Reconstitution of surface lipoprotein translocation through the Slam translocon
Figures
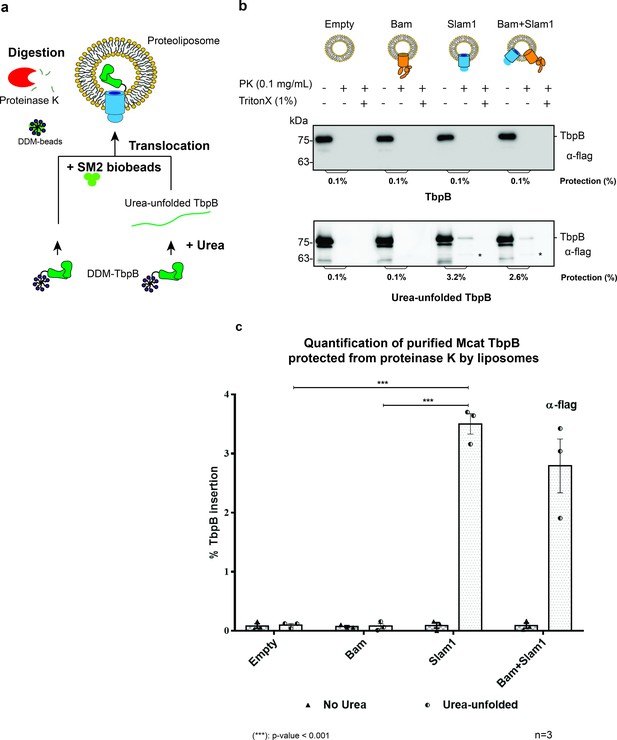
Slam1 is necessary for translocation of unfolded TbpB.
(a) Model of a defined in vitro assay for TbpB translocation. M. catarrhalis TbpB (folded and urea-unfolded) is translocated inside Slam1-containing proteoliposomes. SM2 biobeads were used to remove DDM detergent from TbpB before adding proteoliposomes for translocation. Efficiency of TbpB translocation/insertion was calculated based on percentage of TbpB that was protected from proteinase K. (b) Representative proteinase K protection assay results obtained for Slam1 or Slam1 + Bam incubated with purified TbpB (folded or kept unfolded by 8 M urea). Proteoliposomes containing Empty or Bam were used as controls. Each sample was treated with PK or PK + Triton X-100 and examined by western blot. α-flag antibody western blots were used to quantify the amount of TbpB. Asterisk (*) The lower band in the (+PK and −Triton X) samples in Slam1 and Bam + Slam1 proteoliposomes treatment likely represents incompletely translocated TbpB that has been partially degraded by proteinase K. (c) Quantification of TbpB protection in proteoliposomes through densitometry analysis. The % TbpB insertion was calculated by dividing the protected TbpB of + PK sample by TbpB of the input sample. The plot contains results obtained from three biological replicates. Individual data points were included on the graph. Two-way analysis of variance (ANOVA) test was performed to determine the statistical significance for the translocation of unfolded TbpB by Slam1 proteoliposomes and Slam1 + Bam proteoliposomes treatment versus by the negative controls (empty liposomes and Bam proteoliposomes). Only statistical significance of the unfolded TbpB translocation by Slam1 proteoliposomes against the two negative controls are included on the blot for simplification. (***) represents p-value < 0.001.
-
Figure 1—source data 1
Quantification of in vitro translocation of purified TbpB.
- https://cdn.elifesciences.org/articles/72822/elife-72822-fig1-data1-v2.zip
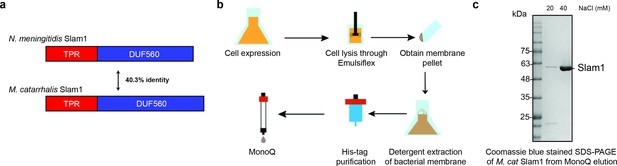
Purification of M. catarrhalis Slam1.
(a) Sequence identity of N. meningitidis Slam1 and M. catarrhalis Slam1. (b) Overall membrane protein expression and purification pipeline. (c) Sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) gels of pure MonoQ fractions from Mcat Slam1 purification. Pure Slam–DDM detergent complex eluted at 50 mM Tris–HCl pH 8, 40 mM NaCl from a MonoQ column. The proteins were used for the downstream functional assays. Proteins that were used for structural studies were passed through S200 gel filtration.
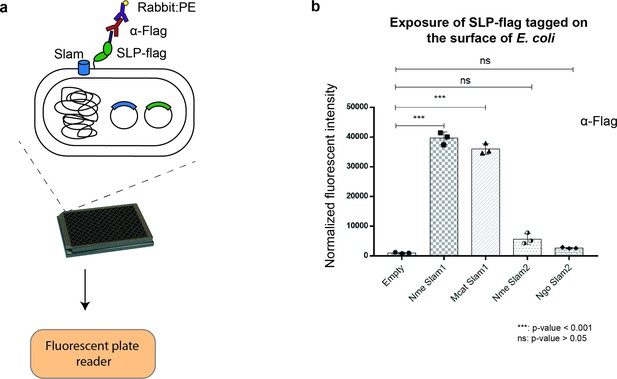
Translocation of Mcat TbpB to the surface of E. coli cells.
