Mapping dopaminergic projections in the human brain with resting-state fMRI
Figures
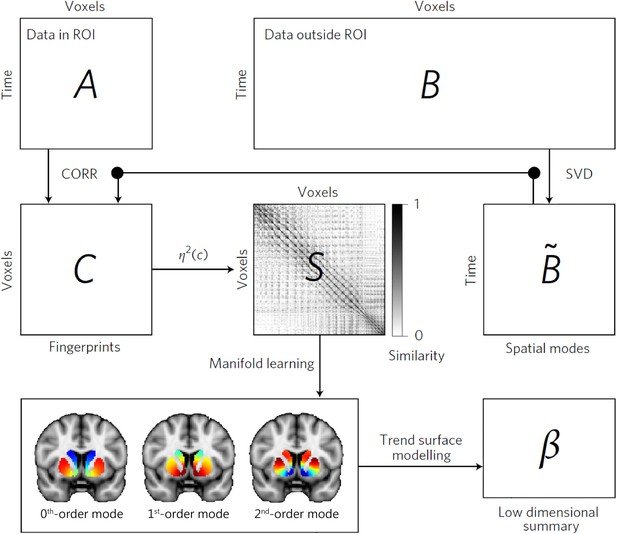
The connectopic mapping pipeline.
The functional MRI (fMRI) time-series data from a predefined region of interest (ROI), here the striatum, are rearranged into a time-by-voxels matrix A, as are the time series from all voxels outside the ROI (matrix B). For reasons of computational tractability, the dimensionality of B is losslessly reduced using singular value decomposition (SVD), yielding ∼B. For every voxel within the ROI, its connectivity fingerprint is computed as the Pearson’s correlation (CORR) between the voxel-wise time-series and the SVD-transformed data, yielding matrix C. Then similarity between voxels is computed using the η2 coefficient, resulting in matrix S. Manifold learning using Laplacian eigenmaps is then applied to this matrix, yielding a set of overlapping, but independent, connection topographies or ‘connectivity modes’ that together describe the functional organization of the striatum. These connection topographies indicate how the connectivity profile with the rest of the brain changes across striatum. Voxels that have similar colours in these connectivity modes have similar connectivity patterns with the rest of the brain. Finally, trend surface modelling is applied to summarize the connectivity modes by fitting a set of trend coefficients (β) that optimally combine a set of spatial polynomial basis functions. See Haak et al., 2018 for further details.
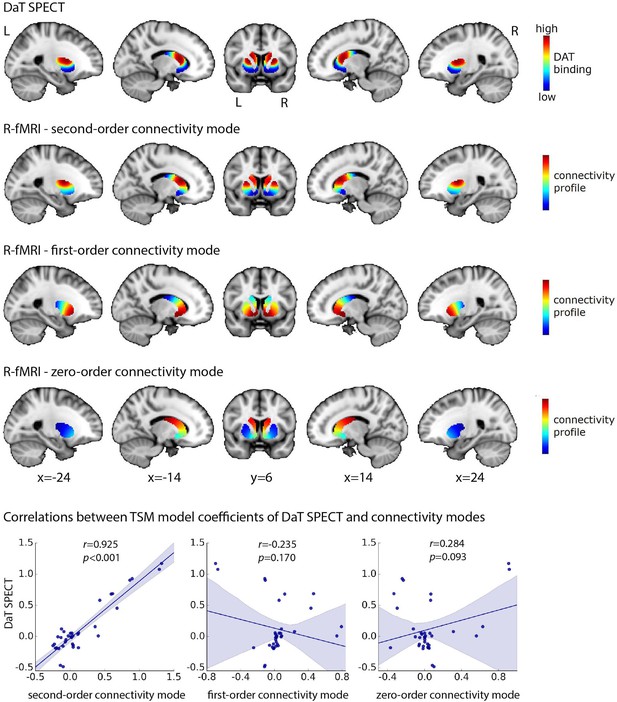
High spatial correspondence between the second-order mode of connectivity in striatum and the DaT SPECT image.
The figure displays the DaT SPECT image averaged across 209 Parkinson’s Progression Markers Initiative (PPMI) controls and the group-level connectivity modes obtained in 839 Human Connectome Project (HCP) subjects. The group-level modes were modelled separately for the left and right putamen and caudate-nucleus accumbens (caudate-NAcc) subregions and have been combined in this figure to aid in visualization. The voxel-wise spatial correlation between the second-order mode of connectivity in striatum and the DaT SPECT image is very high: r = 0.884 (p<0.001). Similarly, the correlation between the orthogonal trend surface model (TSM) coefficients modelling the second-order connectivity mode and the DaT SPECT scan in striatum is very high: r = 0.925 (p<0.001, bottom row). R-fMRI, resting-state fMRI; L, left; R, right.
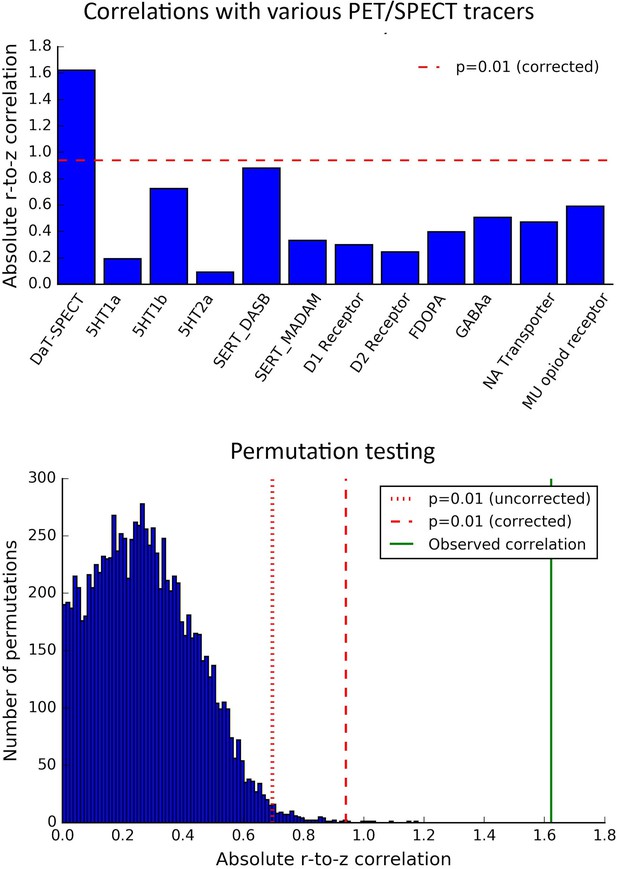
The correlation between trend surface model (TSM) coefficients modelling the second-order connectivity mode and the DaT SPECT scan is highly significant and substantially higher than all other position emission tomography (PET)-derived markers indexing other neurotransmitter systems.
