Glial Nrf2 signaling mediates the neuroprotection exerted by Gastrodia elata Blume in Lrrk2-G2019S Parkinson’s disease
Figures
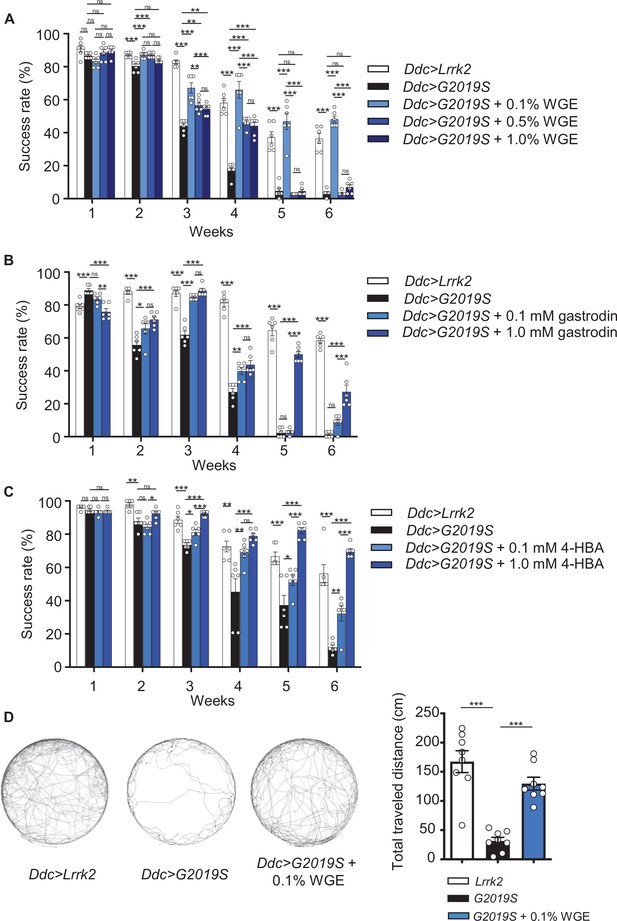
Water extract of Gastrodia elata Blume (WGE) treatment rescues the diminished locomotion of Ddc>G2019S flies.
(A–C) Climbing activities of Ddc>G2019S flies fed on food supplemented with 0.1, 0.5, or 1.0% WGE (A), 0.1 or 1.0 mM gastrodin (B), and 0.1 or 1.0 mM 4-hydroxybenzyl alcohol (4-HBA) (C). Controls are Ddc>Lrrk2 and Ddc>G2019S flies fed regular food. Bar graphs show the percentages of flies (mean ± SEM, N = 6) that climbed above 8 cm within 10 s. (D) Five-minute walking tracks pooled from eight flies each of the Ddc>Lrrk2, Ddc>G2019S, and Ddc>G2019S + 0.1% WGE groups at week 4. Bar graph at right summarizes their walking distances (mean ± SEM, N = 8). One-way analysis of variance (ANOVA) with Tukey’s post-hoc multiple comparison test: *p<0.05, **p<0.01, and ***p<0.001, ns, not significant.
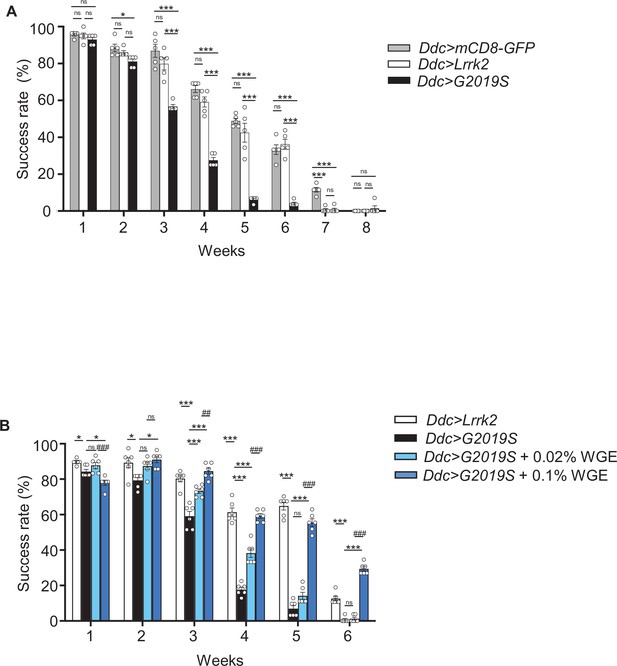
Climbing activity assay of Ddc>mCD8-GFP and Ddc>Lrrk2 flies.
(A) Comparable climbing activities were detected for Ddc>mCD8-GFP and Ddc>Lrrk2 flies. Both exhibited better climbing activities than Ddc>G2019S flies in the climbing assay from weeks 1 to 6. Bar graph shows percentage (mean ± SEM, N = 5) of flies that successfully climbed above 8 cm within 10 s. One-way analysis of variance (ANOVA) and Tukey’s post-hoc multiple comparison test: *p<0.05, ***p<0.001, ns, not significant. (B) Ddc>G2019S flies fed with 0.02 or 0.1% water extract of Gastrodia elata Blume (WGE) exhibited improved climbing activity (mean ± SEM, N = 6). One-way ANOVA and Tukey’s post-hoc multiple comparison test: *p<0.05, ***p<0.001 (relative to Ddc>G2019S); and ##p<0.01, ###p<0.001 (comparing different doses of WGE), ns, not significant.
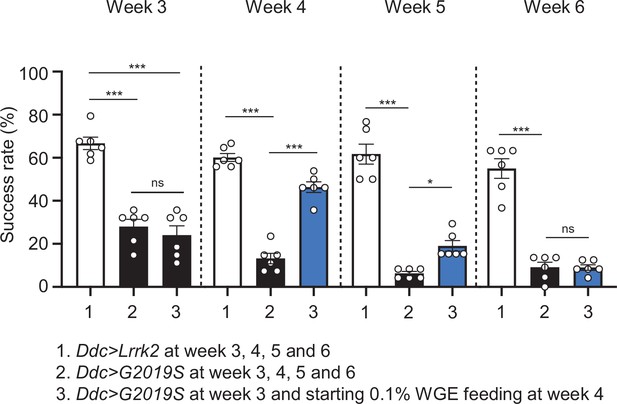
Locomotion improvement of Ddc>G2019S flies starting water extract of Gastrodia elata Blume (WGE) feeding at week 4.
The climbing activities of Ddc>Lrrk2, Ddc>G2019S, and Ddc>G2019S with 0.1% WGE feeding at week 4 were assessed at weeks 3–6. Bar graphs show success rates (mean ± SEM, N = 6) of flies climbing over 8 cm height in 10 s. One-way ANOVA and Tukey’s post-hoc multiple comparison test were performed and statistical significance is shown as ***p<0.001, *p<0.05, ns, not significant.
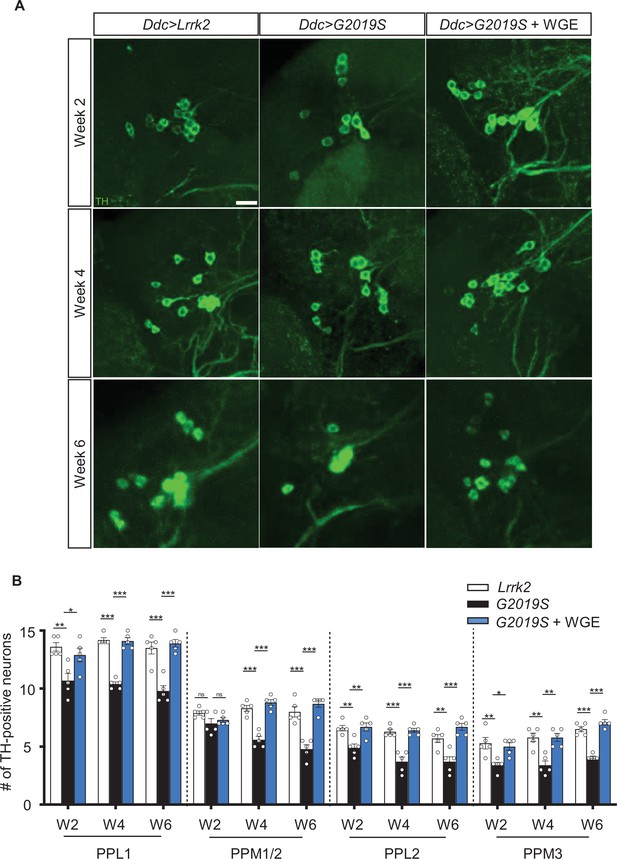
Water extract of Gastrodia elata Blume (WGE) prevents loss of dopaminergic neurons in Ddc>G2019S flies.