(a) Plate reader assay was used to examine the function of Slam homologs. Slams and TbpB were coexpressed in E. coli strain C43(DE3) and probed with α-flag antibodies followed by labeling with the secondary antibody conjugated with fluorescent probe phycoerythrin (PE). The fluorescence was quantified using a plate reader. (b) Quantification of surface display of Mcat TbpB by Slam1 and Slam2 homologs. Slam2 are negative controls as they have different substrate specificity (HpuA) for translocation. Normalized fluorescence values obtained for each of the Slam homologs is shown. The results represent at least three biological replicates and demonstrate that Mcat Slam1 is functional in translocating TbpB in the E. coli model. (***) represents p-value < 0.001 and (ns) represents p-value > 0.05.
-
Figure 1—figure supplement 2—source data 1
Fluorescent signal of TbpB on the surface of E.coli.
- https://cdn.elifesciences.org/articles/72822/elife-72822-fig1-figsupp2-data1-v2.zip
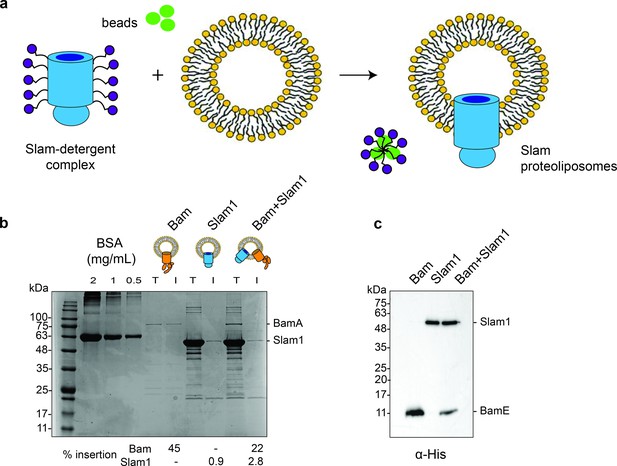
Generation of Slam1 and Bam proteoliposomes.
(a) Protocol used for insertion of outer membrane proteins (OMPs) into liposomes. OMP-DDM protein–detergent complexes were diluted (below the critical micellar concentration [CMC] of DDM) into preformed liposomes. Detergent was further removed using SM2 biobeads. (b) Quantification of Slam1 and BamABCDE insertion using Coomassie staining. Bovine Serum Albumin (BSA) was used as a control for estimating absolute protein quantity. Insertion percentages were calculated by dividing the band intensity of protein inserted in liposomes (l) by total protein incubated with liposomes (T). For the Bam complex, BamA intensity was used to calculate insertion efficiency. (c) Confirmation of Slam1 and BamABCDE insertion using western blots with α-His antibodies.
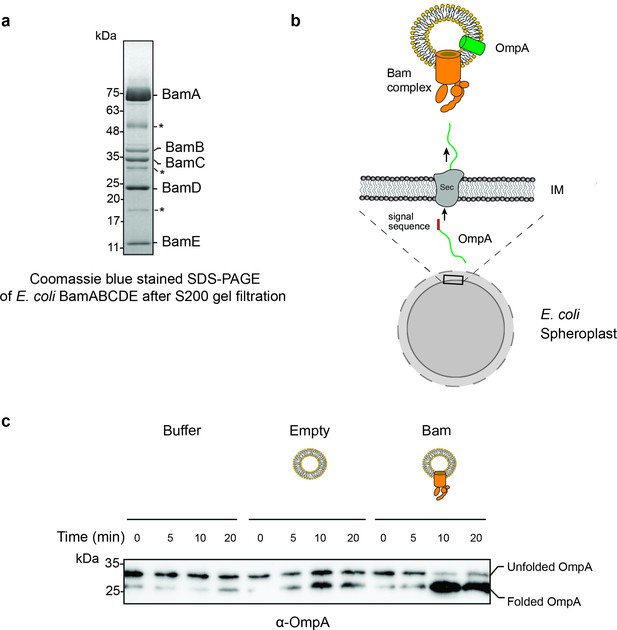
Purification and characterization of E. coli Bam complex.
(a) BamABCDE fractions obtained from a S200 gel filtration column. The BamABCDE complex was obtained using previously described protocols (Hagan et al., 2010). Some non-Bam complex bands (marked in asterisk) were observed, and they most likely correspond to common E. coli proteins that have been reported in previous His-tag purified proteins. (b) Design of an in vitro translocation assay for testing the function of the Bam complex. E. coli spheroplasts secrete porins such as OmpA into the supernatant. When incubated with Bam complex proteoliposomes, secreted OmpA is successfully inserted into Bam proteoliposomes. (c) An α-OmpA western blot to access the folding states of secreted OmpA over time in Tris pH 8 buffer, empty liposome, and Bam proteoliposomes. Approximately 95% of OmpA achieved folded form in the presence of Bam proteoliposomes within the first 10 min of incubation while self-folding in the empty liposome remained at 50%.
