Environmentally sensitive hotspots in the methylome of the early human embryo
Figures
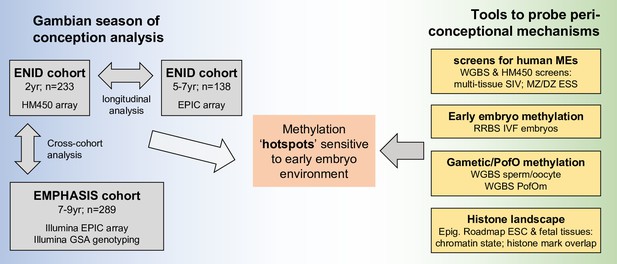
Study design.
DNAm: DNA methylation; EPIC: Illumina Infinium MethylationEPIC BeadChip; Epig. Roadmap: Roadmap Epigenomics Consortium; ESC: embryonic stem cell; ESS: epigenetic supersimilarity; GSA: Global Screening Array; HM450: Illumina Infinium HumanMethylation450 BeadChip; IVF: in vitro fertilisation; MEs: metastable epialleles; MZ/DZ: monozygotic/dizygotic twins; PofO: parent of origin; RRBS: reduced representation bisulfite-seq; SIV: systemic interindividual variation; SoC: season of conception; WGBS: whole-genome bisulfite-seq. See Tables 1 and 3 for further details of Gambian and public datasets used in this analysis.
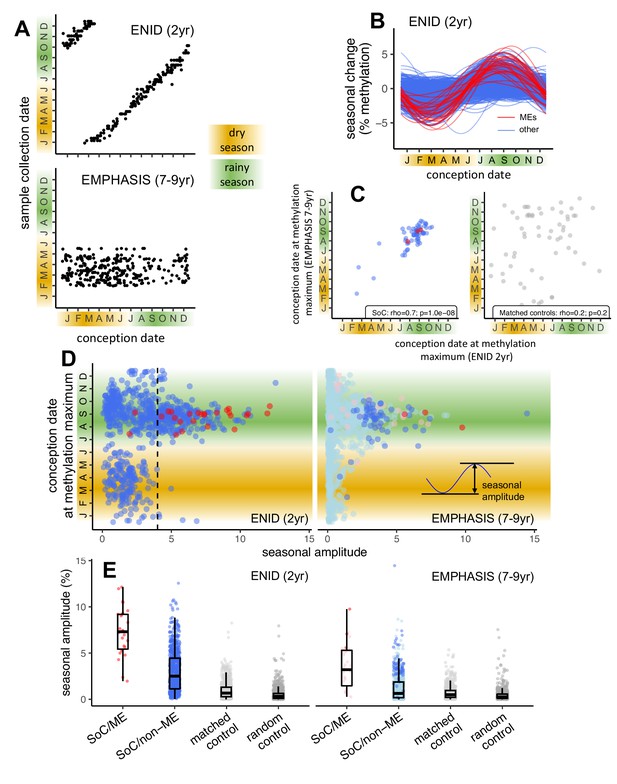
Identification of Gambian season of conception associated CpGs: ENID (2 yr) vs. EMPHASIS (7–9 yr) cross-cohort analysis.
(A) Relationship between date of conception and date of sample collection for ENID (top) and EMPHASIS (bottom) cohorts. (B) Modelled seasonal change in methylation for 768 SoC-associated loci (false discovery rate [FDR] < 5%) in the ENID cohort. 26 ME CpGs are marked in red. (C) Conception date of modelled methylation maximum in each cohort for 61 CpGs significantly associated with SoC in both cohorts (left) and 61 matched controls (right). (D) (Left) Date of modelled DNAm maximum vs. seasonal amplitude in each cohort for 768 CpGs significantly associated with SoC in the ENID cohort. MEs are marked in red. Dashed line indicates SoC amplitude threshold used to identify SoC-CpGs. (Right) Same CpGs as left, but in the older EMPHASIS cohort. Significant SoC associations for this cohort are marked in a darker colour. (E) (Left) Seasonal amplitudes for SoC-associated CpGs that are (red) or are not (blue) MEs; along with amplitudes for 768 matched and random controls (light/dark grey respectively). (Right) As left but in the older EMPHASIS cohort. For EMPHASIS significant SoC associations are marked in a darker colour. Boxes represent the middle 50% of the data (inter-quartile range [IQR]); the line inside the box is the median, and whiskers represent values lying within 1.5 times the IQR.
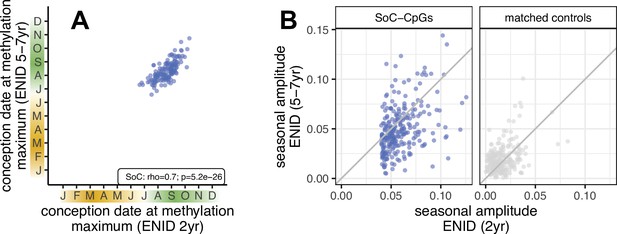
Identification of Gambian season of conception associated CpGs: ENID 2 yr vs. 5–7 yr longitudinal analysis.
