Excitatory neurotransmission activates compartmentalized calcium transients in Müller glia without affecting lateral process motility
Figures
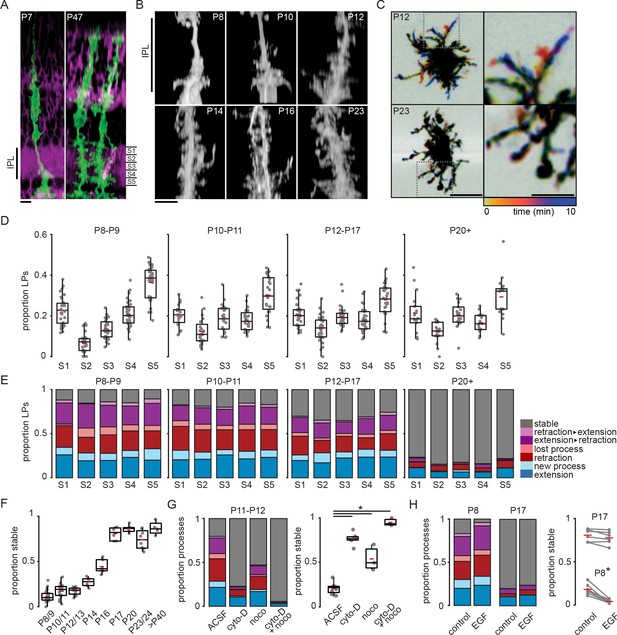
Müller glial lateral processes are highly motile during development.
(A) Orthogonal projections of two-photon volumetric images of Müller glia expressing membrane-GFP (mGFP, green) in sparsely recombined Slc1a3-CreER;mTmG retinas at P7 (left) and P47 (right). Cells without Cre-mediated recombination express membrane-tdTomato (magenta). Scale bar: 10 µm. IPL: inner plexiform layer. (B) Orthogonal projections of mGFP-expressing Müller glia from P8 to P23. Scale bar: 10 µm. (C) Temporally color-coded projections of two-photon Z-stack time series showing motile processes at P12 (top) and stable processes at P23 (bottom). The location and dynamics of lateral processes at each time point can be determined by referencing the color time scale at bottom. Images on right are enlarged insets from images on left to highlight individual processes. Left scale bar: 10 µm; right scale bar: 5 µm. See Figure 1—figure supplement 1 for enlarged images of these cells in sublayers S1, S3, and S5. (D) Distribution of lateral processes across IPL sublayers through development. See Supplementary file 1 for summary statistics and goodness-of-fit test statistics, and Supplementary file 2 for tests for independent proportions across age groups. (E) Proportion of lateral processes in each sublayer that underwent extension (blue), sprouting (light blue), retraction (red), elimination (light red), a combination (purple), or that were stable (gray). (F) Proportion of total stable processes across development. See Supplementary file 3 for statistical comparisons. (G) Proportion of total processes that were motile (left) and stable (right) in the absence and presence of blockers of cytoskeletal turnover at P11/12. cyto-D: cytochalasin-D (5 µM); noco: nocodazole (10 µM). See Supplementary file 4 for statistical comparisons. (H) Proportion of total processes that were motile (left) and stable (right) in the absence and presence of epidermal growth factor (EGF, 1 unit [100 ng]/ml) at P8 and P17. See Supplementary file 5 for statistical comparisons. * corresponds to p < 0.05 for all comparisons. Source data available in Figure 1—source data 1 and Figure 1—source data 2.
-
Figure 1—source data 1
Proportions and counts of motile and stable lateral processes across sublayer and age.
- https://cdn.elifesciences.org/articles/73202/elife-73202-fig1-data1-v2.xlsx
-
Figure 1—source data 2
Proportions and counts of motile and stable lateral processes in the presence and absence of cytoskeletal blockers or EGF.
- https://cdn.elifesciences.org/articles/73202/elife-73202-fig1-data2-v2.xlsx
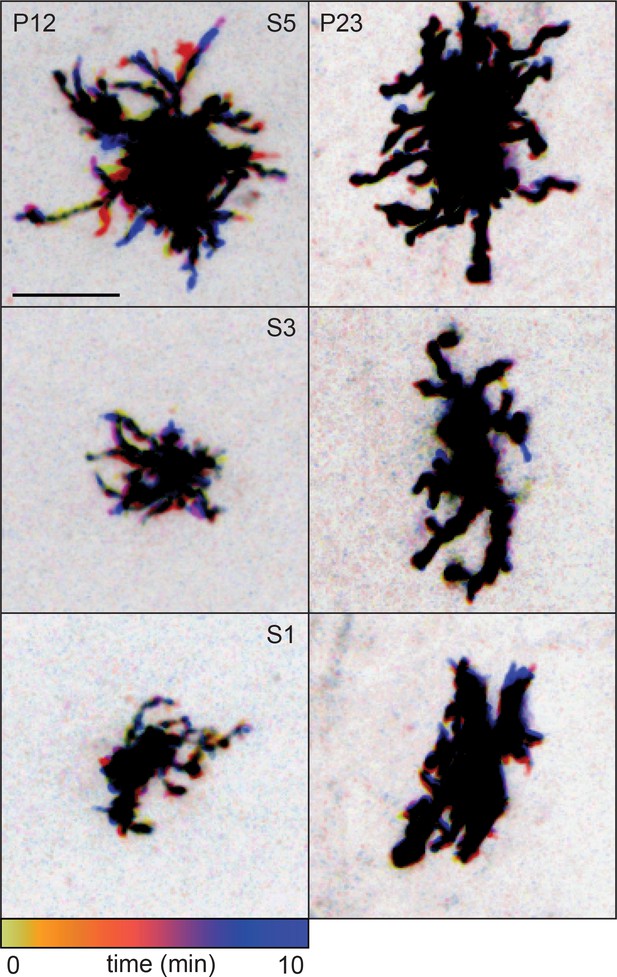
Temporally color-coded projections of the cells shown in Figure 1C, highlighting motile and stable processes in sublayers S5 (top), S3 (middle), and S1 (bottom).