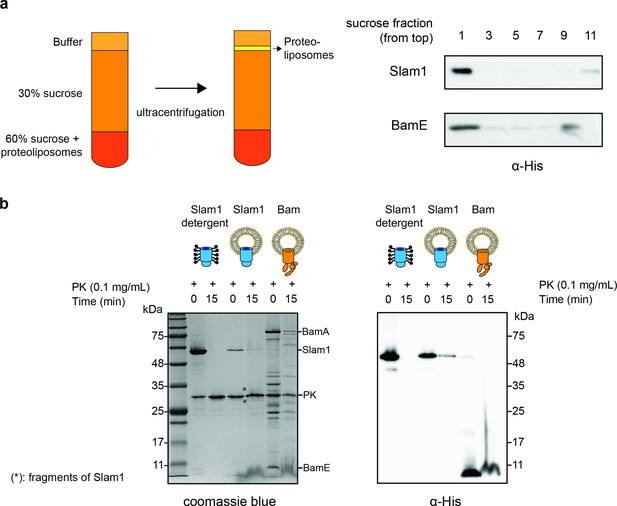
Characterization of Mcat Slam1 and BamABCDE containing proteoliposomes.
(a) Sucrose floatation assay for Slam1 and Bam proteoliposomes. Proteoliposomes were resuspended to a final concentration of 60% sucrose and subsequently layered with 30% sucrose and buffer B (50 mM Tris pH 7, 200 mM NaCl). Ten fractions were collected from the top and alternate fractions were run on an sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) gel. Western blots using α-His antibodies are shown indicating the amount of protein present in each fraction. (b) Proteinase K protection assay on Slam1 and Bam proteoliposomes. Proteoliposomes were incubated with 0.1 mg/ml proteinase K for 15 min at room temperature. Coomassie blue stained gel and α-His western blot were used to assess orientation of the proteins in liposomes. Approximately 18% of Slam1 inserted with N-terminal his-tag residing in the lumen of liposomes and was protected from PK digestion. Percentage protection was calculated using densitometry. Asterisk (*) indicates fragments of Slam1 (potentially C-terminal barrel) remaining after PK shaving.
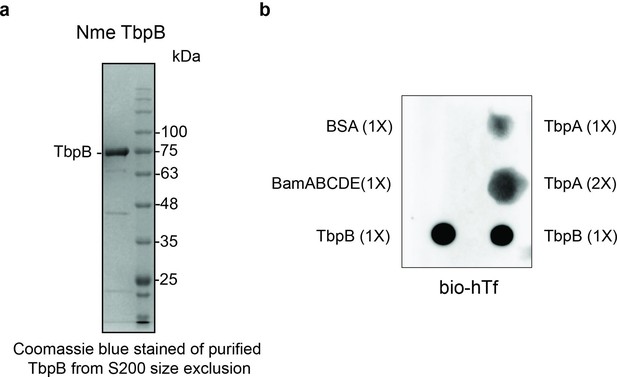
Purification and functional characterization of M. catarrhalis TbpB.
(a) Purified TbpB–DDM complexes obtained from a S200 size chromatography column. The sample was subsequently used for the in vitro translocation assay. (b) Dot blot assay with biotinylated human transferrin (bio-hTf) for detecting the function of TbpB. 0.5 μl TbpB and respective controls (TbpA, BamABCDE, and BSA) were spotted on nitrocellulose membrane and blotted with bio-hTf (50 μg/ml) followed by streptavidin–horseradish peroxidase (HRP). TbpB is bound tightly with bio-hTf indicating it is functional.
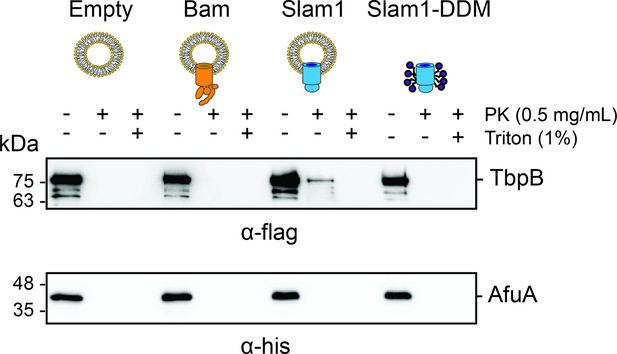
Protection of urea-unfolded TbpB (flag-tag) and negative control urea-unfolded AfuA (his-tag) by liposomes, proteoliposomes, and purified Slam1–DDM complex.
Mixture of urea-unfolded TbpB and urea-unfolded AfuA (1:1 molar ratio) was incubated with empty liposomes, proteoliposomes, or Slam1–DDM complex (1:20 volume ratio), followed by proteinase K treatment (0.1 mg/ml). Reaction was inhibited by 1 mM phenylmethylsulfonyl fluoride (PMSF) before loading on sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) gel and western transferred to PVDF blot. Blot was cut and the two halves were incubated with α-flag (for 75 kDa TbpB) or α-his antibodies (for 38 kDa AfuA) accordingly.
