NHE6 depletion corrects ApoE4-mediated synaptic impairments and reduces amyloid plaque load
Figures
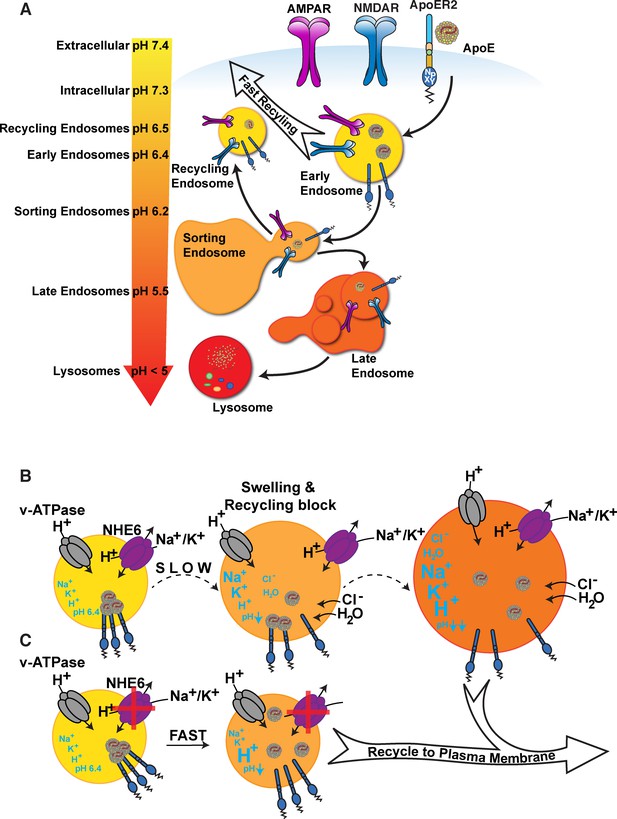
ApoE4 induces endolysosomal trafficking delay.
(A) pH regulation within the endolysosomal pathway. Upon receptor binding, ApoE is endocytosed along with glutamate receptors (AMPA and NMDA receptors). Cargo that has entered the early endosome (EE) can undergo recycling through a fast direct route without further acidification (fast recycling) or through slower sorting pathways that require further acidification (Casey et al., 2010; Naslavsky and Caplan, 2018). While lipid components are targeted to the lysosome, the majority of receptors, as well as ApoE, remain in endosomal compartments at the cellular periphery where they rapidly move back to the surface (Heeren et al., 1999). The increasingly acidic luminal pH is illustrated as a color gradient and depicted on the left. (B) In the presence of ApoE4, early endosomal trafficking and fast recycling are delayed. At the pH of the EE, ApoE4 is near its isoelectric point where solubility is reduced (Wintersteiner and Abramson, 1933), impairing receptor dissociation and resulting in delayed endosomal maturation with a concomitant entrapment of co-endocytosed glutamate receptors. Endosomal pH is regulated by the vesicular ATPase and the counterregulatory action of the proton leakage channel NHE6. NHE6 is an antiporter that exchanges a Na+ or K+ ion for each proton. As the pH decreases, ligands dissociate from their receptors allowing the EE to mature. If dissociation is delayed, as in case of ApoE4, endosomal trafficking is arrested, leading to progressive acidification as Na+, K+, and Cl- ions continue to enter the endosome to maintain charge neutrality while also drawing in water molecules due to osmotic pressure. We thus propose a model in which delayed ApoE4-receptor dissociation prevents early endosomal maturation and causes osmotic swelling while the pH continues to decrease until dissociation occurs. (C) Accelerated endosomal acidification by inhibition of the proton leak channel NHE6 resolves ApoE4 accumulation, promotes rapid receptor dissociation, and promotes the vesicle entry into the lysosomal delivery or recycling pathways.
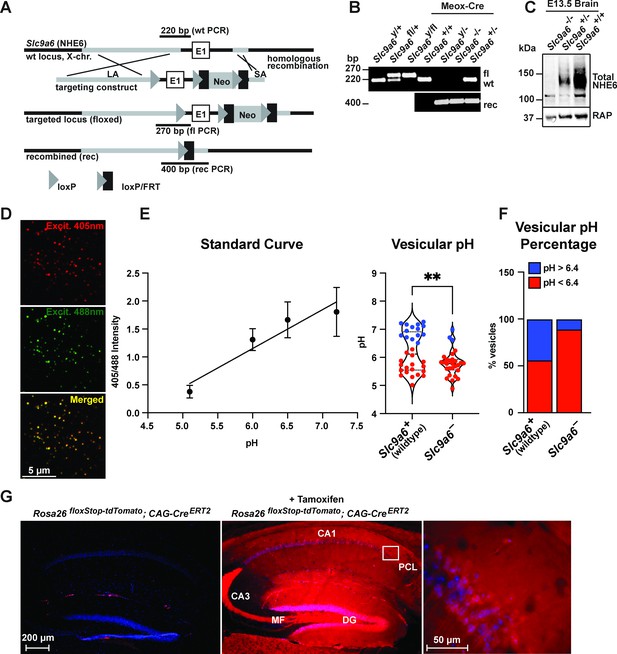
Generation of Slc9a6fl and Slc9a6- mice.
(A) Gene targeting strategy. LoxP sites were introduced to flank the first exon (E1) of Slc9a6 (located on the X-chromosome) by gene targeting in embryonic stem cells. The targeting construct contained a long arm of homology (LA, gray) upstream of the first loxP site and the first exon. An loxP/FRT-flanked neomycin resistance cassette was cloned downstream of the first exon, followed by a short arm of homology (SA, gray). The targeted locus is shown below. Targeted stem cells were used to generate chimeric Slc9a6fl mice. Germline NHE6 knockout mice (NHE6-/- [female], NHE6y/- [male]; rec indicates recombined allele) were generated by breeding the Slc9a6fl line with Meox-Cre mice. (B) Genotyping of wildtype (wt, +), floxed (fl), and recombined (rec, -) NHE6 alleles. The PCR amplified regions are indicated in panel A. The wildtype and floxed allele PCR products differ by 50 bp (270 for floxed, 220 for wildtype). (C) Western blot showing brain lysates (left) of different NHE6 genotypes after Meox-Cre-induced germline recombination. (D) Mouse embryonic fibroblasts from Slc9a6- and control littermate were infected with Vamp3-pHluorin2 and excited at 408 and 488 nm with emission measured at 510 nm. (E) Vesicular pH measured using a standard curve was significantly decreased in Slc9a6- fibroblasts. Data is expressed as mean ± SEM. Statistical analysis was performed using Student’s t-test. (**p < 0.01) (F) The percent of vesicles with pH >6.4 is significantly decreased in Slc9a6- fibroblasts. (G) CAG-CreERT2 activity after tamoxifen application in a reporter mouse line expressing tdTomato. CreERT2 recombination activity without (left panel) or with (middle panel) tamoxifen application in the CAG-CreERT2 line bred with Rosa26floxStop-tdTomato line. After tamoxifen induction, CreERT2 activity led to a robust tdTomato signal in the hippocampus (middle panel). Pyramidal neurons in the CA1 pyramidal cell layer (PCL) (middle panel) are shown magnified in the right panel.
