A nanocompartment system contributes to defense against oxidative stress in Mycobacterium tuberculosis
Figures
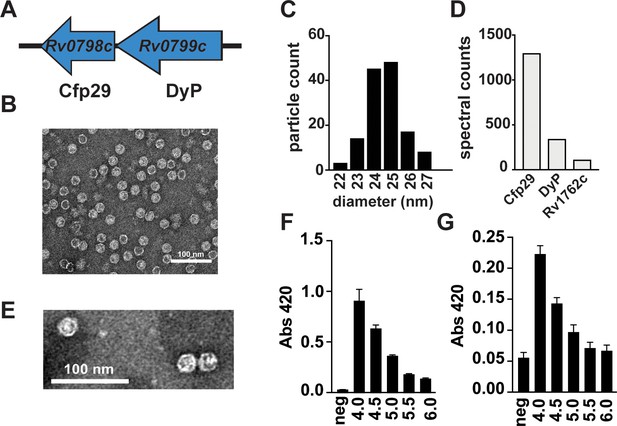
Mtb produces endogenous nanocompartments that package a peroxidase.
(A) Schematic of the nanocompartment operon in Mtb that encodes the encapsulin shell protein (Cfp29) and the dye-decoloring peroxidase cargo protein (DyP). (B) Transmission electron microscopy (TEM) of Cfp29 encapsulin proteins purified following heterologous expression of the Mtb nanocompartment operon in E. coli. (C) Size distribution of Cfp29 protomers purified from E. coli. (D) Peptide counts from mass spectrometry analysis of endogenous nanocompartments purified from Mtb. (E) TEM of endogenous nanocompartments purified from Mtb. Peroxidase activity of (F) unencapsulated and (G) encapsulated DyP (5 nM) using ABTS (480 nM) as a substrate in the presence of H2O2 (480 nM) at varying pH levels (4.0–6.0) as reported by a change in the absorbance at 420 nm. neg, no added enzyme.

Nanocompartment purification and complementation strategies.
(A) Coomassie-stained SDS-PAGE of fractions collected during purification of nanocompartments heterologously expressed in E. coli: (1) ultracentrifugation pellet post-CellLytic B solubilization, (2) size-exclusion chromatography input, (3) lane 1 diluted, (4) lane 2 diluted, and (5) encapsulin fraction from size exclusion. (B) Coomassie-stained SDS-PAGE of sucrose fractions collected during purification of nanocompartments from wild-type Mtb lysates: (1) fraction containing assembled encapsulin nanocompartment complexes, (5) ladder (6), lane 1 boiled in SDS for 30 min to dissociate encapsulin nanocompartment into monomers. (C) DyP samples with or without addition of hemin were analyzed by SDS-PAGE. Samples were loaded either in their unboiled native state (left half) or heat-denatured by boiling at 95°C for 15 min. Addition of hemin yields a tetrameric DyP at 144 kDa. (D) Western blot for Cfp29 (arrow) and Ag85A control (star) from wild-type Mtb (lane 1) and Δoperon mutant (lane 2) lysates. (E) Complementation strategy schematic for DyP::Tn mutants (top). DyP::Tn mutants were transformed with ATc-inducible complementation constructs encoding the unencapsulated cargo protein (DyP), the encapsulin shell protein (Cfp29), or the nanocompartment operon (Operon). Lysates from each strain were used for nanocompartment purification. Sucrose fractions containing high molecular weight Cfp29 protomers were identified in complemented strains expressing the encapsulin shell and the operon (middle) and were analyzed using TEM (bottom).