(A) Relationship between conception date of modelled methylation maximum measured at 2 and 5–7 yr in the same n = 138 individuals from the ENID cohort. n = 157 SoC-CpGs with a significant SoC association (false discovery rate [FDR] < 5%) at 5–7 yr are plotted. (B) Change in seasonal amplitude between 2 and 5–7 yr for n = 259 SoC-CpGs (left) and matched controls (right) with DNAm measured in the same n = 138 individuals from the ENID cohort. There is evidence of SoC effect attenuation with age at SoC-CpGs, but not at matched controls (Wilcoxon signed rank test for difference in SoC amplitude p = 6.7 × 10–12 and p = 1.0, respectively).
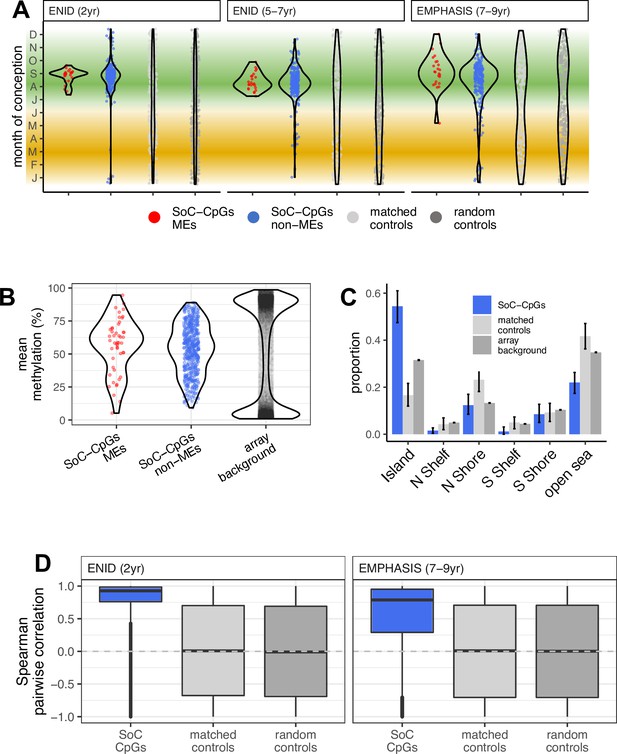
Properties of SoC-CpGs.
(A) Date of conception at modelled methylation maxima for 259 SoC-CpGs and 259 corresponding matched and random controls across all three analysed datasets. Green and yellow bands indicate the extent of the rainy and dry seasons, respectively. (B) SoC-CpGs show increased intermediate methylation compared to array background (data from ENID 2 yr and EMPHASIS cohorts combined). (C) Distribution of 259 SoC-CpGs [including MEs], 259 matched controls and array background with respect to CpG islands. Error bars are bootstrapped 95% CIs. N/S Shore/Shelf: North/South Shore/Shelf, respectively (regions proximal to CpG Islands defined in Illumina manifest). (D) Distribution of pairwise Spearman correlations for CpG sets in the ENID (left) and EMPHASIS (right) cohorts. Boxplot elements as described in Figure 2E.
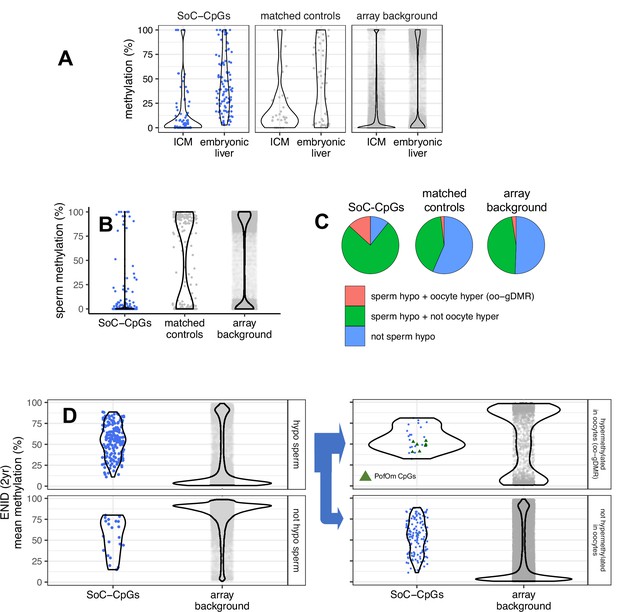
Early stage embryo, gametic, and parent-of-origin-specific methylation (PofOm).
(A) Methylation distribution of SoC-CpGs, matched controls, and array background in pre-gastrulation inner cell mass (ICM) and post-gastrulation embryonic liver, measured in reduced representation bisulfite-seq (RRBS) embryo methylation data from Guo et al., 2014. Data comprises 112,380 CpGs covered at ≥10× in ICM and/or embryonic liver that overlap array background, including 118 SoC-CpGs and 51 matched controls. (B) Methylation distribution of SoC-CpGs, matched controls, and array background in sperm whole-genome bisulfite-seq (WGBS) data from Okae et al., 2014. Data comprises 294,240 CpGs covered at ≥10× including 196 SoC-CpGs and 207 matched controls. (C) Sperm methylation and oocyte germline differentially methylated region (oo-gDMR) status at 196 SoC-CpGs covered at ≥10× in Okae et al. sperm WGBS data. Sperm hypomethylation is defined as methylation ≤25%. oo-gDMRs defined as sperm methylation <25% and oocyte methylation >75% in WGBS analysis by Sanchez-Delgado et al., 2016. (D) Mean methylation at SoC-CpGs and array background measured in n = 233 individuals in the ENID (2 yr) cohort stratified by sperm and oocyte methylation status. Left: Methylation stratified according to sperm hypomethylation status (n = 175 SoC-CpGs hypomethylated in sperm, n = 21 not hypomethylated). Right: As left but with loci hypomethylated in sperm further stratified according to oo-gDMR status (n = 26 SoC-CpGs hypermethylated in oocytes/oo-gDMR, n = 149 not hypermethylated). Sperm/oo-gDMR status thresholds as for Figure 4C. Eight PofOm CpGs (green triangles) are those identified in Zink et al., 2018.