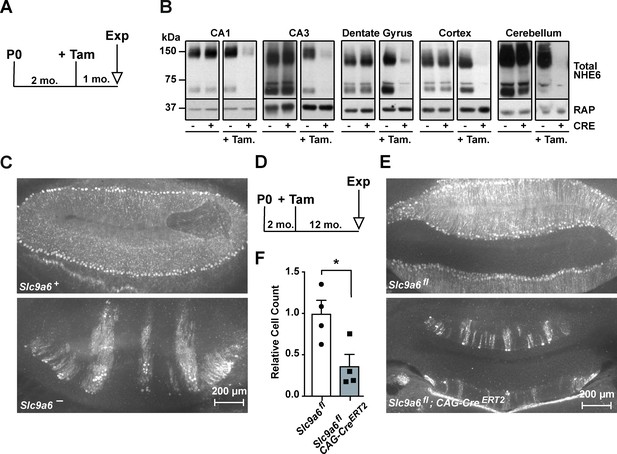
Long-term sodium-hydrogen exchanger 6 (NHE6) deficiency induced after Purkinje cell maturation causes Purkinje cell loss.
(A) Experimental timeline for B, mice were injected with tamoxifen at 2 months; after 1 month the brains were analyzed for NHE6 expression (Tam = tamoxifen, Exp. = experiment, mo. = months). (B) Western blot showing the efficiency of tamoxifen-induced NHE6 knockout in different brain regions (CA1, CA3, dentate gyrus, cortex, and cerebellum). The knockout efficiency differed between brain regions, it was 80% ± 2% in CA1, 82 ± 5.7% in the CA3, 67 ± 6.8% in the dentate gyrus, 65% ± 11.2% in the cortex, and 74% ± 4.7% in the cerebellum. A total of three brains in each group were examined. (C–F) NHE6 deficiency leads to cerebellar Purkinje cell loss in germline (Slc9a6-, C) and conditional (Slc9a6fl;CAG-CreERT2, D–F) knockout mice. Slc9a6+ includes both female wildtypes (Slc9a6+/+) and male wildtypes (Slc9a6y/+) mice. Slc9a6- includes both female knockouts (Slc9a6-/-) and male knockouts (Slc9a6y/-) mice. In addition, Slc9a6fl mice includes both female Slc9a6fl/fl and male Slc9a6y/fl. The timeline shows that Slc9a6fl;CAG-CreERT2 and control mice were tamoxifen-injected at 2 months and analyzed 1 year after (D). Calbindin was fluorescently labeled to highlight Purkinje cells in the cerebellum. Massive loss of Purkinje cells was found in Slc9a6- (C, lower panel), compared to wildtype Slc9a6+ control (C, upper panel). Long-term loss of NHE6, induced after Purkinje cell maturation at 2 months of age, also led to massive Purkinje cells loss when mice were examined 1 year after NHE6 ablation (E, lower panel). (F) Quantification of Purkinje cell loss in the cerebellum of Slc9a6fl;CAG-CreERT2 mice. Values are expressed as mean ± SEM from four independent experiments. Statistical analysis was performed using Student’s t-test. *p < 0.05.
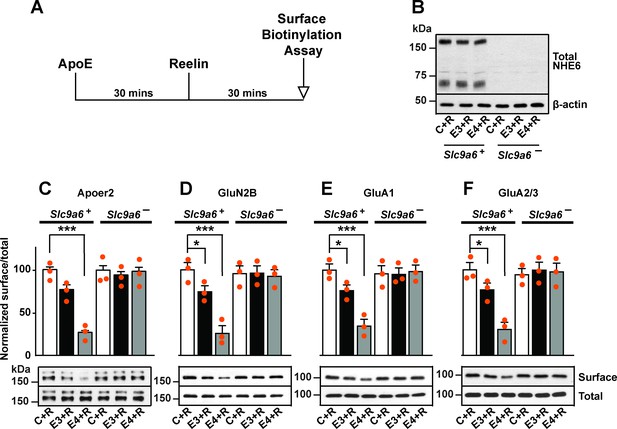
Sodium-hydrogen exchanger 6 (NHE6) deficiency alleviates ApoE4-impaired surface trafficking deficits of Apoer2 and glutamate receptors.
(A) Timeline for the receptor surface expression assay applied for the experiments shown in B–F. Primary neurons were treated with naturally secreted ApoE3 or ApoE4 and/or Reelin before they underwent surface biotinylation. (B–F) Wildtype and Slc9a6- primary neurons were prepared from littermates and used in the receptor surface expression assay described in A. Slc9a6+ includes both female wildtypes (Slc9a6+/+) and male wildtypes (Slc9a6y/+) mice. Slc9a6- includes both female knockouts (Slc9a6-/-) and male knockouts (Slc9a6y/-) mice. (B) NHE6 deficiency was confirmed via Western blot, β-actin was used as loading control. (C–F) ApoE-conditioned media treatment reduces the surface expression of Apoer2 and glutamate receptors in presence of Reelin in primary neurons. Receptor surface levels show a stronger reduction with ApoE4 than ApoE3. NHE6 depletion counteracts the ApoE4-induced reduction of receptor surface expression. Cell surface biotinylation assay was performed for Apoer2 (C), GluN2B (D), GluA1, (E) and GluA2/3 (F). Total levels were analyzed by immunoblotting of whole cell lysates against the same antibodies. β-Actin was used as loading control. Quantitative analysis of immunoblot signals is shown in the lower panels (C–F). All data are expressed as mean ± SEM from three independent experiments. *p < 0.05, **p < 0.01, ***p < 0.005. Statistical analysis was performed using one-way analysis of variance (ANOVA) and Dunnett’s post hoc test (C–F).
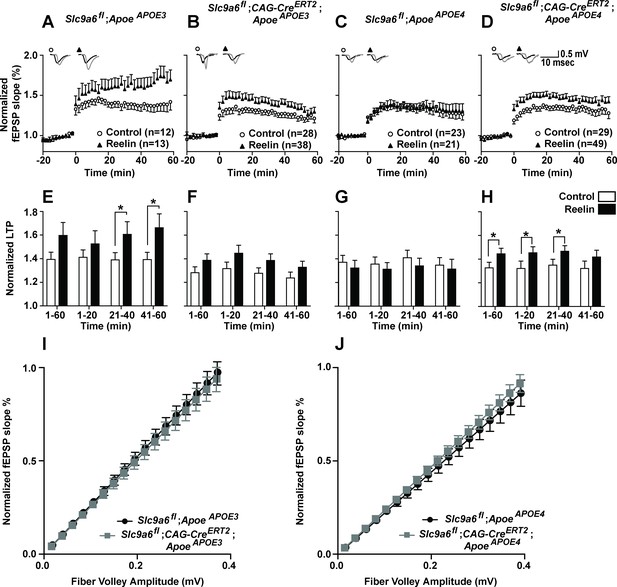
Effect of conditional sodium-hydrogen exchanger 6 (NHE6) knockout on Reelin-potentiated synaptic plasticity.