(A) Representative adult brain images showing TH-positive dopaminergic neurons in the PPL1 cluster of 2-, 4-, and 6-week-old flies of the Ddc>Lrrk2, Ddc>G2019S, and 0.1% WGE-fed Ddc>G2019S groups. Scale bar: 10 μm. (B) Average numbers (mean ± SEM, N = 5) of TH-positive dopaminergic neurons in the PPL1, PPM1/2, PPL2, and PPL3 clusters per brain hemisphere. One-way ANOVA with Tukey’s post-hoc multiple comparison test: *p<0.05, **p<0.01, ***p<0.001, ns, not significant.
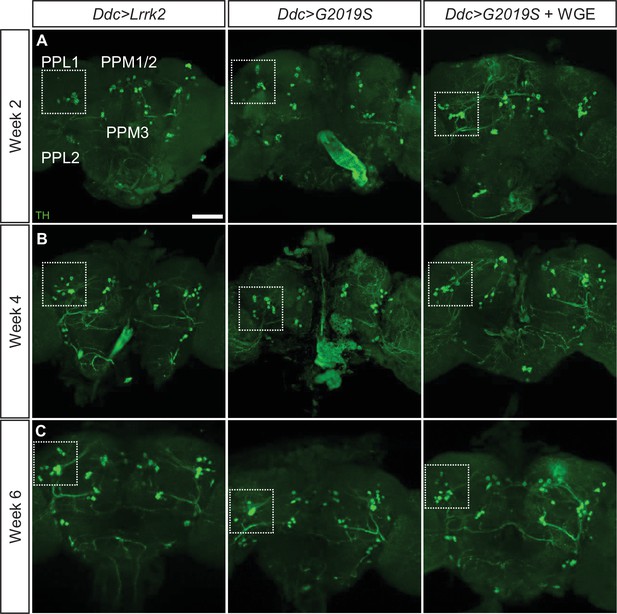
Water extract of Gastrodia elata Blume (WGE) treatment prevents dopaminergic neuron loss in Ddc>G2019S flies.
(A–C) Representative adult whole-brain images for TH staining to reveal dopaminergic neurons in the PPL1, PPL2, PPM1/2, and PPM3 clusters of 2-, 4-, and 6-week-old flies. Scale bar: 40 μm. The images for the PPL1 cluster are shown as enhanced views of the dashed boxes in Figure 2.
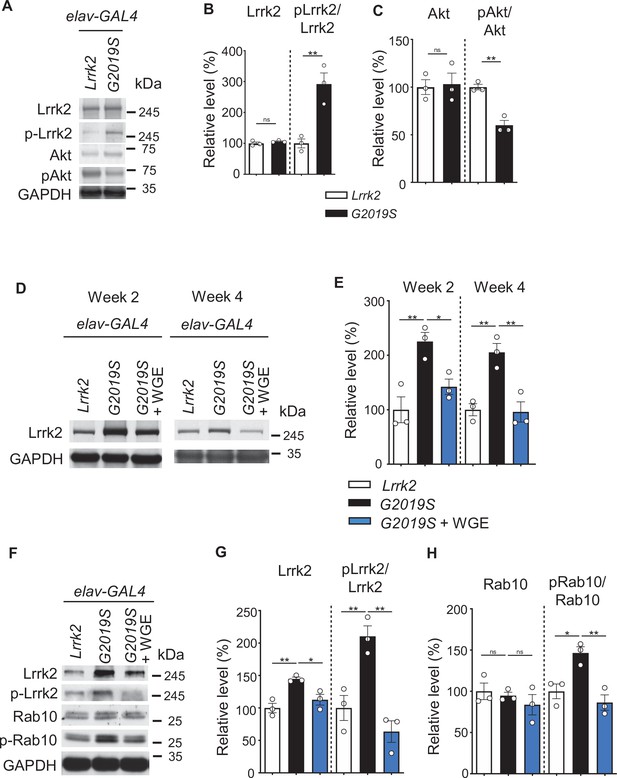
Water extract of Gastrodia elata Blume (WGE) modulates Lrrk2 accumulation and hyperactivation in elav>G2019S flies.
(A) Representative immunoblots of 3-day-old adult brain lysates showing expression levels of Lrrk2, pLrrk2 (phosphorylated at Ser1292), Akt, and pAkt (phosphorylated at Ser505) in elav>Lrrk2 and elav>G2019S flies. (B, C) Quantifications (mean ± SEM, N = 3) of Lrrk2 and pLrrk2/Lrrk2 (B), and Akt and pAkt/Akt (C). (D) Representative immunoblots of 2- and 4-week-old adult brain lysates showing Lrrk2 levels in the fly groups elav>Lrrk2, elav>G2019S, and elav>G2019S fed with 0.1% WGE. (E) Quantification of Lrrk2 levels in 2- and 4-week-old adult brains (mean ± SEM, N = 3). (F) Representative immunoblots of 4-week-old adult brain lysates showing expression levels of Lrrk2, pLrrk2, Rab10, and pRab10 (phosphorylated at Thr73). (G, H) Quantification (mean ± SEM, N = 3) of levels of Lrrk2 and pLrrk2/Lrrk2 (G), and of Rab10 and pRab10/Rab10 (H). One-way ANOVA with Tukey’s post-hoc multiple comparison test: *p<0.05, **p<0.01, ns, not significant.
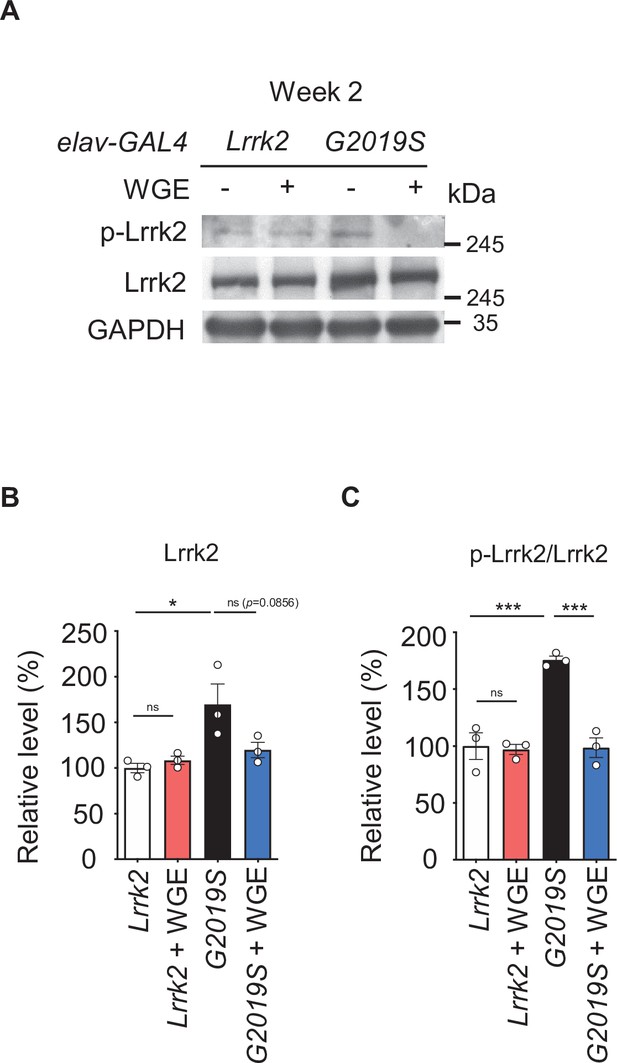
Water extract of Gastrodia elata Blume (WGE) specifically modulates Lrrk2 accumulation and hyperactivation in elav>G2019S but not elav> Lrrk2 flies.