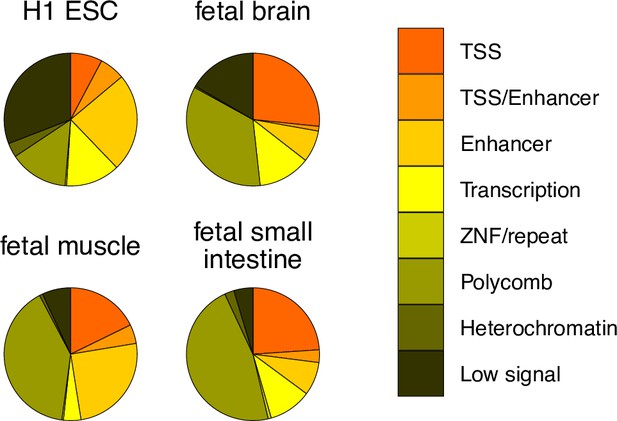
SoC-CpG overlap with predicted chromatin states.
Chromatin states predicted by ChromHMM (Ernst and Kellis, 2012) from chromatin marks in four cell lines and tissues generated by the Roadmap Epigenomics Consortium et al., 2015. Predicted states for all 259 SoC-CpGs are shown. Predictions from the ChromHMM 15-state model are collapsed to eight states for clarity. TSS: active transcription start site/flanking active TSS/bivalent or poisedTSS; TSS/enhancer: flanking bivalent TSS/enhancer; enhancer: enhancer/bivalent enhancer/genic enhancer; transcription: strong/weak transcription/transcription at gene 5’ and 3’; ZNF/repeat: zinc finger genes and repeats; polycomb: repressed/weak repressed polycomb; heterochromatin; low signal: low signal in all marks states used as inputs to ChromHMM.
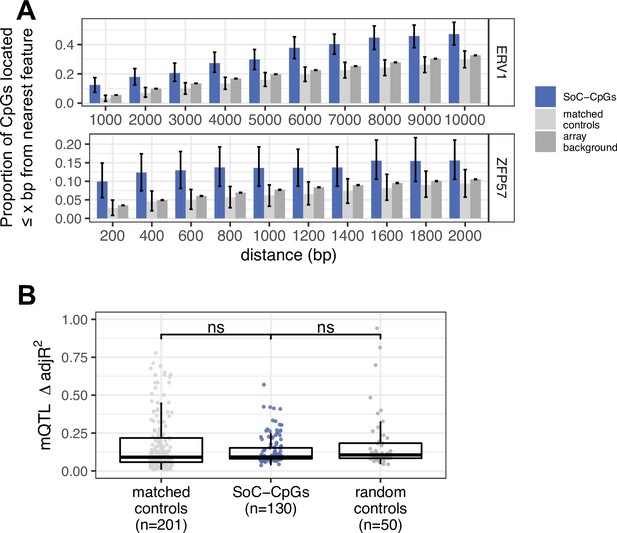
Links between endogenous retroviruses (ERVs), ZFP57 binding sites, genetic variation, and DNAm at season of conception associated loci.
(A) Proportion of SoC-CpGs, matched controls, and array background CpGs proximal to ERV1 endogenous retroviral elements (top) and ZFP57 binding sites (bottom), within the specified distance. CpG clustering effects are removed by sampling a single CpG from each cluster (see Materials and methods). Error bars are bootstrapped 95% CIs. (B) Proportion of methylation variance explained by methylation quantitative trait locus (mQTL) for matched controls, SoC-CpGs and random controls. Only CpGs with at least one significant mQTL are plotted (n = 201, 130, and 50 respectively; see Materials and methods for further details). Boxplot elements as described in Figure 2E.
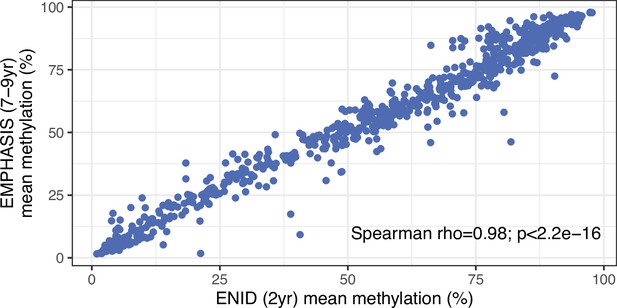
Cross-cohort correlation of mean methylation values at 768 season of conception (SoC)-associated loci.
Data comprises n=233 and n=289 individuals from the ENID (2 yr) and EMPHASIS cohorts, respectively. 768 SoC-associated CpGs (false discovery rate [FDR]<5%) identified in the ENID (2 yr) cohort.
