The interplay of RNA:DNA hybrid structure and G-quadruplexes determines the outcome of R-loop-replisome collisions
Figures
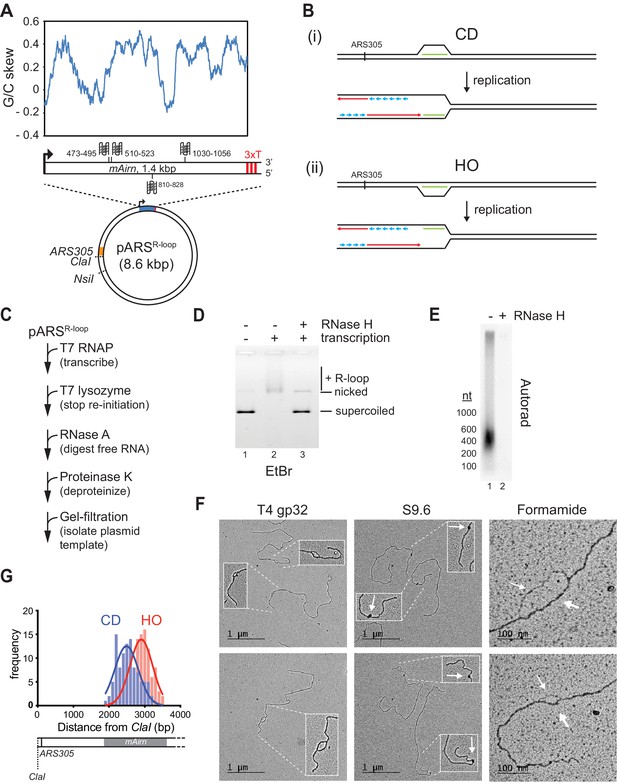
Preparation and characterization of R-loop-containing templates.
(A) Schematic of R-loop plasmid template. Plot shows G/C skew at Airn sequence. Graphic shows positions of potential G-quadruplexes composed of stacks of three G-quartets in non-template (top) and template (bottom) strand. 3× T: T7 terminator tandem repeat. (B) Schematic of co-directional (CD) and head-on (HO) R-loop-replisome collisions in experimental setup. Template strands: black; leading strand: red; lagging strand: blue; RNA: green. (C) Reaction scheme for preparation of R-loop-containing template. (D) Native agarose gel analysis of purified plasmid template. The gel was stained with ethidium bromide. (E) R-loop-containing template harboring 32P-labeled RNA was mock-treated or digested with RNase H and analyzed by denaturing formaldehyde agarose gel-electrophoresis and autoradiography. (F) Electron microscopy (EM) analysis of R-loop templates. White arrows in center panels indicate S9.6-specific density. Thin and thick arrows in right panels indicate displaced non-template or RNA:DNA duplex, respectively. (G) Frequency distribution of R-loop distances from ClaI site in CD and HO orientation.
-
Figure 1—source data 1
Preparation and characterization of R-loop-containing templates.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig1-data1-v2.xlsx
-
Figure 1—source data 2
Preparation and characterization of R-loop-containing templates.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig1-data2-v2.pdf
-
Figure 1—source data 3
Preparation and characterization of R-loop-containing templates.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig1-data3-v2.pdf
-
Figure 1—source data 4
Preparation and characterization of R-loop-containing templates.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig1-data4-v2.pzfx
-
Figure 1—source data 5
Preparation and characterization of R-loop-containing templates.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig1-data5-v2.pzfx
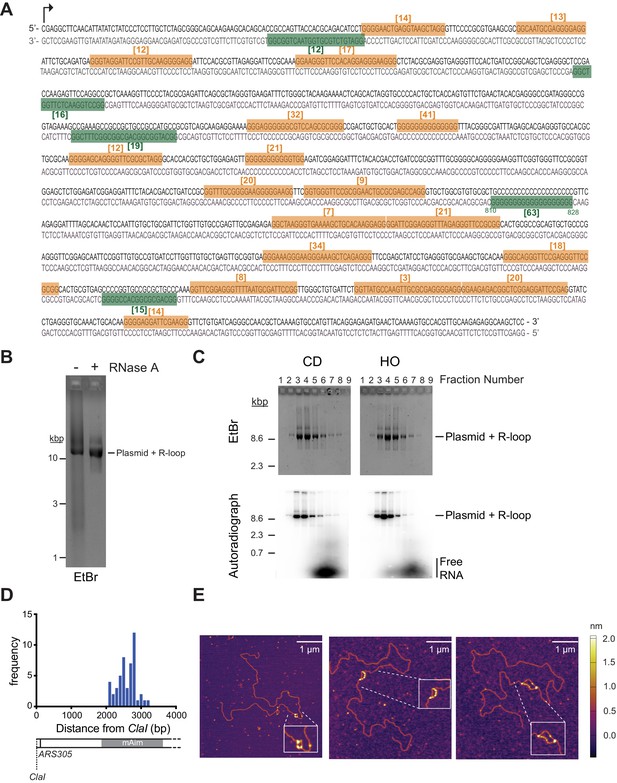
R-loop template preparation and characterization by atomic force microscopy (AFM).
(A) Sequence of Airn sequence element. Top strand: non-template strand; bottom strand: template strand. QGRS Mapper (https://bioinformatics.ramapo.edu/QGRS/index.php) was used to identify sequences with G-quadruplex-forming potential in non-template and template strand, highlighted in orange and green, respectively. Numbers in brackets indicate G-quadruplex-forming potential. (B) pARSR-loop was transcribed, the salt adjusted to 0.4 M NaCl, and either mock-treated or treated with RNase A. Reactions were de-proteinated, phenol/chloroform-extracted, filtered through Illustra MicroSpin G25 Spin column, and 2 μL of each sample analyzed by native agarose gel-electrophoresis and ethidium bromide staining. (C) S1000 gel filtration profiles of R-loop plasmid templates. RNA is 32P-labeled. Fractions were analyzed by ethidium bromide staining (top) or autoradiography (bottom) after native agarose gel-electrophoresis. (D) Frequency distribution of location of S9.6-dependent electron-dense structures relative to ClaI site in CD orientation. (E) Representative AFM images of linearized R-loop templates. Templates were incubated with yeast RPA prior to deposition on mica.
-
Figure 1—figure supplement 1—source data 1
R-loop template preparation and characterization by atomic force microscopy (AFM).
