram showing the behavioral paradigm for the light-evoked freezing response in an arena. (C) Time courses of the locomotion speed of mice before, during, and after light stimulation (10 Hz, 20 ms, 5 mW, 5 s) of the Cbln2+ SC-LPTN pathway expressing ChR2-mCherry (ChR2) or mCherry (Ctrl). (D) Quantitative analysis of the locomotion speed of mice before, during, and after light stimulation of the Cbln2+ SC-LPTN pathway expressing ChR2-mCherry (ChR2) or mCherry (Ctrl). (E) Sample micrographs showing the expression of ChR2-mCherry in the Pitx2+ SC neurons of Pitx2-Cre mice (left) and the optical fiber track above the ChR2-mCherry+ axons in the ZI (right). (F) Schematic diagram showing the behavioral paradigm for prey capture paired with light stimulation of the Pitx2+ SC-ZI pathway. (G) Behavioral ethograms of predatory hunting in mice without (laser OFF) and with (laser ON) light stimulation of the Pitx2+ SC-ZI pathway. (H–J) Quantitative analyses of latency to attack (H), time to capture (I), and attack frequency (J) in mice without (OFF) and with (ON) light stimulation of the Pitx2+ SC-ZI pathway. (K) Schematic diagram showing a mouse encountering a cruising aerial predator or a terrestrial prey in the natural environment. (L) Yin-Yang circuit modules formed by Cbln2+ and Pitx2+ SC neurons and their downstream target areas. The Cbln2+ SC neurons in the ‘Yin’ module detect the sensory features of cruising aerial predators and initiate freezing as a defensive response for the avoidance of predators through the Cbln2+ SC-LPTN pathway. The Pitx2+ SC neurons in the Yang module mediate tactile-triggered prey capture behavior through the Pitx2+ SC-ZI pathway. The data in (C, D, H–J) are presented as mean ± SEM (error bars). The statistical analyses in (D, H–J) were performed using Student’s t-test (n.s. p>0.1; *p<0.05; **p < 0.01; ***p<0.001). For the p-values, see Supplementary file 8. Scale bars are indicated in the graphs.
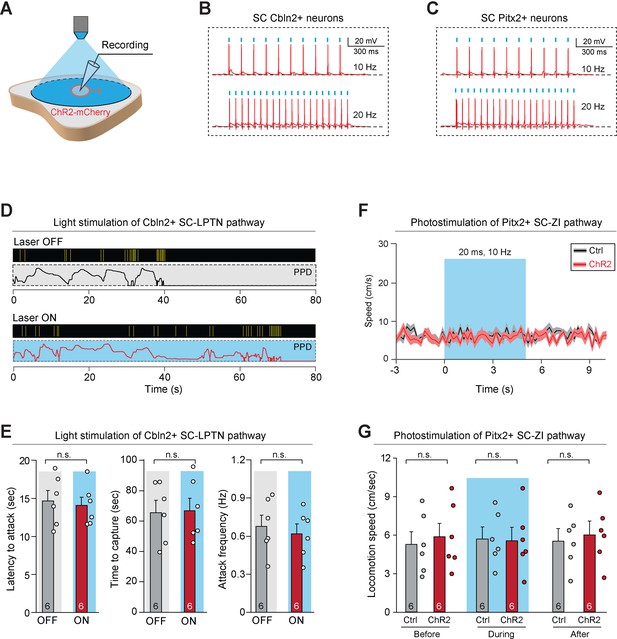
Activation of Cbln2+ superior colliculus (SC)-lateral posterior thalamic nucleus (LPTN) pathway and Pitx2+ SC-zona incerta (ZI) pathway.
(A) Schematic diagram showing whole-cell recording from ChR2-mCherry+ SC neurons in the acute brain slice. (B, C) Example traces of phase-locked spiking activity evoked by light-pulse train (1 ms, 5 mW) in 10 Hz (top) or 20 Hz (bottom) from Cbln2+ SC neurons (B) or Pitx2+ SC neurons (C) that expressed ChR2-mCherry in acute brain slices. (D) Behavioral ethogram of predatory hunting in mice without (laser OFF) and with (laser ON) light stimulation of Cbln2+ SC-LPTN pathway. (E) Quantitative analyses of latency to attack (left), time to capture (middle), and attack frequency (right) in mice without (laser OFF) and with (laser ON) light stimulation of Cbln2+ SC-LPTN pathway. (F) Time courses of locomotion speed before, during, and after light stimulation of Pitx2+ SC-ZI pathway that expressed mCherry (Ctrl) or ChR2-mCherry (ChR2). (G) Quantitative analyses of locomotion speed before, during, and after light stimulation of Pitx2+ SC-ZI pathway that expressed mCherry (Ctrl) or ChR2-mCherry (ChR2). Data in (E–G) are mean ± SEM. Statistical analyses in (E, G) were performed with Student’s t-test (n.s., p>0.1). For p-values, see Supplementary file 8.
An example video showing that light stimulation of ChR2-mCherry+ axon terminals of Cbln2+ SC neurons in the LPTN induced freezing response in mice.
Related to Figure 8B–C.