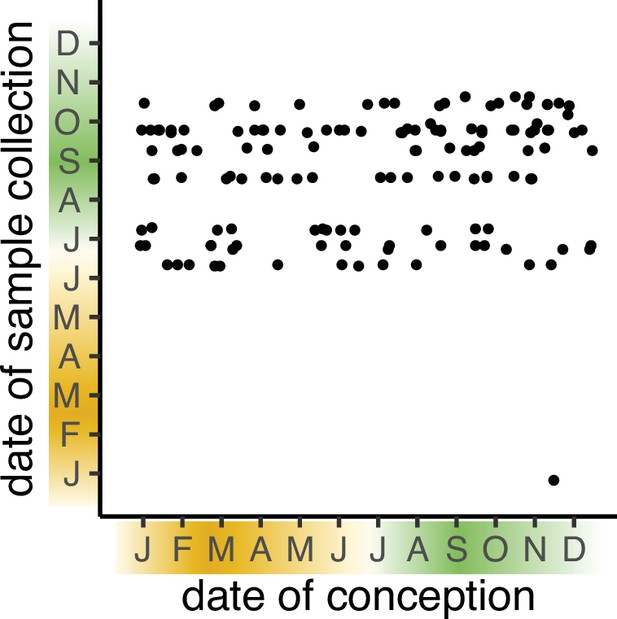
Date of conception vs. date of collection for samples analysed in the ENID longitudinal (5–7 yr) dataset.
Data comprises samples from n=138 ENID participants with methylation data at both 2 yr and 5–7 yr.
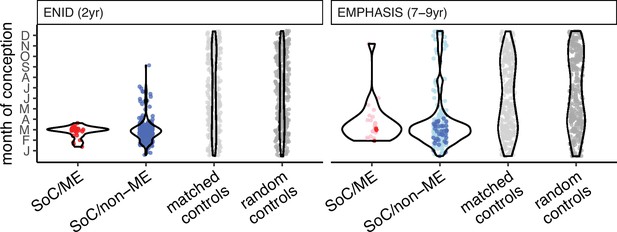
Date of conception at modelled methylation minima.
Conception dates at methylation minima for loci plotted in Figure 4A. Note that since seasonality is modelled by a single pair of Fourier terms, maxima and minima are 6 months apart.
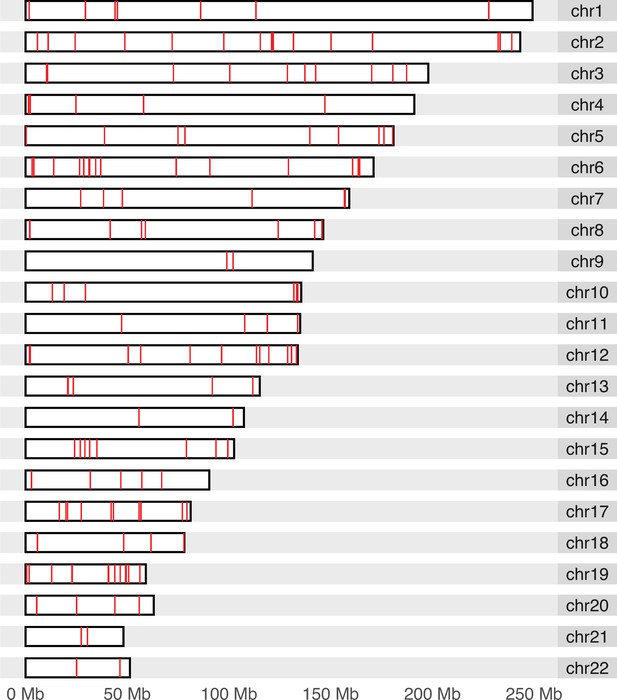
Genomic locations (hg19) of 259 SoC-CpGs.
Data from Supplementary file 1D.
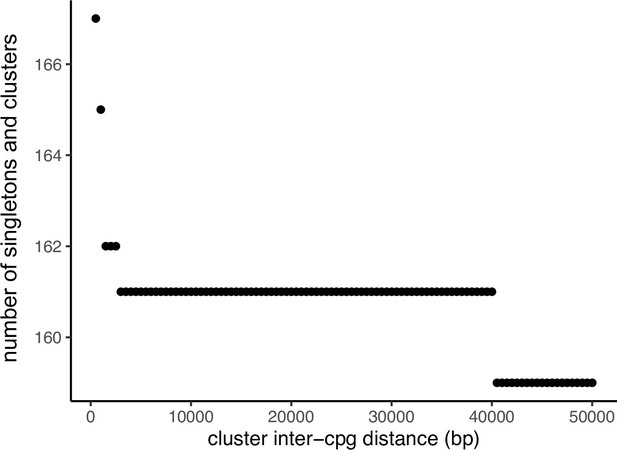
Relationship between the inter-CpG distance used to define season of conception (SoC-CpG clusters) and the number of identified clusters and singletons (CpGs not falling within a cluster).