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig1-figsupp1-data1-v2.pdf
-
Figure 1—figure supplement 1—source data 2
R-loop template preparation and characterization by atomic force microscopy (AFM).
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig1-figsupp1-data2-v2.pdf
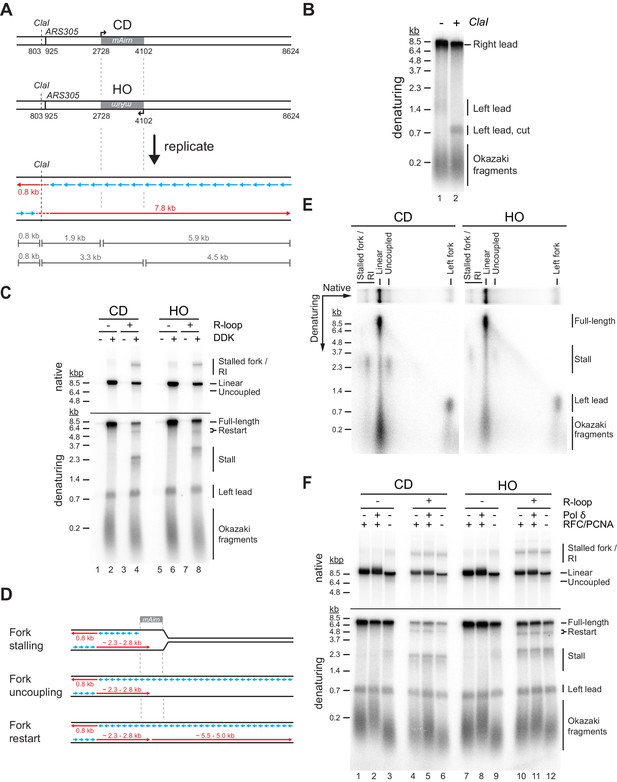
Both co-directional (CD) and head-on (HO) R-loops perturb normal fork progression.
(A) Schematic illustrating expected sizes of replication products. (B) Denaturing agarose gel analysis of replication products obtained on R-loop-free template. Left lead: Leftward leading strands; Right lead: Rightward leading strands. (C) Native (top) and denaturing (bottom) agarose gel analyses of replication products obtained on templates harboring Airn sequence in CD or HO orientation. Stall: Stalled rightward leading strands; Restart: Rightward leading strand restart product; Full-length: full-length rightward leading strand; RI: replication intermediates. (D) Schematic illustrating replication products observed in (C). (E) Two-dimensional gel analysis of replication products obtained in presence of R-loops (corresponding to lanes 4 and 8 in (C)). Products were digested with ClaI. (F) Replication products obtained in the absence or presence of RFC/PCNA or Pol δ.
-
Figure 2—source data 1
Both co-directional (CD) and head-on (HO) R-loops perturb normal fork progression.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig2-data1-v2.pdf
-
Figure 2—source data 2
Both co-directional (CD) and head-on (HO) R-loops perturb normal fork progression.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig2-data2-v2.pdf
-
Figure 2—source data 3
Both co-directional (CD) and head-on (HO) R-loops perturb normal fork progression.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig2-data3-v2.pdf
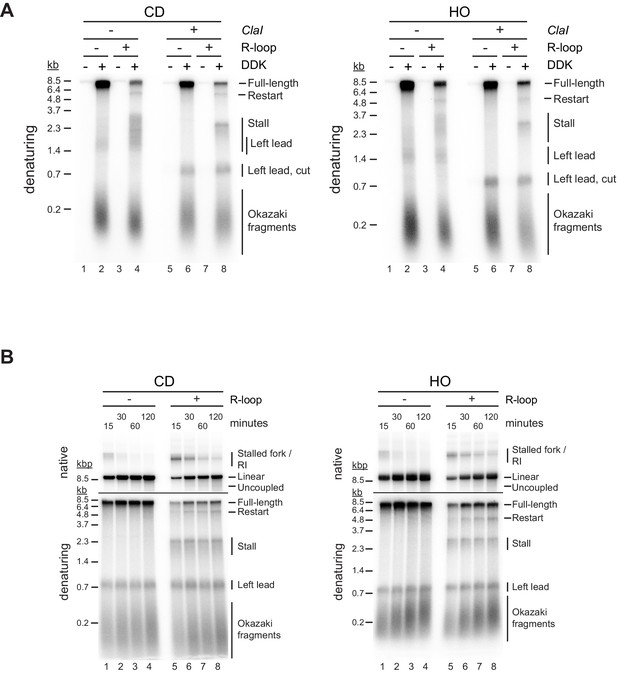
Both co-directional (CD) and head-on (HO) R-loops perturb normal fork progression.
(A) Replication products obtained on CD and HO templates with and without ClaI digestion post replication. (B) Conventional time course analyses of replication reactions on CD and HO templates. Time indicates minutes after addition of Mcm10 (origin firing).
-
Figure 2—figure supplement 1—source data 1
Characterization of replisome encounters with co-directional (CD) and head-on (HO) R-loops.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig2-figsupp1-data1-v2.pdf
-
Figure 2—figure supplement 1—source data 2
Characterization of replisome encounters with co-directional (CD) and head-on (HO) R-loops.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig2-figsupp1-data2-v2.pdf
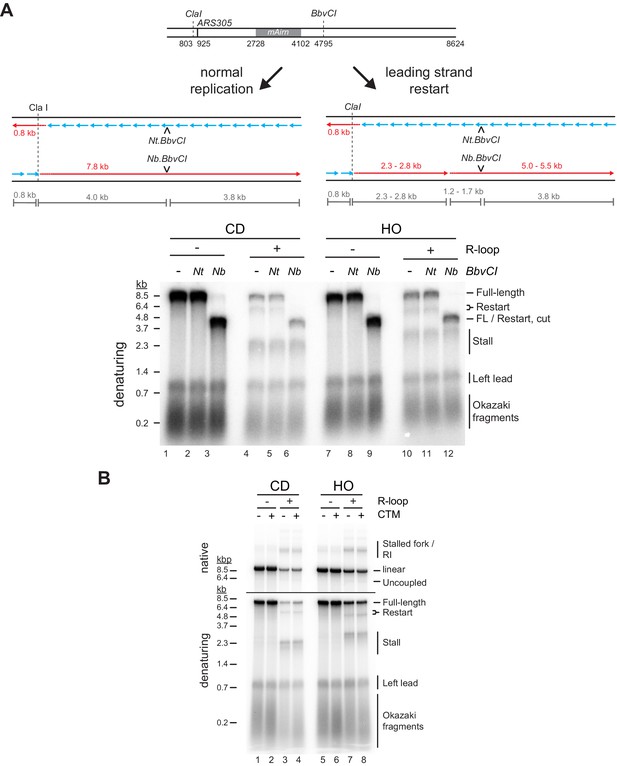
Characterization of replisome encounters with co-directional (CD) and head-on (HO) R-loops.