The top panel displays the absolute, Fisher’s Z normalized correlations between the TSM coefficients modelling the second-order connectivity mode and the TSM coefficients of various SPECT- and PET-derived markers. The dotted lines represent significance values of p=0.01 and p=0.0008 (i.e., p=0.01/12 PET/SPECT scans; Bonferroni-corrected) derived from the null distribution in the bottom panel, which was generated by permuting the TSM coefficients obtained for each of the PET markers (N = 10,000) and computing the absolute (Fisher r-to-z normalized) correlations with the TSM coefficients of the connectivity mode.
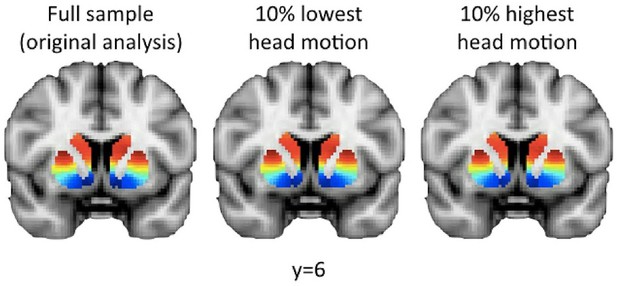
The second-order connectivity mode obtained in the 10% lowest and 10% highest movers of the Human Connectome Project (HCP) dataset is comparable to the mode obtained in the full sample.
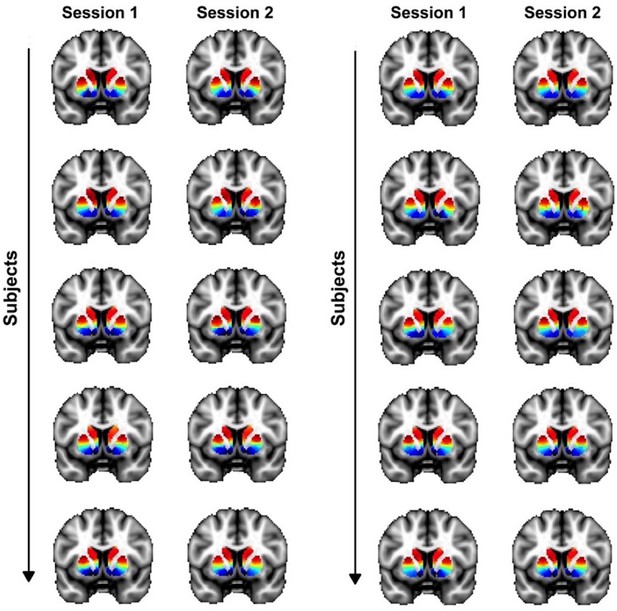
Inter-subject and inter-session (within-subject) variability in the second-order mode of connectivity in striatum.
Individual-subject connectivity modes are shown for 10 randomly selected Human Connectome Project (HCP) subjects (from a total of 839). This figure shows variations between subjects as well as variations between sessions for the same subjects.
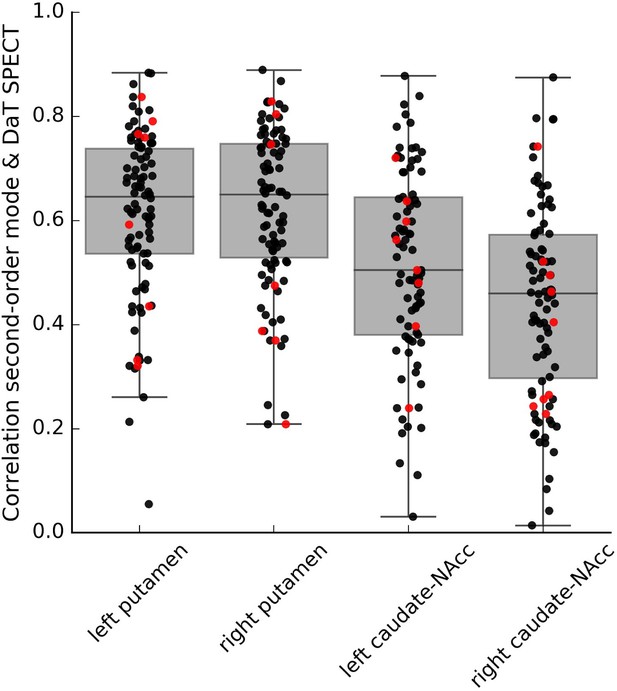
Within-subject correlations between the second-order connectivity mode and the DaT SPECT scan from subjects in the Parkinson’s Progression Markers Initiative (PPMI) cohort where both resting-state functional MRI (fMRI) and DaT SPECT data is available.
These correlations were obtained in a subsample of the PPMI dataset (6–8 datasets from controls and 73–82 datasets from Parkinson’s disease patients depending on the striatal subregion) with good-quality connectivity modes as defined by a high spatial correlation (r > 0.5) with the group-average Human Connectome Project (HCP) connectivity mode. Red dots represent control participants; black dots represent Parkinson’s disease patients.
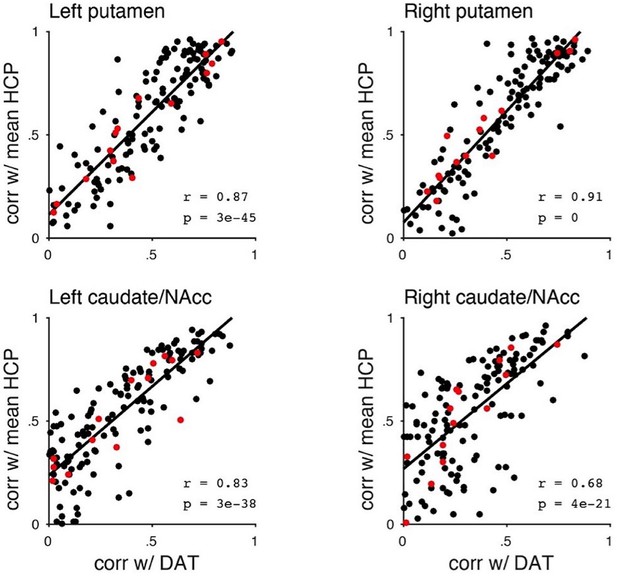
Spatial correlations of the subject-specific second-order connectivity modes with the mean Human Connectome Project (HCP) connectivity mode and DaT SPECT scan.
These plots show that when the connectivity mode of a subject resembles the HCP group-average connectivity mode – assumed to be an index of good quality – a high spatial similarity can be observed between the connectivity mode and the DaT SPECT scan of that subject. Red dots represent control participants, black dots represent Parkinson’s disease patients.
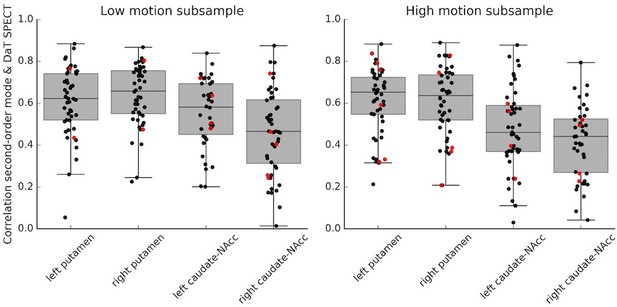
Within-subject correlations between the second-order connectivity mode and the DaT SPECT scan in a low motion and high motion subsample.