An example video showing light stimulation of ChR2-mCherry+ axon terminals of Pitx2+ SC neurons in the ZI promoted predatory hunting behavior in mice, related to Figure 8F–G.
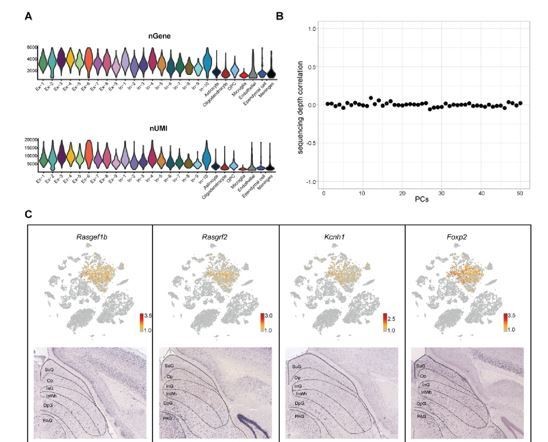
Quality control metrices and spatial distribution of Ex-2.
(A) Violin plots showing the number of detected genes (nGene) and unique molecular identifiers (uUMI) of each cell types identified in the SC. (B) Scatter plot displaying the correlation between computed reduced dimension components (PCs) and sequencing depth. (C) Spatial distribution of the DEGs enriched in Ex2. Upper panel: gene expression among excitatory neurons were projected onto the two-dimensional t-SNE. Cells were colored according to relative gene expression level (gray, low; red, high). Lower panel: In situ hybridization staining of mouse superior colliculus for the identified DEGs (from the Allen Brain Atlas).
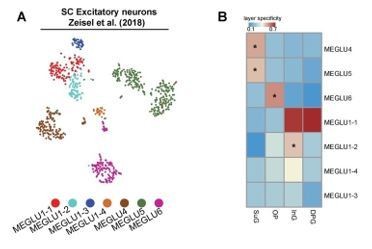
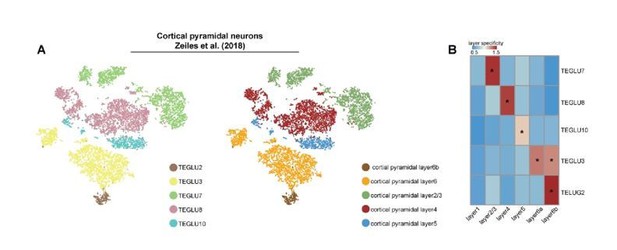
Comparison of mapping results for SC.
(A) tSNE displaying the spatial distribution of excitatory neuron subtypes from Zeisel’s study. (B) Heatmap showing the layer specificity for each subtype computed by SPACED.
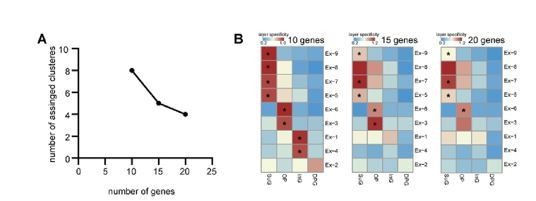
Impact of the number of selected genes on mapping results of SPACED.
(A) Scatter plot showing the relation between number of selected genes and the number of assigned clusters. (B) Heatmap showing the computed layer specificity for each excitatory neuron subtype with 10 DEGs (left), 15 DEGs (middle) and 20 DEGs (right). * p < 0.05.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Mus musculus) | Pitx2-Cre mice | Mutant Mouse Resource Center | Cat# 000126-UCD, RRID:MMRRC_000126-UCD | |
| Strain, strain background (Mus musculus) | vGlut2-IRES-Cre mice | JAX Mice | Cat# JAX:028863, RRID:IMSR_JAX:028863 | |
| Strain, strain background (Mus musculus) | Cbln2-IRES-Cre mice | NIBS | NA | |
| Genetic reagent (virus) | AAV-EF1α-DIO-EGFP-2A-TeNT | Thomas Südhof Lab at Stanford University | NA | |
| Genetic reagent (virus) | pAAV-EF1α-DIO-ChR2-mCherry | Addgene | RRID:Addgene_20297 | |
| Genetic reagent (virus) | AAV-EF1α-DIO-jGCaMP7s | Addgene | RRID:Addgene_104463 | |
| Genetic reagent (virus) | AAV2-retro-hSyn-DIO-EGFP | TaiTool | NA | |
| Genetic reagent (virus) | AAV2/9-hSyn-DIO-EGFP-2A-TeNT | TaiTool | NA | |
| Genetic reagent (virus) | AAV2/9-hSyn-DIO-EGFP | TaiTool | NA | |
| Genetic reagent (virus) | AAV2/9-hSyn-DIO-ChR2-mCherry | TaiTool | NA | |
| Genetic reagent (virus) | AAV2/9-hSyn-DIO-mCherry | TaiTool | NA | |
| Genetic reagent (virus) | AAV2/9-hSyn-DIO-jGCaMP7s | TaiTool | NA | |
| Genetic reagent (virus) | AAV2/9-EF1α-DIO-EGFP-2A-TVA | BrainVTA | NA | |
| Genetic reagent (virus) | AAV2/9-EF1α-DIO-RV-G | BrainVTA | NA | |
| Genetic reagent (virus) | RV-EnvA-ΔG-DsRed | BrainVTA | NA | |
| Antibody | Rabbit polyclonal anti-EGFP | Abcam | Cat# ab290, RRID:AB_303395 | |
| Antibody | Polycolonal anti-mCherry | Abcam | Cat# ab167453, RRID:AB_2571870; Cat# ab205402, RRID:AB_2722769 | |
| Sequence-based reagent | M-Cbln2-cre-upF | Tsingke Biological Technology, China | Primer | 5’-GGTACCTACTGTGTATCGCCAG-3’ |
| Sequence-based reagent | CRE-AS | Tsingke Biological Technology, China | Primer | 5’-CTGTTTCACTATCCAGGTTACG-3’ |
| Sequence-based reagent | CRE-S | Tsingke Biological Technology, China | Primer | 5’-TACTGACGGTGGGAGAATG-3’ |