(A) Top: Schematic of products expected after digestion of full-length or restart replication products with Nt.BbvCI or Nb.BbvCI. Bottom: Denaturing gel analysis of replication products digested as indicated. (B) Replication products obtained on CD and HO templates in the absence or presence of Csm3-Tof1 and Mrc1 (CTM).
-
Figure 2—figure supplement 2—source data 1
Characterization of replisome encounters with co-directional (CD) and head-on (HO) R-loops.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig2-figsupp2-data1-v2.pdf
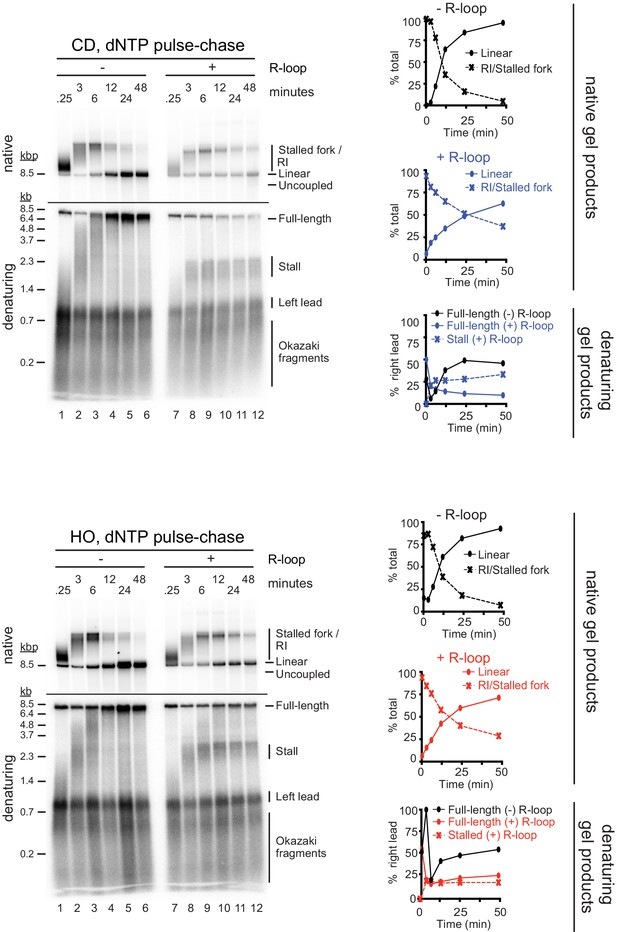
dNTP pulse-chase analysis of fork progression through co-directional (CD) and head-on (HO) R-loops.
Pulse-chase time course analysis of replication reactions on CD (top) and HO (bottom) templates. Signal intensities of replication products were quantified and plotted as percentage of total signal.
-
Figure 3—source data 1
dNTP pulse-chase analysis of fork progression through co-directional (CD) and head-on (HO) R-loops.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig3-data1-v2.pdf
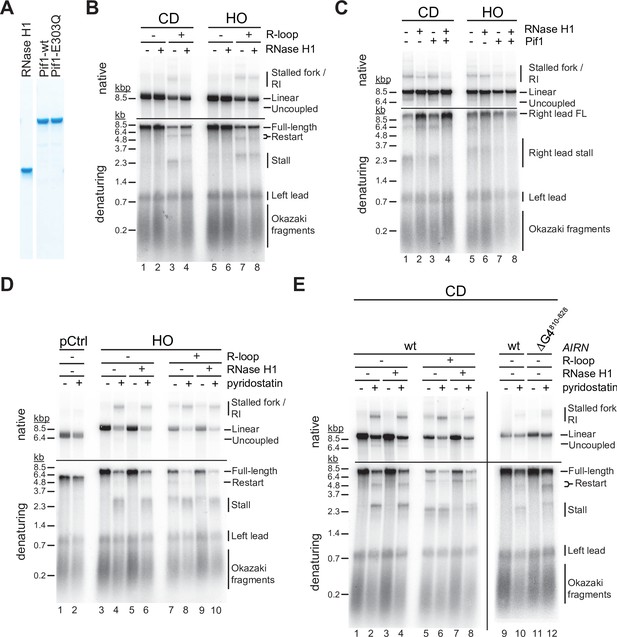
G-quadruplexes (G4s) and RNA:DNA hybrids pose impediment to leading strand synthesis that can be resolved by Pif1 or RNase H1, respectively.
(A) Purified proteins analyzed by SDS-PAGE and Coomassie stain. (B) Replication products obtained on co-directional (CD) or head-on (HO) templates in the absence or presence of RNase H1. (C) Replication products obtained on CD or HO templates in the absence or presence of RNase H1 and Pif1. (D) Replication products obtained on HO templates in the absence or presence of RNase H1 and pyridostatin. (E) Replication products obtained on CD templates in the absence or presence of RNase H1 and pyridostatin. ΔG4810-828: Airn sequence containing deletion of G4 sequence at position 810–828.