These correlations were obtained in a subsample of the Parkinson’s Progression Markers Initiative (PPMI) dataset with good-quality connectivity modes as defined by a high spatial correlation (r > 0.5) with the group-average Human Connectome Project (HCP) connectivity mode. The meanFD cutoff = 0.126, meaning that if the meanFD of a subject was below 0.126 that subject was part of the low motion subsample, if the meanFD was above 0.126, the subject was part of the high motion subsample. Red dots represent control participants; black dots represent Parkinson’s disease patients.
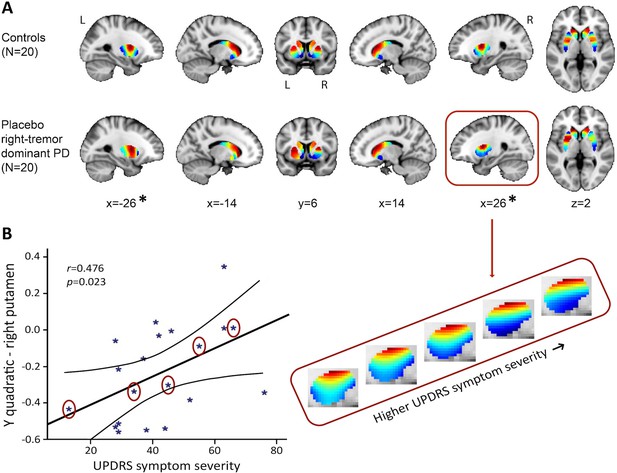
The second-order striatal connectivity mode is altered in right-dominant Parkinson’s disease.
(A) Significant difference between the control group and the right-dominant Parkinson’s disease group under placebo in putamen (omnibus test of all trend surface model [TSM] coefficients for putamen: X2 = 27.17, p=0.007). Images represent the mean connectivity modes across each of the investigated groups. The slices at MNI coordinates x = –26 and x = 26, respectively, show views of the striatal connectivity mode across left and right putamen where the connectivity mode is significantly different between groups (*); the slices at MNI coordinates x = –14 and x = 14, respectively, show views of the mode across left and right caudate-nucleus accumbens (caudate-NAcc) (no significant difference). (B) Trend-level association between the Unified Parkinson’s Disease Rating Scale (UPDRS) symptom severity score and the TSM coefficients modelling the second-order connectivity mode in putamen under placebo across patients in the right-dominant Parkinson’s disease group (general linear model [GLM] omnibus test of all TSM coefficients: X2 = 22.28, p=0.035). Post-hoc Pearson correlations revealed that this effect was driven by the quadratic TSM coefficients modelling the striatal connectivity mode in the right putamen in the Y (i.e., anterior-posterior) direction (right Y2: r = 0.476, p=0.034) and Z (i.e., superior-inferior) direction (right Z2: r = −0.460, p=0.041). The correlation between UPDRS symptom severity scores is displayed for the right Y2 coefficient. To visualize this association, the reconstructed second-order connectivity mode in the right putamen is shown for five Parkinson’s disease patients (data points circled) with increasing UPDRS symptom severity scores.
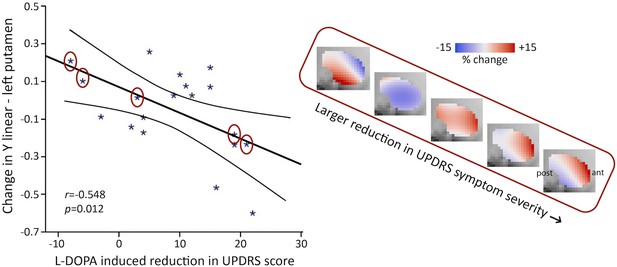
Levodopa-benserazide (L-DOPA)-induced reduction in Unified Parkinson’s Disease Rating Scale (UPDRS) symptom severity score is associated with the L-DOPA-induced change in the second-order mode of connectivity in putamen in right-dominant Parkinson’s disease (PD) (general linear model [GLM] omnibus test of all trend surface model [TSM] coefficients modelling putamen: X2 = 25.48, p=0.012).
Post-hoc Pearson correlations revealed that this effect was driven by the linear TSM coefficient modelling the striatal connectivity mode in the left putamen in the Y direction (left Y1: r = −0.548, p=0.012). To visualize this association, the difference in the reconstructed second-order connectivity modes between the placebo and L-DOPA session is shown for the left putamen (at slice X = –24) for five PD patients (data points circled). Red-coded voxels are hypothesized to map onto an increase in dopaminergic connectivity, blue-coded voxels onto a decrease. A significant effect was also observed for the left-dominant PD group (GLM omnibus test of all TSM coefficients: X2 = 34.07, p=0.001), but as post-hoc Pearson correlations did not reveal significant associations with one of the individual TSM coefficients in this group, this association is not shown. ant, anterior; post, posterior putamen.
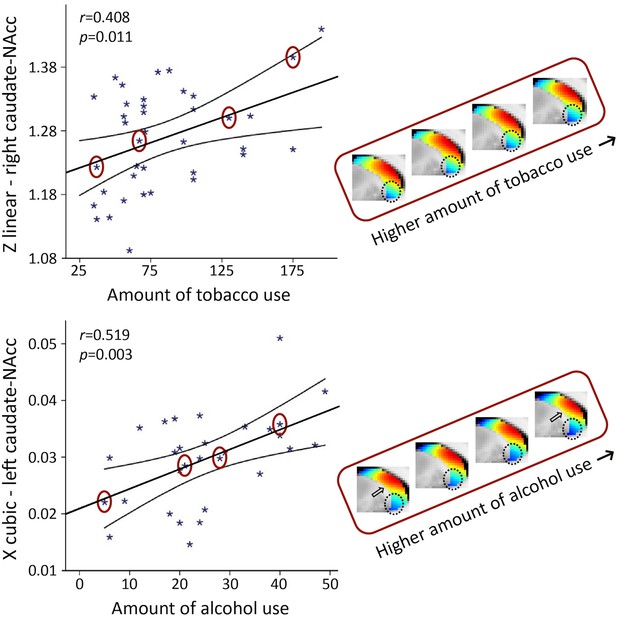
The second-order mode of connectivity in striatum is associated with the amount of tobacco use (top) and alcohol use (bottom).
