Hemodynamic molecular imaging of tumor-associated enzyme activity in the living brain
Figures
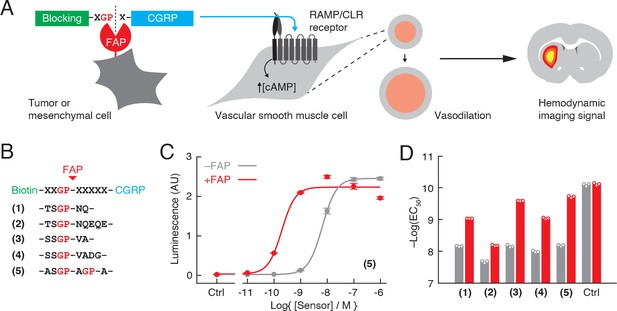
Design of fibroblast activation protein (FAP)-sensitive vasoactive imaging probes.
(A) FAP-sensitive vasoprobes are constructed by fusing the vasoactive peptide calcitonin gene-related peptide (CGRP) (cyan) via a FAP-cleavable linker to a steric blocking domain (green) that prevents CGRP from acting on its receptor when present. Cleavage by cell surface-expressed FAP (red) removes the blocking domain and allows the CGRP domain to stimulate the RAMP/CLR receptor on vascular smooth muscle cells, promoting intracellular cAMP accumulation, vasodilation, and production of consequent hemodynamic imaging signals (right). (B) Candidate FAP sensors designed with biotin as a blocking domain and FAP-sensitive linkers for constructs (1–5) as shown. (C) Luminescence readouts obtained using a cell-based RAMP/CLR activation bioassay during stimulation with varying concentrations of construct (5), with (red) or without (gray) pretreatment by 10 ng/µL FAP. Data points reflect mean and SD (error bars) of three technical replicates each. (D) Midpoints for receptor activation (EC50 values) measured using the bioassay of panel (C) for constructs (1–5) and control peptide (CGRP) with (red) or without (gray) FAP pretreatment.
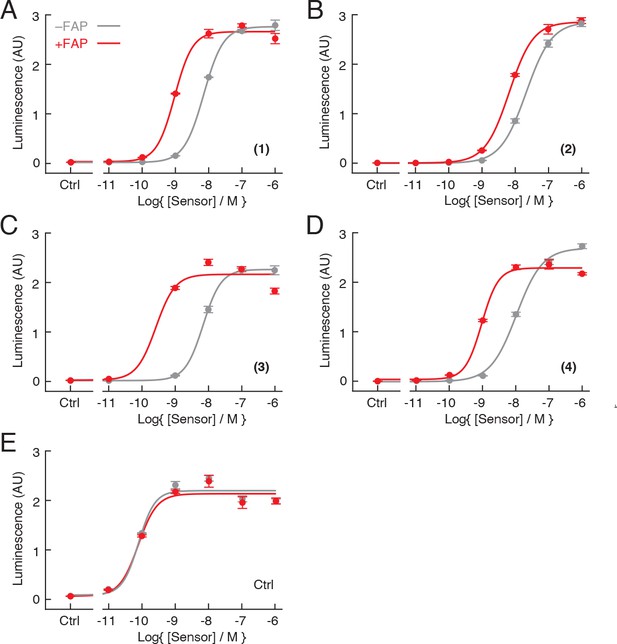
Assessment of candidate fibroblast activation protein (FAP) sensors in vitro.
Measurements of RAMP/CLR receptor activation obtained using the bioassay technique of Figure 1C for candidate FAP sensors (A–D) and α-calcitonin gene-related peptide (α-CGRP) control peptide (E), with (red) or without (gray) pretreatment by 10 ng/µL FAP. Data points reflect mean and SD (error bars) of three technical replicates each.
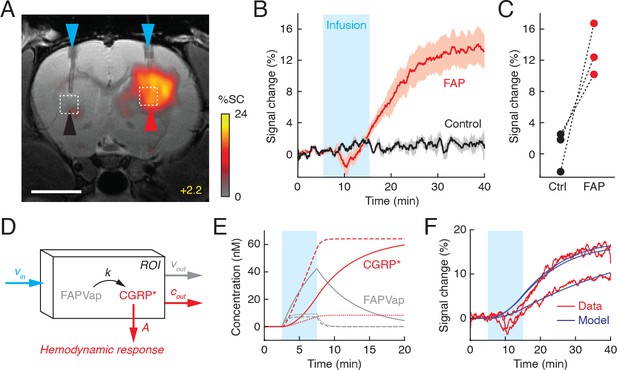
Fibroblast activation protein-sensitive vasoprobe (FAPVap) reports FAP enzyme activity in vivo.