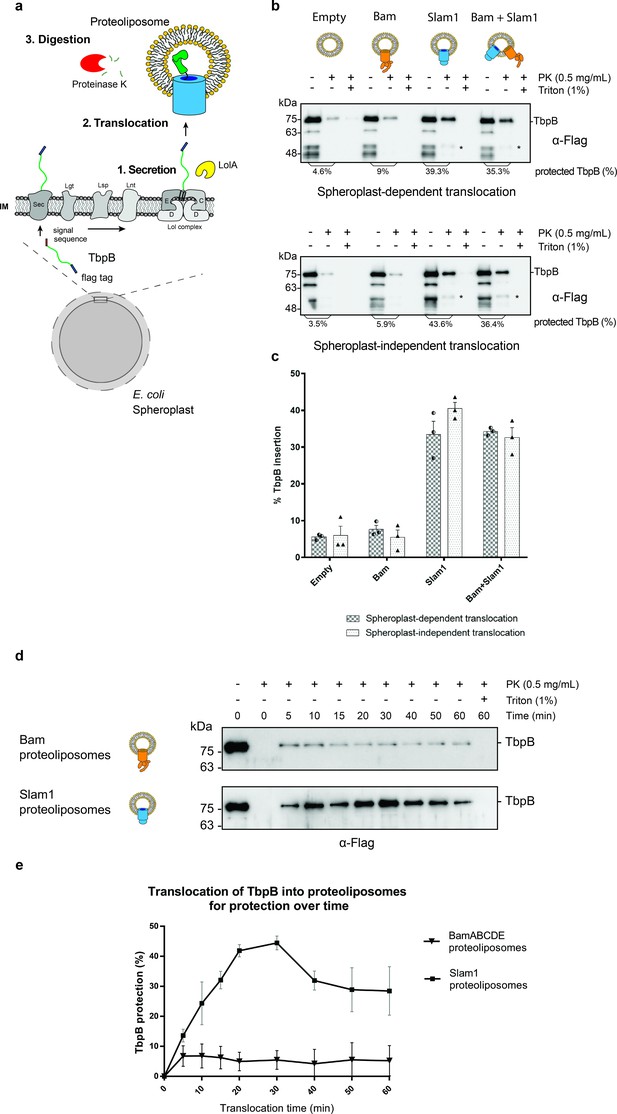
In vitro translocation assay for reconstitution of Slam-dependent SLP translocation.
(a) Model of the proposed in vitro proteoliposomes translocation assay for TbpB secreted directly from E. coli spheroplast. As E. coli cell expressing TbpB, the cells were converted into spheroplast that has an intact inner membrane. Purified LolA was added to release mature TbpB (processed by Lgt, Lsp, and Lnt) from the LolCDE complex in the inner membrane. The secreted TbpB was incubated with proteoliposomes for translocation, followed by PK digestion to quantify the amount of TbpB that had been translocated inside the liposomes. (b) Representative α-flag western blots obtained for the in vitro translocation assay. Slam1 proteoliposomes were incubated either with spheroplasts expressing TbpB (spheroplast-dependent translocation, upper panel) or supernatant of spheroplasts that have been induced for TbpB production (spheroplast-independent translocation, lower panel). Empty liposomes and Bam proteoliposomes were used as controls. Proteoliposomes containing Bam + Slam1 were used to test if the Bam complex plays an accessory role to Slam in TbpB translocation. For each proteoliposome, no proteinase K treatment (−PK), proteinase K treatment (+PK) and proteinase K + Triton X-100 treatment (+PK + T) samples are shown. The % TbpB protection shown was calculated by dividing the intensity of the mature TbpB band (~75 kDa) for each sample by the −PK sample. (*) Partial TbpB fragment which is only seen in the presence of Slam1 proteoliposomes. (c) Quantification of TbpB protection in proteoliposomes through densitometry analysis. The plot represents data obtained from at least three biological replicates for both spheroplast-dependent translocation and spheroplast-independent assay. Individual data points were included on the graph. (d) Representative α-flag western blot of spheroplast-independent TbpB translocation into Bam and Slam1 proteoliposomes over time. Spheroplast-secreted TbpB was incubated with proteoliposomes in 1:1 ratio at room temperature. Samples were collected every 5 or 10 min and left on ice before proteinase K treatment. (e) Quantification of spheroplast-secreted TbpB translocation into Bam proteoliposomes and Slam1 proteoliposomes over the course of 60 min.
-
Figure 2—source data 1
Quantification of in vitro translocation of spheroplast-secreted TbpB.
- https://cdn.elifesciences.org/articles/72822/elife-72822-fig2-data1-v2.csv
-
Figure 2—source data 2
Quantification of time-dependent translocation of spheroplast-secreted TbpB.
- https://cdn.elifesciences.org/articles/72822/elife-72822-fig2-data2-v2.csv
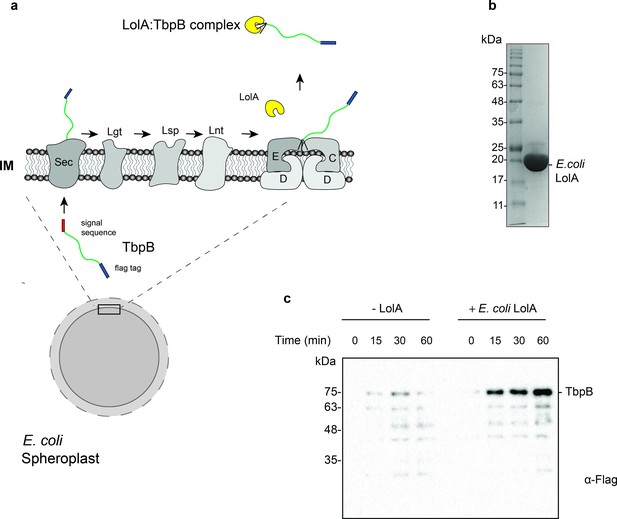
Purification and functional characterization of LolA from E. coli.