-
Figure 4—source data 1
G-quadruplexes (G4s) and RNA:DNA hybrids pose impediment to leading strand synthesis that can be resolved by Pif1 or RNase H1, respectively.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig4-data1-v2.pdf
-
Figure 4—source data 2
G-quadruplexes (G4s) and RNA:DNA hybrids pose impediment to leading strand synthesis that can be resolved by Pif1 or RNase H1, respectively.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig4-data2-v2.pdf
-
Figure 4—source data 3
G-quadruplexes (G4s) and RNA:DNA hybrids pose impediment to leading strand synthesis that can be resolved by Pif1 or RNase H1, respectively.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig4-data3-v2.pdf
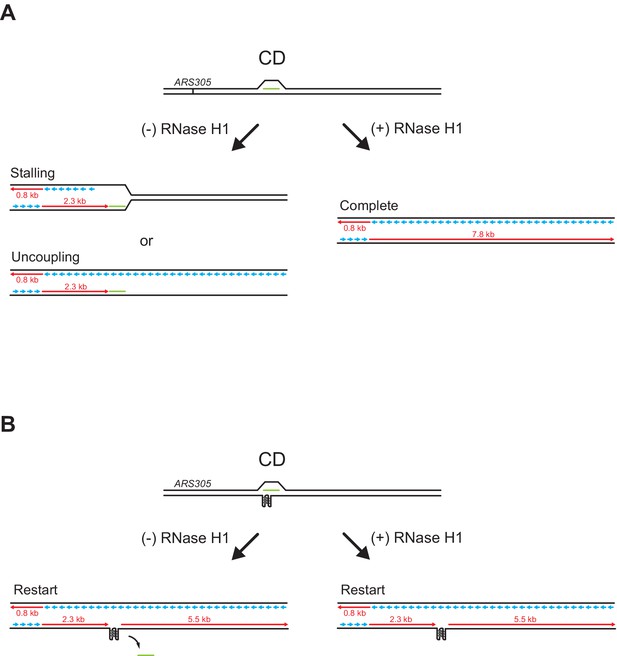
RNase H1 promotes fork passage specifically at co-directional (CD) R-loops.
(A) Schematic of replication products observed after replisome encounters with CD R-loops lacking G-quadruplexes (G4s) in the leading strand template. Treatment with RNase H1 removes the RNA:DNA hybrid, allowing completion of DNA replication. (B) Schematic of replication products observed after replisome bypass of CD R-loops harboring G4s in the leading strand template. Bypass of CD R-loops or fork passage in the presence of RNase H1 results in the uncoupling of leading strand synthesis at G4s, which promotes restart of leading strand synthesis by repriming.
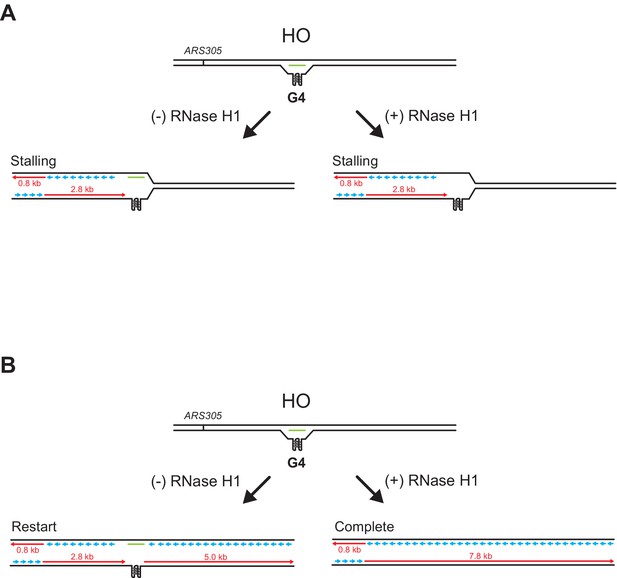
RNase H1 promotes fork passage specifically at co-directional (CD) R-loops.
(A) Schematic of replication products observed after replisome encounters with head-on (HO) R-loops harboring G-quadruplexes (G4s) in the displaced strand that block CMG progression. Stalling persists upon RNase H1 treatment. (B) Schematic of replication products observed after replisome encounters with HO R-loops harboring G4s in the displaced strand that are stabilized in single-stranded DNA, that is, when an RNA:DNA hybrid is present on the template strand, and that block leading strand polymerase but not CMG progression.
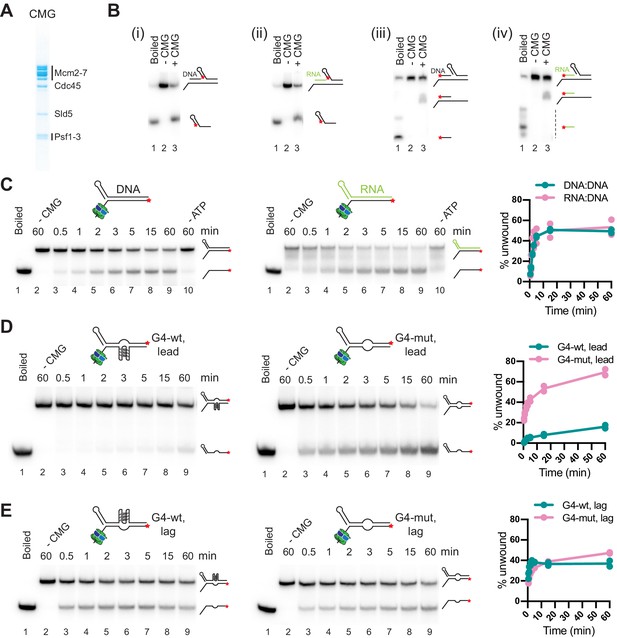
CMG can unwind or translocate on RNA:DNA hybrids, while G-quadruplexes (G4s) can block DNA unwinding by CMG.
(A) Purified CMG. (B) Helicase assays with 40 bp forked DNA duplex preceded by 40 bp DNA (i + iii) or RNA:DNA (ii + iv) duplex. ★ indicates position of 5’-32 P label. Products were analyzed by native PAGE and autoradiography. (C) CMG helicase activity on 60 bp forked DNA (left) or RNA:DNA duplex (right). Plot shows average of two replicates. (D) CMG helicase activity on 60 bp substrate harboring wildtype (left) or mutant (right) G4 sequence on the template strand (‘lead’). (E) As (D), with wildtype (left) or mutant (right) G4 sequence on the non-template strand (‘lag’).