(A–H) Conditional knockout of NHE6 restores Reelin-enhanced long-term potentiation (LTP) in ApoeAPOE4 mice. Reelin facilitated induction of LTP in ApoeAPOE3 (A, E), but not ApoeAPOE4 (C, G) control (Slc9a6fl) mice. Slc9a6 deficiency in ApoeAPOE3 mice caused a reduction in Reelin-enhanced LTP, such that it is not significantly different from the control LTP (B, F). Importantly, in ApoeAPOE4;Slc9a6fl;CAG-CreERT2 mice Reelin was able to enhance theta burst-induced potentiation (D, H). Hippocampal slices were prepared from 3-month-old double mutant mice with either human ApoeAPOE3 or ApoeAPOE4 crossed with Slc9a6 conditional knockout mice (Slc9a6fl;CAG-CreERT2, tamoxifen injections at 6–8 weeks). Extracellular field recordings were performed in slices treated with or without Reelin. Theta burst stimulation (TBS) was performed after 20 min of stable baseline. Representative traces are shown in each panel, before TBS induction (black) and 40 min after TBS (gray). (E–H) Quantitative analysis of normalized fEPSP slopes at time intervals as indicated. (I, J) Input output curves of ApoeAPOE3 (I) and ApoeAPOE4 (J) mice with or without Slc9a6fl;CAG-CreERT2. Slc9a6fl mice includes both female Slc9a6fl/fl and male Slc9a6y/fl mice. Apoe mice are homozygous for APOE3 or APOE4. All data are expressed as mean ± SEM. N-numbers for each genotype group and treatment are indicated in panels A–D. *p < 0.05. Statistical analysis was performed using Student’s t-test.
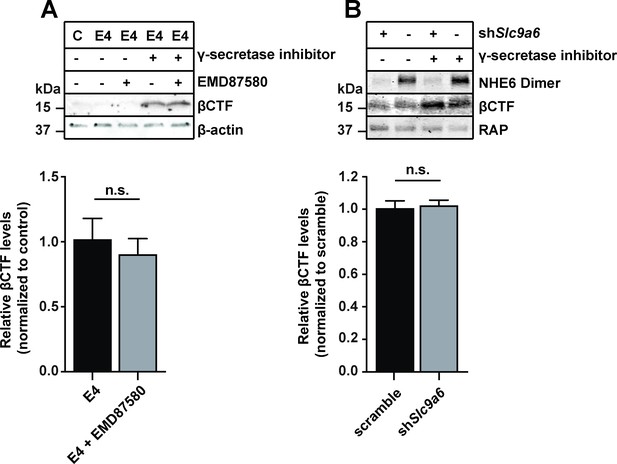
Na+/H+ exchanger (NHE) inhibition or sodium-hydrogen exchanger 6 (NHE6) knockdown does not alter β-site amyloid precursor protein cleaving enzyme 1 (BACE1) activity in primary neurons.
(A, B) Pan-NHE inhibition by EMD87580 or lentiviral knockdown of Slc9a6 (NHE6) did not alter BACE1 activity in primary neurons of AppSwe mice (Tg2576). (A) DIV10 primary neurons were treated with γ-secretase inhibitor L-685458, EMD87580, and/or ApoE4 (as indicated) and harvested for immunoblotting against Aβ-containing C-terminal fragment of APP (β-CTF). β-Actin was blotted as loading control. Bar graph shows the statistics of n = 3 experiments. (B) Primary neurons of AppSwe mice were infected with lentivirus for shRNA expression directed against Slc9a6(NHE6) (shSlc9a6) or a scramble control sequence (-) at DIV7. At DIV13 neurons were treated with L-685458 overnight and harvested for immunoblotting against NHE6 and β-CTF on DIV14. RAP was blotted as loading control. Bar graph shows the statistics of n = 6 experiments. All data are expressed as mean ± SEM. Statistical analysis was performed using Student’s t-test. n.s. = not significant.
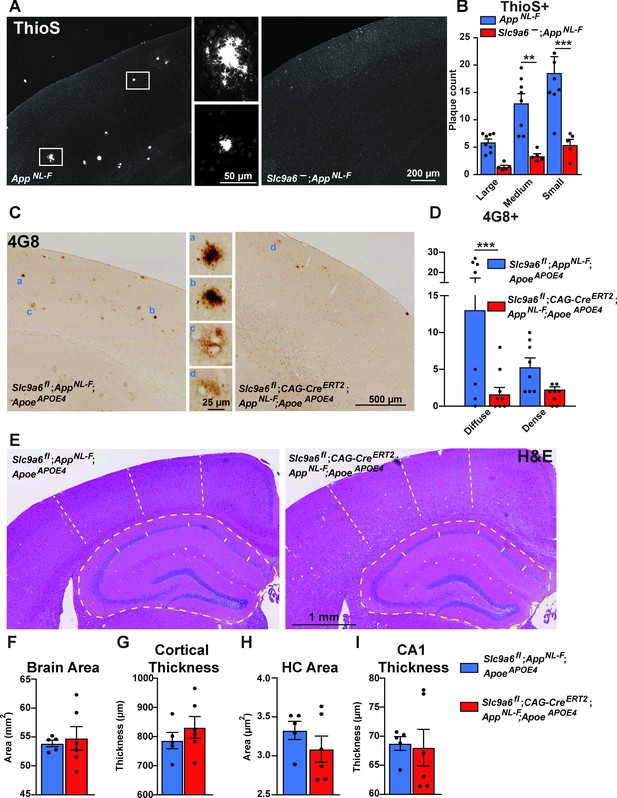
Sodium-hydrogen exchanger 6 (NHE6) deficiency decreases plaque formation in both AppNL-F and AppNL-F;ApoeAPOE4 mice.
