trim-21 promotes proteasomal degradation of CED-1 for apoptotic cell clearance in C. elegans
Figures
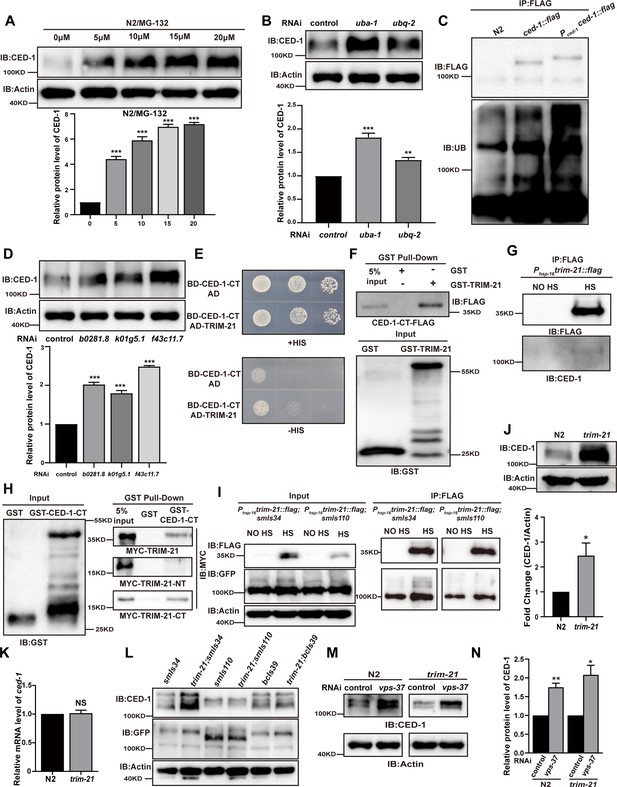
TRIM-21 is the E3 ubiquitin ligase to mediate the degradation of CED-1.
(A, B, D) The endogenous CED-1 was examined by immunoblot analysis in N2 treated with different concentrations of MG-132 (A), control RNAi, uba-1 RNAi, and ubq-2 RNAi (B), control RNAi, b0281.8 RNAi, k01g5.1 RNAi, and f43c11.7 RNAi (D). Graphs show the quantification of the protein level of CED-1. Data were from three independent experiments. (C) Ubiquitination of CED-1 was examined in C. elegans. FLAG IP was performed, followed by detection of ubiquitination with anti-ubiquitin antibodies. CED-1 has four isoforms; Pced-1ced-1::flag overexpressed CED-1 isoform a, and ced-1::flag was inserted as a FLAG tag into the endogenous ced-1 locus. (E–H) The interaction between CED-1-CT and TRIM-21 was examined by yeast two-hybrid analyses (+HIS, the medium lacking Trp and Leu; -HIS, the medium lacking Trp, Leu, and His) (E), GST pull-down assays (F), FLAG IP in vivo (G), and the CED-1-CT-TRIM-21 interaction occurred through the coiled-coil domain of TRIM-21 (H). (I) FLAG-IP of worm lysates were prepared from Phsp-16trim-21::flag strains carrying smIs34(Pced-1ced-1::gfp) and smIs110(Pced-1ced-1DC::gfp). (J) The endogenous CED-1 was examined by immunoblot analysis in N2 and trim-21(xhw12) mutants. The graph shows CED-1 level. Data were from three independent experiments. (K) ced-1 mRNA in N2 and trim-21(xhw12) mutants was determined by qRT-PCR. Data were from three independent experiments. (L) The exo/endogenous CED-1 expression was examined by immunoblot analyses in trim-21(xhw12) mutants carrying smIs34, smIs110, and bcIs39(Plim-7ced-1::gfp). (M) Endogenous CED-1 was examined by immunoblot analysis in N2 and trim-21 treated with control RNAi and vps-37 RNAi. (N) Graph shows CED-1 levels, which were quantified using ImageJ software. Data were from three independent experiments. The unpaired t-test was performed in this figure. *p<0.05, **p<0.01, ***p<0.001, NS, no significance. All bars indicate means and SEM.