-
Figure 5—source data 1
CMG can unwind or translocate on RNA:DNA hybrids, while G-quadruplexes (G4s) can block DNA unwinding by CMG.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig5-data1-v2.pdf
-
Figure 5—source data 2
CMG can unwind or translocate on RNA:DNA hybrids, while G-quadruplexes (G4s) can block DNA unwinding by CMG.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig5-data2-v2.pdf
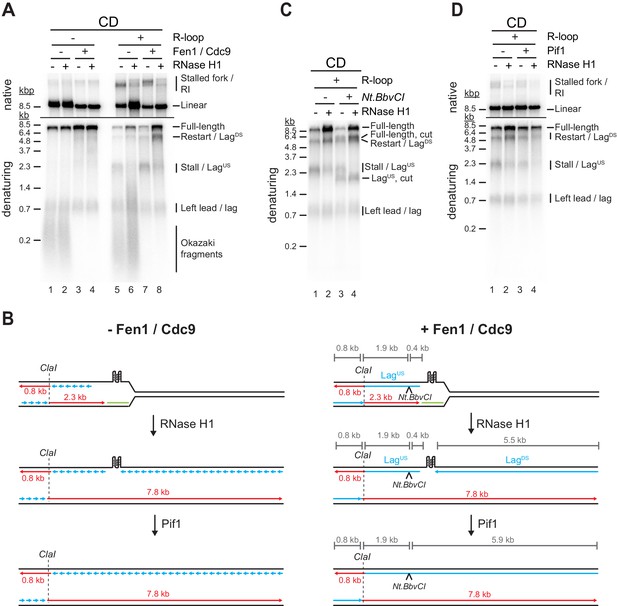
G-quadruplexes (G4s) at co-directional (CD) R-loops can induce lagging strand gaps that can be resolved by Pif1.
(A) Replication products obtained on CD template in the absence or presence of Fen1/Cdc9 and RNase H1. (B) Schematic illustrating replication products observed in (A). (C) Replication products obtained on CD template in the presence of Fen1/Cdc9. RNase H1 was included and products were digested with Nt.BbvCI as indicated. (D) Replication products obtained on CD template in the presence of Fen1/Cdc9. Pif1 and RNase H1 were included as indicated.
-
Figure 6—source data 1
G-quadruplexes (G4s) at co-directional (CD) R-loops can induce lagging strand gaps that can be resolved by Pif1.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig6-data1-v2.pdf
-
Figure 6—source data 2
G-quadruplexes (G4s) at co-directional (CD) R-loops can induce lagging strand gaps that can be resolved by Pif1.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig6-data2-v2.pdf
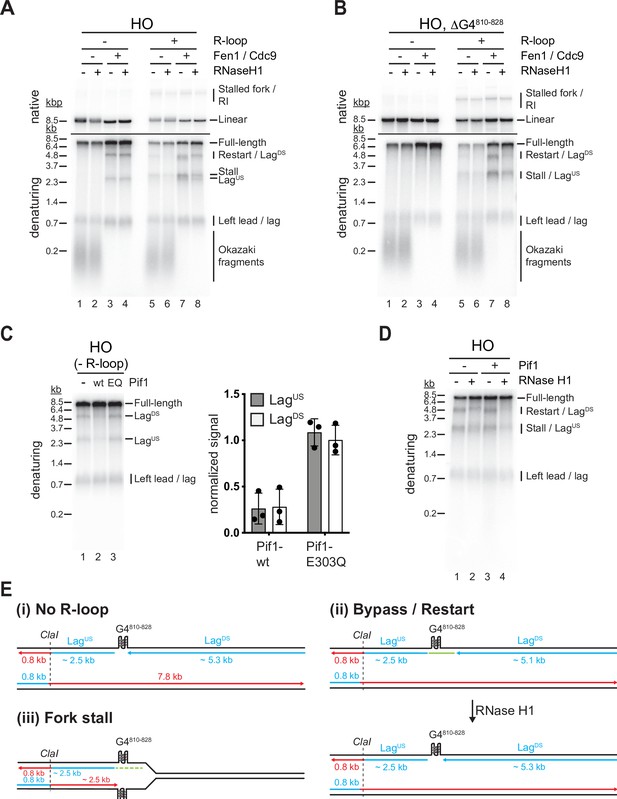
Both G-quadruplexes (G4s) and RNA:DNA hybrids cause lagging strand gaps at head-on (HO) R-loops.
(A) Replication products obtained on HO template in the absence or presence of Fen1/Cdc9 and RNase H1. (B) Replication products obtained on HO template harboring ΔG4810-828 deletion in the absence or presence of Fen1/Cdc9 and RNase H1. (C) HO template lacking R-loop was replicated with Fen1/Cdc9 in the absence or presence of Pif1 or Pif1-E303Q. Signal intensities of LagUS and LagDS were quantified and normalized to reactions without Pif1. (D) HO template replicated with Fen1/Cdc9 in the absence or presence of Pif1. (E) Schematic illustrating replication products observed in A-D.
-
Figure 7—source data 1
Both G-quadruplexes (G4s) and RNA:DNA hybrids cause lagging strand gaps at head-on (HO) R-loops.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig7-data1-v2.pdf
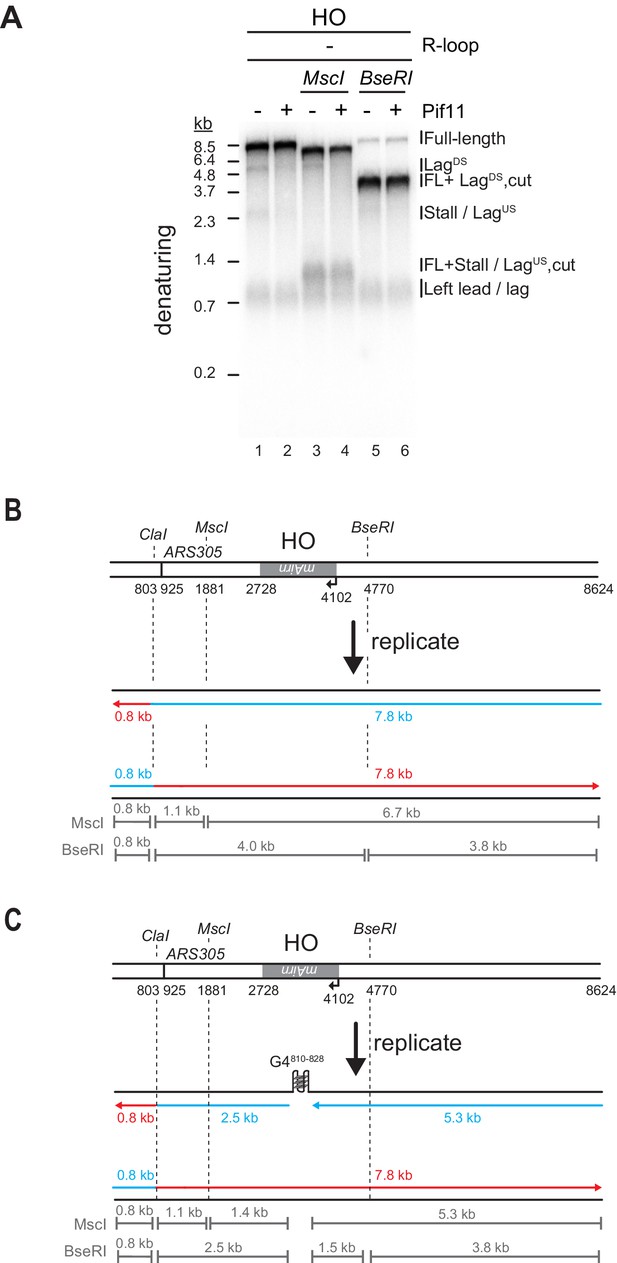
Both G-quadruplexes (G4s) and RNA:DNA hybrids cause lagging strand gaps at head-on (HO) R-loops.