(a) Model of the release of SLPs from spheroplasts upon addition of purified E. coli LolA. SLPs are synthesized in the cytoplasm and transported to the periplasm via the Sec translocon. After the addition of the lipid anchor, SLPs are transferred to the LolCDE complex and released into the periplasm by LolA. In spheroplasts, SLPs are released into the supernatant upon LolA addition. (b) Uncut 6x-His tagged E. coli LolA (~22 kDa) after Ni-NTA affinity chromatography purification. (c) Release of Mcat TbpB in the presence of purified LolA over a time course of 60 min. E. coli cell pellets were converted into spheroplasts and induced for expression of TbpB in the presence or absence of 10 μM LolA. Samples were collected every 15 min, spun down at 13,500 rpm for 5 min and the supernatants were loaded on sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE). The amount of TbpB (~75 kDa) that was released into the supernatant in the presence and absence of LolA was accessed using a α-flag antibody western blot. Lower bands are degradation products of TbpB which also increase over time.
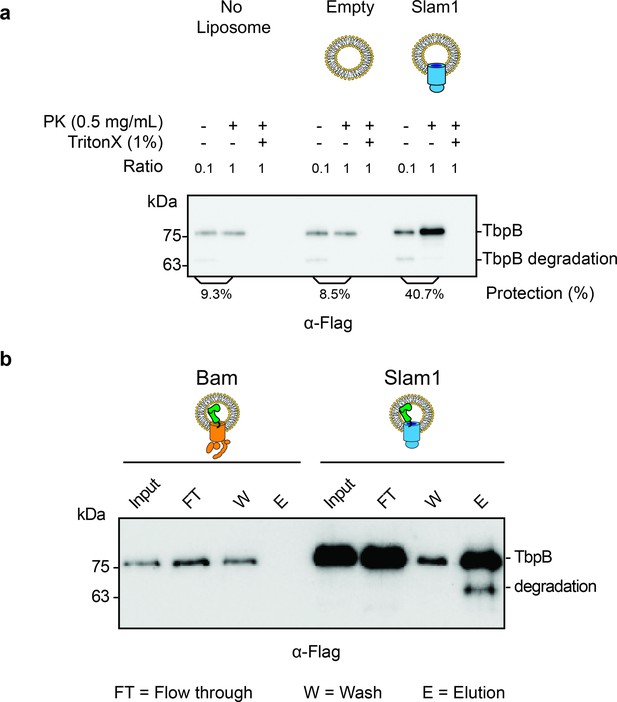
Characterization of spheroplast-secreted TbpB.
(a) Western blot accessing the protection of secreted TbpB in the absence of liposomes. Similar background protection was observed in no liposomes and liposome negative control (8.5% and 9.3%, respectively). Note: The input (−PK/−T) has been diluted by 1:10 to enhance the signal of the background protection for quantification. The protection percentage was calculated using the densitometry method and accounted for the dilution. Protection for TbpB in Slam1 proteoliposomes is still four times higher than the background protection (40.7%). (b) Functionality test of TbpB-containing proteoliposomes after translocation. After inhibiting proteinase K using PMSF, protected TbpB-proteoliposomes samples were treated with 0.1% Triton X and further incubated with human transferrin-conjugated beads to pulldown functional TbpB (flag-tagged). The lower band in Slam1-elution represents partially degraded TbpB. Note: The input of TbpB from Bam proteoliposomes and Slam1 proteoliposomes are different as they are outputs of the PK digestion assay (5% vs. 45% protected TbpB as the input for Bam and Slam1 proteoliposomes respectively).
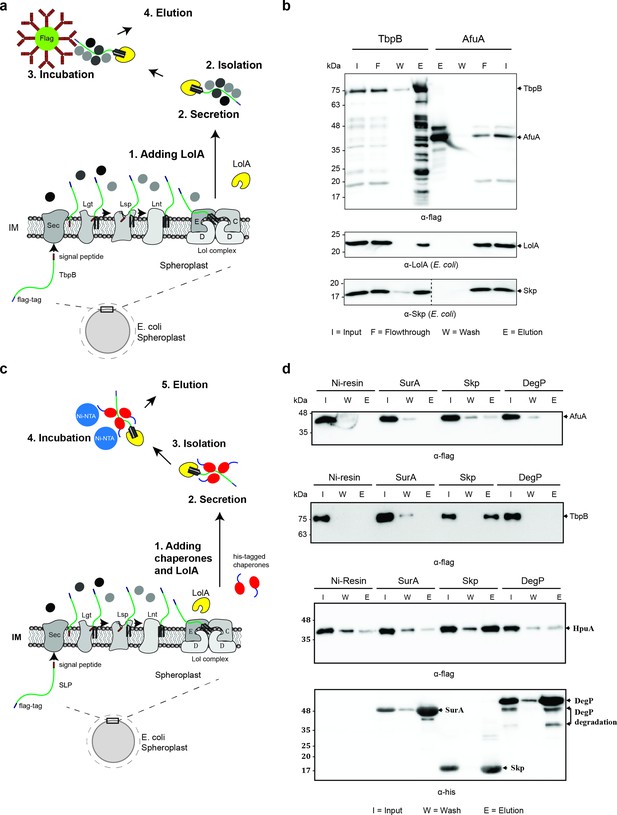
Periplasmic chaperone Skp interacts with surface lipoproteins TbpB and HpuA after being released from the inner membrane.