(A, B) NHE6-deficient AppNL-F and control AppNL-F mice were analyzed for plaque deposition at an age of 12 months. Thioflavin S staining was performed to visualize plaques. Plaques were found more frequently in the control AppNL-F mice (left panel in A), magnifications of the boxed areas are shown in the two middle panels. The plaque load between Slc9a6- mice and control littermates (all AppNL-F) was compared and analyzed. (B) In the Slc9a6- littermates, the plaque number was reduced, when compared to controls. (C–D) Slc9a6fl;CAG-CreERT2;AppNL-F;ApoeAPOE4 and Slc9a6fl;AppNL-F;ApoeAPOE4 mice were analyzed for plaque deposition. NHE6 was ablated at 2 months and brains were analyzed at 13.5–16 months. 4G8-immunolabeling against Aβ was performed to visualize plaques. In AppNL-F;ApoeAPOE4 mice conditional Slc9a6 knockout caused a reduction in plaque load compared to the Slc9a6fl control littermates. (C) Magnifications of the boxed areas in C are shown in the middle. (D) Plaque load was analyzed and compared between Slc9a6fl;CAG-CreERT2 mice and floxed control littermates. (I) Hematoxylin and eosin (H&E) staining was performed to investigate for gross anatomic abnormalities in the Slc9a6fl;CAG-CreERT;AppNL-F;ApoeAPOE4 and Slc9a6fl;AppNL-F;ApoeAPOE4 mice. (F–I) Brain area (F), cortical thickness (G), hippocampal (HC) area (H), and CA1 thickness (I) were analyzed. Student’s t-test did not reveal a significant difference. Plaques were differentiated by size or staining density as described in detail in the supplements (Figure 7—figure supplement 2). Labeled plaques were analyzed by a blinded observer. All data are expressed as mean ± SEM. (B) Slc9a6- n = 5, control n = 8, (C) Slc9a6fl n = 8 (Slc9a6fl;CAG-CreERT2 n = 8), in (F–I) derived from n = 5 (Slc9a6fl) and n = 6 (Slc9a6fl;CAG-CreERT2) animals. *p < 0.05. **p < 0.01, ***p < 0.005. Slc9a6+ represents both female wildtypes (Slc9a6+/+) and male wildtypes (Slc9a6y/+). Slc9a6- represents both female knockouts (Slc9a6-/-) and male knockouts (Slc9a6y/-). In addition, Slc9a6fl mice includes both female Slc9a6fl/fl and male Slc9a6y/fl mice. Apoe mice are homozygous for APOE4 (ApoeAPOE4). AppNL-F mice are homozygous for human NL-F knockin mutation (AppNL-F/NL-F). Statistical analysis was performed using two-way analysis of variance (ANOVA) with Sidak’s post hoc test (B and D) and Student’s t-test (F–I).
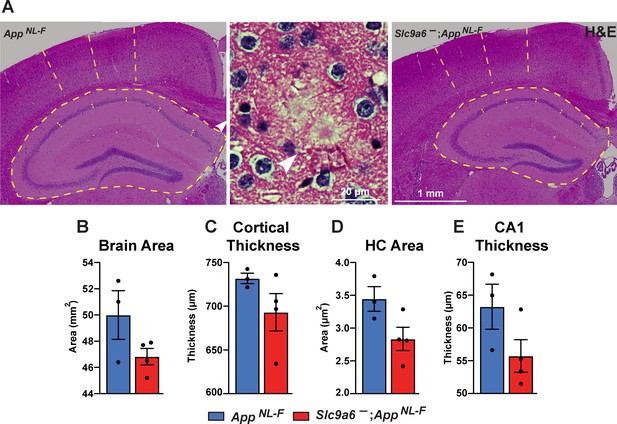
Gross anatomical brain structure in Slc9a6- mice.
(A) Hematoxylin and eosin (H&E) staining was performed to investigate for gross anatomic abnormalities in the Slc9a6-;AppNL-F and AppNL-F mice. Structures representing plaques were found in the AppNL-F control groups (magnified example is shown in the middle panel). (B–E) Brain area (B), cortical thickness (C), hippocampal (HC) area (D), and CA1 thickness (E) were analyzed. All data are expressed as mean ± SEM. Student’s t-test did not reveal significant differences, n = 3 (control) and n = 4 (Slc9a6-). Slc9a6+ represents both female wildtypes (Slc9a6+/+) and male wildtypes (Slc9a6y/+). Slc9a6- represents both female knockouts (Slc9a6-/-) and male knockouts (Slc9a6y/-). AppNL-F mice are homozygous for human NL-F knockin mutation (AppNL-F/NL-F).
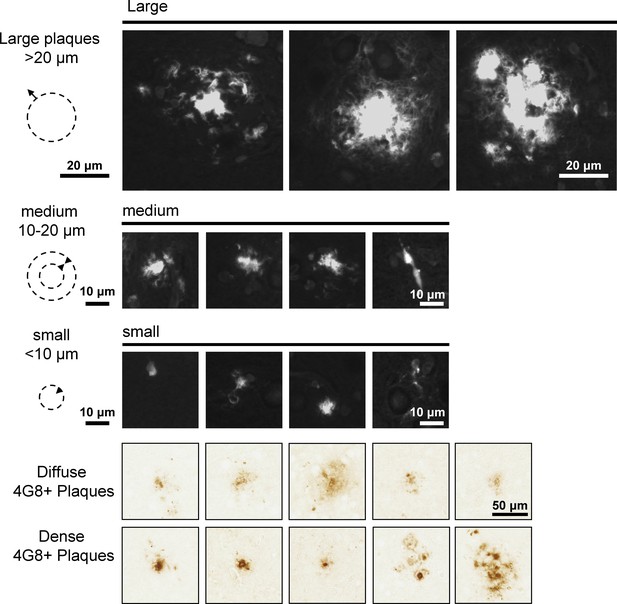
Example of Thioflavin S stained plaques for quantification.
Different types of Thioflavin S stained plaques and 4G8-immunoreactive accumulations in AppNL-F brains are shown. Different sizes of plaques were grouped together for quantification Thioflavin S-labeled plaques (Figure 7 and Figure 7—figure supplement 3). Plaques bigger in diameter than 20 µm with a dense core were defined as big. Medium sized plaques had a diameter between 10 and 20 µm with a dense core. Small plaques were smaller than 10 µm and often represented individual cells. 4G8-labeled plaques were differentiated by diffuse or dense appearance as depicted in the examples.
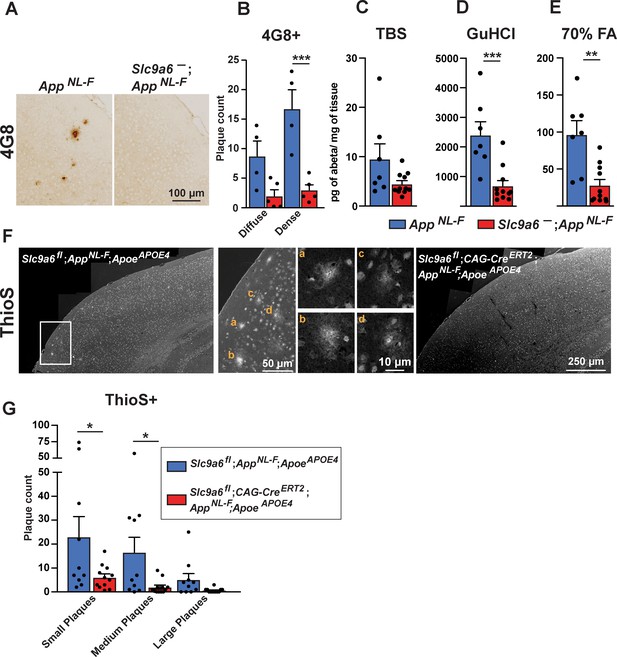
Sodium-hydrogen exchanger 6 (NHE6) deficiency decreases plaque formation in both AppNL-F and AppNL-F;ApoeAPOE4 mice.