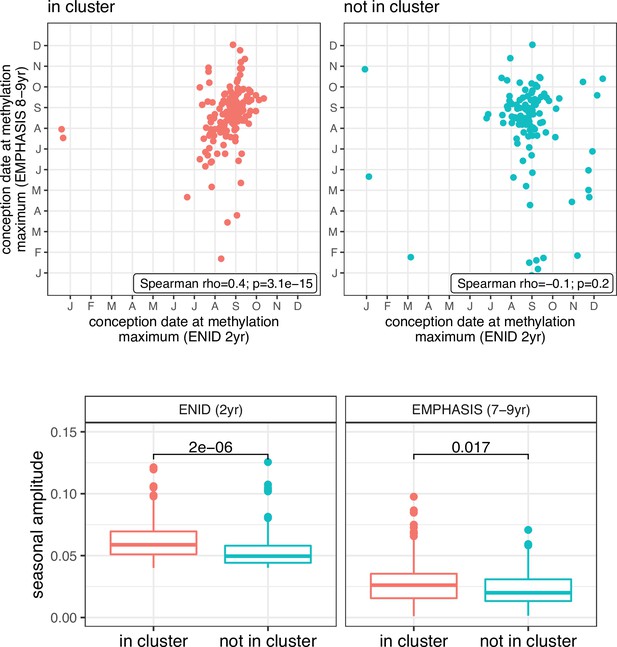
Comparison of SoC effects at SoC-CpGs within and outside of SoC-CpG clusters.
Top: Conception date at modelled methylation maxima for ENID 2 yr (x-axis) and EMPHASIS (y-axis) cohorts, according to whether locus falls within (n=154; left-red), or outside (n=105; right-blue) of an SoC-CpG cluster. Bottom: Seasonal effect size (amplitude) for ENID 2 yr (left) and EMPHASIS (cohorts), according to whether locus falls within (red), or outside (blue) of an SoC-CpG cluster. Numbers are p-values for two-sided Wilcoxon rank sum tests under the null hypothesis of no difference between the two distributions.
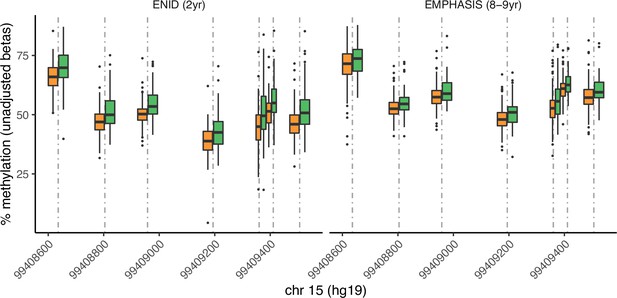
Seasonal variation at the IGF1R locus in ENID (2 yr) and EMPHASIS cohorts.
Unadjusted methylation beta values are plotted. Boxes represent inter-quartile ranges (IQRs) for season-specific DNAm at each locus. Note that for visualisation purposes Gambian seasons are dichotomised (dry: Jan-Jun; rainy: Jul-Dec), whereas seasonality is modelled as a continuous cyclical variable in Fourier regression models used in the main analysis.
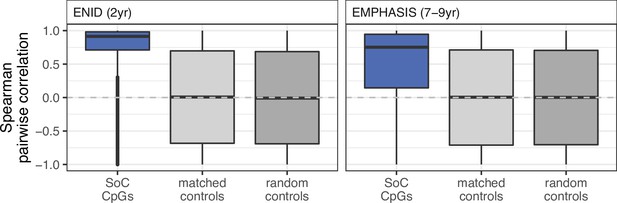
Cluster-adjusted pairwise inter-CpG methylation correlations for CpG sets in ENID 2 yr (left) and EMPHASIS (right) cohorts.
This is the same as Figure 4D, but with a single CpG randomly sampled from each CpG cluster so that CpGs in each set are a minimum distance of 5000 bp apart. This shows that pairwise correlation distributions are not disproportionately driven by large numbers of pairwise correlations between highly correlated neighbours.
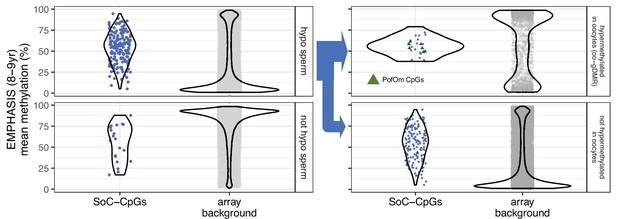
Mean methylation at SoC-CpGs and array background measured in n=289 individuals in the EMPHASIS (7–9 yr) cohort stratified by sperm and oocyte methylation status.
(Left): Methylation stratified according to sperm methylation status reported in Sugden et al., 2020. Sperm hypomethylation is defined as methylation ≤ 25%. (Right) As left but with loci hypomethylated in sperm further stratified according to oocyte germline differentially methylated region (oo-gDMR) status reported in Zink et al., 2018. Parent-of-origin-specific methylation (PofOm) CpGs are those identified in James et al., 2018b. oo-gDMR have oocyte methylation >75%.