(A) MRI data from an individual rat implanted with FAP-expressing or control tumor cells (red and black arrowheads, respectively) in the dorsal striatum at bregma +2.2 mm. 100 nM FAPVap was infused over the tumors via cannulae (blue arrowheads), resulting in signal changes observed in the presence but not the absence of FAP expression. Color scale denotes percent T2-weighted MRI signal changes (%SC) recorded after infusion with respect to the pre-infusion baseline. Grayscale image underlay is an anatomical scan. Scale bar = 4 mm. (B) Mean time courses of MRI signal expressed as percent change observed before, during (cyan box), and after FAPVap infusion from regions of interest (ROIs) defined with respect to FAP-expressing (red) or control (black) xenograft locations in three animals (cf. dashed boxes in A). Shading denotes SEM (n = 3). (C) Magnitudes of FAPVap responses observed in paired probe injections over control (Ctrl) and FAP-expressing tumors. (D) Diagram of a kinetic model accounting for rates of FAPVap injection (vin), FAP-catalyzed conversion from FAPVap to CGRP* (k), and efflux of FAPVap and CGRP* (vout and cout) from an ROI. Hemodynamic responses are related to CGRP* concentration by a constant of proportionality, A. (E) Concentration dynamics of FAPVap (gray) and CGRP* (red) simulated using the model of (D). Calculations assume that 100 nM FAPVap is infused into an ROI containing 1 nM (solid lines) or 10 nM FAP (dashed lines) with vout = 0, or containing 1 nM FAP with vout = 0.01 s–1 (dotted lines). See text for further details. (F) Fitting of model curves (blue) to ROI-averaged FAPVap imaging time courses (red) from three animals, showing that broad features of the MRI signal are closely approximated, with the exception of suspected pressure artifacts during the infusion period.
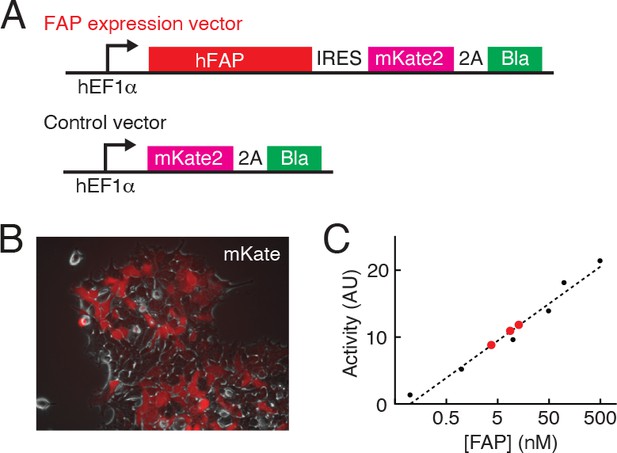
Fibroblast activation protein (FAP) expression in transfected cells.
(A) Constructs for directing FAP expression (top) or control mKate expression (bottom) in stably transfected HEK293FT cells. (B) Visualization of mKate expression in FAP-expressing cells. (C) Quantification of FAP expression from three aliquots of transfected cells (red data points) with respect to calibration standards (black data points). Assay wells contained ~105 cells in a 100 µL volume. FAP expression measurements correspond to free enzyme concentrations of 3–13 nM.
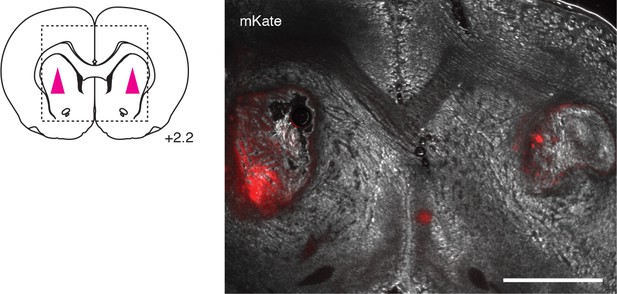
Microscopic visualization of brain xenografts.
Position of xenografts for the experiments of Figure 2 indicated at left in a brain atlas slice at bregma = 2.2 mm. Micrograph at right shows xenografts in situ, with mKate expression reflected by red fluorescence. Scale bar = 2 mm.
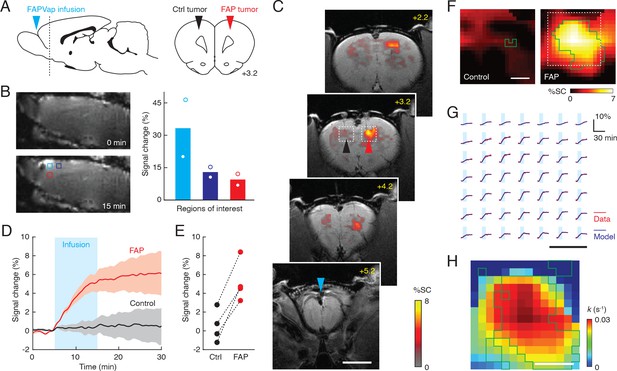
Wide-field imaging of fibroblast activation protein (FAP)-expressing tumors in vivo.