| Sequence-based reagent | M-Cbln2-ires-cre-doR | Tsingke Biological Technology, China | Primer | 5’-GTTTGAAGCTGCACTGAGAGAG-3’ |
| Chemical compound, drug | D-AP5/CNQX | Tocris | Cat# 0106 / 0190 | |
| Chemical compound, drug | Picrotoxin/TTX | Tocris | Cat# 1128 / 1078 | |
| Chemical compound, drug | 4-AP | Sigma | Cat# 275875 | |
| Chemical compound, drug | DAPI | Sigma | Cat# D8417 | |
| Software, algorithm | GraphPad Prism 9.0.0 | GraphPad, 2015 | https://www.graphpad.com/scientific-software/prism/ | |
| Software, algorithm | Cell ranger 3.0.2 | Zheng et al., 2017 | http://10xgenomics.com | |
| Software, algorithm | R version 3.6.1 | R Development Core Team, 2020 | https://www.r-project.org | |
| Software, algorithm | Seurat 3.1.0 | Stuart et al., 2019 | https://satijalab.org/seurat/ | |
| Software, algorithm | Scrublet 0.2.1 | Wolock et al., 2019 | https://github.com/swolock/scrublet | |
| Software, algorithm | batchelor 1.0.1 | Haghverdi et al., 2018 | https://github.com/MarioniLab/MNN2017/ | |
| Software, algorithm | pheatmap 1.0.12 | Kolde, 2019 | https://github.com/raivokolde/pheatmap | |
| Software, algorithm | rgl 0.100.54 | Adler and Murdoch, 2020 | https://github.com/dmurdoch/rgl | |
| Software, algorithm | metascape | Zhou et al., 2019 | https://metascape.org/gp/index.html#/main/step1 | |
| Software, algorithm | Image J v1.48h3 | Schneider et al., 2012 | https://imagej.nih.gov/ij/ | |
| Software, algorithm | MATLAB 2019b | MATLAB, 2018 | https://www.mathworks.com | |
| Software, algorithm | Brainrender 2.0 | Claudi et al., 2020 | https://github.com/brainglobe/brainrender |
Additional files
-
Supplementary file 1
Sample information for high-throughput single-nucleus RNA-sequencing (snRNA-seq) of nucleus from superior colliculus (SC) and top 50 differentially expressed genes among nine major cell types in SC, related to Figure 1A.
- https://cdn.elifesciences.org/articles/69825/elife-69825-supp1-v2.xlsx
-
Supplementary file 2
Differentially expressed genes among 9 excitatory neuron subtypes and 10 inhibitory neuron subtypes, related to Figure 1C–D.
- https://cdn.elifesciences.org/articles/69825/elife-69825-supp2-v2.xlsx
-
Supplementary file 3
Sample information for patch-seq of neurons from superior colliculus (SC), related to Figure 2H.
- https://cdn.elifesciences.org/articles/69825/elife-69825-supp3-v2.xlsx
-
Supplementary file 4
Differentially expressed genes between zona incerta (ZI)- and lateral posterior thalamic nucleus (LPTN)-projecting neurons in superior colliculus (SC), related to Figure 2I.
- https://cdn.elifesciences.org/articles/69825/elife-69825-supp4-v2.xlsx
-
Supplementary file 5
Differentially expressed genes between excitatory neuron subtype Ex-3 and Ex-6, and between Ex-1 and Ex-4, related to Figure 2L–M, respectively.
- https://cdn.elifesciences.org/articles/69825/elife-69825-supp5-v2.xlsx
-
Supplementary file 6
Summary of all experimental designs.
- https://cdn.elifesciences.org/articles/69825/elife-69825-supp6-v2.xlsx
-
Supplementary file 7
Summary of cell-counting strategies.
- https://cdn.elifesciences.org/articles/69825/elife-69825-supp7-v2.xlsx
-
Supplementary file 8
Summary of statistical analyses.
- https://cdn.elifesciences.org/articles/69825/elife-69825-supp8-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/69825/elife-69825-transrepform-v2.docx