Strong associations were observed between the trend surface model (TSM) coefficients modelling the connectivity mode in the caudate-nucleus accumbens (caudate-NAcc) region and the total amount of tobacco use as well as alcohol use over the past week (general linear model [GLM] omnibus test tobacco use: X2 = 49.55, p=0.002; alcohol use: X2 = 64.45, p<0.001). To visualize these relationships, Pearson correlations between one of the significant TSM coefficients and the amount of use are shown as well as the reconstructed second-order connectivity mode in the right caudate-NAcc (at slice X = 14) for four tobacco users and in the left caudate-NAcc (at slice X = –14) for four alcohol users with increasing of amounts of use (data points circled). Circles and arrows indicate where in the connectivity mode tobacco and alcohol use-related changes can be observed. Correlation plots for the other TSM coefficients can be found in Figure 6—figure supplements 1 and 2.
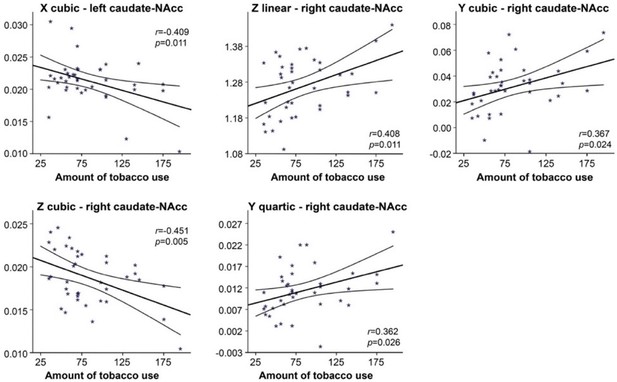
The second-order mode of connectivity in striatum is associated with the amount of tobacco use.
A strong association was observed between the trend surface model (TSM) coefficients modelling the connectivity mode in the caudate-nucleus accumbens (caudate-NAcc) region and the total amount of tobacco use over the past week (general linear model [GLM] omnibus test: X2 = 49.55, p=0.002). To visualize this relationship, Pearson correlations between the individual TSM coefficients and the amount of use were computed, and the correlations reaching significance (p<0.05) are shown in this figure.
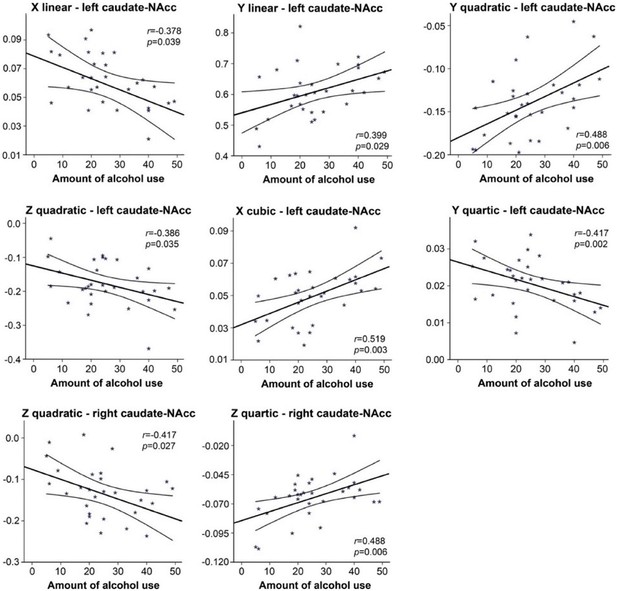
The second-order mode of connectivity in striatum is associated with the amount of alcohol use.
A strong association was observed between the trend surface model (TSM) coefficients modelling the connectivity mode in the caudate-nucleus accumbens (caudate-NAcc) region and the total number of alcoholic drinks over the past week (general linear model [GLM] omnibus test: X2 = 64.45, p<0.001). To visualize this relationship, Pearson correlations between the individual TSM coefficients and the amount of use were computed, and the correlations reaching significance (p<0.05) are shown in this figure.
Tables
Interclass correlation coefficients (ICCs) between the two scanning sessions and the session 1 to session 2 within-subject and between-subject spatial correlations.
| Striatal subregion | ICC[bootstrapped 95% CI] | Within-subject correlation | Between-subject correlation | Within vs. between permutation test(Nperm = 10,000) |
|---|---|---|---|---|
| Left putamen | 0.960 [0.951–0.965] | 0.969 | 0.965 | p<0.0001 |
| Right putamen | 0.961 [0.952–0.967] | 0.970 | 0.966 | p<0.0001 |
| Left caudate-NAcc | 0.974 [0.968–0.978] | 0.981 | 0.976 | p<0.0001 |
| Right caudate-NAcc | 0.974 [0.968–0.978] | 0.981 | 0.977 | p<0.0001 |
-
CI = confidence interval; NAcc = nucleus accumbens.
Participant characteristics.
| Demographic information(mean, SD) | ControlsN = 20 | PDN = 39 | Test statistic | |||
|---|---|---|---|---|---|---|
| Age, years | 61.9 | 10.4 | 60.9 | 10.7 | t(57) = 0.337 | NS |
| Sex, male (number, %) | 11 | 55.0% | 16 | 41.0% | X2(1) = 0.086 | NS |
| FAB, total score | 17.6 | 0.67 | 17.3 | 0.97 | t(57) = 0.23 | NS |
| Disease duration, years | NA | 3.96 | 4.57 | NA | ||
| L-DOPA equivalent at home (mg/day) | NA | 467.8 | 227.3 (range: 0–1100) | NA | ||
| UPDRS total score (mean, SD) | ||||||
| Placebo session | NA | 40.5 | 16.4 | t(38) = 5.58 | p<0.001 | |
| L-DOPA session | NA | 31.9 | 13.1 | |||
-
For the FAB, lower scores indicate worse functioning; for the UPDRS, higher scores indicate worse functioning. The FAB score was evaluated off medication.
-
FAB = frontal assessment battery (score 0–18); UPDRS = Unified Parkinson’s Disease Rating Scale part III (score 0–132); NS = not significant; NA = not applicable; PD = Parkinson’s disease; L-DOPA = levodopa-benserazide.
-
Table 2—source data 1
Source data for participant characterists listed in Table 2.
- https://cdn.elifesciences.org/articles/71846/elife-71846-table2-data1-v2.xlsx
Post-hoc analyses of age and sex.
| Original analysis | Original analysis +age and sex | Age and sex only | ||||
|---|---|---|---|---|---|---|
| X2 | p-Value | X2 | p-Value | X2 | p-Value | |
| Putamen | ||||||
| Patients vs. controlsRight tremor-dominant Parkinson’s disease | 27.17 | 0.007 | 27.21 | 0.018 | 0.48 | 0.786 |
| UPDRS symptom severityRight tremor-dominant Parkinson’s disease | 22.28 | 0.035 | 23.46 | 0.053 | 2.38 | 0.305 |
| L-DOPA-placebo differenceLeft tremor-dominant Parkinson’s disease | 34.07 | 0.001 | 46.14 | <0.001 | 2.42 | 0.299 |
| L-DOPA-placebo differenceRight tremor-dominant Parkinson’s disease | 25.48 | 0.012 | 37.53 | 0.001 | 7.18 | 0.028 |
| Caudate-NAcc | ||||||
| Tobacco useHCP dataset | 49.55 | 0.002 | 53.56 | 0.001 | 1.04 | 0.594 |
| Alcohol useHCP dataset | 64.45 | <0.001 | 174.87 | <0.001 | 9.26 | 0.010 |
-
UPDRS = Unified Parkinson’s Disease Rating Scale; L-DOPA = levodopa-benserazide; HCP = Human Connectome Project; NAcc = nucleus accumbens.