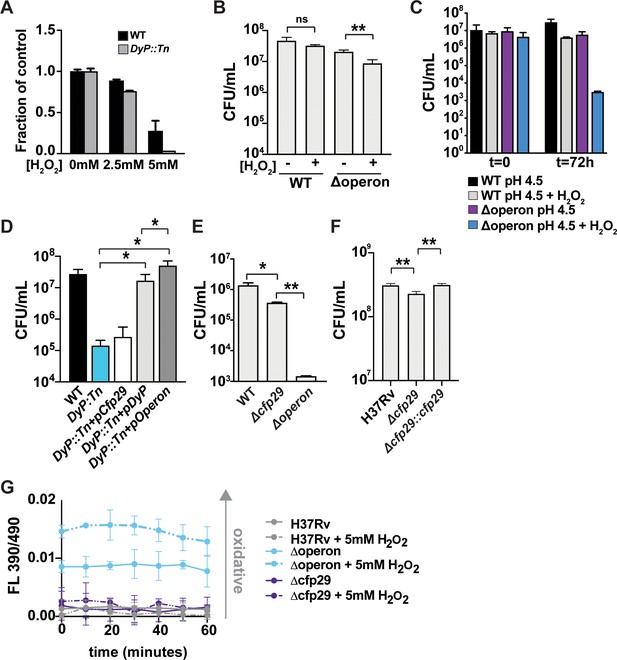
Nanocompartments protect Mtb from oxidative stress in acidic environments.
(A) OD600 measurements of wild-type and DyP::Tn Mtb grown in 7H9 medium following exposure to H2O2 for 96 hr. Values reported are normalized to the untreated controls. CFU enumeration of wild-type and Mtb nanocompartment mutants grown in (B) standard 7H9 medium (pH 6.5) and (C–F) acidified 7H9 medium (pH 4.5) following exposure to oxidative stress (2.5 mM H2O2) for 72 hr. (G) Fluorescence emissions of wild-type and Δoperon Mtb expressing mrx1-roGFP exposed to 5 mM H2O2 at pH 4.5 in 7H9 medium for 60 min. Data are reported as a ratio of fluorescence emissions following excitation at 490 nm and 390 nm. Figures are representative of at least two (E, F) or three (A–D, G) independent experiments. p-Values were determined using unpaired t-test. *p<0.05, **p<0.01.
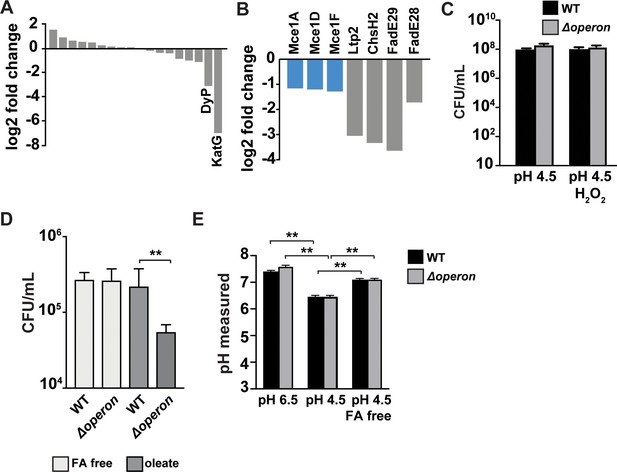
Susceptibility of Mtb nanocompartment mutants to oxidative and acid stress is mediated by free fatty acids.
(A) Transposon sequencing (Tn-seq) data showing normalized sequence reads per gene for all putative Mtb peroxidases, catalases, and superoxide dismutases and (B) lipid and cholesterol metabolism Mtb mutants that were significantly attenuated following 72 hr exposure to 2.5 mM H2O2 at pH 4.5. (C) CFU enumeration of wild-type Mtb and Δoperon mutants following 24 hr exposure to 2.5 mM H2O2 at pH 4.5 in Sauton’s minimal medium and (D) 72 hr exposure to 2.5 mM H2O2 at pH 4.5 in 7H9 medium prepared using fatty acid (FA)-free bovine serum albumin (BSA) ± oleic acid (150 µM). (E) Intrabacterial pH measurements of wild-type and Δoperon Mtb expressing pUV15-pHGFP following 20 min exposure to 5 mM H2O2 at pH 6.5 or pH 4.5. 7H9 medium was prepared with standard BSA or FA-free BSA. Figures are representative of at least two (D) or three (A–C, E) independent experiments. p-Values were determined using an unpaired t-test. *p<0.05, **p<0.01.