(A) Denaturing gel analysis of replication products obtained in the presence of Fen1/Cdc9 on HO templates lacking R-loops were digested with ClaI (lanes 1 + 2), ClaI and MscI (lanes 3 + 4), or ClaI and BseRI (lanes 5 + 6). LagUS and LagDS are sensitive to MscI and BseRI, respectively. (B) Schematic of expected products generated by digestion of full-length replication products. (C) Schematic of expected products generated by digestion of replication products containing a lagging strand gap.
-
Figure 7—figure supplement 1—source data 1
Both G-quadruplexes (G4s) and RNA:DNA hybrids cause lagging strand gaps at head-on (HO) R-loops.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig7-figsupp1-data1-v2.pdf
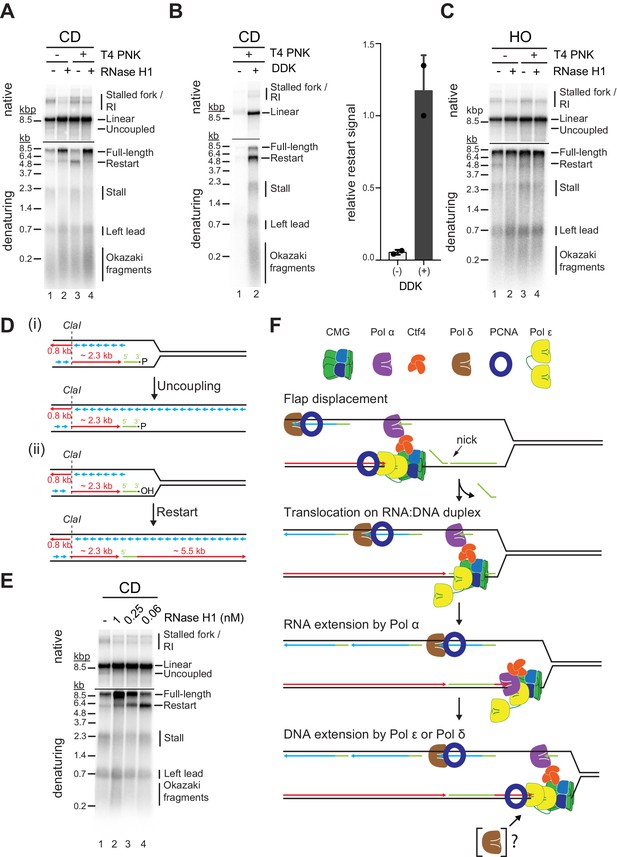
R-loop transcripts can prime leading strand restart after co-directional (CD) R-loop-replisome collisions.
(A) Replication products obtained on mock- or T4 polynucleotide kinase (PNK)-treated CD R-loop-containing templates in absence or presence RNase H1. (B) Products obtained on CD T4 PNK-treated R-loop templates in absence or presence of DDK. Relative signal intensity for restart product is quantified on the right. (C) Replication products obtained on mock- or T4 PNK-treated CD R-loop-containing templates in absence or presence RNase H1. (D) Schematic illustrating replication products observed in A. (E) RNase H1 titration into reactions with CD R-loop template. (F) Model for leading strand restart at R-loop transcript after replisome encounter with CD R-loop harboring 5’ RNA flap and RNA nick.
-
Figure 8—source data 1
R-loop transcripts can prime leading strand restart after co-directional (CD) R-loop-replisome collisions.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig8-data1-v2.pdf
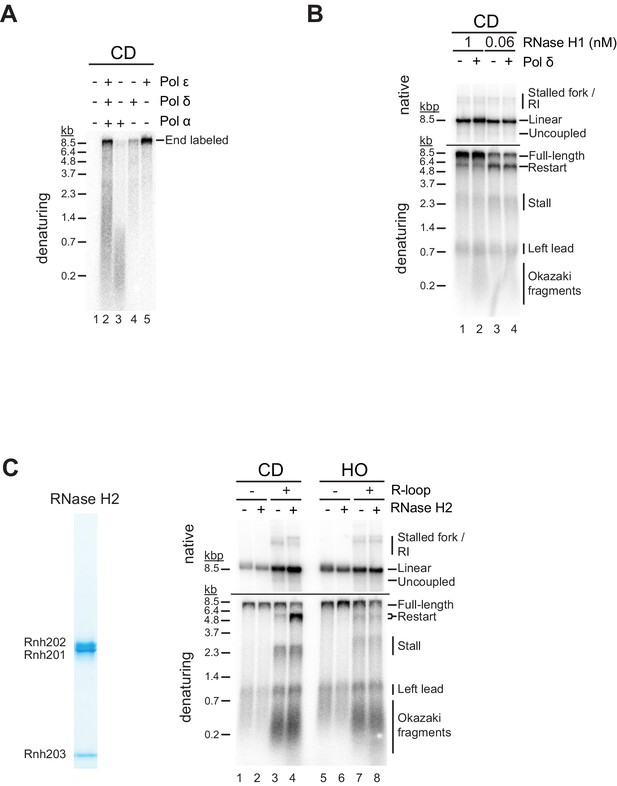
R-loop transcripts can prime leading strand restart after co-directional (CD) R-loop-replisome collisions.