The location and dynamics of lateral processes at each time point can be determined by referencing the color time scale at bottom. Scale bar: 10 µm.
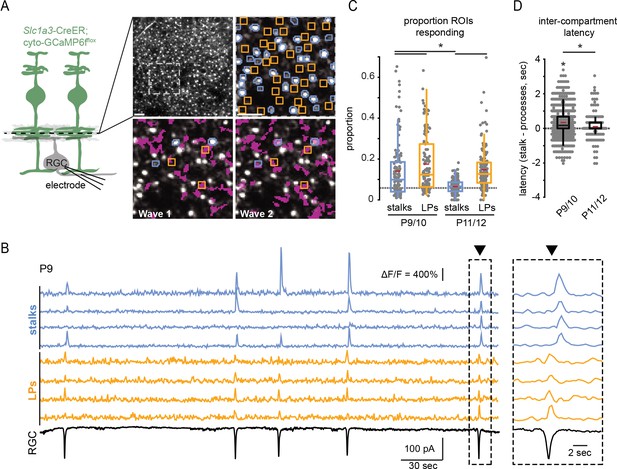
Retinal wave-associated calcium transients are compartmentalized within Müller glial stalks and lateral processes.
(A) Left, diagram of experimental setup; cytosolic GCaMP6f was expressed in Müller glia (green) via Slc1a3-CreER, and a two-photon microscope was used to image calcium transients in glial stalks and lateral processes in the IPL (dashed line indicates focal depth in whole-mount retina). Excitatory currents were recorded from a retinal ganglion cell (RGC) to detect retinal waves. Right, average projection of a field of view (FOV) showing GCaMP6f-expressing Müller glia in retinal whole mount. Upper left panel is the full FOV; upper right panel is a magnified portion of this FOV overlayed with regions of interest (ROIs) for stalks (blue) and lateral processes (yellow); bottom panels show the extent of Müller glial activation by two retinal waves (active regions overlayed in magenta). (B) Example traces of fractional change in fluorescence of GCaMP6f-expressing Müller glia showing wave-associated calcium transients in stalks (blue) and processes (yellow) and wave-associated EPSCs recorded from an RGC (black). Right, Müller glial calcium response during a retinal wave, expanded in time to highlight temporal delay of stalk transients relative to lateral process transients. Black trace is a voltage-clamp recording from an RGC held at –60 mV. (C) Proportion of total stalk and lateral process ROIs that responded to retinal waves at P9/10 and P11/12. Gray dashed line indicates the proportion of ROIs undergoing spontaneous calcium transients at randomly selected times. See Supplementary file 6 for summary statistics. (D) Intercompartment response latency, calculated as latency of stalk responses from the median lateral process response time for retinal waves at P9/10 and P11/12. See Supplementary file 7 for summary statistics. Image scale bars: 10 µm. Source data available in Figure 2—source data 1.
-
Figure 2—source data 1
Proportion of total ROIs responding per wave and intercompartment latency among wave-associated calcium transients.
- https://cdn.elifesciences.org/articles/73202/elife-73202-fig2-data1-v2.xlsx
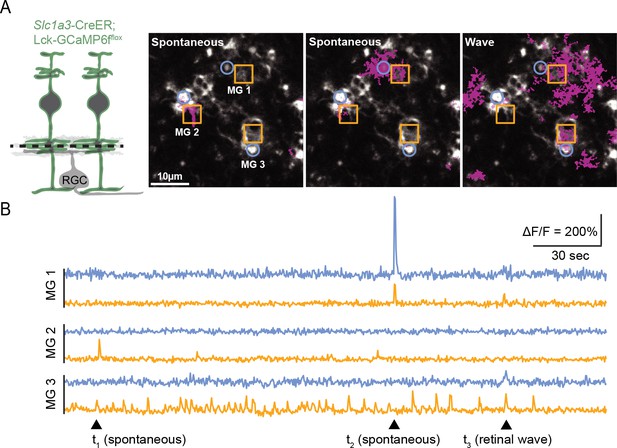
Visualization of calcium compartments using Slc1a3-CreER;Lck-GCaMP6fflox.
(A) Left, diagram of Slc1a3-CreER;Lck-GCaMP6fflox retina used for imaging calcium in glial processes. Black dashed line indicates two-photon imaging focal plane. Right, example FOV showing two spontaneous calcium events (left and middle; transients denoted as magenta regions) and one wave-associated calcium event (right) in Müller glia (MG) at P10. Three stalks (blue circles) and three neighboring lateral process ROIs (yellow squares) have been denoted as MG 1–3. (B) Normalized fluorescence traces showing calcium activity in MG 1–3. Arrowheads indicate time points shown in images in (A). FOV, field of view; ROIs, regions of interest.