(A) Representative immunoblots of 2-week-old adult brain lysates showing levels of Lrrk2 and pLrrk2 (Ser1292) in elav>Lrrk2, WGE-fed elav>Lrrk2, elav>G2019S, and WGE-fed elav>G2019S flies. (B, C) Quantification (mean ± SEM, N = 3) of Lrrk2 (B) and pLrrk2/Lrrk2 (C) levels. One-way ANOVA and Tukey’s post-hoc multiple comparison test: *p<0.05, ***p<0.001, ns, not significant.
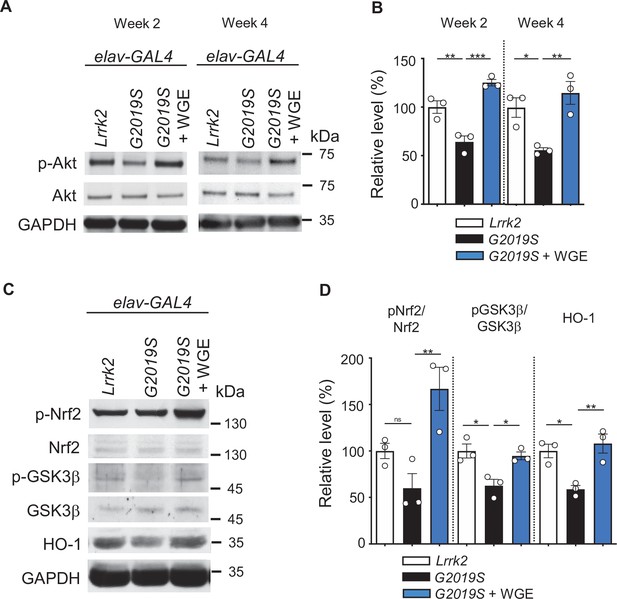
Water extract of Gastrodia elata Blume (WGE) activates the Akt-Nrf2 pathway in elav>G2019S flies.
(A) Representative immunoblots of 2- and 4-week-old adult brain lysates showing levels of Akt and pAkt in brain extracts of elav>Lrrk2, elav>G2019S, and WGE-fed elav>G2019S flies. (B) Quantification (mean ± SEM, N = 3) of pAkt/Akt levels in 2- and 4-week-old adult brains. (C) Representative immunoblots of 4-week-old adult brain lysates showing levels of Nrf2, pNrf2 (phosphorylated at Ser40), GSK3β, pGSK3β (phosphorylated at Ser9), and HO-1 in elav>Lrrk2, elav>G2019S, and WGE-fed elav>G2019S flies. GADPH acted as a loading control in (A) and (C). (D) Quantification (mean ± SEM, N = 3) of relative protein levels to respective Nrf2, GSK3β, and HO-1. One-way ANOVA with Tukey’s post-hoc multiple comparison test (relative to elav>G2019S): *p<0.05, **p<0.01, ***p<0.001, ns, not significant.
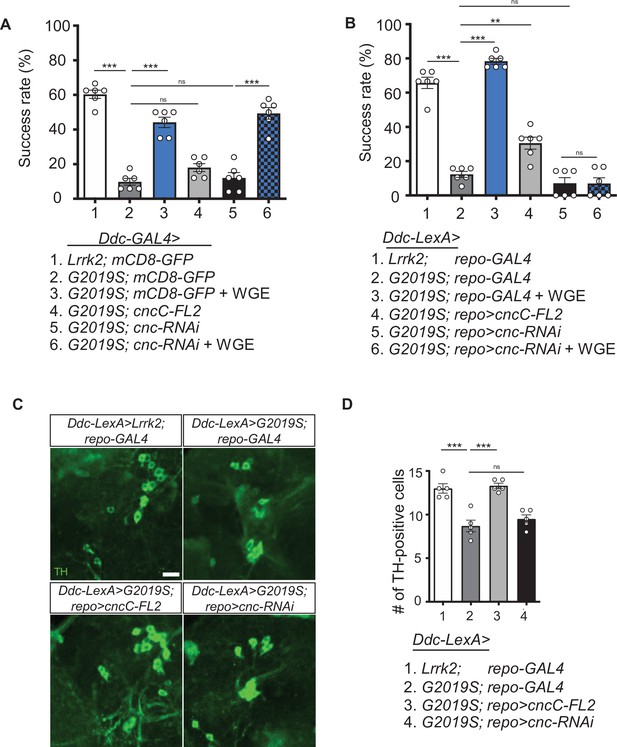
Activation of Nrf2 in glia rescues locomotion defects in Ddc>G2019S flies.
(A, B) Requirement of Nrf2 in glia but not neurons for water extract of Gastrodia elata Blume (WGE)-improved Ddc>G2019S climbing activity. (A) Climbing success rates of flies in which Ddc-GAL4 drives coexpression of Lrrk2-G2019S and mCD8-GFP, cncC-FL2 or cnc-RNAi. As a control line, Ddc-GAL4 drives coexpression of Lrrk2 and mCD8-GFP. (B) Climbing success rates of flies in which Ddc-LexA drives wild-type Lrrk2 or Lrrk2-G2019S expression and repo-GAL4 drives cncC-FL2 or cnc-RNAi expression (mean ± SEM, N = 6 for A and B). WGE was added to food at a concentration of 0.1% (w/w). (C) Adult brain images showing TH-positive dopaminergic neurons in the PPL1 clusters of 6-week-old Ddc-LexA>Lrrk2 or Ddc-LexA>G2019S flies with repo-GAL4 control or repo-GAL4-driven cncC-FL2 or cnc-RNAi expression. Scale bar: 10 μm. (D) Average numbers (mean ± SEM, N = 5) of TH-positive dopaminergic neurons in PPL1 clusters per brain hemisphere are shown. One-way ANOVA with Tukey’s post-hoc multiple comparison test (relative to Ddc>G2019S [A] or Ddc-LexA>G2019S; repo-GAL4 [B, D]): **p<0.01, ***p<0.001, ns, not significant.
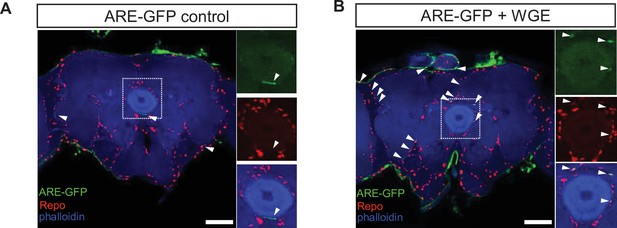
Water extract of Gastrodia elata Blume (WGE) treatment specifically activates glial Nrf2 signals in the ARE-GFP reporter flies.
(A, B) WGE treatment activates glial Nrf2 signals in 1-week-old ARE-GFP reporter flies. ARE-GFP (green) and glial Repo (red) in whole-mount adult brains without (A) or with 0.1% WGE treatment (B) for 5 days. Phalloidin (in blue) reveals brain structures. White arrowheads indicate GFP signals. Scale bar: 50 μm. Dotted boxes are enhanced views and shown as separate channels at right.