Post-hoc analyses using different thresholds for tobacco use.
| Original analysis:≥5× tobacco use a dayN = 38 | ≥2× tobacco use a dayN = 62 | ≥8× tobacco use a dayN = 30 | ||||
|---|---|---|---|---|---|---|
| X2 | p-Value | X2 | p-Value | X2 | p-Value | |
| Tobacco useHCP dataset caudate-NAcc | 49.55 | 0.002 | 37.96 | 0.035 | 70.54 | <0.001 |
-
HCP = Human Connectome Project; NAcc = nucleus accumbens.
Post-hoc analyses using different thresholds for alcohol use.
| Original analysis:≥3× light alcoholic and/or ≥1× hard liquor drinks a dayN = 30 | ≥1× alcoholic drinks a day (light and/or hard liquor)N = 103 | ≥3× alcoholic drinks a day (light and/or hard liquor) *N = 26 | ||||
|---|---|---|---|---|---|---|
| X2 | p-Value | X2 | p-Value | X2 | p-Value | |
| Alcohol useHCP dataset caudate-NAcc | 64.45 | <0.001 | 29.94 | 0.187 | 196.57 | <0.001 |
-
HCP = Human Connectome Project; NAcc = nucleus accumbens.
Subject IDs from the 839 HCP subjects used in the connectopic mapping analysis.
| 100206 | 129129 | 155635 | 181636 | 212823 | 385450 | 580044 | 784565 |
| 100610 | 129331 | 155938 | 182032 | 213017 | 386250 | 580347 | 788674 |
| 101006 | 129533 | 156031 | 182436 | 213421 | 387959 | 580650 | 789373 |
| 101107 | 129634 | 156435 | 183034 | 213522 | 389357 | 580751 | 792766 |
| 101309 | 129937 | 156536 | 183337 | 214524 | 391748 | 581450 | 792867 |
| 101410 | 130114 | 157437 | 183741 | 214625 | 392447 | 583858 | 793465 |
| 101915 | 130316 | 157942 | 185341 | 214726 | 392750 | 585256 | 800941 |
| 102008 | 130417 | 158136 | 185442 | 217126 | 393247 | 587664 | 802844 |
| 102109 | 130619 | 158338 | 185846 | 219231 | 393550 | 588565 | 803240 |
| 102311 | 130720 | 158843 | 185947 | 220721 | 394956 | 589567 | 804646 |
| 102513 | 130821 | 159138 | 186040 | 221218 | 395251 | 590047 | 809252 |
| 102614 | 131217 | 159340 | 186141 | 223929 | 395756 | 592455 | 810439 |
| 102715 | 131419 | 159441 | 186545 | 227432 | 395958 | 594156 | 810843 |
| 103010 | 131722 | 159744 | 186848 | 227533 | 397154 | 597869 | 812746 |
| 103111 | 131823 | 159845 | 187143 | 228434 | 397861 | 599065 | 814548 |
| 103212 | 132017 | 159946 | 187345 | 231928 | 406432 | 599469 | 814649 |
| 104012 | 132118 | 160729 | 187547 | 233326 | 406836 | 599671 | 815247 |
| 104416 | 133019 | 160830 | 187850 | 236130 | 412528 | 601127 | 816653 |
| 104820 | 134021 | 160931 | 188145 | 237334 | 413934 | 604537 | 818455 |
| 105014 | 134223 | 161630 | 188347 | 238033 | 419239 | 609143 | 818859 |
| 105620 | 134425 | 161832 | 188448 | 239136 | 421226 | 611938 | 820745 |
| 105923 | 134627 | 162026 | 188549 | 248339 | 422632 | 613235 | 822244 |
| 106016 | 134829 | 162228 | 188751 | 250932 | 424939 | 613538 | 825048 |
| 106521 | 135124 | 162733 | 189349 | 255740 | 432332 | 615441 | 825553 |
| 106824 | 135225 | 162935 | 189450 | 256540 | 436239 | 615744 | 825654 |
| 107018 | 135528 | 163129 | 191033 | 257542 | 436845 | 616645 | 826454 |
| 107220 | 135629 | 163331 | 191235 | 257845 | 441939 | 617748 | 827052 |
| 107321 | 135730 | 163836 | 191336 | 257946 | 445543 | 618952 | 828862 |
| 107422 | 136126 | 164030 | 191841 | 263436 | 449753 | 620434 | 832651 |
| 107725 | 136227 | 164131 | 191942 | 268749 | 453441 | 622236 | 833148 |
| 108020 | 136631 | 164636 | 192035 | 268850 | 453542 | 623137 | 833249 |
| 108121 | 136732 | 164939 | 192136 | 270332 | 454140 | 623844 | 835657 |
| 108222 | 137027 | 165032 | 192237 | 274542 | 456346 | 626648 | 837560 |
| 108323 | 137229 | 165234 | 192641 | 275645 | 459453 | 627852 | 837964 |
| 108525 | 137431 | 165436 | 192843 | 280739 | 461743 | 633847 | 841349 |
| 108828 | 137532 | 165638 | 193441 | 280941 | 463040 | 634748 | 843151 |
| 109123 | 137633 | 165941 | 193845 | 281135 | 467351 | 635245 | 844961 |
| 109325 | 137936 | 166438 | 194443 | 283543 | 468050 | 644044 | 845458 |
| 109830 | 138130 | 166640 | 194645 | 285345 | 473952 | 645450 | 849264 |
| 110007 | 138332 | 167036 | 194746 | 285446 | 475855 | 647858 | 849971 |
| 111211 | 138837 | 167238 | 194847 | 286347 | 479762 | 654350 | 852455 |
| 111413 | 139233 | 167440 | 195041 | 286650 | 480141 | 654552 | 856463 |
| 112112 | 139435 | 168240 | 195445 | 287248 | 481042 | 656253 | 856968 |
| 112314 | 139839 | 168341 | 195950 | 289555 | 481951 | 657659 | 867468 |
| 112516 | 140117 | 168745 | 196346 | 290136 | 486759 | 660951 | 869472 |
| 112920 | 140319 | 168947 | 196851 | 295146 | 492754 | 662551 | 870861 |