(A–B) NHE6-deficient AppNL-F and control AppNL-F mice were analyzed for plaque deposition at an age of 12 months. 4G8-immunolabeling against Aβ (A) visualized more plaques in the control AppNL-F mice. The plaque load between Slc9a6- mice and control littermates (all AppNL-F) was compared and analyzed. (B) In the Slc9a6- littermates the plaque number was reduced, when compared to controls. (C–E) Soluble (TBS) and insoluble (GuHCl and 70% FA) Aβ fractions of cortical lysates were analyzed by commercial ELISA; 1.5-year-old Slc9a6- mice showed less insoluble Aβ than their control littermates (all AppNL-F). (F–G) Slc9a6fl;CAG-CreERT2;AppNL-F;ApoeAPOE4 and Slc9a6fl;AppNL-F;ApoeAPOE4 mice were analyzed for plaque deposition. NHE6 was ablated at 2 months and brains were analyzed at 13.5–16 months. Thioflavin S staining was performed to visualize plaques. With AppNL-F;ApoeAPOE4 background, the Slc9a6fl;CAG-CreERT2 mice had a reduced plaque load compared to the Slc9a6fl control mice (left panel in F). Magnifications of the boxed areas in left panel are shown in the middle. (G) Plaque load was analyzed and compared between Slc9a6fl;CAG-CreERT2 mice and floxed control littermates. Plaques were differentiated by size or staining density as described in detail in the supplements (Figure 7—figure supplement 2). Labeled plaques were analyzed by a blinded observer (B, G). All data (immunohistochemistry: Slc9a6- n = 5, control n = 4, Slc9a6fl n = 10, Slc9a6fl;CAG-CreERT2 n = 12; biochemistry: Slc9a6- n = 11, control n = 7) are expressed as mean ± SEM. *p < 0.05. **p < 0.01, ***p < 0.005. Slc9a6+ represents both female wildtypes (Slc9a6+/+) and male wildtypes (Slc9a6y/+). Slc9a6- represents both female knockouts (Slc9a6-/-) and male knockouts (Slc9a6y/-). In addition, Slc9a6fl mice includes both female Slc9a6fl/fl and male Slc9a6y/fl mice. Apoe mice are homozygous for APOE4. AppNL-F mice are homozygous for human NL-F knockin mutation (AppNL-F/NL-F). Statistical analysis was performed using two-way analysis of variance (ANOVA) with Sidak’s post hoc test.
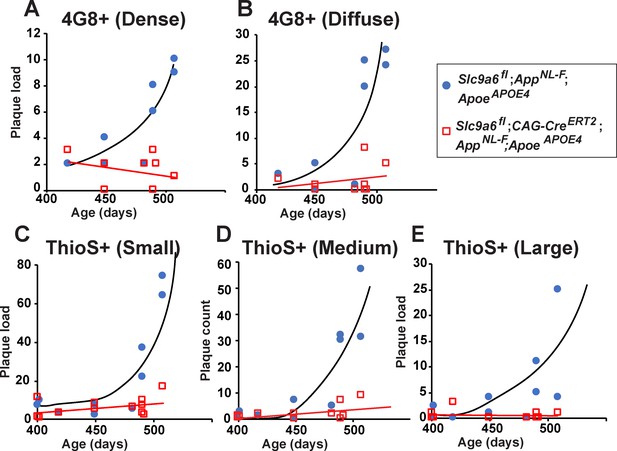
Age-dependent increase in plaque load is abolished in Slc9a6fl;CAG-CreERT2;AppNL-F;ApoeAPOE4 mice.
(A–E) Slc9a6fl;CAG-CreERT2;AppNL-F;ApoeAPOE4 and Slc9a6fl;AppNL-F;ApoeAPOE4 mice were analyzed for plaque deposition. Sodium-hydrogen exchanger 6 (NHE6) was ablated at 2 months and brains were analyzed at 13.5–16 months. 4G8-immunolabeling against Aβ (A,B) and Thioflavin S staining (C–E) were performed to visualize plaques (Figure 7C and Figure 7—figure supplement 3F). Plaque load was analyzed and compared between Slc9a6fl;CAG-CreERT2 mice and floxed control littermates. Plaques were differentiated by staining intensity (A, B) or size (C–E) as described in the supplements (Figure 7—figure supplement 2). In the time range analyzed, plaque load increased by age in control, but not in Slc9a6fl;CAG-CreERT2 mice. Plaques were analyzed by a blinded observer. Plaque count (Slc9a6fl;CAG-CreERT2 n = 8, Slc9a6fl n = 8 for A) (B); Slc9a6fl;CAG-CreERT2 n = 12; Slc9a6fl n = 10 in (C–E) is pis plotted against age of mice. Slc9a6fl mice includes both female Slc9a6fl/fl and male Slc9a6y/fl mice. Apoe mice are homozygous for APOE4. AppNL-F mice are homozygous for human NL-F knockin mutation (AppNL-F/NL-F).
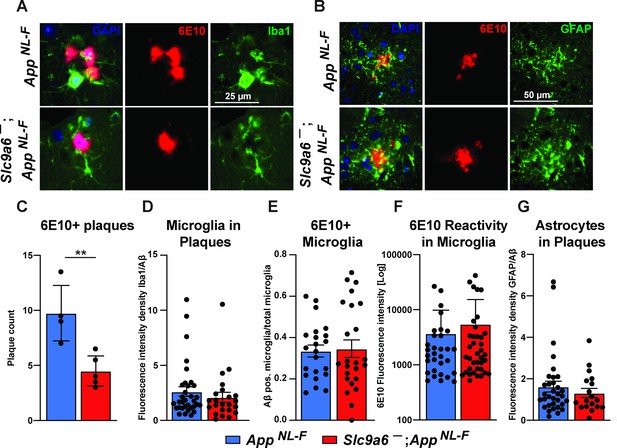
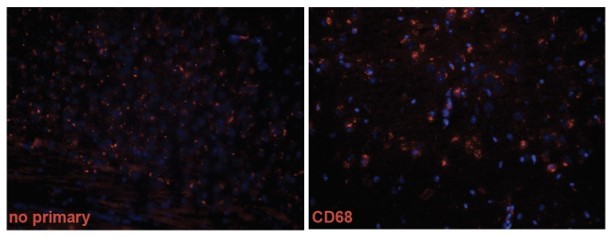
Microglia and astrocytes surround plaques in both AppNL-F control and AppNL-F;Slc9a6- brains.