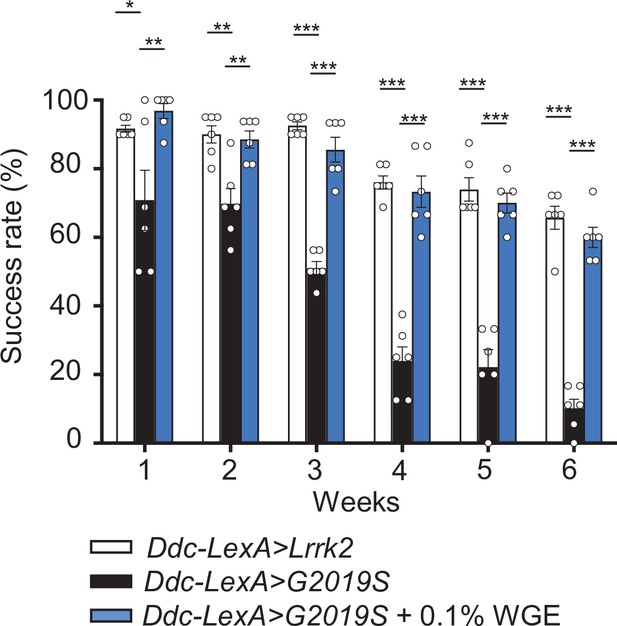
Water extract of Gastrodia elata Blume (WGE) rescues the locomotion defect displayed by Ddc-LexA>G2019S flies.
WGE treatment improves the climbing ability of Ddc-LexA>G2019S flies. Bar graph shows percentage (mean ± SEM, N = 6) of flies that successfully climbed above 8 cm within 10 s. One-way ANOVA and Tukey’s post-hoc multiple comparison test (relative to Ddc-LexA>G2019S): *p<0.05, **p<0.01, ***p<0.001.
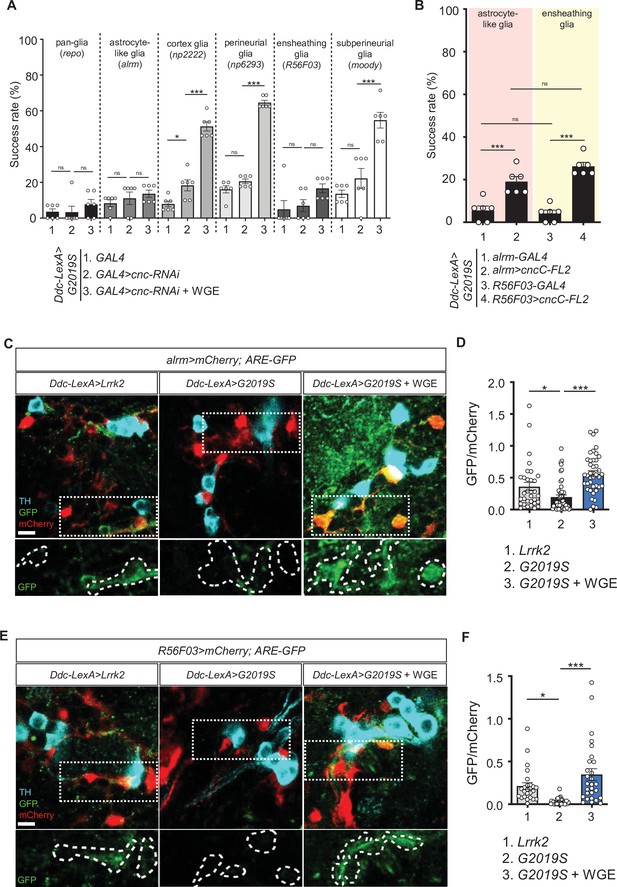
Nrf2 in astrocyte-like and ensheathing glia mediates the effect of water extract of Gastrodia elata Blume (WGE) treatment in Ddc>G2019S flies.
(A) Nrf2 knockdown in astrocyte-like and ensheathing glia abolishes the improved locomotion elicited by WGE treatment in Ddc>G2019S flies. Composite bar graph shows climbing success rates for 6-week-old Ddc-LexA>G2019S flies with cnc-RNAi driven by repo-GAL4 in all glia, alrm-GAL4 in astrocyte-like glia, np2222 in cortex glia, np6293 in perineurial glia, R56F03 in ensheathing glia, and moody-GAL4 in subperineurial glia. (B) Composite bar graph shows climbing success rates (mean ± SEM, N = 6) for 6-week-old Ddc-LexA>G2019S flies with overexpression of Nrf2 in astrocyte-like glia (alrm>cncC-FL2) or ensheathing glia (R56F03>cncC-FL2). One-way ANOVA and Tukey’s post-hoc multiple comparison test (relative to Ddc-LexA>G2019S; GAL4>cnc-RNAi [A] or Ddc-LexA>G2019S; GAL4 [B]): *p<0.05, ***p<0.001, ns, not significant. (C, E) Representative images showing expression of ARE-GFP in astrocyte-like glia (alrm>mCherry) (C) and ensheathing glia (R56F03>mCherry) (E), together with TH-positive dopaminergic neurons in the PPL1 clusters of 6-week-old Ddc-LexA>Lrrk2, Ddc-LexA>G2019S, or WGE (0.1% w/w)-fed Ddc-LexA>G2019S flies. Bar: 5 μm. GFP channels in the dashed boxes are shown as enhanced views in the lower panel, with glial signals labeled by dashed lines. (D, F) Quantifications (mean ± SEM, n > 25 for each genotype) of GFP intensities in astrocyte-like glia (D) or ensheathing glia (F). GFP intensities in glia have been outlined manually using the mCherry-positive signals. One-way ANOVA and Tukey’s post-hoc multiple comparison test (relative to Ddc-LexA>G2019S; alrm>mCherry [D] or Ddc-LexA>G2019S; R56F03>mCherry [F]): *p<0.05, ***p<0.001, ns, not significant.
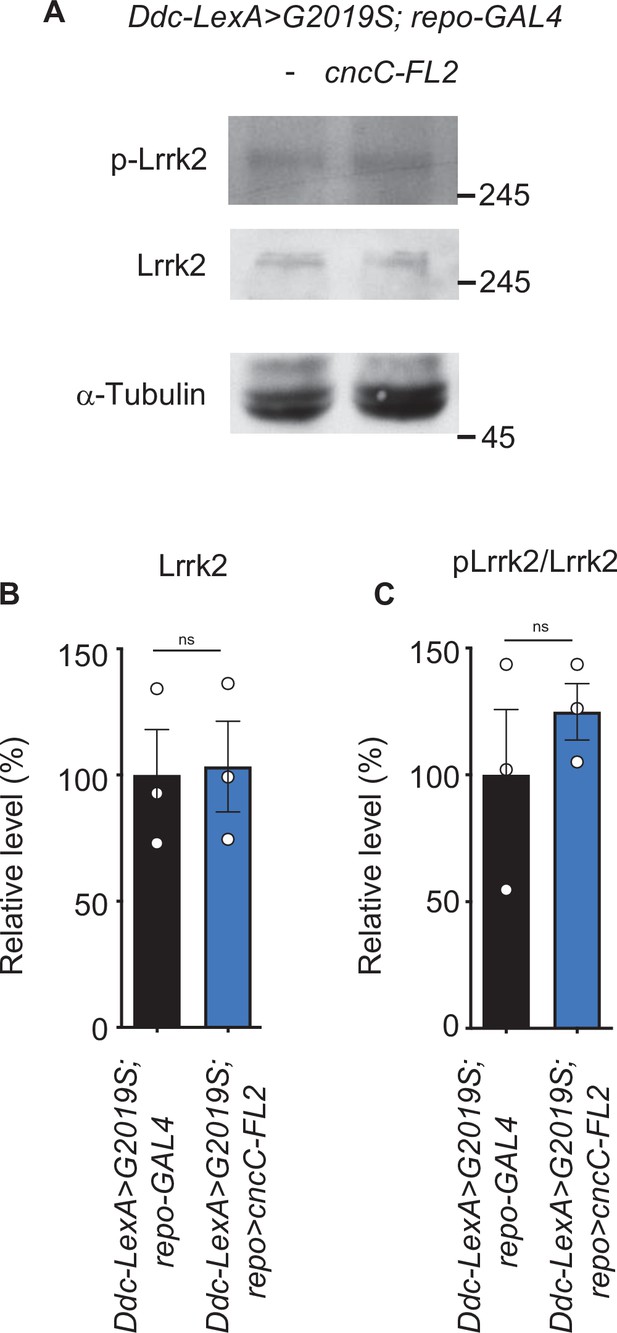
Lrrk2 and pLrrk2 levels are maintained upon glial Nrf2 overexpression.