| 113316 | 140824 | 169040 | 196952 | 297655 | 495255 | 663755 | 871762 |
| 113922 | 140925 | 169444 | 197348 | 298455 | 497865 | 664757 | 872562 |
| 114116 | 141119 | 169545 | 197651 | 299154 | 500222 | 667056 | 873968 |
| 114217 | 141422 | 169747 | 198047 | 299760 | 506234 | 668361 | 877269 |
| 114318 | 141826 | 169949 | 198249 | 300618 | 510225 | 671855 | 878776 |
| 114419 | 142424 | 170631 | 198350 | 300719 | 510326 | 673455 | 878877 |
| 114621 | 143224 | 170934 | 198653 | 303119 | 512835 | 675661 | 880157 |
| 114823 | 143426 | 171128 | 198855 | 303624 | 513130 | 677766 | 882161 |
| 115017 | 144125 | 171330 | 199352 | 304727 | 513736 | 679568 | 884064 |
| 115219 | 144731 | 171431 | 199453 | 305830 | 516742 | 679770 | 886674 |
| 115724 | 144832 | 171532 | 200008 | 308129 | 517239 | 680250 | 888678 |
| 115825 | 144933 | 171633 | 200109 | 308331 | 518746 | 680452 | 891667 |
| 116221 | 145127 | 171734 | 200311 | 309636 | 519647 | 683256 | 894067 |
| 116423 | 145531 | 172029 | 200513 | 310621 | 519950 | 686969 | 894774 |
| 116524 | 145632 | 172130 | 200917 | 311320 | 520228 | 687163 | 898176 |
| 116726 | 145834 | 172433 | 201414 | 314225 | 521331 | 689470 | 901038 |
| 117021 | 146129 | 172534 | 201717 | 316633 | 522434 | 690152 | 901442 |
| 117728 | 146331 | 172635 | 201818 | 316835 | 523032 | 692964 | 902242 |
| 117930 | 146432 | 172938 | 202113 | 317332 | 524135 | 693764 | 905147 |
| 118023 | 146533 | 173132 | 202719 | 318637 | 525541 | 694362 | 907656 |
| 118124 | 146634 | 173334 | 203418 | 320826 | 529549 | 695768 | 908860 |
| 118225 | 146735 | 173435 | 203923 | 321323 | 529953 | 698168 | 910241 |
| 118528 | 146937 | 173536 | 204016 | 322224 | 531536 | 700634 | 910443 |
| 118831 | 147030 | 173637 | 204319 | 325129 | 536647 | 701535 | 911849 |
| 119025 | 147636 | 173738 | 204420 | 329844 | 540436 | 706040 | 912447 |
| 119126 | 147737 | 173839 | 204521 | 330324 | 541640 | 707749 | 917558 |
| 119732 | 148133 | 173940 | 204622 | 333330 | 545345 | 715041 | 919966 |
| 120414 | 148335 | 174841 | 205220 | 334635 | 547046 | 715950 | 922854 |
| 120515 | 148436 | 175136 | 206222 | 339847 | 548250 | 720337 | 923755 |
| 120717 | 148941 | 175237 | 206323 | 341834 | 549757 | 724446 | 926862 |
| 121315 | 149236 | 175338 | 206525 | 342129 | 550439 | 725751 | 927359 |
| 121416 | 149741 | 175540 | 206727 | 346137 | 552241 | 727553 | 929464 |
| 121618 | 149842 | 175742 | 206828 | 346945 | 553344 | 727654 | 930449 |
| 121921 | 150625 | 176037 | 206929 | 348545 | 555348 | 728454 | 933253 |
| 122317 | 150726 | 176441 | 207123 | 349244 | 555651 | 729254 | 942658 |
| 122418 | 150928 | 176744 | 207426 | 350330 | 555954 | 731140 | 947668 |
| 122620 | 151021 | 176845 | 208024 | 352132 | 557857 | 734247 | 952863 |
| 122822 | 151324 | 177140 | 208125 | 352738 | 558657 | 735148 | 953764 |
| 123420 | 151425 | 177241 | 208327 | 353740 | 558960 | 737960 | 955465 |
| 123521 | 151728 | 177342 | 208428 | 355239 | 559457 | 742549 | 957974 |
| 123723 | 151829 | 177645 | 208630 | 356948 | 561444 | 744553 | 958976 |
| 123824 | 151930 | 178142 | 209127 | 358144 | 561949 | 748662 | 962058 |
| 123925 | 152225 | 178243 | 209228 | 360030 | 562345 | 749058 | 965771 |
| 124220 | 152427 | 178647 | 209329 | 361234 | 562446 | 751550 | 966975 |
| 124624 | 152831 | 178748 | 209531 | 361941 | 565452 | 753150 | 970764 |
| 124826 | 153025 | 178849 | 209834 | 362034 | 566454 | 757764 | 971160 |
| 125222 | 153126 | 178950 | 210011 | 365343 | 567052 | 759869 | 972566 |
| 125424 | 153227 | 179245 | 210112 | 366042 | 567961 | 760551 | 973770 |
| 126426 | 153631 | 179346 | 210415 | 368551 | 568963 | 763557 | 978578 |
| 126628 | 153732 | 179952 | 211114 | 368753 | 569965 | 765864 | 979984 |
| 127226 | 153833 | 180129 | 211215 | 376247 | 571144 | 766563 | 983773 |
| 127327 | 153934 | 180230 | 211316 | 377451 | 571548 | 769064 | 987074 |
| 127630 | 154229 | 180432 | 211619 | 378756 | 572045 | 770352 | 989987 |
| 127731 | 154330 | 180533 | 211821 | 378857 | 573249 | 771354 | 990366 |
| 127832 | 154532 | 180735 | 211922 | 379657 | 573451 | 773257 | 991267 |
| 128026 | 154734 | 180836 | 212015 | 380036 | 576255 | 774663 | 992673 |
| 128127 | 154835 | 180937 | 212116 | 381038 | 578057 | 779370 | 993675 |
| 128329 | 154936 | 181131 | 212217 | 381543 | 578158 | 782561 | 996782 |
| 128935 | 155231 | 181232 | 212419 | 382242 | 579867 | 783462 | 788674 |
-
HCP = Human Connectome Project.