(A) Polymerase assay with T4 polynucleotide kinase (PNK)-treated CD R-loop template in presence of Pol α, Pol ε, or Pol δ, as indicated. All reactions include RFC, PCNA, RPA, dNTPs, NTPs, and α-[32P]-dATP. The concentrations of reaction components are equivalent to those used in the replication assay. (B) Replication products obtained on RNase H1-treated CD R-loop templates with or without Pol δ. Reactions were performed either in the presence of 1 nM (lanes 1 + 2) or 0.06 nM (lanes 3 + 4) RNase H1 to induce resolution or nicking of the RNA:DNA hybrid, respectively. (C) Left: Purified RNase H2. Right: Replication products obtained on CD and head-on (HO) templates in the presence of sub-saturating levels of RNase H2.
-
Figure 8—figure supplement 1—source data 1
R-loop transcripts can prime leading strand restart after co-directional (CD) R-loop-replisome collisions.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig8-figsupp1-data1-v2.pdf
-
Figure 8—figure supplement 1—source data 2
R-loop transcripts can prime leading strand restart after co-directional (CD) R-loop-replisome collisions.
- https://cdn.elifesciences.org/articles/72286/elife-72286-fig8-figsupp1-data2-v2.pdf
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Saccharomyces cerevisiae) | YDR35 | This study | See Table 1 for genotype | Overexpression and purification of CMG |
| Strain, strain background (Saccharomyces cerevisiae) | YDR154 | This study | See Table 1 for genotype | Overexpression and purification of RNase H2 |
| Strain, strain background (Saccharomyces cerevisiae) | ySD-ORC | PMID:23474987 | Overexpression and purification of ORC | |
| Strain, strain background (Saccharomyces cerevisiae) | YJF38 | PMID:23474987 | Overexpression and purification of Cdt1⋅Mcm2-7 | |
| Strain, strain background (Saccharomyces cerevisiae) | YSA35 | PMID:32701054 | Overexpression and purification of DDK | |
| Strain, strain background (Saccharomyces cerevisiae) | YSD15 | PMID:27989437 | Overexpression and purification of Cdc45 | |
| Strain, strain background (Saccharomyces cerevisiae) | YDR105 | PIMD:27989437 | Overexpression and purification of Clb5⋅Cdk1 | |
| Strain, strain background (Saccharomyces cerevisiae) | YDR109 | PIMD:27989437 | Overexpression and purification of GINS | |
| Strain, strain background (Saccharomyces cerevisiae) | YSD116 | PIMD:27989437 | Overexpression and purification of Pol ε | |
| Strain, strain background (Saccharomyces cerevisiae) | YDR110 | PIMD:27989437 | Overexpression and purification of Dpb11 | |
| Strain, strain background (Saccharomyces cerevisiae) | YSD13 | PIMD:27989437 | Overexpression and purification of Sld2 | |
| Strain, strain background (Saccharomyces cerevisiae) | YSD16 | PIMD:27989437 | Overexpression and purification of Pol α | |
| Strain, strain background (Saccharomyces cerevisiae) | YIW389 | PIMD:27989437 | Overexpression and purification of RFC | |
| Strain, strain background (Saccharomyces cerevisiae) | YDR131 | PIMD:27989437 | Overexpression and purification of Pol δ | |
| Strain, strain background (Saccharomyces cerevisiae) | YDR128 | PIMD:27989437 | Overexpression and purification of Top1 | |
| Strain, strain background (Saccharomyces cerevisiae) | YDR163 | PIMD:32341532 | Overexpression and purification of Mrc1 | |
| Strain, strain background (Saccharomyces cerevisiae) | YDR137 | PIMD:32341532 | Overexpression and purification of Csm3-Tof1 | |
| Antibody | S9.6(mouse monoclonal) | Sigma Millipore | MABE1095 | (1 μg per 1 μg) of DNA |
| Recombinant DNA reagent | pFC57 | PMID:22387027 | AIRN -containing plasmid | |
| Recombinant DNA reagent | p470 | PMID:27989437 | ARS305 -containing plasmid | |
| Recombinant DNA reagent | p1216 | This paper | p470 with secondary origin sequences deleted, without AIRN | |
| Recombinant DNA reagent | P1214 | This paper | p1216 with AIRN in CD orientation | |
| Recombinant DNA reagent | p1215 | This paper | p1216 with AIRN in HO orientation | |
| Recombinant DNA reagent | p1290 | This paper | p1214 with BbvCI site 2075 bp downstream of AIRN | |
| Recombinant DNA reagent | p1285 | This paper | p1215 with BbvCI site 2075 bp downstream of AIRN | |
| Recombinant DNA reagent | p1291 | This paper | p1214 with BbvCI site 11 bp upstream of AIRN | |
| Recombinant DNA reagent | p1292 | This paper | p1215 with BbvCI site 11 bp upstream of AIRN | |
| Recombinant DNA reagent | p1293 | This paper | CD, AIRNΔ810–828 | |
| Recombinant DNA reagent | p1284 | This paper | HO, AIRNΔ810–828 | |
| Recombinant DNA reagent | p1134 | This paper | pET15b-6x-His-RNH1 | Overexpression of His-tagged Rnh1 |
| Recombinant DNA reagent | p1254 | This paper | pET15b-T7 Lysozyme-6x-His | Overexpression of C-terminally His-tagged T7 lysozyme |
| Recombinant DNA reagent | p1102 | This paper | pET15b-PIF1-N | Overexpression of His-tagged Pif1-N |