(A) Representative immunoblots of 2-week-old adult brain lysates showing protein expression levels of Lrrk2 and pLrrk2 (Ser1292) in Ddc-LexA>G2019S; repo-GAL4 and Ddc-LexA>G2019S; repo>cncC-FL2 flies. (B, C) Quantification of Lrrk2 (B) and pLrrk2/Lrrk2 (C) (mean ± SEM, N = 3). Student t-test: ns, not significant.
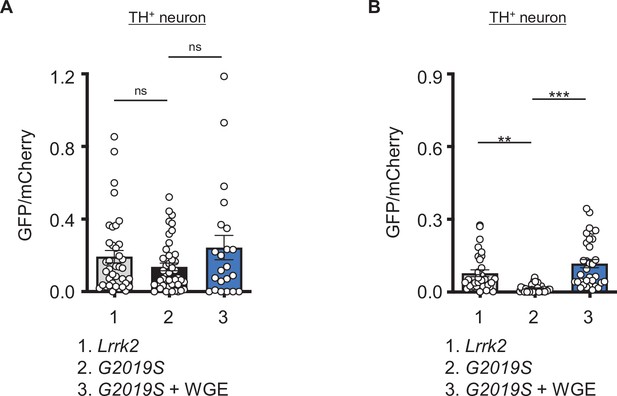
Water extract of Gastrodia elata Blume (WGE) treatment induces a mild Nrf2 activation in dopaminergic neurons in Ddc>G2019S flies.
(A, B) Quantifications (mean ± SEM, n > 20 for each genotype) of GFP intensities in TH-positive dopaminergic neurons nearby the astrocyte-like glia (A) or ensheathing glia (B). GFP intensities were measured within dopaminergic neurons outlined by the TH-positive signals and were normalized to mCherry intensities. One-way ANOVA and Tukey’s post-hoc multiple comparison test with reference to Ddc-LexA>G2019S; alrm>mCherry (A) or Ddc-LexA>G2019S; R56F03>mCherry (B) were performed and shown as **p<0.01, ***p<0.001, ns, not significanct.
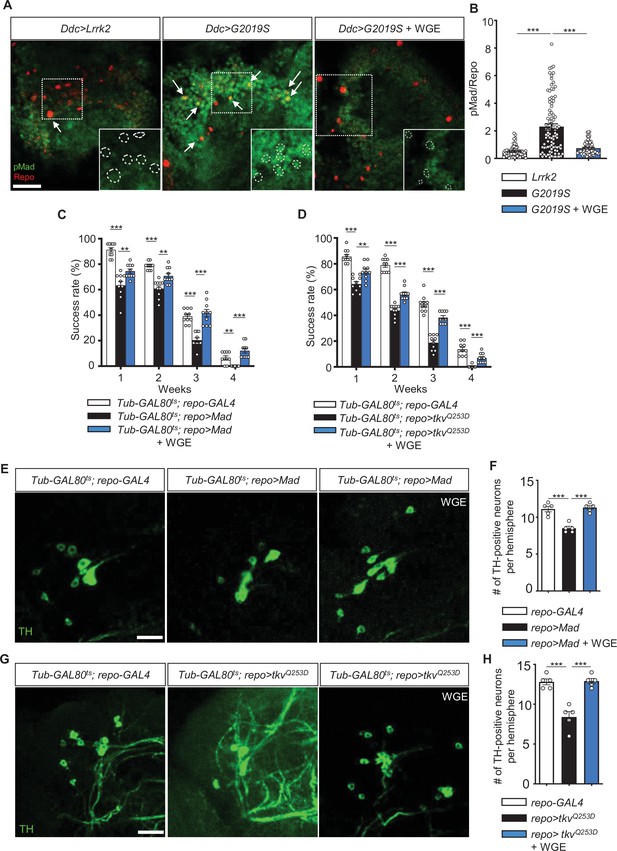
Water extract of Gastrodia elata Blume (WGE) downregulates G2019S-induced BMP/Mad signaling.
(A) Representative images of glial pMad staining in the adult PPL1 clusters of Ddc>Lrrk2, Ddc>G2019S, and WGE-fed Ddc>G2019S flies. White arrows indicate pMad signals co-localized with Repo, with single channels for pMad signals shown as insets. Bar: 10 μm. (B) Quantification (mean ± SEM, n > 60 for each genotype) of pMad signals normalized to Repo levels in glia of the indicated genotypes. One-way ANOVA and Tukey’s post-hoc multiple comparison test (relative to Ddc>G2019S): ***p<0.001. (C, D) Climbing success rates (mean ± SEM, N = 10) at weeks 1–4 demonstrating that WGE treatment rescues locomotion deficits induced by glial overexpression of Mad (C) or tkvQ253D (D) in Tub-GAL80ts; repo-GAL4 flies. One-way ANOVA and Tukey’s post-hoc multiple comparison test (relative to repo>Mad or repo>tkvQQ253D): **p<0.01, ***p<0.001. (E, G) Representative images of 4-week-old adult brain showing TH staining of the PPL1 clusters of Tub-GAL80ts; repo-GAL4, Tub-GAL80ts; repo>Mad, and WGE-fed Tub-GAL80ts; repo>Mad flies (E), and Tub-GAL80ts; repo-GAL4, Tub-GAL80ts; repo>tkvQQ253D, and WGE-fed Tub-GAL80ts; repo>tkvQQ253D flies (G). Bars: 12.5 μm. (F, H) Bar graphs show mean ± SEM (N = 5) of TH-positive dopaminergic neurons in the PPL1 clusters of 4-week-old flies. One-way ANOVA and Tukey’s post-hoc multiple comparison test (relative to Tub-GAL80ts; repo>Mad [F] or Tub-GAL80ts; repo>tkvQQ253D [H]): ***p<0.001.
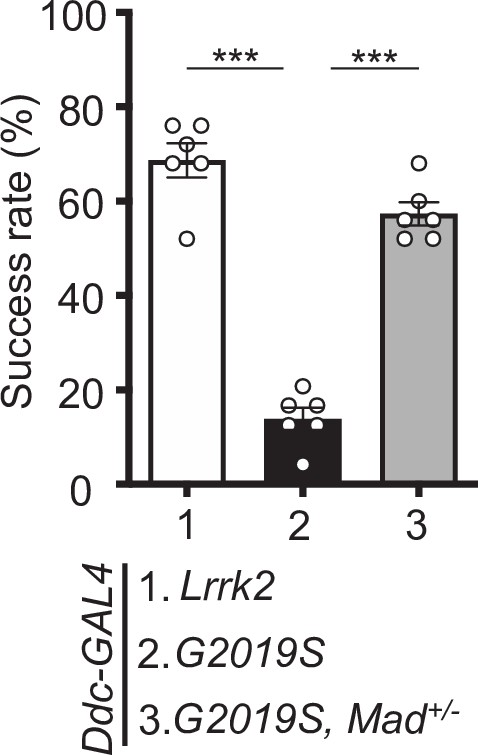
Mad heterozygosity rescues the impaired locomotion of Ddc>G2019S flies.
Removing one copy of Mad improves Ddc>G2019S climbing activity. Bar graphs show climbing success rates (mean ± SEM, N = 6) of 6-week-old Ddc>Lrrk2, Ddc>G2019S, and Ddc>G2019S, Mad+/− flies. One-way ANOVA and Tukey’s post-hoc multiple comparison test (relative to Ddc>G2019S): ***p<0.001.