Subject IDs from the 209 PPMI controls with DaT SPECT data used in our analysis.
| PPMIsubject ID | Image IDDaT SPECT | PPMIsubject ID | Image IDDaT SPECT | PPMIsubject ID | Image IDDaT SPECT |
|---|---|---|---|---|---|
| 3000 | 323662 | 3350 | 339901 | 3637 | 388521 |
| 3004 | 341194 | 3351 | 339902 | 3639 | 388523 |
| 3008 | 341195 | 3353 | 339904 | 3651 | 339008 |
| 3009 | 341196 | 3355 | 341236 | 3651 | 355956 |
| 3011 | 341198 | 3357 | 339907 | 3656 | 339014 |
| 3013 | 341200 | 3358 | 339908 | 3658 | 339016 |
| 3016 | 341202 | 3361 | 339911 | 3662 | 355221 |
| 3029 | 388468 | 3362 | 339912 | 3668 | 388528 |
| 3053 | 341207 | 3363 | 338780 | 3750 | 388535 |
| 3055 | 341209 | 3368 | 339917 | 3754 | 360616 |
| 3057 | 341211 | 3369 | 339918 | 3756 | 360617 |
| 3064 | 341217 | 3370 | 339919 | 3759 | 363950 |
| 3069 | 341221 | 3389 | 388504 | 3765 | 363951 |
| 3070 | 341222 | 3390 | 388505 | 3767 | 388536 |
| 3071 | 341223 | 3401 | 340345 | 3768 | 363952 |
| 3072 | 341224 | 3404 | 340346 | 3769 | 360618 |
| 3073 | 341225 | 3405 | 340347 | 3779 | 453700 |
| 3074 | 341226 | 3410 | 340351 | 3794 | 388545 |
| 3075 | 341227 | 3411 | 340352 | 3796 | 388147 |
| 3085 | 388470 | 3414 | 340354 | 3803 | 355230 |
| 3087 | 388472 | 3424 | 340363 | 3804 | 354344 |
| 3100 | 341230 | 3438 | 340388 | 3805 | 354345 |
| 3103 | 341233 | 3450 | 340398 | 3806 | 354346 |
| 3104 | 339536 | 3452 | 339923 | 3807 | 355231 |
| 3106 | 340418 | 3453 | 339924 | 3811 | 360620 |
| 3109 | 340423 | 3457 | 339928 | 3812 | 355232 |
| 3112 | 340426 | 3458 | 339929 | 3813 | 355233 |
| 3114 | 340430 | 3460 | 341243 | 3816 | 363953 |
| 3115 | 340431 | 3464 | 341245 | 3817 | 388148 |
| 3151 | 341018 | 3466 | 339932 | 3850 | 337832 |
| 3156 | 341021 | 3468 | 339934 | 3851 | 337833 |
| 3157 | 341022 | 3478 | 360613 | 3852 | 337834 |
| 3160 | 341023 | 3479 | 363945 | 3853 | 337835 |
| 3161 | 341024 | 3480 | 388509 | 3854 | 337836 |
| 3165 | 341027 | 3481 | 388510 | 3855 | 337445 |
| 3169 | 341031 | 3503 | 340400 | 3857 | 337837 |
| 3171 | 341033 | 3515 | 340408 | 3859 | 337839 |
| 3172 | 341034 | 3517 | 341248 | 3907 | 388556 |
| 3188 | 388483 | 3518 | 339537 | 3908 | 363957 |
| 3191 | 388486 | 3521 | 339539 | 3917 | 388563 |
| 3200 | 341036 | 3523 | 339541 | 3950 | 341083 |
| 3201 | 341037 | 3524 | 339542 | 3952 | 341085 |
| 3202 | 341038 | 3525 | 339543 | 3955 | 388565 |
| 3204 | 341040 | 3526 | 339544 | 3959 | 355241 |
| 3206 | 341042 | 3527 | 339545 | 3965 | 388573 |
| 3208 | 341044 | 3541 | 355215 | 3966 | 388574 |
| 3213 | 341049 | 3543 | 363946 | 3967 | 388576 |
| 3215 | 341051 | 3544 | 388514 | 3968 | 388577 |
| 3216 | 341052 | 3551 | 339550 | 3969 | 388578 |
| 3217 | 341053 | 3554 | 339552 | 4004 | 339032 |
| 3219 | 341055 | 3554 | 358138 | 4007 | 339035 |
| 3221 | 341057 | 3555 | 339553 | 4008 | 339036 |
| 3222 | 341058 | 3563 | 339559 | 4009 | 339037 |
| 3235 | 388488 | 3565 | 339561 | 4010 | 339038 |
| 3237 | 388490 | 3569 | 339564 | 4014 | 389268 |
| 3257 | 341067 | 3570 | 339565 | 4018 | 339045 |
| 3260 | 341068 | 3571 | 389245 | 4032 | 388583 |
| 3264 | 341070 | 3572 | 338781 | 4063 | 355246 |
| 3270 | 341074 | 3576 | 338785 | 4067 | 388593 |
| 3271 | 341075 | 3600 | 338788 | 4079 | 388596 |
| 3274 | 341077 | 3611 | 338797 | 4090 | 343886 |
| 3276 | 341079 | 3613 | 338799 | 4095 | 354353 |
| 3277 | 388491 | 3614 | 338800 | 4100 | 360623 |
| 3286 | 388494 | 3615 | 338801 | 4104 | 363963 |
| 3300 | 339889 | 3619 | 339001 | 4105 | 388600 |
| 3301 | 339890 | 3620 | 339002 | 4116 | 388613 |
| 3310 | 339896 | 3624 | 341251 | 4118 | 388615 |
| 3316 | 342187 | 3627 | 342204 | 4139 | 388627 |
| 3318 | 342189 | 3635 | 388519 | 4140 | 388628 |
| 3320 | 342191 | 3636 | 388520 |
-
PPMI = Parkinson’s Progression Markers Initiative; DaT = dopamine transporter; SPECT = single-photon emission computed tomography.
Subject IDs from PD patients and controls with resting-state fMRI data and DaT SPECT data from the PPMI dataset used in our analysis.