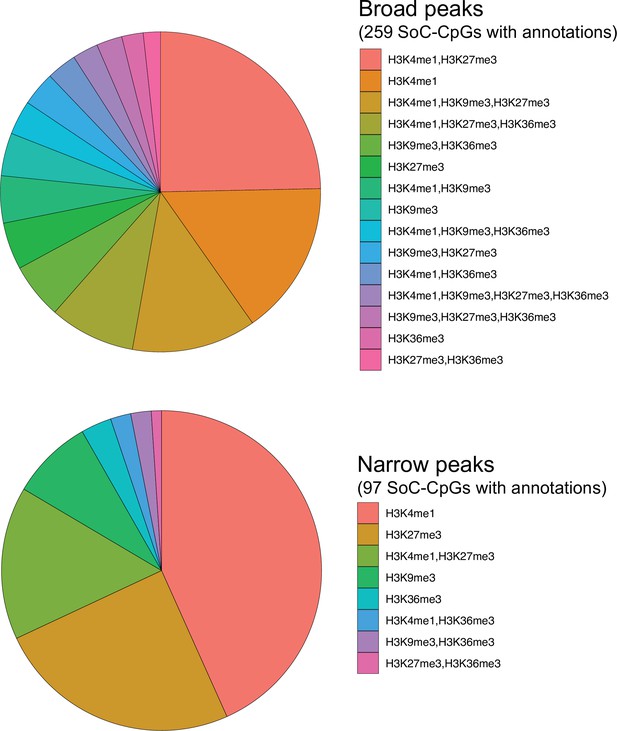
Histone marks overlapping SoC-CpGs in H1 embryonic stem cells.
Histone marks or combinations thereof are ordered by abundance. H3 marks were generated by the Roadmap Epigenomics Consortium Sanchez-Delgado et al., 2016 and downloaded using the annotatr (v1.10.0) package in R. Broad peaks overlapping all 259 SoC-CpGs (top) and (more sharply defined) narrow peaks overlapping 97 SoC-CpGs (bottom) correspond to different thresholds used by the MACS peak-calling algorithm. See Sanchez-Delgado et al., 2016 for further details.
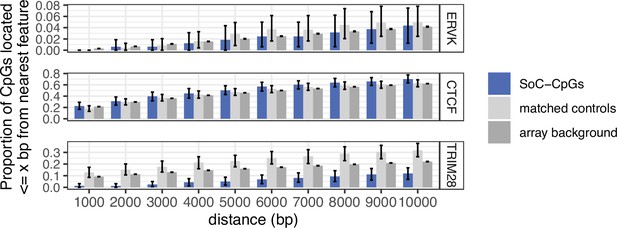
Links between ERVK endogenous retroviral elements, ZFP57 and TRIM28 binding sites, and DNAm at SoC-CpGs.
(A) Proportion of SoC-CpGs, matched controls, and array background CpGs proximal to ERVK endogenous retroviral elements (top), and ZFP57 and TRIM28 binding sites (bottom), within the specified distance. CpG clustering effects are removed by sampling a single CpG from each cluster (see Materials and methods). Error bars are bootstrapped 95% CIs.
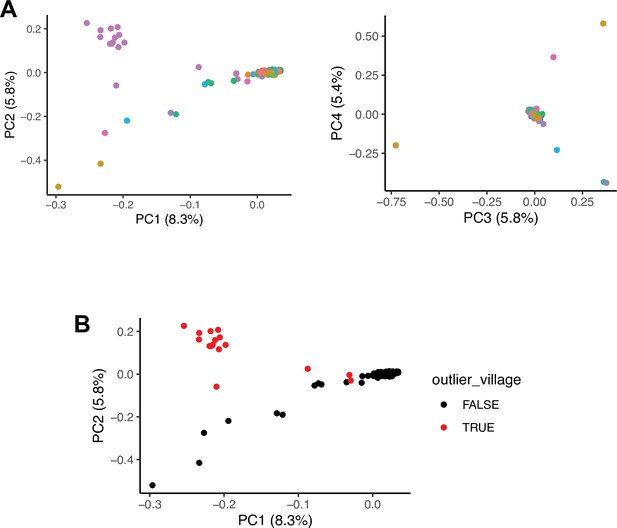
Population structure in the EMPHASIS cohort.
(A) First and second (left) and second and third (right) principal components (PCs) from a principal component analysis (PCA) of genome-wide genetic data from n=294 individuals from the EMPHASIS cohort. Figures in brackets give % variance explained by each PC. Colours correspond to the 24 villages from the Kiang West region which are covered in this cohort. A majority of individuals are of Mandinka origin and villages that are predominantly Mandinka are tightly clustered in the PCA plot. The distinct cluster in the top left of the first plot corresponds to individuals from a single village which is predominantly Fula. (B) Same as A left, but with individuals from the ‘outlier village’ plotted as a distinct colour. This illustrates that all 16 individuals from this village are distinct from the main cluster.

Season of conception analysis adjusted for ethnicity.
This figure replicates Figure 4A in the main paper for the ENID 2 yr and EMPHASIS cohorts, but with additional adjustment for ethnicity. Here, we adjust for ethnicity in the EMPHASIS cohort using the first two genetic principal components as covariates in the Fourier regression models (Appendix 1—figure 12A). For the ENID cohort where we do not have genetic data, we adjust for ethnicity using an additional covariate dichotomised according to whether an individual was one of the nine who came from the predominantly Fula ‘outlier village’ identified from the principal component analysis (PCA) using genetic data in the EMPHASIS cohort (Appendix 1—figure 12B).
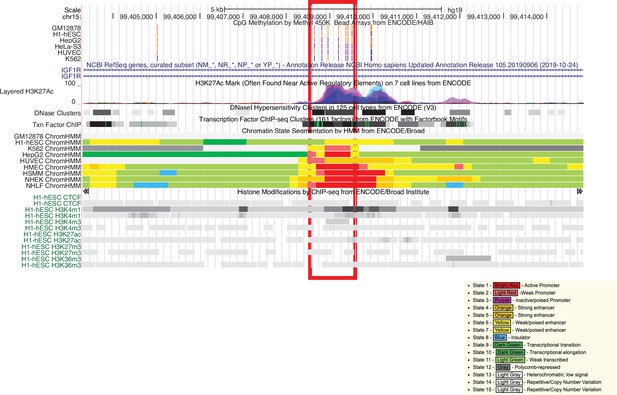
UCSC genome browser plot of the IGF1R SoC-CpG cluster.