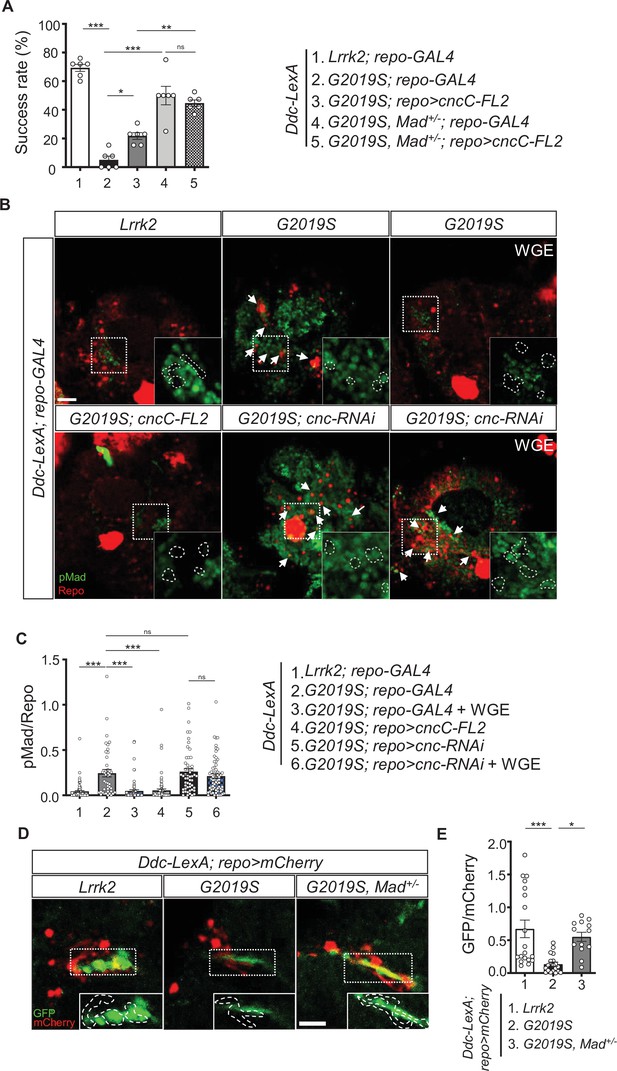
Nrf2 antagonizes BMP/Mad signaling in glia.
(A) Heterozygosity of Mad suppresses G2019S mutation-induced locomotion impairment in a negative geotaxis assay. Bar graph shows percentages (mean ± SEM, N = 6) of 6-week-old flies that climbed above 8 cm within 10 s. One-way ANOVA and Tukey’s post-hoc multiple comparison test: *p<0.05, **p<0.01, ***p<0.001, ns, not significant. (B) Representative images of pMad staining in the adult brains of Ddc-LexA>Lrrk2 or Ddc-LexA>G2019S flies in which repo-GAL4 drives expression of cncC-FL2 or cnc-RNAi. Impact of WGE treatment is shown in the rightmost panels. Arrows indicate pMad and Repo dual-positive cells. Insets are enlarged images of the dashed boxes, and dashed lines encompass Repo-positive cells. Bar: 20 μm. (C) Quantification of the ratio of pMad to Repo (mean ± SEM, N > 40). One-way ANOVA and Tukey’s post-hoc multiple comparison test: ***p<0.001, ns, not significant. (D) Images show pan-glial ARE-GFP expression (repo>mCherry) in Ddc-LexA>Lrrk2, Ddc-LexA>G2019S, and Ddc-LexA>G2019S, Mad+/- fly brains. GFP channels within the dashed boxes are shown as enhanced views in the inset, with glial signals outlined by dashed lines. Bar: 20 μm. (E) Quantification for GFP expression levels (mean ± SEM, n > 13). One-way ANOVA and Tukey’s post-hoc multiple comparison test: *p<0.05, ***p<0.001.
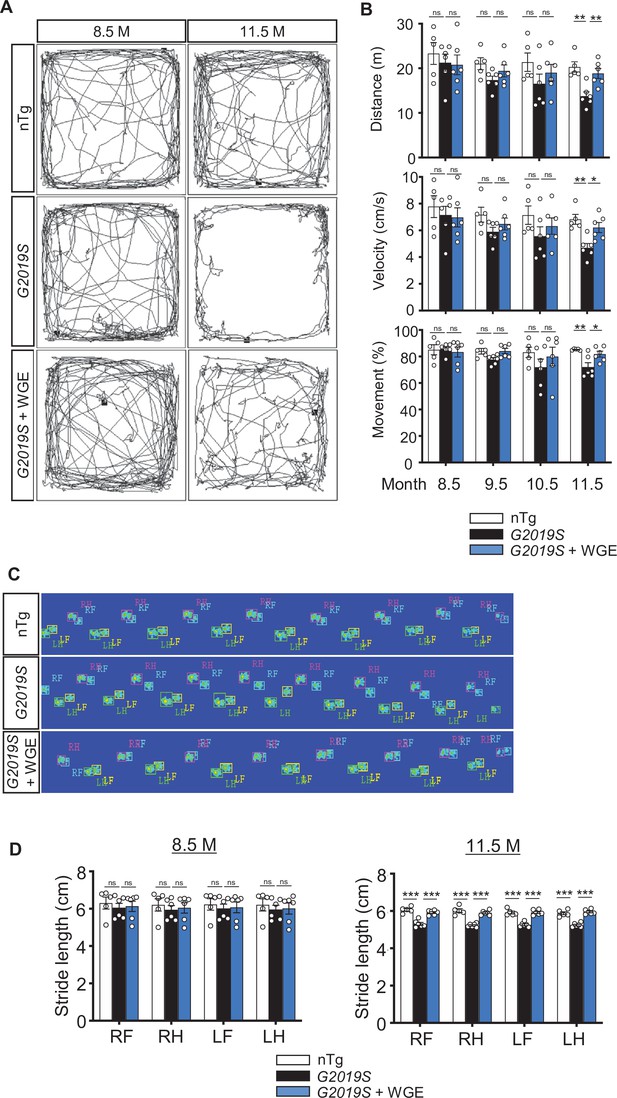
Water extract of Gastrodia elata Blume (WGE) treatment rescues impaired locomotion of LRRK2-G2019S mice.
(A) Video-tracked paths for nTg, LRRK2-G2019S, and WGE-fed LRRK2-G2019S mice (8.5 and 11.5 months old) during the open-field test. (B) Quantification of total distance (m), average velocity (cm/s), and percentage moving time for 8.5-, 9.5-, 10.5-, and 11.5-month-old mice. (C) Captured and converted images of single stance for each paw of 11.5-month-old nTg, LRRK2-G2019S, and WGE-fed LRRK2-G2019S mice in a catwalk analysis. (D) Quantification of stride length for each paw of 8.5- and 11.5-month-old mice. Data in (B) and (D) are presented as mean ± SEM (nTg; N = 5, G2019S; N = 6, WGE-fed G2019S; N = 6). One-way ANOVA and Tukey’s post-hoc multiple comparison test: *p<0.05, **p<0.01, ***p<0.001, ns, not significant. RF, right front; RH, right hind; LF, left front; LH, left hind.