| PPMI subject ID | Image ID DaT SPECT | Image ID MRI | Diagnosis |
|---|---|---|---|
| 3310 | 339896 | 369414 | Control |
| 3318 | 342189 | 374882 | Control |
| 3350 | 339901 | 515208 | Control |
| 3351 | 339902 | 508245 | Control |
| 3353 | 339904 | 515216 | Control |
| 3361 | 339911 | 581042 | Control |
| 3369 | 339918 | 544617 | Control |
| 3389 | 388504 | 367349 | Control |
| 3551 | 339550 | 548987 | Control |
| 3563 | 339559 | 548989 | Control |
| 3565 | 339561 | 560369 | Control |
| 3769 | 360618 | 362609 | Control |
| 4018 | 339045 | 365285 | Control |
| 4032 | 388583 | 367390 | Control |
| 3107 | 419849 | 378215 | PD |
| 3108 | 419850 | 378223 | PD |
| 3116 | 418649 | 366137 | PD |
| 3116 | 419854 | 417052 | PD |
| 3118 | 418470 | 362555 | PD |
| 3118 | 446107 | 430138 | PD |
| 3119 | 418650 | 382277 | PD |
| 3119 | 446108 | 430147 | PD |
| 3120 | 418651 | 374854 | PD |
| 3122 | 419241 | 382284 | PD |
| 3123 | 418652 | 382289 | PD |
| 3123 | 449008 | 440114 | PD |
| 3124 | 418653 | 387304 | PD |
| 3124 | 449009 | 440118 | PD |
| 3125 | 418654 | 387314 | PD |
| 3125 | 449010 | 440128 | PD |
| 3126 | 418655 | 397752 | PD |
| 3126 | 449011 | 440131 | PD |
| 3128 | 419553 | 395434 | PD |
| 3128 | 504427 | 466848 | PD |
| 3130 | 360608 | 355962 | PD |
| 3130 | 419554 | 417000 | PD |
| 3132 | 436066 | 423718 | PD |
| 3132 | 504428 | 498892 | PD |
| 3134 | 388480 | 369013 | PD |
| 3134 | 436067 | 436351 | PD |
| 3327 | 389212 | 362478 | PD |
| 3327 | 486550 | 412180 | PD |
| 3332 | 388500 | 378540 | PD |
| 3352 | 418905 | 372319 | PD |
| 3354 | 418906 | 372327 | PD |
| 3359 | 419866 | 397593 | PD |
| 3360 | 419867 | 393662 | PD |
| 3364 | 419868 | 393672 | PD |
| 3365 | 419659 | 397597 | PD |
| 3366 | 419869 | 397624 | PD |
| 3367 | 419870 | 393674 | PD |
| 3371 | 418673 | 365166 | PD |
| 3372 | 436070 | 369487 | PD |
| 3372 | 446121 | 420330 | PD |
| 3373 | 418674 | 387316 | PD |
| 3373 | 449019 | 440174 | PD |
| 3374 | 418675 | 393614 | PD |
| 3374 | 446122 | 430165 | PD |
| 3377 | 418677 | 393628 | PD |
| 3377 | 449020 | 440186 | PD |
| 3378 | 418678 | 387324 | PD |
| 3380 | 418679 | 393636 | PD |
| 3380 | 468270 | 449575 | PD |
| 3383 | 355208 | 351070 | PD |
| 3383 | 419560 | 415707 | PD |
| 3385 | 360612 | 353398 | PD |
| 3385 | 436861 | 415713 | PD |
| 3386 | 388502 | 369048 | PD |
| 3387 | 389214 | 357590 | PD |
| 3387 | 436071 | 417033 | PD |
| 3392 | 388507 | 372995 | PD |
| 3392 | 442969 | 436390 | PD |
| 3552 | 418922 | 378354 | PD |
| 3556 | 418923 | 372348 | PD |
| 3556 | 504848 | 482323 | PD |
| 3557 | 504849 | 482329 | PD |
| 3559 | 418926 | 372359 | PD |
| 3559 | 504850 | 491605 | PD |
| 3574 | 419676 | 414623 | PD |
| 3575 | 419677 | 581115 | PD |
| 3575 | 418690 | 365225 | PD |
| 3585 | 449026 | 440198 | PD |
| 3586 | 468275 | 449581 | PD |
| 3587 | 468276 | 449584 | PD |
| 3591 | 388516 | 373018 | PD |
| 3591 | 504435 | 491626 | PD |
| 3592 | 388517 | 373035 | PD |
| 3592 | 442973 | 436404 | PD |
| 3593 | 388518 | 369096 | PD |
| 3593 | 436073 | 430199 | PD |
| 3593 | 504436 | 507400 | PD |
| 3758 | 418698 | 374893 | PD |
| 3758 | 419880 | 402067 | PD |
| 3760 | 418499 | 362591 | PD |
| 3787 | 419576 | 412194 | PD |
| 3800 | 389258 | 393684 | PD |
| 3808 | 419885 | 402071 | PD |
| 3815 | 419886 | 581145 | PD |
| 3818 | 446139 | 440242 | PD |
| 3819 | 419270 | 395448 | PD |
| 3822 | 419271 | 382366 | PD |
| 3822 | 449035 | 440262 | PD |
| 3823 | 419272 | 395585 | PD |
| 3823 | 449036 | 440267 | PD |
| 3824 | 419579 | 395592 | PD |
| 3824 | 468279 | 449614 | PD |
| 3825 | 419273 | 393639 | PD |
| 3825 | 504450 | 549048 | PD |
| 3826 | 419274 | 395600 | PD |
| 3826 | 468280 | 449625 | PD |
| 3828 | 419580 | 395605 | PD |
| 3828 | 468281 | 449661 | PD |
| 3829 | 419581 | 395614 | PD |
| 3830 | 419582 | 412202 | PD |
| 3830 | 495006 | 468929 | PD |
| 3831 | 419583 | 402267 | PD |
| 3832 | 419584 | 412209 | PD |
| 3832 | 495007 | 468935 | PD |
| 3834 | 419585 | 415724 | PD |
| 3834 | 504454 | 473094 | PD |
| 3835 | 436875 | 415731 | PD |
| 3838 | 436075 | 423748 | PD |
| 3838 | 504456 | 515249 | PD |
| 3869 | 436077 | 415744 | PD |
| 3870 | 363956 | 395313 | PD |
| 3870 | 486557 | 415751 | PD |
| 4005 | 419890 | 397646 | PD |
| 4011 | 418504 | 402285 | PD |
| 4019 | 418710 | 362640 | PD |
| 4019 | 446143 | 417057 | PD |
| 4021 | 419277 | 430178 | PD |
| 4022 | 418712 | 365294 | PD |
| 4022 | 446145 | 417065 | PD |
| 4029 | 468288 | 468943 | PD |
| 4030 | 363959 | 356036 | PD |
| 4030 | 419596 | 415756 | PD |
| 4030 | 495322 | 468949 | PD |
| 4034 | 388585 | 367425 | PD |
| 4034 | 436083 | 423755 | PD |
| 4035 | 388587 | 369183 | PD |
| 4035 | 436084 | 423762 | PD |
| 4035 | 504466 | 475680 | PD |
| 4038 | 388590 | 367446 | PD |
| 4038 | 436085 | 430210 | PD |
-
PD = Parkinson’s disease; DaT = dopamine transporter; fMRI = functional MRI; SPECT = single-photon emission computed tomography; PPMI = Parkinson’s Progression Markers Initiative.
| meanFD | ||||
|---|---|---|---|---|
| N | Mean | SD | Range | |
| HCP | 839 | 0.0871 | 0.0362 | 0.0375-0.3155 |
| PPMI all DaT SPECT controls | 209 | Not available for DaT SPECT scan | ||
| PPMI within-subject analysis, full sample | 144 | 0.1308 | 0.0620 | 0.0438-0.3124 |
| PPMI within-subject analysis, patients | 130 | 0.1292 | 0.0609 | 0.0438-0.3124 |
| PPMI within-subject analysis, controls | 14 | 0.1463 | 0.0541 | 0.0722-0.2778 |
| Local PD dataset, full sample placebo | 59 | 0.1104 | 0.0535 | 0.0352-0.2762 |
| Local PD dataset, patients placebo | 39 | 0.1165 | 0.0492 | 0.0490-0.2363 |
| Local PD dataset, controls placebo | 20 | 0.0987 | 0.0604 | 0.0352-0.2762 |
| Local PD dataset, patients LDOPA | 39 | 0.1277 | 0.0825 | 0.0376-0.5252* |
| *There was 1 subject with a meanFD of 0.5252, but also after excluding this subject results remained significant. All other subjects had a meanFD<0.3. |