(A) Geometry employed for wide-field molecular imaging of FAP activity in rat brain tumors. Blue arrowhead denotes probe infusion site. Horizontal dashed line in sagittal view (left) denotes position of coronal section (right) showing location of implanted FAP-expressing and control tumors with respect to brain atlas sections (Paxinos and Watson, 2009). (B) Infusion of gadolinium-diethylenetriamine pentaacetic acid (Gd-DTPA) via the intra-cerebrospinal fluid (intra-CSF) route diagrammed in (A) results in broad signal enhancements when comparing T1-weighted MRI scans obtained before (top left) and after (bottom left) contrast agent infusion. Plot at right shows signal changes observed in the three color-coded regions of interest (ROIs) shown at bottom left, indicating inverse dependence of signal enhancements on distance from the infusion site (n = 2). (C) Coronal scans of a rat showing infusion location (bregma +5.2 mm) and profiles of T2*-weighted MRI signal change (color) following FAP-sensitive vasoprobe (FAPVap) infusion in the presence of control (black arrowhead) and FAP-expressing (red arrowhead) xenografts. The gray underlay is an anatomical scan and the scale bar = 4 mm. (D) Mean signal change time courses with respect to baseline averaged over ROIs defined with respect to xenografts in four animals. Cyan box denotes the FAPVap infusion period, and shaded intervals denote SEM (n = 4). (E) FAPVap response amplitudes corresponding to the 15 min time point in (D). (F) Maps of mean MRI signal changes observed near FAP-expressing and control tumors (n = 4 each), corresponding to boxed regions as in panel (C). Green outlines denote voxels with responses significantly different from zero (t-test p≤0.05, n = 4). Scale bar = 0.5 mm. (G) Analysis of time courses in the boxed region of (F) using the kinetic model of Figure 2D, showing comparison of fitted models (blue) to data (red) in a regular grid of individual voxels evenly spaced over the dashed box in panel (F). Scale bar = 0.5 mm. (H) Values of the FAP catalysis rate k obtained by fitting the FAPVap kinetic model to the data in the dashed box of panel (F). Green outlines denote values with t-test p≤0.05 (n ≥ 2). Scale bar = 0.5 mm.
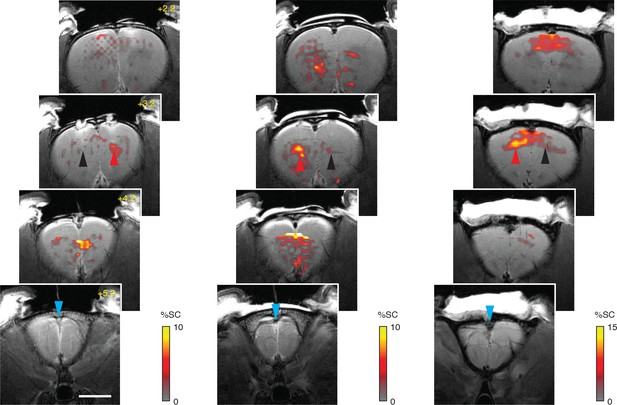
Wide-field molecular imaging of fibroblast activation protein (FAP) activity in rat brain.
Sets of four coronal scans from three rats, analogous to the data in Figure 3C, with cyan arrowheads indicating infusion location (bregma +5.2 mm) and profiles of MRI signal change (color) following FAP-sensitive vasoprobe (FAPVap) infusion in the presence of control (black arrowhead) and FAP-expressing (red arrowhead) xenografts. Gray underlays are corresponding anatomical scans and the scale bar = 4 mm.
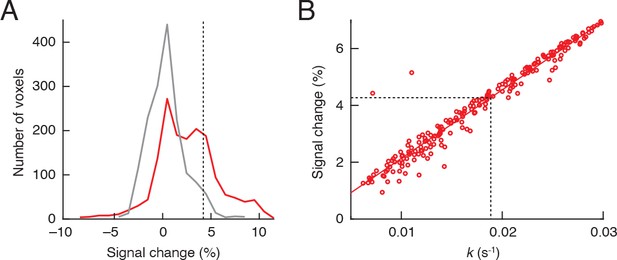
Fibroblast activation protein (FAP) activity detection sensitivity.