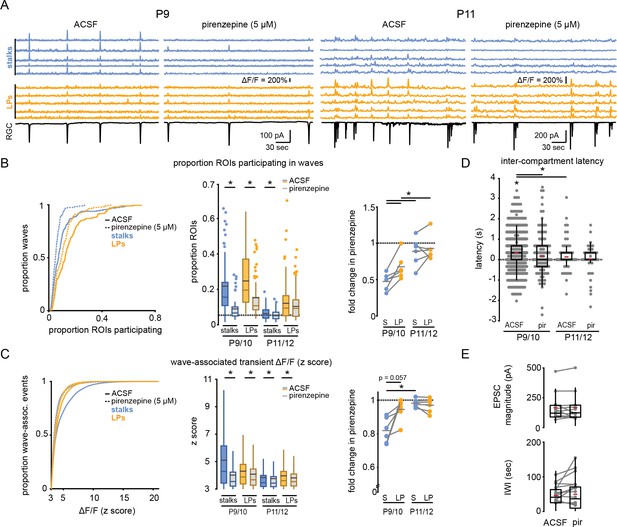
M1 mAChRs mediate retinal wave-associated calcium transients preferentially localized to Müller glial stalks.
(A) Example traces of fractional change in fluorescence of GCaMP6f-expressing Müller glia (top) showing wave-associated calcium transients in stalks (blue) and processes (yellow), and wave-associated EPSCs recorded from a retinal ganglion cell (RGC, bottom) in the absence and presence of pirenzepine (5 µM) at P9 (left) and P11 (right). (B) Left, cumulative distribution of proportion of stalks and lateral processes that exhibited wave-associated calcium transients in absence (solid line) and presence of pirenzepine (dotted line). Middle, box plots showing proportion of ROIs that exhibited wave-associated calcium transients. Gray dashed line indicates the proportion of ROIs that exhibited spontaneous calcium transients at randomly selected times. See Supplementary file 8 for summary statistics. Right, fold-change in proportion of ROIs that exhibited wave-associated calcium transients in presence of pirenzepine. See Supplementary file 9 for summary statistics. (C) Left, cumulative distribution of the z scored ΔF/F amplitude of wave-associated calcium transients in the absence and presence of pirenzepine. See Supplementary file 10 for summary statistics. Middle, box plots showing z scored ΔF/F amplitudes separated by age. Right, fold-change in wave-associated ΔF/F amplitude in pirenzepine. See Supplementary file 11 for summary statistics. (D) Intercompartment response latency for each age and condition. See Supplementary file 12 for summary statistics. (E) Summary data for magnitude of compound excitatory post-synaptic currents (EPSC; top) and interwave interval (IWI; bottom) during retinal waves. See Supplementary file 13 for summary statistics. Source data available in Figure 3—source data 1.
-
Figure 3—source data 1
Retinal wave response properties among Müller glia and recorded RGCs in ACSF and pirenzepine.
- https://cdn.elifesciences.org/articles/73202/elife-73202-fig3-data1-v2.xlsx
-
Figure 3—source data 2
Retinal wave response properties among Müller glia in ACSF and DNQX.
- https://cdn.elifesciences.org/articles/73202/elife-73202-fig3-data2-v2.xlsx
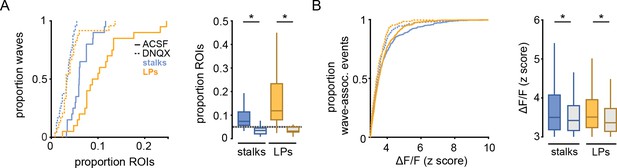
DNQX reduces glial participation in waves among stalks and lateral processes.
(A) Left, cumulative distribution of proportion stalk ROIs (blue) or lateral process ROIs (yellow, LPs) undergoing wave-associated calcium transients for all waves in control (solid lines) and 20 µM DNQX (dotted lines). Right, box plots showing median and 25–75% interquartile range for data shown in left. Colored boxes indicate control, while gray boxes indicate DNQX. Gray dashed line indicates proportion of ROIs undergoing calcium transients at randomly selected times. N (waves): 20, ACSF; 24, DNQX across two mice each. * indicates p < 0.05, unpaired t-test. (B) Left, cumulative distribution of Z scored wave-associated ΔF/F amplitude in stalks and LPs in control and DNQX. Right, box plots showing median and 25–75% interquartile range for data shown in left. N (wave-associated transients): 466 in stalks/ACSF; 658 in LPs/ACSF; 274 in stalks/DNQX; 259 in LPs/DNQX. * indicates p < 0.05, unpaired t-test. Source data available in Figure 3—source data 2.