-
Figure 1—source data 1
Comparison of the levels of proteins in different samples.
Including uncropped Western blot images and raw statistics.
- https://cdn.elifesciences.org/articles/76436/elife-76436-fig1-data1-v2.zip
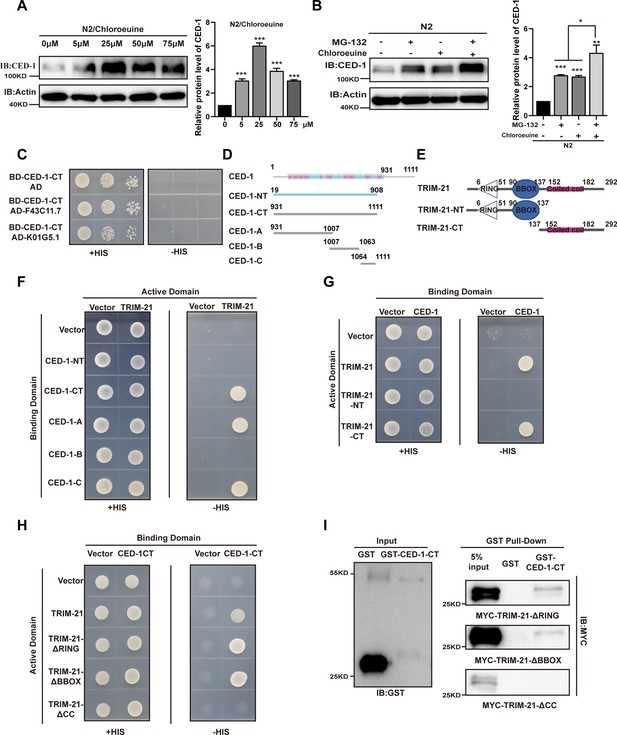
The coiled-coil domain of TRIM-21 interacts with the intracellular domain of CED-1.
(A, B) The endogenous CED-1 was examined by immunoblot analysis in N2 treated with different concentrations of chloroeuine (A), 5 µM MG-132, 5 µM chloroeuine, and 5 µM MG-132 + 5 µM chloroeuine (B). The graph shows the quantification of the protein level of CED-1. Data were from three independent experiments. (C) The E3 ubiquitin ligase F43C11.7 and K01G5.1 did not interact with CED-1-CT in the yeast two-hybrid assay. (D, E) Schematic diagram showing domains and truncations of CED-1 (D) and TRIM-21 (E). (F–I) The interaction between CED-1 and TRIM-21 was examined by yeast two-hybrid analyses (F–H) and GST pull-down assays (I). The interaction between CED-1 and TRIM-21 occurs through the CED-1 intracellular domain (F) and the coiled-coil domain of TRIM-21 (G–I). An unpaired t-test was performed in this figure. *p<0.05, **p<0.01, ***p<0.001. All bars indicate means and SEM.
-
Figure 1—figure supplement 1—source data 1
Variation in protein levels of CED-1 after N2 treatment of chloroquine or MG-132 and CED-1-CT interacting with TRIM-21 CC domain.
Including uncropped Western blot images and raw statistics.