(a) Model of pulldown assay using the flag-tag on the C-terminus of TbpB. Samples were analyzed using mass spectrometry (summarized in Table 1) and examined on western blots. (b) Representative western blots for TbpB pulldown assay followed by mass spectrometry. LolA and periplasmic chaperone Skp were detected in the TbpB eluted fraction. No LolA and Skp were eluted in negative control AfuA experiment. Note: Upper and lower panels are from the same blot. Bottom panel was run on a second blot and the dashed line in the middle to indicate the exclusion of lanes (bead lanes – not shown) between the control (AfuA) and treatment (TbpB). (c) Model of reciprocal pulldown assay using the his-tag on the N-terminus of chaperones. Purified his-tagged chaperones (SurA, Skp, and DegP) were added to the spheroplast before the induced secretion of SLPs. (d) Representative western blots of the reciprocal pulldown assay. Only periplasmic chaperone Skp (17 kDa) was found to pull down lipoprotein TbpB and HpuA while no AfuA was found in the Skp elution fraction. All three proteins AfuA (negative control), TbpB and HpuA (Slam-dependent lipoproteins) were not found in the elution fraction of SurA and DegP pulldown. Note: α-his blot was used to detect chaperones and α-flag was used to detect HpuA samples within the same experiment. Elution volumes were 100 μl (1:10 input). DegP has molecular weight at 54 kDa. Lower band is the result of self degradation.
-
Figure 3—source data 1
Mass spectrometry of pulldown samples.
- https://cdn.elifesciences.org/articles/72822/elife-72822-fig3-data1-v2.rar
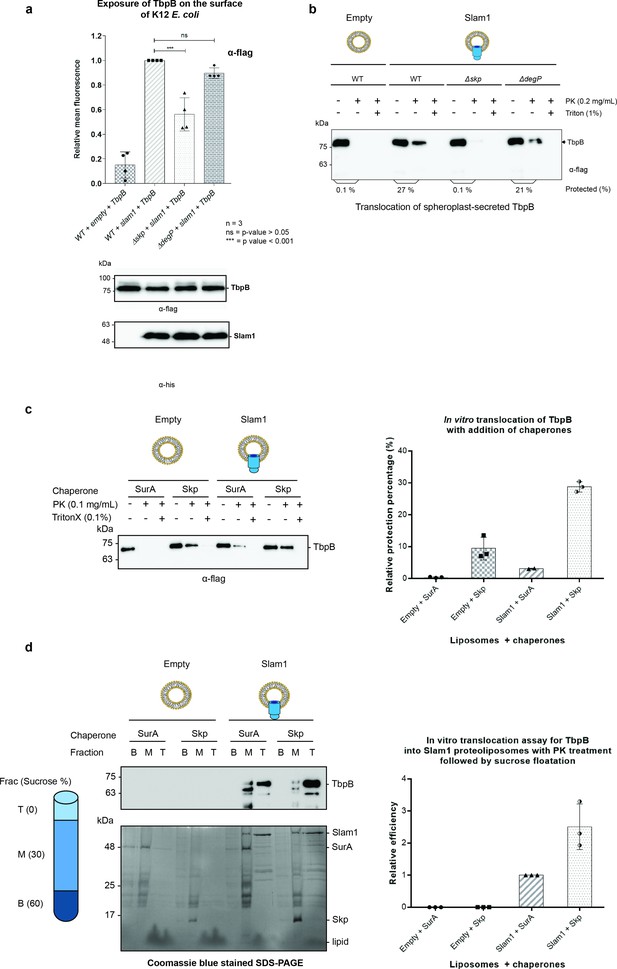
Periplasmic chaperone Skp is required for Slam1–TbpB translocation in the reconstitution systems.
(a) Translocation of TbpB via Slam1 to the surface of E. coli K12 mutants. Depletion of Skp significantly reduces the translocation of TbpB to the surface (by 50% – detecting by using α-flag antibody). Bottom panel – western blots of whole cell lysates detecting the expression of TbpB and Slam1 in K12 E. coli mutant strains. The processed TbpB (high intensity lower band at 75kDa) was unaffected in the mutant E. coli indicating that absence of one of the chaperones did not affect the maturation of TbpB in the periplasm. (***) represents p-value < 0.001 and (ns) represent p-value > 0.05. (b) Representative western blot of the in vitro proteoliposomes translocation for TbpB secreted from K12 E. coli spheroplast mutants. TbpB secreted from Δskp spheroplast fails to translocate into the Slam1 proteoliposomes for protection against proteinase K. (c) Representative western blot (left panel) and quantification (right panel) of the in vitro translocation of purified TbpB into Slam1 proteoliposomes in addition of purified chaperones. Full length lipidated TbpB was unfolded in urea followed by incubation with LolA and either SurA (negative control) or Skp before incubating with empty or Slam1 proteoliposomes and proteinase K digestion. TbpB–Skp complex provided extra protection for TbpB even in the absence of Slam1. (d) Representative western blot (left panel) and quantification (right panel) of the protected TbpB by the liposomes which were isolated using sucrose flotation assay after proteinase K digestion. Translocation of TbpB into Slam1 proteoliposomes increased by 2.5-fold in the presence of Skp in comparison with the Slam1 proteoliposomes + SurA (positive control). Results are from at least three biological replicates. Individual data points were included on the graph.
-
Figure 4—source data 1
Fluorescence of TbpB on surface of E. coli.
- https://cdn.elifesciences.org/articles/72822/elife-72822-fig4-data1-v2.csv
-
Figure 4—source data 2
Quantification of in vitro translocatio.