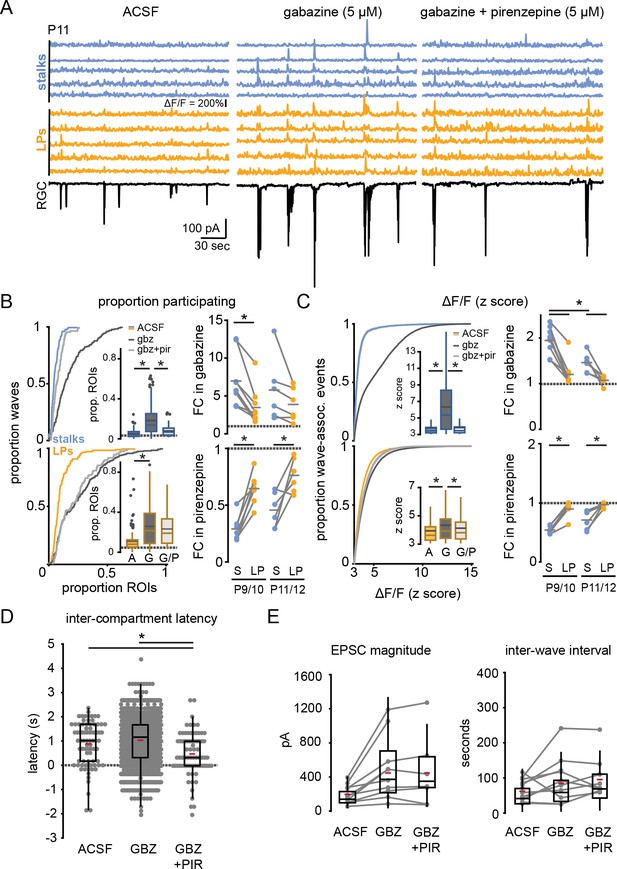
GABAA receptor block potentiates wave responses in stalks via M1 mAChR activation.
(A) Example traces of fractional change in fluorescence of GCaMP6f-expressing Müller glia showing wave-associated calcium responses (top) and wave-associated excitatory post-synaptic currents (EPSCs) recorded from an RGC (bottom) in control (left), 5 µM gabazine (middle), and gabazine plus 5 µM pirenzepine (right) at P11. (B) Left, cumulative distributions and inset box plots of proportion of stalk ROIs (top) and lateral process ROIs (bottom) that exhibited wave-associated calcium transients in control (solid line, A), gabazine (dotted line, G), and gabazine/ pirenzepine (dashed line, G/P) from P9 to P12. Gray dashed line in box plots indicates the proportion of ROIs undergoing spontaneous calcium transients at randomly selected times. Right, fold-change (FC) in proportion ROIs responding in gabazine versus ACSF (top) and in pirenzepine vs. gabazine (bottom). Gray dashed line indicates FC of 1. See Supplementary files 14 and 15 for summary statistics. (C) Left, cumulative distributions of normalized wave-associated calcium transient amplitudes in ACSF, gabazine, and gabazine/pirenzepine from P9 to P12, with inset box plots generated from the same data. Right, FC in normalized calcium response amplitude after application of gabazine (top) and then pirenzepine (bottom). See Supplementary files 16 and 17 for summary statistics. (D) Intercompartment latency in wave-associated calcium transients in control, gabazine (gbz), and gabazine/pirenzepine (gbz+pir), from P9 to P12. See Supplementary file 18 for summary statistics. (E) Summary data for magnitude of (top) and interval between (bottom) compound EPSCs associated with retinal waves in control, gabazine, and gabazine/pirenzepine for paired FOVs from P9 to P12. See Supplementary file 19 for summary statistics. Source data available in Figure 4—source data 1. FOV, field of view; RGC, retinal ganglion cell; ROIs, regions of interest.
-
Figure 4—source data 1
Retinal wave response properties among Müller glia and recorded RGCs in ACSF, gabazine, and gabazine plus pirenzepine.
- https://cdn.elifesciences.org/articles/73202/elife-73202-fig4-data1-v2.xlsx
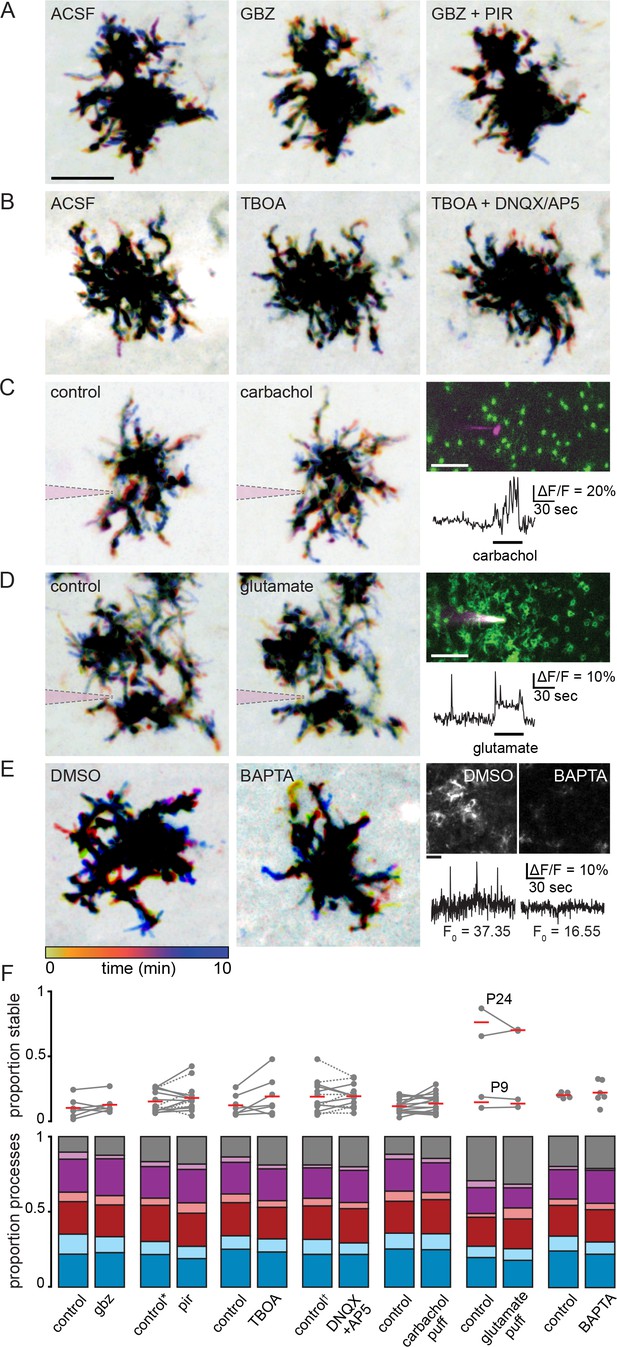
Müller glial lateral process motility is unaffected by manipulations of retinal waves.