- https://cdn.elifesciences.org/articles/76436/elife-76436-fig1-figsupp1-data1-v2.zip
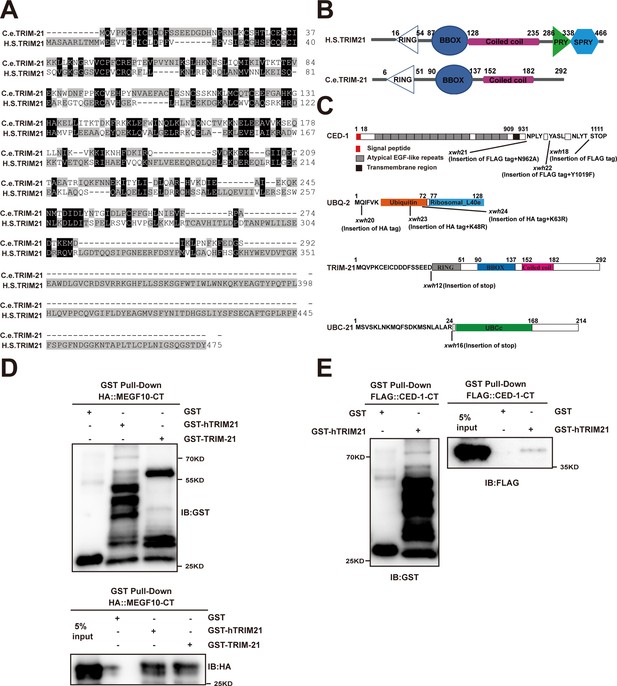
Sequence alignment of C. elegans TRIM-21 and human TRIM21, and schematic illustration of the mutation and tag insertions generated by CRISPR-Cas9.
(A) Sequence alignment of C. elegans (C.e) TRIM-21 and human (H.S) TRIM21. Identical residues are shaded in black, and similar ones in gray. (B) Schematic diagram showing domains and truncations of C. elegans (C.e) TRIM-21 and human (H.S) TRIM21. (C) Schematic illustration of the mutation and tag insertions generated by CRISPR-Cas9 editing of the endogenous ced-1, ubq-2, trim-21, and ubc-21 loci. The amino acids near the insertion or mutation sites are indicated. (D, E) The interactions between hTRIM21 or TRIM-21 with MEGF10-CT (D) and hTRIM21 with CED-1-CT (E) were examined by GST pull-down assays. The hTRIM21 was mean Homo sapiens TRIM21.
-
Figure 1—figure supplement 2—source data 1
The interactions between hTRIM21 or TRIM-21 with MEGF10-CT and hTRIM21 with CED-1-CT.
Including uncropped Western blot images.
- https://cdn.elifesciences.org/articles/76436/elife-76436-fig1-figsupp2-data1-v2.zip
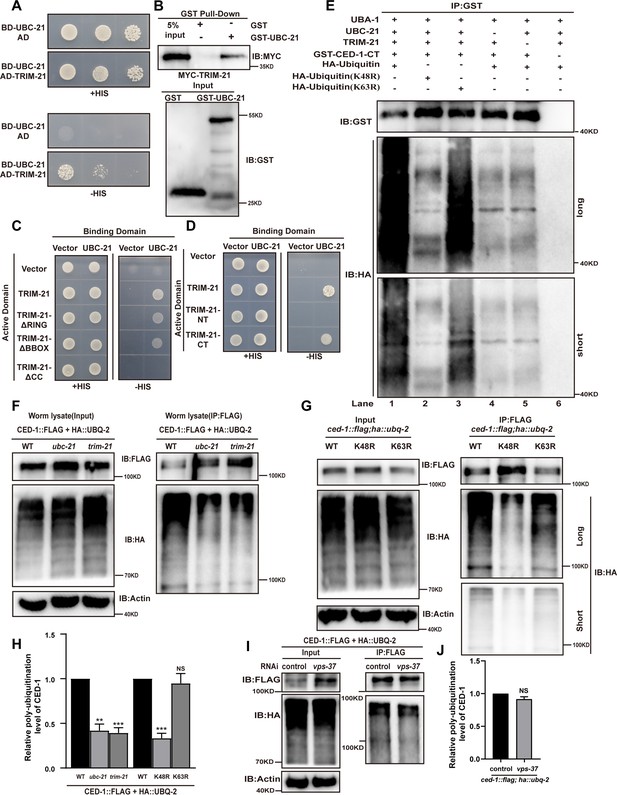
UBC-21 and TRIM-21 mediate the poly-ubiquitination of CED-1.