(A) Comparison of voxel-level signal changes following FAP-sensitive vasoprobe (FAPVap) injection over control (gray) and FAP-expressing (red) HEK cell xenografts. Histograms are computed from individual animals that contributed to mean signal changes shown in the peri-xenograft regions of interest (ROIs) of Figure 3F. Dashed line indicates 2 standard deviations above the mean value of control signal changes, approximately 4.3%. (B) Comparison of k values with corresponding mean signal changes for voxels with k > 0.005 s–1 in the data of Figure 3H, indicating a largely linear relationship with regression line denoted in red. Black dashed lines indicate signal change and corresponding minimum k value that could be recognizable at a single voxel level in individual animals, according to the criterion defined in panel (A) without correction for multiple hypotheses. The resulting k value of 0.019 s–1 corresponds to a FAP concentration of approximately 13 nM, according to kinetic constants reported by Edosada et al., 2006 and the relationship that k = kcat[FAP]/Km.
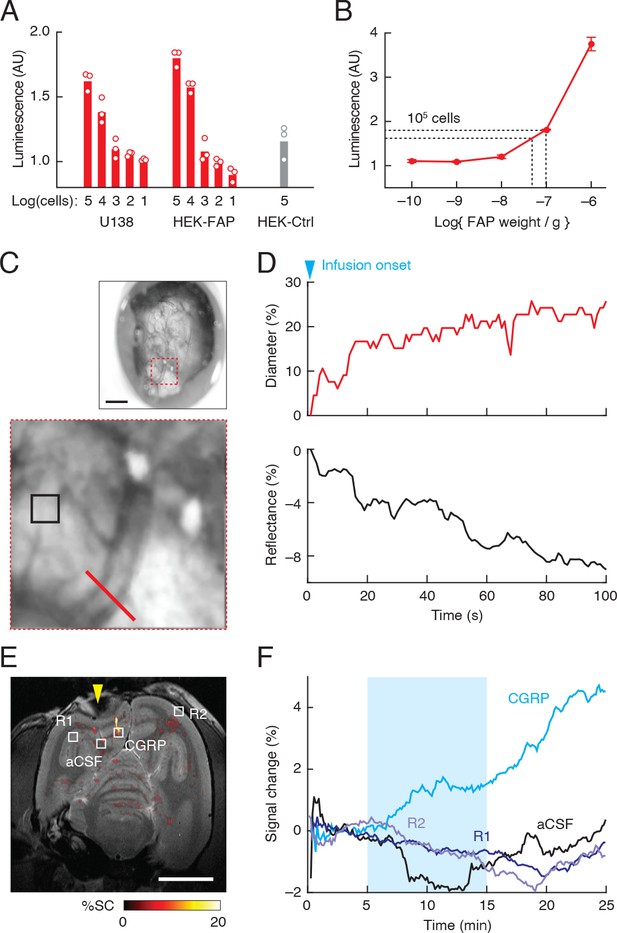
Clinical applicability of fibroblast activation protein-sensitive vasoprobe (FAPVap)-based imaging.
(A) Luminescence response measured after addition of FAPVap to varying numbers of U138 human glioma cells, FAP-transfected HEK cells (HEK-FAP) and control untransfected HEK cells (HEK-Ctrl), using the bioassay of Figure 1. Values normalized to the response to uncleaved FAPVap without cells. (B) FAPVap processing by known amounts of FAP under the same assay conditions as (A). Dashed lines indicate mean assay results obtained from 105 U138 and HEK-FAP cells, indicating the presence of roughly 1 pg FAP/cell, according to the standard curve. (C) Optical imaging of responses to vasoprobe infusion into a cranial chamber (top, Figure 4—figure supplement 1) implanted over the cortex of a squirrel monkey. Scale bar = 1 mm. Close-up (bottom) corresponds to red dashed square at top and shows a single blood vessel (red bar), as well as reflective parenchymal tissue (black square). (D) Top: diameter of the vessel labeled in (C) over 100 s immediately following onset of perfusion with 100 nM calcitonin gene-related peptide (CGRP), equivalent to cleaved FAPVap. Bottom: reflectance from the parenchymal region of interest (ROI) labeled in (E) over the same time period. (E) Map of MRI signal changes elicited by injection of CGRP or artificial cerebrospinal fluid (aCSF) into the squirrel monkey brain. R1 and R2 denote two uninjected control regions. Yellow arrowhead denotes the position of the cranial chamber in (C). Scale bar = 1 cm. (F) Time courses of MRI signal change observed in the four regions designated in (E), before, during (cyan shading), and after vasoprobe infusion.
Additional files
-
Supplementary file 1
Sequences of candidate fibroblast activation protein (FAP) probes.
- https://cdn.elifesciences.org/articles/70237/elife-70237-supp1-v1.docx
-
Supplementary file 2
Fibroblast activation protein (FAP) expression plasmid sequence.
- https://cdn.elifesciences.org/articles/70237/elife-70237-supp2-v1.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/70237/elife-70237-transrepform1-v1.docx