(A) Temporally color-coded projections of two-photon Z stack time series showing motile and stable processes in control (ACSF), gabazine (5 µM), and gabazine plus pirenzepine (5 µM) at P11. Scale bar: 10 µm. (B) Same as A in control (ACSF), TBOA (25 µM), and TBOA plus DNQX (20 µM) and AP5 (50 µM) as pharmacological manipulations of retinal waves. (C and D) show motility (left, middle) and calcium responses (right, scale bar: 20 µm) to iontophoretically applied carbachol (P10) and glutamate (P9). A sharp electrode (indicated in magenta) was filled with 10 mM agonist and current continuously applied to eject agonist into the IPL near mGFP-expressing processes to image motility, or near GCaMP6f-expressing processes to image calcium responses. (E) Left, motility persisted at P12 even when intracellular calcium was chelated via 200 µM BAPTA-AM bath application, which greatly reduced baseline calcium (F0) and eliminated neuronal activity (see Figure 5—figure supplement 1). Scale bars: 10 µm. Right, average projections (top) and normalized traces of Lck-GCaMP6f fluorescence (bottom) in DMSO and BAPTA. (F) Summary data showing proportion of total processes exhibiting stability (top) and other categories of motility (bottom) for drug conditions shown in (A–E). * gabazine was included in the control condition for a subset of experiments using pirenzepine, as in (A). † TBOA was included in the control condition for a subset of experiments using DNQX/AP5, as in (B). These trials are denoted by dotted lines connecting control and drug conditions. See Figure Supplementary file 20 for summary statistics. Source data available in Figure 5—source data 1.
-
Figure 5—source data 1
Proportions and counts of motile and stable lateral processes during perturbations of neuronal activity or intracellular calcium.
- https://cdn.elifesciences.org/articles/73202/elife-73202-fig5-data1-v2.xlsx
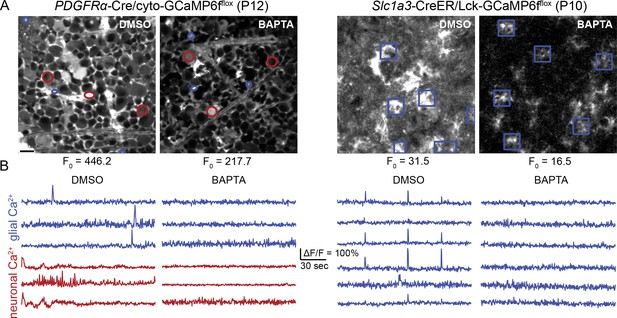
Bath loading with BAPTA-AM during development eliminates glial and neuronal calcium activity and retinal waves.
(A) Left, average projection of GCL in PDGFRα;cyto-GCaMP6fflox retinas at P12 following incubation in vehicle control or 200 µM BAPTA-AM. Cyto-GCaMP6f fluorescence is localized to Müller glia and a subset of neurons. Scale bar: 10 µm. Right, average projection of IPL in Slc1a3-CreER;Lck-GCaMP6fflox retinas at P10 following incubation in vehicle or 200 µM BAPTA-AM. Lck-GCaMP6f fluorescence is localized to Müller glial processes. Baseline fluorescence (F0) is greatly reduced in the presence of BAPTA. (B) Cytosolic calcium transients are eliminated in neurons and glia (left), and membrane-localized calcium transients are eliminated in glia (right) following incubation in BAPTA-AM. Related to Figure 5E.
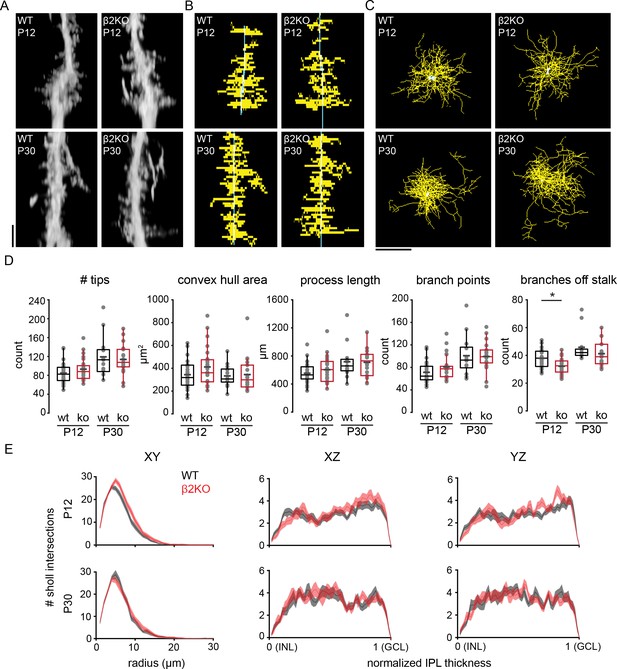
Chronic perturbation of retinal waves does not impact distribution, length, or complexity of Müller glial lateral processes.