(A–B) Co-labeling of microglia (Iba1, green, A) or astrocytes (GFAP, green, B) with Aβ (6E10, red) in brain slices of AppNL-F and AppNL-F;Slc9a6- mice. (C) Quantification of plaques in control and Slc9a6- brain slices. (D) Bar graph showing the intensity density of Iba1/6E10 as quantitative measure of microglia surrounding plaques. (E) Statistical analysis of 6E10 positive microglia and (F) the intensity of 6E10 signal within microglia. (G) Bar graph showing the intensity density of GFAP/Aβ as quantitative measure of astrocytes surrounding plaques. Data were analyzed by a blinded observer. All data are expressed as mean ± SEM. Data were obtained from n = 4 (control) and n = 5 (Slc9a6-) mice (A–G).(D) n = 33 (control) and n = 23 (Slc9a6-) plaques were analyzed, in (E) n = 22 (control) and n = 24 (Slc9a6-) microscopical pictures were analyzed, in (F) n = 31 (control) and n = 38 (Slc9a6-) 6E10 positive (defined as signal intensity above 500) microglia were analyzed, in (G) n = 33 (control) and n = 18 (Slc9a6-) plaques were analyzed. Student’s t-test revealed a difference in C (**p < 0.01) and did not reveal significant differences in (D–G). Slc9a6+ represents both female wildtypes (Slc9a6+/+) and male wildtypes (Slc9a6y/+). Slc9a6- represents both female knockouts (Slc9a6-/-) and male knockouts (Slc9a6y/-). AppNL-F mice are homozygous for human NL-F knockin mutation (AppNL-F/NL-F).
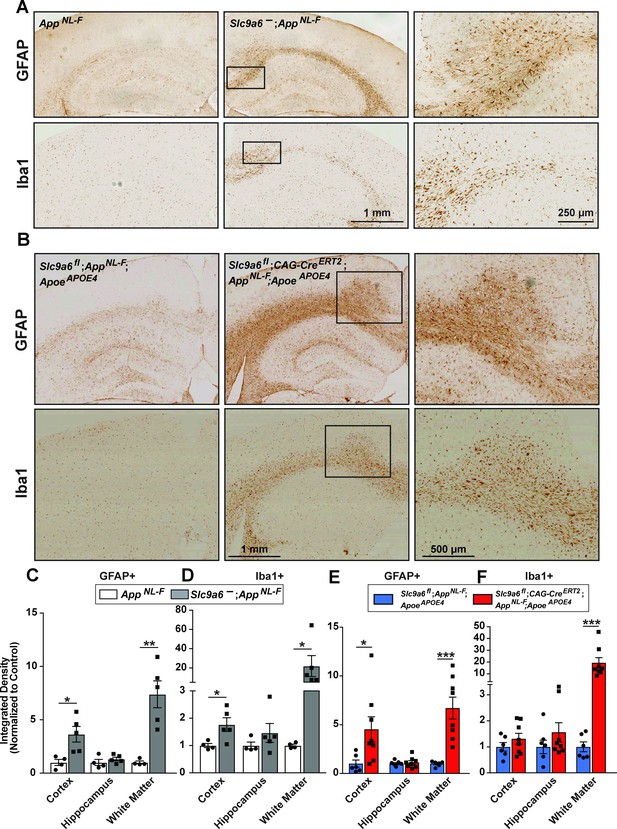
Sodium-hydrogen exchanger 6 (NHE6) deficiency causes an increase in Iba1 and glial fibrillary acidic protein (GFAP) immunoreactivity in both AppNL-F and AppNL-F;ApoeAPOE4 mice.
(A) Immunohistochemistry against GFAP and ionized calcium-binding adapter molecule (Iba1) was performed on brain slices obtained from 1-year-old AppNL-F mice deficient for NHE6 (AppNL-F;Slc9a6-) and control littermates (AppNL-F). GFAP (upper panels) and Iba1 (lower panels) immunoreactivity was increased in NHE6-deficient AppNL-F mice when compared to NHE6 expressing controls. (B) Slc9a6fl;CAG-CreERT2;AppNL-F;ApoeAPOE4 and Slc9a6fl;AppNL-F;ApoeAPOE4 mice were analyzed for immunoreactivity GFAP and Iba1. Mice were injected with tamoxifen at 2 months and brain slices obtained from 13.5- to 16-month-old mice. GFAP (upper panels) and Iba1 (lower panels) immunoreactivity was increased in NHE6-deficient AppNL-F;ApoeAPOE4 mice when compared to NHE6-expressing controls. (C–F) Intensity of the staining in various areas was compared for GFAP (C) and Iba1 (D) in AppNL-F;Slc9a6- and AppNL-F mice. Intensity of the staining for GFAP (E) and Iba1 (F) in Slc9a6fl;CAG-CreERT2;AppNL-F;ApoeAPOE4 and Slc9a6fl;AppNL-F;ApoeAPOE4 mice. Analysis was performed by a blinded observer. ‘White matter’ comprises corpus callosum, cingulum, and external capsule. All data (Slc9a6- n = 5; control n = 4; Slc9a6fl;CAG-CreERT2 n = 6; Slc9a6fl n = 8) are expressed as mean ± SEM. *p < 0.05. **p < 0.01, ***p < 0.005. Slc9a6+ represents both female wildtypes (Slc9a6+/+) and male wildtypes (Slc9a6y/+). Slc9a6- represents both female knockouts (Slc9a6-/-) and male knockouts (Slc9a6y/-). In addition, Slc9a6fl mice includes both female Slc9a6fl/fl and male Slc9a6y/fl mice. For the Cag-CreERT2, Cre+ + the absence of Cre (Slc9a6fl alone) and Cag-CreERT2 indicates the presence of Cre (Slc9a6fl;CAG-CreERT2), which are both injected with Tamoxifen at 2 months of age. Apoe mice are homozygous for APOE4. AppNL-F mice are homozygous for human NL-F knockin mutation (AppNL-F/NL-F). Statistical analysis was performed using Student’s t-test.
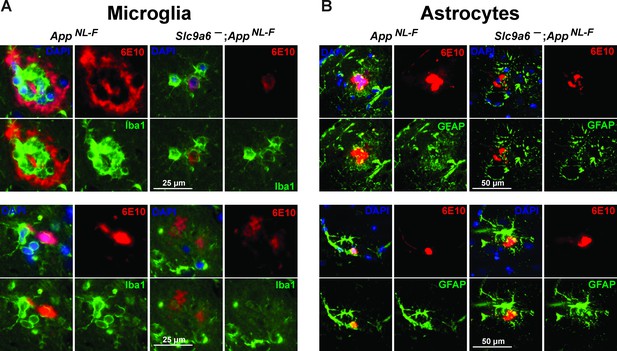
Examples of plaques surrounded by microglia and astrocytes.
(A) Co-labeling of microglia (Iba1, green) and Aβ (6E10, red) in brain slices of AppNL-F and AppNL-F;Slc9a6- mice. (B) Co-labeling of astrocytes (glial fibrillary acidic protein [GFAP], green) and Aβ (6E10, red) in brain slices of AppNL-F and AppNL-F; Slc9a6- mice. Slc9a6+ represents both female wildtypes (Slc9a6+/+) and male wildtypes (Slc9a6y/+). Slc9a6- represents both female knockouts (Slc9a6-/-) and male knockouts (Slc9a6y/-). AppNL-F mice are homozygous for human NL-F knockin mutation (AppNL-F/NL-F).