(A–D) The interaction between TRIM-21 and UBC-21 was examined by yeast two-hybrid analyses (A), GST pull-down assays (B), and the UBC-21-TRIM-21 interaction occurs through the coiled-coil domain of TRIM-21 (C, D). (E) Ubiquitination of recombinant GST-CED-1-CT was examined in vitro using different forms of HA-ubiquitin (WT, K48R, K63R). GST IP was performed, followed by detection of ubiquitination with anti-HA antibodies. Ubiquitination of CED-1-CT was observed (compare lane 1 with lanes 4–6), and inclusion of K48R- but not K63R-ubiquitin disrupted poly-ubiquitination of CED-1-CT (compare lanes 1 and 3 with lane 2). Long and short designate long exposure time and short exposure time, respectively. (F) Ubiquitination of CED-1 was examined in WT, ubc-21(xwh15), and trim-21(xwh12) mutant worms carrying ced-1::flag and ha::ubq-2. FLAG IP was performed, followed by detection of ubiquitination with anti-HA antibodies. (G) Ubiquitination of CED-1 was examined in worms carrying both ced-1::flag and ha::ubq-2(WT, K48R, and K63R). Long and short designate long exposure time and short exposure time, respectively. (H) Graph of the ubiquitination level (F, G) quantified using ImageJ software. The ratio of ubiquitin versus CED-1 was determined and normalized to onefold in the WT. Data were from three independent experiments. (I) Ubiquitination of CED-1 was examined for both ced-1::flag and ha::ubq-2 treated with control or vps-37 RNAi. FLAG IP was performed, followed by detection of ubiquitination with anti-HA antibodies. (J) Graph shows level of ubiquitination. The ratio of ubiquitin versus CED-1 was determined and normalized to onefold in the control. Data were from three independent experiments. An unpaired t-test was performed in this figure. **p<0.01, ***p<0.001. All bars indicate means and SEM.
-
Figure 2—source data 1
The interaction between TRIM-21 and UBC-21, and the relative poly-ubiquitination level of CED-1 in vitro and in vivo.
Including uncropped Western blot images and raw statistics.
- https://cdn.elifesciences.org/articles/76436/elife-76436-fig2-data1-v2.zip
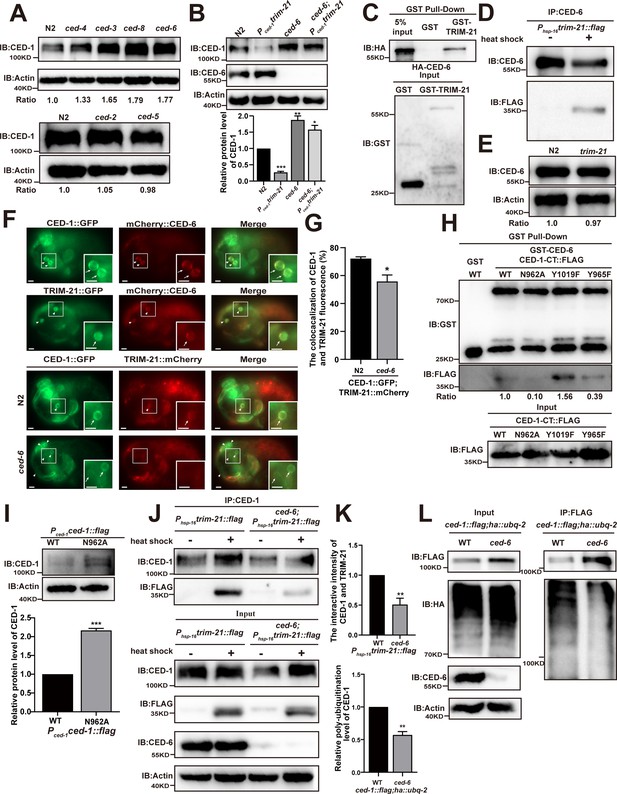
CED-6 mediates the degradation of CED-1 by recruitment of TRIM-21 to CED-1.