(A) Orthogonal projections of two-photon Z stacks obtained after sharp-filling single Müller glial cells with Alexa-488 at P12 (top) and at P30 (bottom) in wild-type (WT, left) and in β2 nicotinic acetylcholine receptor knockout retinas (β2KO, right). Image scale bar is 10 µm. (B) XZ projections of skeletons from cells shown in (A). (C) XY projections of skeletons from cells shown in (A). (D) Morphological measurements obtained from traced and skeletonized Müller glia. See Supplementary file 21 for summary statistics. Source data available in Figure 6—source data 1. (E) Sholl intersection profiles for WT (black) and β2KO (red) at P12 (top) and P30-39 (bottom). Left plots are Sholl profiles carried out on XY projections, middle plots are from XZ projections, and right plots are from YZ projections. For XZ and YZ projections, the center of the Sholl radius was placed on portion of the stalk closest to the inner nuclear layer (INL). Two-way mixed ANOVA revealed no significant effect of genotype on number of intersections at any Sholl radii for both ages. GCL, ganglion cell layer. Source data for Sholl analyses available in Figure 6—source data 2.
-
Figure 6—source data 1
Comparisons of morphological properties between wild type and β2-nAChR-KO Müller glia.
- https://cdn.elifesciences.org/articles/73202/elife-73202-fig6-data1-v2.xlsx
-
Figure 6—source data 2
Sholl analysis profiles for wild type and β2-nAChR-KO Müller glial lateral processes.
- https://cdn.elifesciences.org/articles/73202/elife-73202-fig6-data2-v2.xlsx
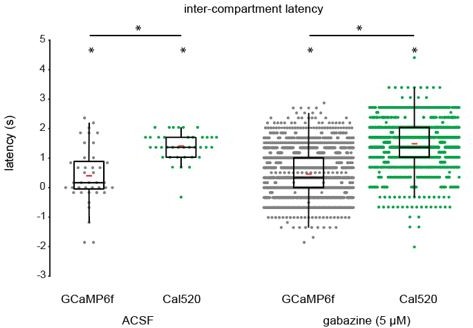
Comparison of intercompartment latency of retinal wave-associated calcium transients between GCaMP6f-expressing and Cal520-loaded Muller glia.
Although intercompartment latency was significantly greater than zero using both GCaMP6f and Cal520, Cal520-loaded Muller glia exhibited a significantly higher latency than GCaMP6f- expressing Muller glia during retinal waves. Median [1st quartile/3rd quartile], ACSF (GCaMP6f, Cal520; seconds): 0.169 [0.0/0.85], 1.35 [1.10/1.69]. Gabazine (GCaMP6f, Cal520; seconds): 0.34 [0.0/1.01]; 1.35 [1.01/2.03]. Red bars indicate mean latency for each group. * indicates p < 0.05. Differences in latency between indicators were tested using Wilcoxon rank-sum, and differences in latency from zero were tested using Wilcoxon signed-rank tests.
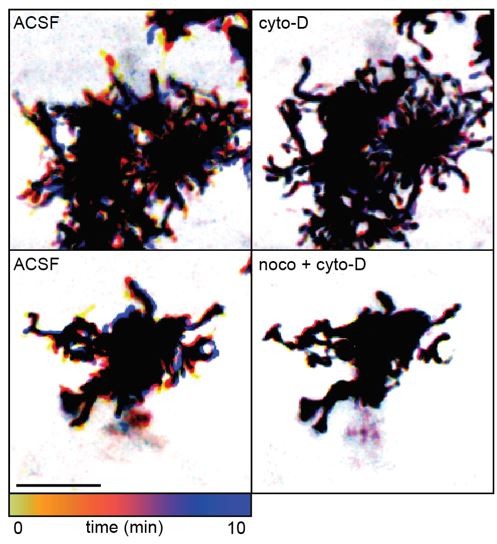
Temporally color-coded images of 2-photon Z stack time series through the IPL of GLAST/mTmG retinas.