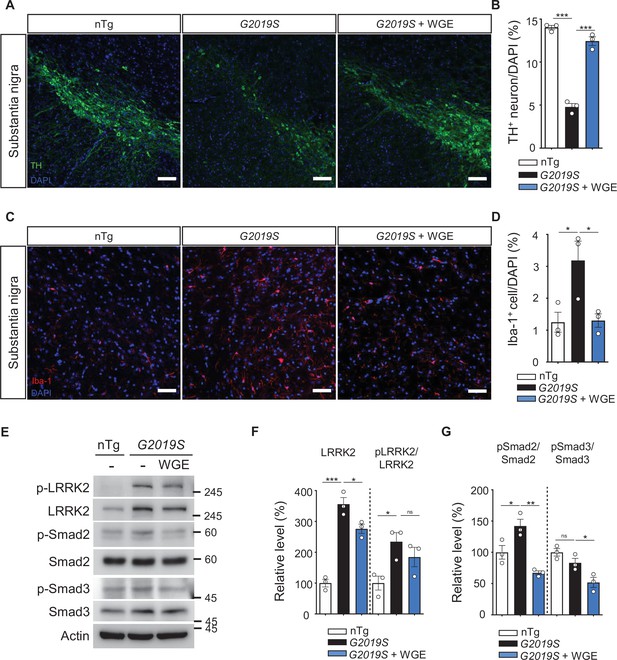
Water extract of Gastrodia elata Blume (WGE) prevents dopaminergic neuron loss, microglial activation, and phosphorylation of LRRK2, Smad2, and Smad3.
(A, C) Representative images showing TH-positive dopaminergic neurons (A) or Iba-1-positive microglia (C) in the substantia nigra of 11.5-month-old nTg, LRRK2-G2019S, and WGE-fed LRRK2-G2019S mice. Bars: 100 μm in (A) and 50 μm in (C). (B, D) Quantification of numbers of TH-positive (B) or Iba-1-positive cells (D) relative to DAPI cells. (E) Representative immunoblots of nigrostriatal lysates prepared from 11.5-month-old nTg, LRRK2-G2019S, and WGE-fed LRRK2-G2019S mice reveal expression levels of LRRK2, pLRRK2 (Ser1292), Smad2, pSmad2 (Ser465 and Ser467), Smad3, and pSmad3 (Ser423 and Ser425). Actin acted as a loading control. (F, G) Quantifications of LRRK2 and pLRRK2/LRRK2 (F), as well as pSmad2/Smad2 and pSmad3/Smad3 (G). Data in (B, D, F, G) are presented as mean ± SEM (N = 3). One-way ANOVA and Tukey’s post-hoc multiple comparison test: *p<0.05, **p<0.01, ***p<0.001, ns, not significant.
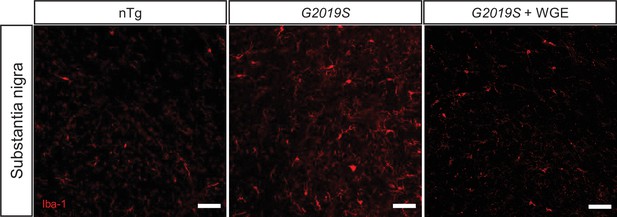
Water extract of Gastrodia elata Blume (WGE) suppresses microglia activation in LRRK2-G2019S mice.
Single-channel images of Figure 10C show Iba-1-positive microglia in the substantia nigra of 11.5-month-old nTg, LRRK2-G2019S, and WGE-fed LRRK2-G2019S mice. Bars: 50 μm.
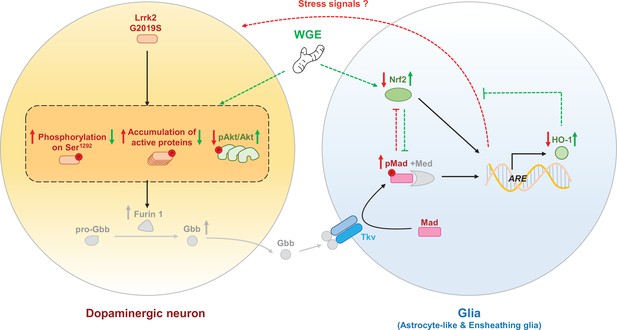
The proposed model of water extract of Gastrodia elata Blume (WGE) in the G2019S-induced neurodegeneration.
Accumulation of the hyperactivated G2019S mutant protein enhances the BMP ligand (Gbb) maturation via upregulation of Furin 1 translation in dopaminergic neurons. Secreted Gbb binds to the BMP receptor, Tkv, and turns on Mad signaling in glia. The G2019S mutation also decreases the Nrf2 activity in the brain, particularly in glia. Both upregulated Mad and downregulated Nrf2 pathways contribute to neurodegeneration. WGE feeding suppresses G2019S hyperactivation in neurons and restores Nrf2 activity mostly in the astrocyte-like and ensheathing glia. WGE-elevated Nrf2 activity in glia antagonizes the BMP/Mad signaling and initiates Nrf2/HO-1 axis in glia, attenuating the stress signals from glia and promoting neuroprotection. Red and green solid arrows (→) indicate the observed effects exerted by G2019S overexpression and WGE feeding, respectively, in the present study. Red and green dashed arrows (-->) indicate the proposed actions trigged by Mad signaling and WGE feeding, respectively. Red and green blunt-ended lines (---|) indicate the proposed inhibitions by Mad and Nrf2 overexpression, respectively. (P) Indicates phosphorylation. The Furin 1-mediated Gbb pathway labeled in gray is modified from Figure 6 of Maksoud et al., 2019.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (Drosophila melanogaster) | UAS-mCD8-GFP | Bloomington Drosophila Stock Center | BDSC Cat# 5137; RRID:BDSC_5137 | |
| Genetic reagent (D. melanogaster) | UAS-cncTRiP | Bloomington Drosophila Stock Center | BDSC Cat# 25984; RRID:BDSC_25984 | |
| Genetic reagent (D. melanogaster) | UAS-tkvQ253D | Bloomington Drosophila Stock Center | BDSC Cat# 36536; RRID:BDSC_36536 | |
| Genetic reagent (D. melanogaster) | UAS-Flag- LRRK2-WT | Lin et al., 2010 | N/A | |
| Genetic reagent (D. melanogaster) | UAS-Flag- LRRK2-G2019S | Lin et al., 2010 | N/A | |
| Genetic reagent (D. melanogaster) | UAS-cncC-FL2 | Sykiotis and Bohmann, 2008 | N/A | |
| Genetic reagent (D. melanogaster) | UAS-Mad | Takaesu et al., 2006 | N/A | |
| Genetic reagent (D. melanogaster) | elav-GAL4 | Bloomington Drosophila Stock Center | BDSC Cat# 8760; RRID:BDSC_8760 | |
| Genetic reagent (D. melanogaster) | repo-GAL4 | Bloomington Drosophila Stock Center | BDSC Cat# 7215; RRID:BDSC_7215 | |
| Genetic reagent (D. melanogaster) | alrm-GAL4 | Bloomington Drosophila Stock Center | BDSC Cat# 67032; RRID:BDSC_67032 | |
| Genetic reagent (D. melanogaster) | R56F03-GAL4 | Bloomington Drosophila Stock Center | BDSC Cat# 39157; RRID:BDSC_39157 | |
| Genetic reagent (D. melanogaster) | NP2222-GAL4 | Bloomington Drosophila Stock Center | BDSC Cat# 112830; RRID:BDSC_112830 | |
| Genetic reagent (D. melanogaster) | NP6293-GAL4 | Kyoto Stock Center | DGRC Cat# 105188;RRID:Kyoto Stock Center_105188 | |
| Genetic reagent (D. melanogaster) | Ddc-GAL4 | Sang et al., 2007 | N/A | |
| Genetic reagent (D. melanogaster) | moody-GAL4 | Bainton et al., 2005 | N/A | |
| Genetic reagent (D. melanogaster) | Tub-GAL80ts | Bloomington Drosophila Stock Center | BDSC Cat# 7108; RRID:BDSC_7108 | |
| Genetic reagent (D. melanogaster) | Ddc-LexA | Bloomington Drosophila Stock Center | BDSC Cat# 54218; RRID:BDSC_54218 | |
| Genetic reagent (D. melanogaster) | MadK00237 | Bloomington Drosophila Stock Center | BDSC Cat# 10474; RRID:BDSC_10474 | |
| Genetic reagent (D. melanogaster) | ARE-GFP | Sykiotis and Bohmann, 2008 | N/A | |
| Genetic reagent (D. melanogaster) | LexAop-LRRK2-WT | This paper | N/A | See Materials and methods, ‘Drosophila stocks and maintenance’ |
| Genetic reagent (D. melanogaster) | LexAop-LRRK2- G2019S | This paper | N/A | See Materials and methods, ‘Drosophila stocks and maintenance’ |
| Genetic reagent (Mus musculus) | FVB/NJ | The Jackson Laboratory | JAX stock #001800 | |