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Mus musculus) | Mouse/Slc9a6fl | This study | Refer to Materials and methods section for detailed description of mouse model production | |
| Strain, strain background (Mus musculus) | Mouse/Slc9a6- | This study | Refer to Materials and methods section for detailed description of mouse model production | |
| Strain, strain background (Mus musculus) | Mouse/ApoeAPOE3 | Sullivan et al., 1997 | IMSR_TAC:2,542 | ApoeAPOE3 |
| Strain, strain background (Mus musculus) | Mouse/ApoeAPOE4 | Knouff et al., 1999 | IMSR_TAC:3,518 | ApoeAPOE4 |
| Strain, strain background (Mus musculus) | Mouse/B6.Cg-Gt(ROSA)26Sortm9(CAG-tdTomato)Hze/J | The Jackson Laboratory Madisen et al., 2010 | JAX #007909 | ROSAfloxedStop-tdTomato |
| Strain, strain background (Mus musculus) | Mouse/CAG-cre/Esr15Amc/J | The Jackson Laboratory Hayashi and McMahon, 2002 | JAX #004682 | CAG-CreERT2 |
| Strain, strain background (Mus musculus) | Mouse/B6.129S4-Meox2tm1(cre)Sor/J | The Jackson Laboratory Tallquist and Soriano, 2000 | JAX 003755 | Meox-Cre |
| Strain, strain background (Mus musculus) | AppNL-F | Saito et al., 2014 | AppNL-F | |
| Strain, strain background (Mus musculus) | Tg2576 | Charles River Hsiao et al., 1996 | Charles River Tg2576 | Tg2576, APPSwe |
| Strain, strain background (Rattus norvegicus) | SD rat | Charles River | SC:400 | |
| Cell line (Homo sapiens) | HEK293 | Thermo Fisher | R70507, RRID:CVCL_0045 | |
| Cell line (Homo sapiens) | HEK293-T | ATCC | CRL-3216 | |
| Cell line (Mus musculus) | Neuro-2a | ATCC | CCL-131 | |
| Cell line (Mus musculus) | NHE6-KO (Slc9a6-) mouse embryonic fibroblasts (MEFs) | This study | Refer to Materials and methods section for detailed description of MEF production | |
| Cell line (Mus musculus) | Slc9a6+ MEFs (Slc9a6- littermate) | This study | Refer to Materials and methods section for detailed description of MEF production | |
| Antibody | Anti-Aβ (clone 6E10) (mouse monoclonal) | Covance | SIG-39320 RRID:AB_662798 | WB and IHC (1:1000) |
| Antibody | Anti-Aβ (clone 4 G8) (mouse monoclonal) | Covance | SIG-39220 RRID:AB_10175152 | IHC (1:1000) |
| Antibody | Anti-phospho tyrosine (clone 4 G10) (mouse monoclonal) | EMD Millipore | Millipore Cat# 05–321, RRID:AB_309678 | WB (1:1000) |
| Antibody | Anti-Apoer2 (rabbit polyclonal) | Herz Lab, #2561, Trommsdorff et al., 1999 | WB (1:1000) | |
| Antibody | Anti-β-Actin (rabbit polyclonal) | Abcam | Ab8227, RRID:AB_23051 86 | WB (1:3000) |
| Antibody | Anti-Calbindin D-28k (mouse monoclonal) | Swant | Swant Cat# 300, RRID:AB_10000347 | IHC (1:1000) |
| Antibody | Anti-GFAP (rabbit polyclonal) | Abcam | Abcam Cat# ab7260, RRID:AB_305808 | IHC (1:2000) |
| Antibody | Anti-GluA1 (rabbit polyclonal) | Abcam | ab31232, RRID:AB_2113447 | WB (1:1000) |
| Antibody | Anti-GluA2/3 (rabbit polyclonal) | EMD Millipore | 07–598, RRID:AB_31074 1 | WB (1:1000) |
| Antibody | Anti-GluN2B (rabbit polyclonal) | Cell Signaling Technology | 4,207 S, RRID:AB_12642 23 | WB (1:1000) |
| Antibody | Anti-Iba1 (rabbit polyclonal) | Wako | 019–19741, RRID:AB_839504 | IHC (1:1000) |
| Antibody | Anti-NHE6 (C-terminus) (rabbit polyclonal) | Herz Lab, Xian et al., 2018 | WB (1:1000) | |
| Antibody | Anti-mouse-IgG AF594 (goat polyclonal) | Thermo Fisher | A-11032, RRID:AB_2534091 | IHC (1:500) |
| Antibody | Anti-rabbit-IgG AF488 (goat polyclonal) | Thermo Fisher | A-11034, RRID:AB_2576217 | IHC (1:500) |
| Commercial assay or kit | Anti-mouse-IgG staining kit | Vector | MP-7602, RRID:AB_2336532 | |
| Commercial assay or kit | Anti-rabbit-IgG staining kit | Vector | MP-7601, RRID:AB_2336533 | |
| Chemical compound, drug | Antigen retrieval citrate buffer | BioGenex, Cat | HK086-9K | |
| Chemical compound, drug | B-27 Supplement (50×), serum free | Thermo Fisher | 17504044 | |
| Chemical compound, drug | Cytoseal 60 | Thermo Fisher | 8310 | |
| Chemical compound, drug | DMEM | Sigma-Aldrich | D6046 | |
| Chemical compound, drug | FuGENE | Promega | E2311 | |
| Chemical compound, drug | HBSS (1×) | Gibco | 14175 | |
| Chemical compound, drug | L-Glutamic acid (glutamate) | Sigma-Aldrich | G1251 | |
| Chemical compound, drug | γ-Secretase inhibitor L-685458 | Tocris Bioscience | 2627 | |
| Chemical compound, drug | Penicillin-streptomycin solution, 100× | Corning | 30–002 CI | |
| Chemical compound, drug | Neurobasal Medium (1×) liquid without Phenol Red | Thermo Fisher | 12348017 | |
| Chemical compound, drug | NeutrAvidin Agarose | Thermo Fisher | 29201 | |
| Chemical compound, drug | Nonidet P-40 Alternative | EMD Millipore | 492016 | |
| Chemical compound, drug | 32% Paraformaldehyde AQ solution | Fisher Scientific | 15714 S | |
| Chemical compound, drug | PBS (1×) | Sigma-Aldrich | D8537 | |
| Chemical compound, drug | Penisillin-streptomycin | Corning | 30–002 CI | |
| Chemical compound, drug | Phosphatase inhibitor cocktail | Thermo Fisher | 78420 | |
| Chemical compound, drug | Poly-D-lysine | Sigma-Aldrich | A-003-M | |
| Chemical compound, drug | Protein A-Sepharose 4B | Thermo Fisher | 101042 | |