Cytoskeletal manipulations using 5 µM cytochalasin-D (cyto-D) and 10 µM nocodazole (noco) blocked motility in lateral processes without inducing a morphologically reactive phenotype at P11. Scale bar 10 µm. Related to Figure 1G.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Mus musculus; male & female) | C57BL/6J (wild-type) | The Jackson Laboratory | RRID:IMSR_JAX:000664 | Background strain used for mouse crosses |
| Strain, strain background (M. musculus; male and female) | Slc1a3-CreER; Tg(Slc1a3-cre/ERT)1Nat/J; also known as GLAST-CreER | The Jackson Laboratory | RRID:IMSR_JAX:012586 | Expresses tamoxifen-inducible Cre/ERT in Müller glia |
| Strain, strain background (M. musculus; male and female) | mTmG; mT/mG; B6.129(Cg)- Gt(ROSA)26 Sortm4(ACTB-tdTomato,-EGFP)Luo/J | The Jackson Laboratory | RRID:IMSR_JAX:007676 | Used to measure morphology/motility in Müller glia |
| Strain, strain background (M. musculus; male and female) | GCaMP6fflox; B6.129S(Cg)-Gt(ROSA) 26Sortm95.1(CAG-GCaMP6f)Hze/J | The Jackson Laboratory | RRID:IMSR_JAX:024105 | Used to measure calcium dynamics in Müller glial stalks and processes |
| Strain, strain background (M. musculus; male and female) | Lck-GCaMP6fflox; C57BL/6N- Gt(ROSA) 26Sortm1(CAG-GCaMP6f)Khakh/J | The Jackson Laboratory | RRID:IMSR_JAX:029626 | Used to measure calcium dynamics in Müller glial processes |
| Strain, strain background (M. musculus; male and female) | PDGFRa-Cre; C57BL/6- Tg(Pdgfra-cre) 1Clc/J | The Jackson Laboratory | RRID:IMSR_JAX:013148 | Used to verify loss of calcium transients following BAPTA-AM application |
| Strain, strain background (M. musculus; male and female) | β2-nAChR-KO; C57BL/6J- Chrnb2tm1Mdb | Xu et al., 1999; Bansal et al., J Neurosci, 2000. doi: https://doi.org/10.1523/JNEUROSCI.20-20-07672.2000 | RRID:MGI:2663180 | Targeted deletion generated by Xu et al. (J Neurosci, 1999) and maintained in our lab since |
| Chemical compound, drug | 4-hydroxytamoxifen (4-OHT, 50:50 E:Z isomers) | Sigma-Aldrich | T176 | Injected at variable doses to induce uniform or sparse Cre-mediated recombination in Müller glia |
| Chemical compound, drug | Cytochalasin-D (from Zygosporium mansoni) | Avantor/VWR | Supplier no.: 250255catalog number: 80055-214 | Disrupts actin filaments and inhibits actin polymerization |
| Chemical compound, drug | Nocodazole | Sigma-Aldrich | M1404 | Disrupts microtubule assembly/disassembly |
| Chemical compound, drug | Epidermal growth factor (EGF) (Rac/Cdc42 Activator II) | Cytoskeleton, Inc | CN02 | EGF receptor agonist |
| Chemical compound, drug | Pirenzepine dihydrochloride | Tocris | 1071 | M1 mAChR antagonist |
| Chemical compound, drug | Gabazine (SR 95531 hydrobromide) | Tocris | 1262 | GABAA receptor antagonist |
| Chemical compound, drug | DL-TBOA | Tocris | 1223 | Glutamate transporter (EAAT) antagonist |
| Chemical compound, drug | DNQX disodium salt | Tocris | 2312 | AMPAR antagonist |
| Chemical compound, drug | DL-AP5 | Tocris | 3693 | NMDAR antagonist |
| Chemical compound, drug | BAPTA-AM | Tocris | 2787 | Intracellular calcium chelator |
| Chemical compound, drug | Carbachol (carbamoylcholine chloride) | Tocris | 2810 | Cholinergic agonist |
| Chemical compound, drug | L-glutamic acid (Glu) | Sigma-Aldrich | 56-86-0 | Glutamatergic agonist |
| Chemical compound, drug | Alexa fluor 488/594 hydrazide, sodium salt (Alexa-488/594) | Thermo Fisher Scientific | A10436/A10438 | Used for filling Müller glia to visualize processes, and as a counterdye for focal agonist application |
| Software, algorithm | ScanImage | Vidrio Technologies, LLC | RRID:SCR_014307 | Two-photon image acquisition software |
| Software, algorithm | FIJI/ImageJ | Created at National Institutes of Health; Schindelin et al., Nature Methods, 2012. doi:10.1038/nmeth.2019 | RRID:SCR_002285 | Used for image processing and analysis |
| Software, algorithm | MATLAB | MathWorks | RRID:SCR_001622 | Custom scripts used for data processing, analysis |
| Software, algorithm | NormCorre: Non-Rigid Motion Correction | Flatiron Institute, Simons Foundation; Pnevmatikakis and Giovannucci, J Neurosci Methods, 2017. doi: https://doi.org/10.1016/j.jneumeth.2017.07.031 | For 2D registration of calcium imaging movies | |
| Software, algorithm | FIJI plugin: Distance Transform Watershed | Legland et al., Bioinformatics, 2016. doi: 10.1093/bioinformatics/btw413 | For segmentation of Müller glial stalks in calcium imaging movies | |
| Software, algorithm | FIJI plugin: Correct 3D Drift | Parslow et al., J Vis Exp, 2014. doi: 10.3791/51086. | For 3D registration of glial morphology time courses | |
| Software, algorithm | FIJI plugin: Simple Neurite Tracer | Arshadi et al., Nat Methods, 2021. doi:10.1038/s41592-021-01105-7 | RRID:SCR_016566 | For tracing of dye-filled glial processes and stalks |
| Software, algorithm | R; RStudio | R Project for Statistical Computing; RStudio | RRID:SCR_001905; RRID:SCR_000432 | Used for statistical analyses |
Comparison of intercompartment latency between GCaMP6f-expressing and Cal520-loaded Muller glia.
| Indicator | GCaMP6f | Cal520 | |
|---|---|---|---|
| n wave-assoc. stalk transients | ACSF: 41gabazine: 1953 | ACSF: 38gabazine: 1630 | Pairwise comparison across indicators (Wilcoxon rank-sum) |
| ACSF latency; signed-rank p-value | 0.17 [0.0/0.85]p = 9.0 x 10-3 | 1.35 [1.10/1.69]p = 9.5 x 10-8 | p = 2.8 x 10-6 |
| Gabazine latency; signed-rank p-value | 0.34 [0.0/1.01]p = 3.7 x 10-117 | 1.35 [1.01/2.03]p = 2.0 x 10-259 | p = 8.8 x 10-267 |
Author response table 2.