(A, B) Endogenous CED-1 was examined by immunoblot analysis in N2 and different null alleles mutants (A), and in N2 and indicated strains (B). Endogenous CED-1 levels are shown at the bottom. Data were from three independent experiments. (C, D) The interaction between CED-6 and TRIM-21 was detected by GST pull-down assays (C), and CED-6 IP in vivo using xwhIs28(Phsp-16trim-21::flag) worms (D). (E) The endogenous CED-6 was examined by immunoblot analysis in N2 and trim-21(xwh12) mutants. The endogenous CED-6 levels were quantified at the bottom. Data were from three independent experiments. (F) Co-localization of CED-1::GFP and mCherry::CED-6, TRIM-21::GFP and mCherry::CED-6 in N2 embryos, CED-1::GFP and TRIM-21:: in N2 and null alleles mutant ced-6(xwh25) embryos. Boxed regions are magnified (2×) in insets. Bars, 2 µm. (G) Quantification of CED-1::GFP and TRIM-21::mCherry co-localization on cell corpses in N2 and ced-6 mutant embryos. At least 100 cell corpses were scored for each strain and the data were repeated three times. The percentage referred to the ratio of TRIM-21::mCherry to CED-1::GFP. (H) The interactions between CED-6 and CED-1-CT (WT, N962A, Y1019F, and Y965F) were examined by GST pull-down assays. The quantity of GST pull-down FLAG-CED-1-CT/input FLAG-CED-1-CT is shown at the bottom. Data were from three independent experiments. (I) The exogenous CED-1 level in null alleles mutant ced-1(e1735) carrying Pced-1ced-1::flag(xwhEx34) and Pced-1ced-1(N962A)::flag(xwhEx35) is shown. The graph shows the quantification of the CED-1 protein level. Data were from three independent experiments. (J) CED-1 IP was performed, followed by detection of interaction between CED-1 and TRIM-21. The interaction was observed in N2 and null alleles mutant ced-6(xwh25). (K, above) The graph shows the quantification of the protein level of TRIM-21/CED-1. The ratio of TRIM-21 versus CED-1 was determined and normalized to onefold in N2. (L) FLAG IP was performed, followed by detection of ubiquitination in WT and null alleles mutant ced-6(xwh25) carrying both ced-1::flag and ha::ubq-2 with anti-HA antibodies. (K, below) The graph shows the relative poly-ubiquitination of CED-1. The ratio of ubiquitin versus CED-1 was determined and normalized to onefold in WT. Data were from three independent experiments. The unpaired t-test was performed in this figure. **p<0.01, ***p<0.001. All bars indicate means and SEM.
-
Figure 3—source data 1
Related protein levels in indicated strains and related proteins interactions.
Including uncropped Western blot images and raw statistics.
- https://cdn.elifesciences.org/articles/76436/elife-76436-fig3-data1-v2.zip
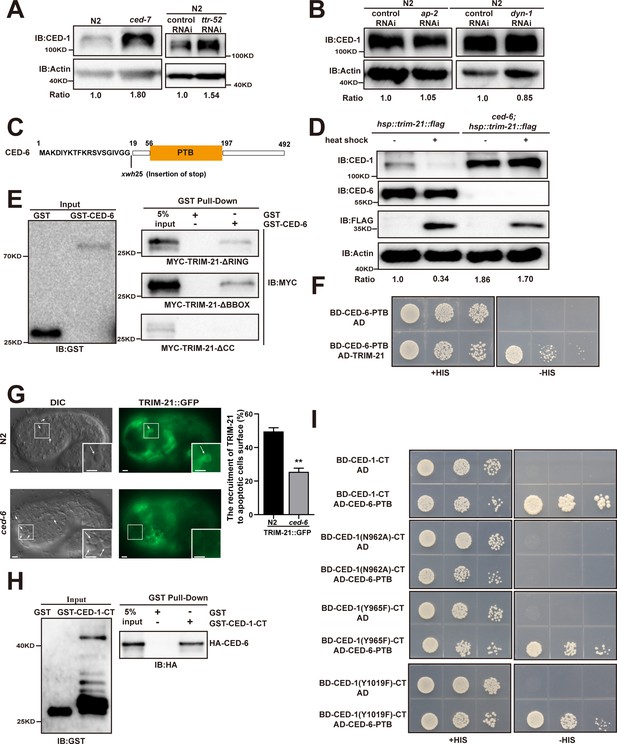
The phosphotyrosine-binding domain (PTB) domain of CED-6 interacts with coiled-coil domain of TRIM-21 and NPXY motif of CED-1.