- https://cdn.elifesciences.org/articles/72822/elife-72822-fig4-data2-v2.csv
-
Figure 4—source data 3
Quantification of in vitro translocation following sucrose floatation assay.
- https://cdn.elifesciences.org/articles/72822/elife-72822-fig4-data3-v2.csv
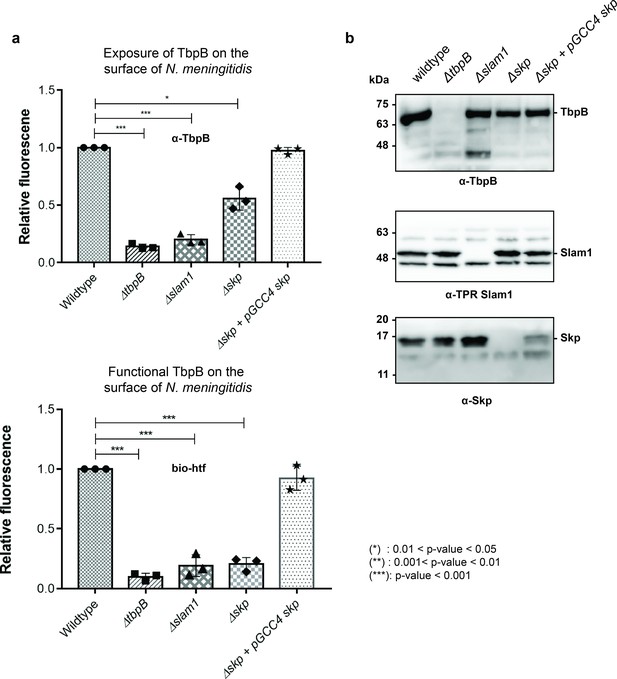
Periplasmic chaperone Skp is important for translocation of TbpB to the surface of N. meningitidis.
(a) Relative fluorescence intensity accessing the presence of TbpB (65 kDa) on the surface of N. meningitidis mutants using α-TbpB antibody (exposure – top) and biotinylated human transferrin (functional – bottom). Individual data points were included on the graph. Depletion of Skp decreased surface exposed TbpB by 50% and the translocated TbpB is nonfunctional (not binding to biotinylated human transferrin). Complementation of skp from pGCC4 vector and 0.1 mM IPTG successfully restored the translocation of TbpB and its function. (*) represent 0.01 < p-value < 0.05 and (***) represent p-value < 0.001. (b) Representative western blots to access the expression of TbpB, Slam1, and Skp in the N. meningitidis strains examined. Depletion of Skp did not affect the expression of outer membrane protein (OMP) Slam1 or TbpB (induced by 0.1 mM deferoxamine). Fluorescent assay results are combined from three biological replicates and statistically analyzed by one-way analysis of variance (ANOVA) test.
-
Figure 5—source data 1
Fluorescent signal of TbpB on the surface of N. meningitidis.
- https://cdn.elifesciences.org/articles/72822/elife-72822-fig5-data1-v2.csv
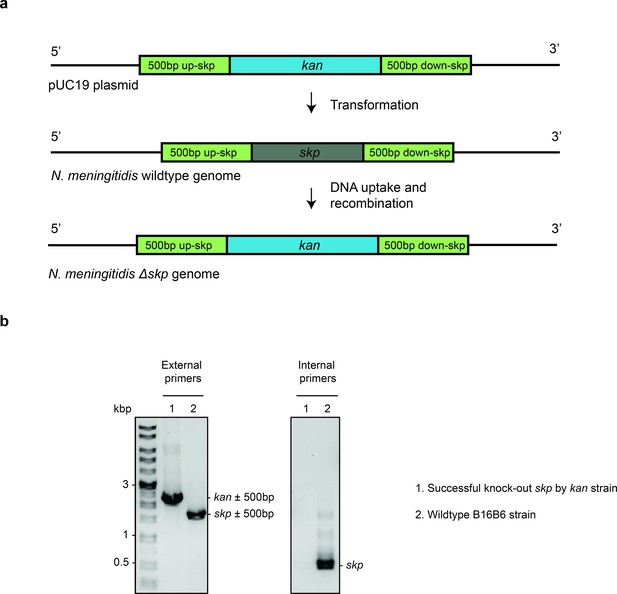
Deletion of Skp in N. meningitidis.
(a) The complete knockout process using spot transformation. Gene skp in N. meningitidis B16B6 is replaced by kan cassette. (b) Agarose gel of single colony PCR using external primers (targeting DNA 500 bp upstream and downstream of skp) and internal primers (targeting DNA within skp gene) to confirm the successful replacement of kan for skp gene.
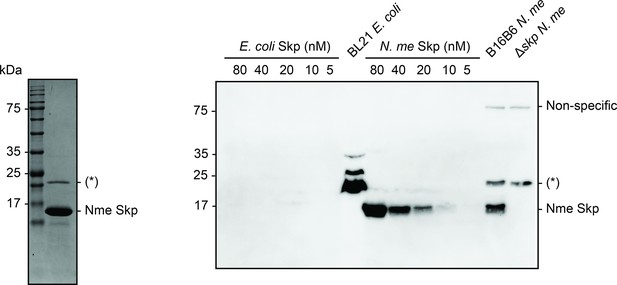
Purification of Nme Skp for antibody production and antibody test.