Seven SoC-CpGs on the Illumina 450k array are highlighted. This region falls within intron 2 and bears the hallmarks of being a promoter and/or active or poised enhancer in multiple cell lines. The key lists colour coding for predicted regulatory regions using ChromHMM.
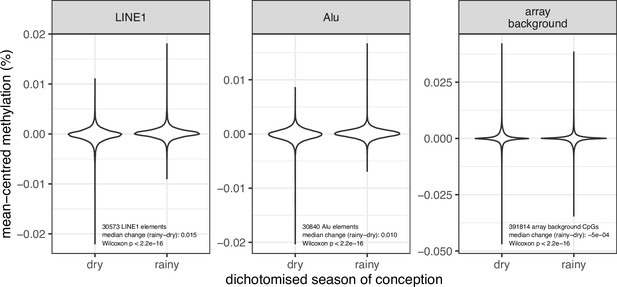
Seasonal differences in methylation at 30,573 LINE1 and 30,840 Alu elements, and 391,814 array background CpGs.
Methylation values at each locus are centred to have mean zero to enable comparison across loci. LINE1 and Alu methylation values are predicted using REMP (v1.16.0) (see Materials and methods). Season of conception is dichotomised to dry: Jan-June; rainy: July-December.
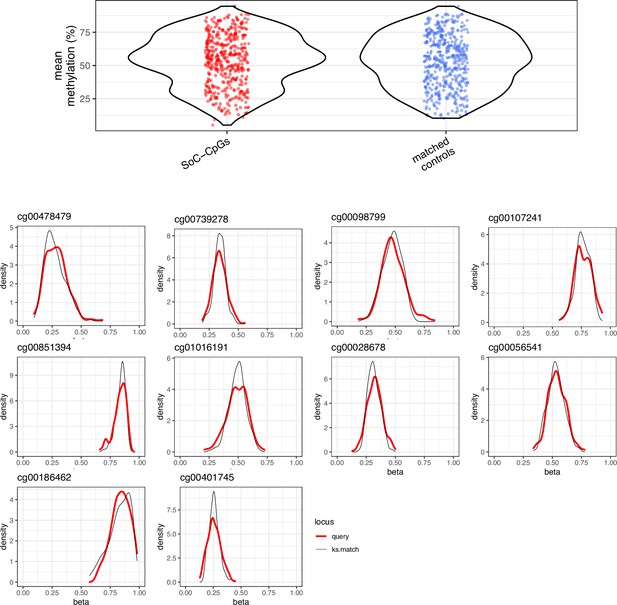
Selection of matched controls.
Matched controls are selected from array background using Kolmogorov-Smirnov (KS) tests to identify CpGs with similar methylation distributions to SoC-CpGs (see Materials and methods). Top: Methylation mean distribution at 259 SoC-CpGs (left) vs. 259 matched controls (right). Bottom: Methylation distribution of 10 SoC-CpGs selected at random (thick red lines) and their corresponding KS-matched controls (thin black lines).
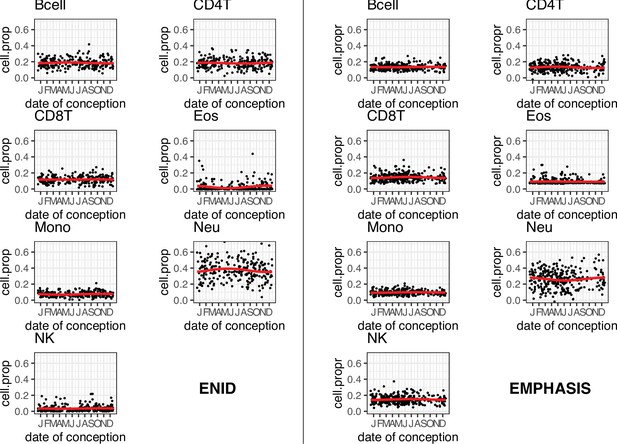
Seasonal variation in estimated cell composition.
Cell proportions for all samples from each cohort are estimated using the estimateCellCounts function from the minfi package in R. Seasonal variation (red curves) for each cell type is determined from Fourier regression models adjusted for sex (Early Nutrition and Immune Development [ENID]) or age and sex (Epigenetic Mechanisms linking Pre-conceptional nutrition and Health Assessed in India and Sub-Saharan Africa [EMPHASIS]). See Materials and methods for further details.
Tables
Gambian seasonality-methylation analysis: cohort characteristics.
| Cohort | Sample size | Age | % male | Tissue | Methylation array |
|---|---|---|---|---|---|
| ENID (2 yr) | 233 | 2 years | 50.6 | Peripheral blood | Illumina Infinium HM450 |
| ENID (5–7 yr) | 138 | 5–7 years | 56.5 | Illumina Infinium MethylationEPIC | |
| EMPHASIS | 289 | 7–9 years | 54.3 | Peripheral blood | Illumina Infinium MethylationEPIC |
-
Note: ENID: Early Nutrition and Immune Development Trial (Moore et al., 2012); EMPHASIS: Epigenetic Mechanisms linking Pre-conceptional nutrition and Health Assessed in India and Sub-Saharan Africa (Chandak et al., 2017). Individuals with ENID longitudinal (5–7 yr) methylation data are a subset of those with methylation at 2 yr. There is no overlap between individuals included in the ENID and EMPHASIS cohorts.