(A, B, D) The endogenous CED-1 was detected by immunoblot analysis in N2, ced-7(n1892), N2 treated with control or ttr-52 RNAi, (A) N2 treated with control, ap-2, and dyn-1 RNAi, (B) and indicated transgenic worms (heat shock at 33°C for 1 hr or not) (D). The endogenous CED-6 levels were quantified at the bottom. Data were from three independent experiments. (C) Schematic illustration of the mutation generated by CRISPR-Cas9 editing of the endogenous ced-6 loci. The amino acids near the mutation sites are indicated. (E, F and H, I) The interactions between CED-6-TRIM-21 and CED-1-CT-CED-6 were examined by GST pull-down assays (E, H) and yeast two-hybrid analyses (F, I). The CED-6 and TRIM-21 interaction occurs through the coiled-coil domain of TRIM-21 (E) and the CED-6 PTB domain (F). The CED-6 and CED-1-CT interaction depends on CED-1-CT NPXY motif and the CED-6 PTB domain (H, I). (G) Representative images (left) and quantification (right) of TRIM-21::GFP localization on cell corpses in N2 and ced-6 mutant embryos. Arrows indicate cell corpses. At least 100 cell corpses were scored for each strain and the data were repeated three times. DIC, differential interference contrast. Bars, 2 µM. The unpaired t-test was performed in this figure. **p<0.01. All bars indicate means and SEM.
-
Figure 3—figure supplement 1—source data 1
The protein level of CED-1 in indicated strains and related proteins interactions.
Including uncropped Western blot images and raw statistics.
- https://cdn.elifesciences.org/articles/76436/elife-76436-fig3-figsupp1-data1-v2.zip
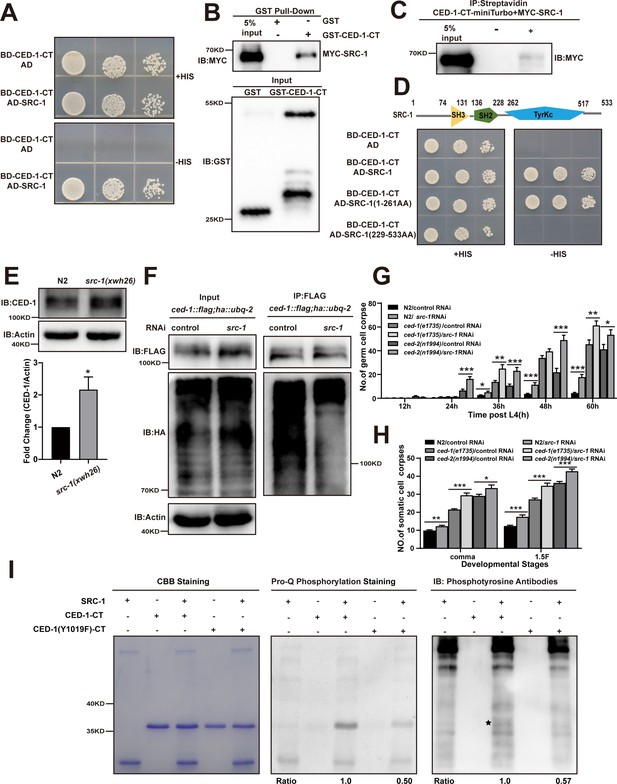
The phosphorylation of YXXL motif in CED-1 by SRC-1 is required for CED-1 degradation.