| Genetic reagent (M. musculus) | FVB/N-Tg(LRRK2* G2019S)1Cjli/J | The Jackson Laboratory | JAX stock #009609 | |
| Antibody | Anti-TH(mouse monoclonal) | Immunostar | Cat# 22941; RRID:AB_572268 | IF (1:1000) |
| Antibody | Anti-TH(rabbit polyclonal) | Millipore | Cat# AB152;RRID:AB_390204 | Mouse-IHC (1:200) |
| Antibody | Anti-Repo(mouse monoclonal) | Hybridoma Bank DSHB | Cat# 8D12; RRID:AB_528448 | IF (1:500) |
| Antibody | Anti-GFP(chicken polyclonal) | Abcam | Cat# ab13970; RRID:AB_300798 | IF (1:10000) |
| Antibody | Anti-LRRK2(rabbit monoclonal) | Abcam | Cat# ab133474;RRID:AB_2713963 | Fly-WB (1:1000) mouse-WB(1:5000) |
| Antibody | Anti-LRRK2 (phospho Ser1292)(rabbit monoclonal) | Abcam | Cat# ab203181 | Fly-WB (1:500) mouse-WB(1:1000) |
| Antibody | Anti-Akt(rabbit monoclonal) | Cell Signaling | Cat# 4691;RRID:AB_915783 | WB (1:1000) |
| Antibody | Anti-phospho- Drosophila Akt (Ser505)(rabbit polyclonal) | Cell Signaling | Cat# 4054;RRID:AB_331414 | WB (1:500) |
| Antibody | Anti-Nrf2(rabbit polyclonal) | ThermoFisherScientific | Cat# 710574;RRID:AB_2532742 | WB (1:1000) |
| Antibody | Anti-phospho- Nrf2 (Ser40)(rabbit polyclonal) | ThermoFisherScientific | Cat# PA5-67520;RRID:AB_2691678 | WB (1:1000) |
| Antibody | Anti-HO-1-1(mouse monoclonal) | ThermoFisherScientific | Cat# MA1-112;RRID:AB_2536823 | WB (1:1000) |
| Antibody | Anti-GAPDH(rabbit polyclonal) | GeneTex | Cat# GTX100118;RRID:AB_1080976 | WB (1:5000) |
| Antibody | Anti-alpha tubulin(rabbit polyclonal) | Cell Signaling | Cat# 2144;RRID:AB_2210548 | WB (1:10,000) |
| Antibody | Anti-Smad2(rabbit monoclonal) | Cell Signaling | Cat# 5339;RRID:AB_10626777 | Mouse-WB (1:1000) |
| Antibody | Anti-phospho-Smad2 (Ser465/467)(rabbit monoclonal) | Cell Signaling | Cat# 3108;RRID:AB_490941 | Mouse-WB (1:1000) |
| Antibody | Anti-Smad3(rabbit monoclonal) | Cell Signaling | Cat# 9523;RRID:AB_2193182 | Mouse-WB (1:1000) |
| Antibody | Anti-Smad3 (phospho S423 + S425)(rabbit monoclonal) | Abcam | Cat# ab52903; RRID:AB_882596 | IF (1:250) mouse-WB (1:1000) |
| Antibody | Anti-beta actin(mouse monoclonal) | Sigma-Aldrich | Cat# A5441;RRID:AB_476744 | Mouse-WB (1:5000) |
| Antibody | Anti-Iba-1(rabbit polyclonal) | GeneTex | Cat# GTX100042;RRID:AB_1240434 | Mouse-IHC (1:200) |
| Antibody | Anti-mouse Alexa 488(goat polyclonal) | Invitrogen | Cat# A28175;RRID:AB_2536161 | IF (1:500) |
| Antibody | Anti-rabbit Alexa 488(goat polyclonal) | Invitrogen | Cat# A27034;RRID:AB_2536097 | IF (1:500) |
| Antibody | Anti-rabbit DyLight 488(goat polyclonal) | ThermoFisherScientific | Cat# 35552;RRID:AB_844398 | Mouse-IHC (1:300) |
| Antibody | Anti-rabbit Alexa 546(goat polyclonal) | ThermoFisherScientific | Cat# A-11035;RRID:AB_2534093 | Mouse-IHC (1:200) |
| Antibody | Anti-mouse FITC(goat polyclonal) | Jackson ImmunoResearch | Cat#115-095-003;RRID:AB_2338589 | IF (1:500) |
| Antibody | Anti-mouse Cy3(goat polyclonal) | Jackson ImmunoResearch | Cat# 115-165-003;RRID:AB_2338680 | IF (1:500) |
| Antibody | Anti-rat IgG Cy5(goat polyclonal) | Invitrogen | Cat# A10525;RRID:AB_2534034 | IF (1:500) |
| Antibody | Anti-rabbit peroxidase(goat polyclonal) | Jackson ImmunoResearch | Cat# 111-035-144;RRID:AB_2307391 | WB (1:7500) |
| Antibody | Anti-mouse peroxidase (goat polyclonal) | Jackson ImmunoResearch | Cat# 115-035-003;RRID:AB_10015289 | WB (1:7500) |
| Antibody | Anti-mouse peroxidase(goat polyclonal) | GeneTex | Cat# GTX213111-01;RRID:AB_10618076 | Mouse-WB (1:5000) |
| Antibody | Anti-rabbit peroxidase(goat polyclonal) | GeneTex | Cat# GTX213110-01;RRID:AB_10618573 | Mouse-WB (1:5000) |
| Chemical compound, drug | Phalloidin-TRITC | Sigma-Aldrich | Cat# P1951;RRID:AB_2315148 | IF (1:5000) |
| Chemical compound, drug | WGE | Lin et al., 2016a;Lin et al., 2018 | N/A | KO DA Pharmaceutical Co. Ltd. |
| Chemical compound, drug | Gastrodin | Lin et al., 2016b | N/A | Wuhan YC Fine Chemical Co. |
| Chemical compound, drug | 4-HBA | Sigma-Aldrich | Cat# H20806-10G | |
| Recombinant DNA reagent(Homo sapiens) | pDEST53-LRRK2-WT (plasmid) | Addgene | Addgene plasmid # 25044;RRID:Addgene_25044 | |
| Recombinant DNA reagent(H. sapiens) | pDEST53-LRRK2-G2019S (plasmid) | Addgene | Addgene plasmid # 25045; RRID:Addgene_25045 | |
| Recombinant DNA reagent(D. melanogaster) | pJFRC19-13XLexAop2-IVS- myr::GFP (plasmid) | Addgene | Addgene plasmid # 26224; RRID:Addgene_26224 | |
| Software, algorithm | ImageJ | PMID:22930834 | https://imagej.nih.gov/ij;RRID:SCR_003070 | |
| Software, algorithm | Ctrax | PMID:19412169 | N/A | |
| Software, algorithm | CatWalk XT | Lin et al., 2020 | N/A | Noldus Information Technology |
| Software, algorithm | Prism 6 | GraphPad | RRID:SCR_002798 | |
| Other | DAPI | GeneTex | Cat# GTX30920 |
Additional files
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/73753/elife-73753-transrepform1-v2.pdf
-
Source data 1
Statistical information.
The file includes all data and the statistical analyses in this article.
- https://cdn.elifesciences.org/articles/73753/elife-73753-supp1-v2.xlsx
-
Source data 2
Data—western blotting.
The file includes the uncropped images of the western blotting in this article.
- https://cdn.elifesciences.org/articles/73753/elife-73753-supp2-v2.pdf