| Chemical compound, drug | Proteinase Inhibitor Cocktail | Sigma-Aldrich | P8340 | |
| Chemical compound, drug | Sulfo-NHS-SS-biotin | Pierce | 21331 | |
| Chemical compound, drug | Triton X-100 | Sigma-Aldrich | CAS9002-93-1 | |
| Chemical compound, drug | Tween 20 | Sigma | P1379 | |
| Other | Vectashield with DAPI | Vector Labs | H-1200 | (DAPI 1.5 µg/ml) |
| Transfected construct (Mus musculus) | pCrl, Reelin expression vector | D’Arcangelo et al., 1997 | N/A | |
| Transfected construct (Homo sapiens) | pcDNA3.1-ApoE3 | Chen et al., 2010 | N/A | Progenitor pcDNA3.1-Zeo |
| Transfected construct (Homo sapiens) | pcDNA3.1-ApoE4 | Chen et al., 2010 | N/A | Progenitor pcDNA3.1-Zeo |
| Transfected construct (Mus musculus) | pLKO.1 scramble shRNA | Xian et al., 2018 | N/A | |
| Transfected construct (Mus musculus) | pLKO.1 shNHE6 | Open Biosystem Xian et al., 2018 | TRCN0000068828 | Refer to shNHE6-a |
| Transfected construct (Mus musculus) | psPAX2 | Addgene | 12260 | Plasmid was a gift from Didier Trono |
| Transfected construct (Mus musculus) | pMD2.G | Addgene | 12259 | Plasmid was a gift from Didier Trono |
| Transfected construct (Mus musculus) | pJB-NHE6 targeting vector | This study | N/A | Refer to Materials and methods section for detailed description |
| Recombinant DNA reagent | pJB1 (plasmid) | Braybrooke et al., 2000 | N/A | |
| Recombinant DNA reagent | pCR4-TOPO (plasmid) | Thermo Fisher | K457502 | |
| Recombinant DNA reagent | pLVCMVfull (plasmid) | Xian et al., 2018 | N/A | |
| Recombinant DNA reagent | pME (plasmid) | Stawicki et al., 2014 | Addgene #73794 | Plasmid was a gift from David Raible |
| Recombinant DNA reagent | pLVCMV Vamp3pHluorin2 (plasmid) | This study | N/A | Refer to Materials and methods section for detailed description |
| Recombinant DNA reagent | BAC containing murine NHE6 sequence (bacterial artificial chromosome) | BACPAC Resources Center | RP23 364 F14 | |
| Software, algorithm | Adobe Creative Cloud | Adobe | RRID:SCR_010279 | |
| Software, algorithm | GraphPad Prism 7.0 | GraphPad Software | RRID:SCR_002798 | |
| Software, algorithm | Fiji/ImageJ | NIH | RRID:SCR_002285 | |
| Software, algorithm | LabView7.0 | National Instruments | RRID:SCR_014325 | |
| Software, algorithm | NDP.view2 | Hamamatsu Photonics | ||
| Software, algorithm | Odyssey Imaging System | LI-COR | RRID:SCR_014579 | |
| Software, algorithm | Clustal Omega | EMBL-EBI | RRID:SCR_001591 | |
| Software, algorithm | Leica TCS SPE | Leica | RRID:SCR_002140 | |
| Sequence-based reagent | SA forward | IDT | GGATCCGTGT GTGTGTTGGG GGAGGGA | |
| Sequence-based reagent | SA reverse | Integrated DNA Technology | CTCGAGCTCAC AATCAGCCCTTT AAATATGCC | |
| Sequence-based reagent | GAP repair US forward | Integrated DNA Technology | AAGCTTGCGGCC GCTTCAATTTCTG TCCTTGCTACTG | |
| Sequence-based reagent | GAP repair US reverse | Integrated DNA Technology | AGATCTCAAGAA AGTTAGCTAGA AGTGTGTC | |
| Sequence-based reagent | GAP repair DS forward | Integrated DNA Technology | AGATCTGTAGA GGATGTGGGA AAGAGAG | |
| Sequence-based reagent | GAP repair DS reverse | Integrated DNA Technology | GTCGACGCGG CCGACACACA CAGATAAATAA CCTCAAAAG | |
| Sequence-based reagent | 5’ flanking 1st LoxP fragment forward | Integrated DNA Technology | GCTTCTCTCG AGCAAGAGTCAAC | |
| Sequence-based reagent | 5’ flanking 1st LoxP fragment reverse | Integrated DNA Technology | GATATCAGCA GGTACCACCAA GATCTCAACCT TATTGTCCTATA TGCACAAAC | |
| Sequence-based reagent | 3’ flanking 1st LoxP fragment forward | Integrated DNA Technology | GTCTTGTTGGTA CCTGATGAAATG GACTACCTCCACTTG | |
| Sequence-based reagent | 3’ flanking 1st LoxP fragment reverse | Integrated DNA Technology | ATCGATCTTCA TAACCCATCTGGATA | |
| Sequence-based reagent | LoxP Oligo forward | Integrated DNA Technology | GATCTGCTCAGC ATAACTTCGTATAG CATACATTATACG AAGTTATGGTAC | |
| Sequence-based reagent | LoxP Oligo reverse | Integrated DNA Technology | CATAACTTCGTA TAATGTATGCTAT ACGAAGTTATGC TGAGCAGATC | |
| Sequence-based reagent | Genotyping NHE6-floxed and wt forward | Integrated DNA Technology | GAGGAAGC AAAGTGTCA GCTCC | |
| Sequence-based reagent | Genotyping NHE6-floxed and wt reverse | Integrated DNA Technology | CTAATCCCCTC GGATGCTGCTC | |
| Sequence-based reagent | Genotyping NHE6-KO forward | Integrated DNA Technology | GAGGAAGC AAAGTGTCA GCTCC | |
| Sequence-based reagent | Genotyping NHE6-KO reverse | Integrated DNA Technology | CCTCACAAGACT AGAGAAATGGTTC | |
| Sequence-based reagent | Vamp3 forward | Integrated DNA Technology | TTCAAGCTTCAC CATGTCTACAGG TGTGCCTTCGGGGTC | |
| Sequence-based reagent | Vamp3 reverse | Integrated DNA Technology | CATTGTCATCAT CATCATCGTGTG GTGTGTCTCTAA GCTGAGCAACAG CGCCGTGGACGGC ACCGCCGGCCCCG GCAGCATCGCCAC CAAGCTTAAC | |
| Sequence-based reagent | pHluorin2 forward | Integrated DNA Technology | CCGGTCCCAAGCTT ATGGTGAGCAAGG GCGAGGAGCTGTTC | |
| Sequence-based reagent | pHluorin2 reverse | Integrated DNA Technology | GCCCTCTTCTAGAG AATTCACTTGTACAG CTCGTCCATGCCGTG |