CpG sets considered in this analysis.
| CpG set | Number of CpGs | Notes |
|---|---|---|
| Array background | 391,814 | Intersection of CpGs on Illumina HM450 (ENID 2 yr) and EPIC (EMPHASIS) cohort arrays, post QC |
| SoC-CpGs | 259 | SoC-associated CpGs with SoC effect size (SoC methylation amplitude) > 4% in the ENID 2 yr dataset |
| Matched controls | 259 | CpGs with similar methylation distributions to SoC-CpGs in the ENID 2 yr dataset* |
| Random controls | 259 | Random sample from array background |
-
*
Matching methylation distributions determined by Kolmogorov-Smirnov tests (see Appendix 1—figure 16). QC: quality control; LRT: likelihood ratio test. See Materials and methods for further details.
External datasets considered in this analysis.
| CpG set | Notes |
|---|---|
| Putative metastable epialleles (MEs) | 1881 ME/SIV/ESS CpGs overlapping array background identified in multi-tissue and MZ/DZ screens in Van Baak et al., 2018 and Kessler et al., 2018. |
| Parent-of-origin-specific methylation (PofOm) | 699 Parent-of-origin-specific methylation loci identified in peripheral blood in Zink et al., 2018, overlapping array background. |
| Embryo DNAm data | RRBS data for inner cell mass and embryonic liver (<10 weeks’ gestation) from Guo et al., 2014. |
| Sperm DNAm data | WGBS data from Okae et al., 2014. |
| Germline DMRs (gDMRs) | Regions differentially methylated in sperm and oocytes identified in WGBS data by Sanchez-Delgado et al., 2016. |
| Transposons (ERVs) | ERVs determined by RepeatMasker were downloaded from the UCSC h19 annotations repository. |
| Transcription factor ChIP-seq | ZFP57, TRIM28, and CTCF transcription factor binding sites identified from ChIP-seq in human embryonic kidney and hESCs are described in Kessler et al., 2018. |
| Chromatin state predictions and histone three marks | Chromatin state predictions for H1 ESCs, fetal brain, fetal muscle, and fetal small intestine generated using Ernst and Kellis, 2012, from Roadmap Epigenomics Consortium et al., 2015. Histone mark data are from the same source. |
-
ME: metastable epiallele; SIV: systemic interindividual variation; ESS: epigenetic supersimilarity; MZ/DZ: monozygotic/dizygotic twins; PofOm: parent-of-origin methylation; RRBS: reduced representation bisulfite-seq; DMR: differentially methylated region; ERV: endogenous retrovirus; ESCs: embryonic stem cells. See materials and methods for further details.
Methylation quantitative trait loci (mQTL) associated with SoC-CpGs and controls.
| CpG set | Number of CpGs with mQTL | Number of mQTL (cis/trans) | Median number of mQTL per CpG (IQR) | Methylation variance explained* |
|---|---|---|---|---|
| SoC-CpGs | 130 (50%) | 2771 (2549/222) | 6 (2–30) | 0.09 (0.08–0.15) |
| Matched controls | 201 (78%) | 7886 (7417/469) | 15 (4–50) | 0.09 (0.06–0.21) |
| Random controls | 50 (19%) | 1512 (1476/36) | 7 (2–35) | 0.1 (0.08–0.18) |
-
*
delta adjusted R2 (see Materials and methods); IQR: inter-quartile range.
Additional files
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/72031/elife-72031-transrepform1-v2.docx
-
Supplementary file 1
Supplementary tables.
(a) ENID SoC-associated CpGs (no amplitude filter applied). (b) Seasonal amplitude tests. (c) Inter-cohort change in SoC amplitude at ENID SoC-associated CpGs. (d) SoC-CpGs (ENID SoC-associated CpGs with SoC methylation amplitude >=4%). (e) SoC-CpG clusters and singletons. (f) SoC-CpG cluster sizes. (g) SoC-CpG enrichment for MEs and overlap with ME sub-classes. (h) SoC-CpG enrichment for gDMRs and parent-of-origin specific methylation. (i) SoC-CpGs overlapping maternally methylated germline DMRs. (j) SoC-CpGs mQTL and their association with SoC. (k) Candidate genes previously associated with periconception / gestational exposures overlapping array background. (l) Look-up of SoC-CpGs in the EWAS Catalog. (m) EWAS Catalog data grouped by trait. (n) SoC-CpG associations with selected variables measured in the ENID 2yr dataset. (o) Look-up of SoC-CpGs in the GWAS Catalog. (p) ENID (2yr): Association test p-values to detect potential residual confounding. (q) EMPHASIS (7-9yr): Association test p-values to detect potential residual confounding. (r) ENID 5-7yr: Association test p-values to detect potential residual confounding. (s) ENID (2yr): Cell count, batch, village sensitivity analyses.
- https://cdn.elifesciences.org/articles/72031/elife-72031-supp1-v2.xlsx