Coomassie blue stained sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) of Nme Skp after S75 gel filtration to access its purity prior to antibody production (left panel) and α-Nme Skp antibody western blot to validate the antibody and confirm the knock-out of Skp (17 kDa) in B16B6 N. meningitidis (right panel). (*) is the contamination from BL21 E. coli. There is no cross-reactivity of the Nme Skp antibody with E. coli Skp.
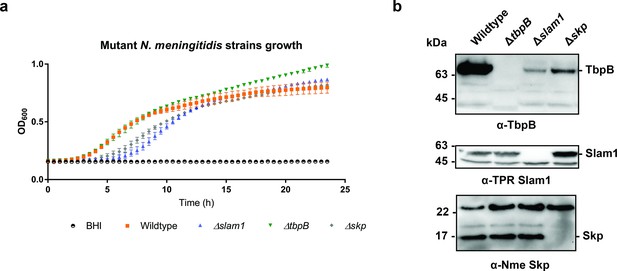
Growth of B16B6 N. meningitidis strains in BHI media.
(a) OD600 was recorded every 30 min over a 24-hr period. Δskp and Δslam1 mutants were lagging behind but reached OD600 of 0.7 eventually. (b) Representative western blots detecting the expression of Nme TbpB (65 kDa), Nme Slam1 (48 kDa), and Nme Skp (17 kDa). Knockout of slam1 or skp affected the quantity of TbpB in whole cells.
-
Figure 5—figure supplement 3—source data 1
Growth of N. meningitidis.
- https://cdn.elifesciences.org/articles/72822/elife-72822-fig5-figsupp3-data1-v2.csv
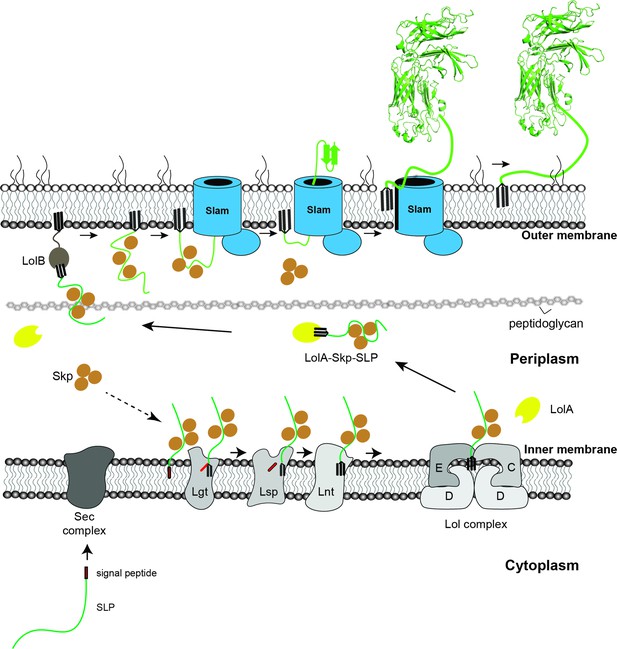
Proposed model of localization for Slam-dependent surface lipoproteins in Gram-negative bacteria.
Once the surface lipoproteins emerged from the Sec complex, periplasmic chaperone Skp binds to the SLPs to prevent early folding while their N-terminus is modified and lipidated by Lgt, Lsp, and Lnt before getting passed to the Lol complex. LolA then releases lipidated SLPs to the periplasm while Skp stays bound to prevent SLPs folding. LolB in the outer membrane (OM) might serve as the receiver for the LolA–SLP–Skp complex. The ‘specificity motif’ present at the C-terminus of the unfolded SLP is recognized by Slam and then transported across the outer membrane through the Slam channel and folds on the other side. Once the entire length of the SLP is transported, the Slam lateral gate allows the lipid anchor to ‘flip’ from the inner leaflet to the outer leaflet of the outer membrane.
Tables
Summary of mass spectrometry for flag-tagged TbpB pulldown.
| AfuA | TbpB | |||||||
|---|---|---|---|---|---|---|---|---|
| Proteins | Total peptides | % Coverage | Location | Proteins | Total peptides | % Coverage | Location | |
| 1 | AfuA | 380 | 78% | P | TbpB | 265 | 31% | – |
| 2 | TufA | 119 | 70% | IM | LolA | 146 | 73% | P |
| 3 | LolA | 36 | 72% | P | TufA | 57 | 59% | IM |
| 4 | OmpF | 12 | 34% | OM | OmpF | 21 | 38% | OM |
| 5 | DegP | 10 | 30% | P | DegP | 15 | 32% | P |
| 6 | FlgH | 12 | 34% | P | OmpA | 15 | 34% | OM |
| 7 | DegQ | 6 | 14% | P | Skp | 9 | 27% | P |
| 8 | OmpA | 5 | 22% | OM | FlgH | 8 | 26% | P |
| 9 | DegQ | 5 | 13% | P | ||||
Additional files
-
Supplementary file 1
Strains, plasmids, and antibodies used in this study.
Raw images for figures and figure supplements.
- https://cdn.elifesciences.org/articles/72822/elife-72822-supp1-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/72822/elife-72822-transrepform1-v2.pdf
-
Source data 1
Mass spectrometry results.
- https://cdn.elifesciences.org/articles/72822/elife-72822-data1-v2.zip
-
Source data 2
Original gels and blots used for the figures.
- https://cdn.elifesciences.org/articles/72822/elife-72822-data2-v2.zip