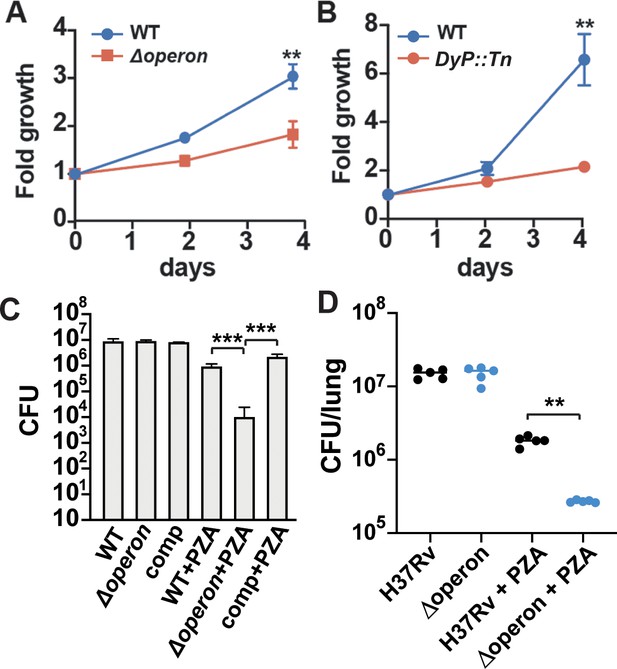
Nanocompartment mutants are attenuated for survival in macrophages and are more susceptible to pyrazinamide (PZA) treatment.
CFU enumeration of wild-type Mtb and (A) Δoperon or (B) DyP::Tn mutants during infection of murine bone marrow-derived macrophages. Macrophages were infected with a bacterial MOI of 1, and CFUs were enumerated immediately following phagocytosis and at days 2 and 4. Error bars are SD from four replicate wells. (C) CFU enumeration of wild-type and Mtb Δoperon mutants following 72 hr exposure to PZA (24 μg/mL) and H2O2 (2.5 mM) in acidified 7H9 medium (pH 5.5). Comp = Δoperon + pOperon. (D) Infection of BALB/C mice with WT and Δoperon mutant with and without treatment with 150 mg/kg PZA. CFU at 35 days post infection in the lung is shown. (A) and (B) are representative of at least five independent experiments; (C) is representative of three experiments and (D) is representative of two experiments. p-Values were determined using an unpaired t-test. **p<0.01; ***p<0.001.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Mycobacterium tuberculosis) | Dyp | GenBank | Gene ID: 885388, Rv0799c | |
| Gene (My. tuberculosis) | Cfp29 | GenBank | Gene ID: 885460, Rv0798c | |
| Strain, strain background (Mus musculus) | BALB/C | The Jackson Laboratory | Stock no: 000651 | |
| Strain, strain background (M. tuberculosis) | H37Rv | Eric Rubin Lab, Harvard School of Public Health | ||
| Strain, strain background (Escherichia coli) | BL21 (DE3) LOBSTR | kerafast | Cat# EC1002 | |
| Genetic reagent (M. tuberculosis) | DyP::Tn | Broad Institute, Hung Lab | Rv0799c::Tn | |
| Genetic reagent (M. tuberculosis) | Δoperon | This study | ΔRv0799c-Rv0798c | See Materials and methods |
| Genetic reagent (M. tuberculosis) | ΔCfp29 | This study | ΔRv0798c | See Materials and methods |
| Genetic reagent (M. tuberculosis) | ΔDyP | This study | ΔRv0799c | See Materials and methods |
| Recombinant DNA reagent | Pet14b | Novagen | Cat# 69660-3 | |
| Recombinant DNA reagent | pUV15tetORm | Addgene | Cat# 17975 | AHT-inducible construct for all complementation |
| Recombinant DNA reagent | pKL4 | This study | Rv0798c cloned into pUV15tetORm (with KanR) | |
| Recombinant DNA reagent | pKL5 | This study | Rv0799c cloned into pUV15tetORm (with KanR) | |
| Recombinant DNA reagent | pKL6 | This study | Operon cloned into pUV15tetORm (with KanR) | |
| Recombinant DNA reagent | pKL14 | This study | Operon cloned into pUV15tetORm (with HygR) | |
| Recombinant DNA reagent | pKL15 | This study | Rv0798c cloned into pUV15tetORm (with HygR) | |
| Recombinant DNA reagent | pKL16 | This study | Rv0799c cloned into pUV15tetORm (with HygR) | |
| Recombinant DNA reagent | pUV15 pHGFP HygR: | Addgene | Cat# 70045 | Rv0799c cloned into pUV15tetORm (with HygR) |
| Recombinant DNA reagent | pMV762-mrx1-roGFP2 | Amit Singh, ICGEB, India | PMC3907381 | |
| Antibody | Anti-Mtb Cfp29 | Rabbit polyclonal | Produced by GenScript USA, see Materials and methods | 1:10,000 |
| Antibody | HRP | Goat anti-rabbit polyclonal | Santa Cruz Biotechnology sc-2030 | 1:5000 |
| Chemical compound, drug | 3-Ethylbenzothia zoline-6-sulfonic acid | Millipore Sigma | Cat# 10102946001 |
Additional files