Analysis of variance (ANOVA) of morphological properties between wild type and β2-nAChR-KO Müller glia. 2-way ANOVA was used to test effects of and interaction between age and genotype on morphological properties of Muller glia in wild type (WT) and β2-nAChR-KO retinas.
| Effects tested | Main: age | Main: genotype | Interaction: genotype*age |
|---|---|---|---|
| # branch points | F = 18.11p = 0.0001 | F = 0.5p = 0.48 | F = 94p = 0.33 |
| stalk length (µm) | F = 0.06p = 0.81 | F = 0.0p = 0.96 | F = 0.0p = 0.99 |
| process length (µm) | F = 9.61p = 0.0027 | F = 0.24p = 0.63 | F = 0.47p = 0.49 |
| # tips | F = 16.69p = 0.0001 | F = 0.04p = 0.84 | F = 1.05p = 0.31 |
| convex hull area (µm2) | F = 1.39p = 0.24 | F = 1.46p = 0.23 | F = 0.73p = 0.40 |
| # branches off stalk | F = 22.08p = 1.12 x 10-5 | F = 8.18p = 0.0055 | F = 0.11p = 0.75 |
Author response table 3.
F-test for unequal variances of morphological properties between wild type and β2-nAChR-KO Müller glia. F-test for unequal variance between groups was used to compare variance of morphological properties of Muller glia at P12 and P30 between wild type and β2-nAChR-KO retinas.
| Effects tested | P12 (F-statistic; p-value) | P30 (F-statistic; p-value) |
|---|---|---|
| # branch points | F = 0.73p = 0.44 | F = 1.84p = 0.24 |
| stalk length (µm) | F = 0.99p = 0.98 | F = 3.14p = 0.06 |
| process length (µm) | F = 0.62p = 0.23 | F = 1.60p = 0.37 |
| # tips | F = 0.89p = 0.77 | F = 1.87p = 0.23 |
| convex hull area (µm2) | F = 0.50p = 0.09 | F = 0.39p = 0.10 |
| # branches off stalk | F = 1.49p = 0.34 | F = 1.43p = 0.49 |
Additional files
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/73202/elife-73202-transrepform1-v2.docx
-
Supplementary file 1
Analysis of differences in lateral process outgrowth between sublayers; separate tests for each age group.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp1-v2.xlsx
-
Supplementary file 2
Analysis of age-dependent differences in lateral process outgrowth within each sublayer.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp2-v2.xlsx
-
Supplementary file 3
Analysis of age-dependent differences in proportion stable processes.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp3-v2.xlsx
-
Supplementary file 4
Analysis of motility in blockers of cytoskeletal rearrangements.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp4-v2.xlsx
-
Supplementary file 5
Analysis of motility in exogenous EGF.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp5-v2.xlsx
-
Supplementary file 6
Analysis of calcium compartmentalization: proportion of ROIs participating in waves.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp6-v2.xlsx
-
Supplementary file 7
Analysis of intercompartment latency in ACSF.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp7-v2.xlsx
-
Supplementary file 8
Proportion of total ROIs participating in retinal waves: ACSF versus pirenzepine.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp8-v2.xlsx
-
Supplementary file 9
Fold-change in proportion ROIs participating in retinal waves: ACSF versus pirenzepine.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp9-v2.xlsx
-
Supplementary file 10
Z scored wave-associated ΔF/F amplitude: ACSF versus pirenzepine.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp10-v2.xlsx
-
Supplementary file 11
Fold-change in wave-associated ΔF/F amplitude: ACSF versus pirenzepine.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp11-v2.xlsx
-
Supplementary file 12
Intercompartment latency in ACSF versus pirenzepine.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp12-v2.xlsx
-
Supplementary file 13
Wave properties in ACSF versus pirenzepine.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp13-v2.xlsx
-
Supplementary file 14
Proportion ROIs participating in retinal waves in ACSF, gabazine, and gabazine + pirenzepine.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp14-v2.xlsx
-
Supplementary file 15
Fold-change in proportion ROIs participating in waves in ACSF versus gabazine, and in gabazine versus gabazine + pirenzepine.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp15-v2.xlsx
-
Supplementary file 16
Z scored wave-associated ΔF/F amplitude in ACSF, gabazine, and gabazine + pirenzepine.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp16-v2.xlsx
-
Supplementary file 17
Fold-change in wave-associated transient amplitude: ACSF versus gabazine; gabazine versus gabazine+ pirenzepine.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp17-v2.xlsx
-
Supplementary file 18
Intercompartment latency in ACSF, gabazine, and gabazine + pirenzepine.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp18-v2.xlsx
-
Supplementary file 19
Wave properties in ACSF, gabazine, and gabazine + pirenzepine.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp19-v2.xlsx
-
Supplementary file 20
Comparisons of proportion stable processes across conditions.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp20-v2.xlsx
-
Supplementary file 21
Comparisons of morphological properties between wild-type (WT) and β2-nAChR-KO Müller glia.
- https://cdn.elifesciences.org/articles/73202/elife-73202-supp21-v2.xlsx