| Recombinant DNA reagent | p1130 | This paper | pET15b-PIF1-N-E303Q | Overexpression of His-tagged Pif1-N-E303Q |
| Recombinant DNA reagent | pET15b-His-CDC6 | PMID:24566988 | Overexpression of His-tagged Cdc6 | |
| Recombinant DNA reagent | p399 | PIMD:27989437 | pSmt3-SLD3 | Overexpression of His-tagged Sld3 |
| Recombinant DNA reagent | P468 | PIMD:27989437 | pGEX-6P-1-SLD7 | Overexpression of GST-tagged - Sld7 |
| Recombinant DNA reagent | pJM126 | PIMD:27989437 | Overexpression of RPA | |
| Recombinant DNA reagent | pCK6 | PIMD:27989437 | pET15b-His-CTF4 | Overexpression of His-tagged Ctf4 |
| Recombinant DNA reagent | p1012 | PIMD:27989437 | pET28a-PCNA | Overexpression of PCNA |
| Recombinant DNA reagent | p619 | PIMD:27989437 | pET15b-His-MCM10 | Overexpression of His-tagged Mcm10 |
| Recombinant DNA reagent | p1137 | PMID:32341532 | pET28a-FEN1-6xHis | Overexpression of C-terminally His-tagged Fen1 |
| Recombinant DNA reagent | p1019 | PMID:32341532 | pET28a-CDC9 | Overexpression of His-tagged Cdc9 |
| Sequence-based reagent | DNA oligo-nucleotides | IDT | (SeeTable 2 Table 3) | To generate templates for helicase assays |
| Chemical compound, drug | Pyridostatin | Sigma Aldrich | SML2690 | |
| Chemical compound, drug | ATP | Thermo Scientific | R1441 | |
| Chemical compound, drug | Formamide | ThermoFisherScientific | 17899 | |
| Chemical compound, drug | Benzalkonium chloride | MilliporeSigma | B6285 | |
| Peptide, recombinant protein | RNase H | New England Biolabs | M0297S | |
| Peptide, recombinant protein | T7 RNA polymerase | Promega | P2075 | |
| Peptide, recombinant protein | RNase A | ThermoFisher Scientific | EN0531 | |
| Peptide, recombinant protein | Proteinase K | New England Biolabs | P8107S | |
| Peptide, recombinant protein | T4 polynucleotide kinase | New England Biolabs | M0201S | |
| Software, algorithm | GraphPad Prism | GraphPad Prism (https://graphpad.com) | RRID:SCR_015807 | Statistical analysis and data plotting |
| Software, algorithm | ImageJ | ImageJ(https://imagej.nih.gov/ij/) | RRID:SCR_003070 | Image analysis |
| Software, algorithm | QGRS mapper | QGRS mapper (https://bioinformatics.ramapo.edu/QGRS/analyze.php) | Prediction of G4 sequences |
Yeast strains.
| Strain name | Protein | Genotype |
|---|---|---|
| YSD35 | CMG | MAT a/α ade2-1 ura3-1 his3-11,15 trp1-1 leu2-3,112 can1-100 pep4::kanMX bar1::hphNAT1 Gal-Gal4 (HIS3) Gal-Mcm5/Mcm4 (TRP1) Gal-Mcm2/FLAG-Mcm3 (URA3) Gal-Mcm7/Mcm6 (LEU2) Gal1-10 Cdc45 (TRP1) Gal1-10 Psf2++/Psf3++ (URA3) Gal1-10 Psf1++/CBP-Sld5++ (LEU2) |
| YDR154 | RNase H2 | MATa ade2-1 ura3-1 his3-11,15 trp1-1 leu2-3,112 can1-100 pep4::kanMX bar::hphNAT1 Gal-Gal4 (HIS3) GAL-CBP-RNH201++ (LEU2) GAL-RNH202++ / RNH203++ (URA3) |
List of oligonucleotides used to prepare templates for helicase assays.
| Name | Sequence 5’ to 3’ |
|---|---|
| A | GGCTCGTTTTACAACGTCGTGCTGAGGTGATATCTGCTGAGGCAATGGGAATTCGCCAACCTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT |
| B | GGCAGGCAGGCAGGCAGGCAGGCAGGCAGGCAGGCAGGCAGGTTGGCGAATTCCCATTGCCTCAGCAGATATCACCTCAGCACGACGTTGTAAAACGAG |
| C | GGCAGGCAGGCAGGCAGGCAGGCAGGCAGGCAGGCAGGCAGGUUGGCGAAUUCCCAUUGCCUCAGCAGAUAUCACCUCAGCACGACGUUGUAAAACGAG |
| D | GGCAGGCAGGCAGGCAGGCAGGCAGGCAGGCAGGCAGGCAGATTAAGTTGGGTAACGCCAGGGTTTTCCCAGTCACGAC |
| E | GTCGTGACTGGGAAAACCCTGGCGTTACCCAACTTAATCGCCTTGCAGCACATCCCCCTTTCGCCAGCTGGCGTAATAGTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT |
| F | CTATTACGCCAGCTGGCGAAAGGGGGATGTGCTGCAAGGC |
| G | CUAUUACGCCAGCUGGCGAAAGGGGGAUGUGCUGCAAGGC |
| H | GGCAGGCAGGCAGGCAGGCAGGCAGGCAGGCAGGCAGGCAGGTTGGCGAATTCCCTTTTTTTTTTTTTTTTTTTCCTCAGCACGACGTTGTAAAACGAG |
| I | GGCTCGTTTTACAACGTCGTGCTGAGGTTGGGTGGGTGGGTGGGTTGGGAATTCGCCAACCTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT |
| J | GGCTCGTTTTACAACGTCGTGCTGAGGTTGGGTGGGTCCCTGGGTTGGGAATTCGCCAACCTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT |
| K | GGCTCGTTTTACAACGTCGTGCTGAGGTTTTTTTTTTTTTTTTTTTGGGAATTCGCCAACCTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT |
| L | GGCAGGCAGGCAGGCAGGCAGGCAGGCAGGCAGGCAGGCAGGTTGGCGAATTCCCTTGGGTGGGTGGGTGGGTTCCTCAGCACGACGTTGTAAAACGAG |
| M | GGCAGGCAGGCAGGCAGGCAGGCAGGCAGGCAGGCAGGCAGGTTGGCGAATTCCCTTGGGTGGGTCCCTGGGTTCCTCAGCACGACGTTGTAAAACGAG |
List of templates used in helicase assays.
| Template | Oligonucleotides | Figure | 32P-labeled oligonucleotide |
|---|---|---|---|
| Forked DNA duplex | A + B | 4 C, left | A |
| Forked RNA-DNA duplex | A + C | 4 C, right | A |
| DNA-DNA 1 | D + E + F | 4B(i) | D |
| DNA-DNA 2 | D + E + F | 4B(iii) | F |
| RNA-DNA 1 | D + E + G | 4B(ii) | D |
| RNA-DNA 2 | D + E + G | 4B(iv) | G |
| G-quad wt, lead | H + I | 4D, left | H |
| G-quad mut, lead | H + J | 4D, right | H |
| G-quad wt, lag | K + L | 4E, left | K |
| G-quad mut, lag | K + M | 4E, right | K |