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/74358/elife-74358-transrepform1-v2.docx
-
Supplementary file 1
Mtb transposon sequencing screen in acidified broth with perxoide stress.
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp1-v2.xlsx
-
Source data 1
Original scanned image of the gel depicting Figure 1—figure supplement 1A.
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp2-v2.zip
-
Source data 2
Scanned image of the gel depicting Figure 1—figure supplement 1A.
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp3-v2.zip
-
Source data 3
Original scanned image of the gel depicting Figure 1—figure supplement 1B.
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp4-v2.zip
-
Source data 4
Original scanned image of the gel depicting Figure 1—figure supplement 1B.
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp5-v2.zip
-
Source data 5
Original scanned image of the gel depicting Figure 1—figure supplement 1C.
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp6-v2.zip
-
Source data 6
Original image capture of PVDF membrane to show the molecular weight ladder (Figure 1—figure supplement 1D).
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp7-v2.zip
-
Source data 7
Original fluorescence image of western blot with Ag85B antibody (Figure 1—figure supplement 1D).
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp8-v2.zip
-
Source data 8
Original fluorescence image of western blot with Cfp29 antibody (Figure 1—figure supplement 1D).
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp9-v2.zip
-
Source data 9
Western images from Figure 1—figure supplement 1D.
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp10-v2.zip
-
Source data 10
Original scanned image of the gel depicting both Tn::DypB+DypB and Tn::DypB+Cfp29 (Figure 1—figure supplement 1E).
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp11-v2.zip
-
Source data 11
Original scanned image of the gel depicting Tn::DypB+Operon (Figure 1—figure supplement 1E).
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp12-v2.zip
-
Source data 12
This image depicts Tn::DypB+DypB, Tn::DypB+Cfp29, and Tn::DypB+Operon with molecular weight markers, lane labels, and the section of the image that is depicted in the main figure outlined in blue (Figure 1—figure supplement 1E).
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp13-v2.zip
-
Source data 13
Wig file for control (library only) replicate 1 (Figure 3A and B).
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp14-v2.zip
-
Source data 14
Wig file for control (library only) replicate 2 (Figure 3A and B).
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp15-v2.zip
-
Source data 15
Wig file for control (library only) replicate 3 (Figure 3A and B).
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp16-v2.zip
-
Source data 16
Wig file for experimental (H2O2_low pH) replicate 1 (Figure 3A and B).
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp17-v2.zip
-
Source data 17
Wig file for experimental (H2O2_low pH) replicate 2 (Figure 3A and B).
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp18-v2.zip
-
Source data 18
Wig file for experimental (H2O2_low pH) replicate 3 (Figure 3A and B).
- https://cdn.elifesciences.org/articles/74358/elife-74358-supp19-v2.zip