(A–D) The interaction between CED-1-CT 1617 and SRC-1 was examined by yeast two-hybrid (Y2H) analyses (A), GST pull-down assays (B), co-IP by 0.5 mM biotin in 293T cells (C), and the CED-1-SRC-1 interaction occurs through the SH3 and SH2 domain of SRC-1 in Y2H (D). (E) The endogenous CED-1 level was detected in N2 and src-1(xwh26). The graph shows quantification of the protein level of CED-1. (F) Ubiquitination of CED-1 was examined in worms carrying both ced-1::flag and ha::ubq-2 treated with control or src-1 RNAi. FLAG IP was performed, followed by detection of ubiquitination with anti-HA antibodies. (G, H) Different stages (hr post L4) of germ cell corpses (G) and comma, 1.5F stage embryo corpses (H) were quantified (mean ± SEM) in indicated strains, in which ced-1(e1735) and ced-2(n1994) are null alleles mutants. Fifteen adult worms or embryos were scored at each stage for each strain. (I) Phosphorylation of CED-1-CT (WT or Y1019F) by SRC-1 was analyzed using CBB staining (left), Pro-Q phosphorylation staining (middle) and phosphotyrosine antibody (right). * shows phosphotyrosine bands. Quantities of Pro-Q phosphorylation staining CED-1-CT level/CBB staining CED-1-CT level and anti-phosphotyrosine CED-1 level/CBB staining CED-1-CT level are shown at the bottom. Data were from three independent experiments. An unpaired t-test was performed. *p< 0.05, **p<0.01, ***p<0.001. All bars indicate means and SEM.
-
Figure 4—source data 1
Related protein levels in indicated strains, related proteins interactions and phosphorylation of CED-1-CT in vitro.
Including uncropped Western blot images and raw statistics.
- https://cdn.elifesciences.org/articles/76436/elife-76436-fig4-data1-v2.zip
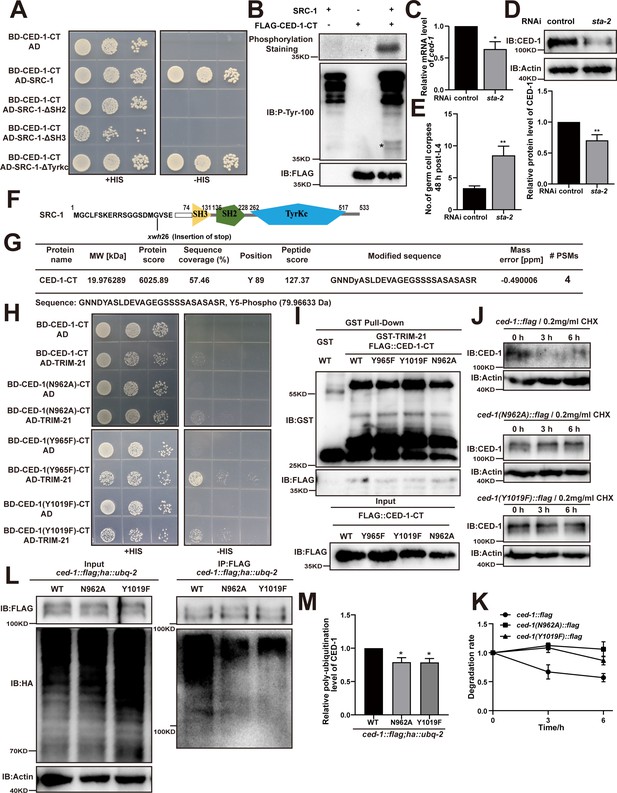
The YXXL motif in CED-1 is phosphorylated by SRC-1.
(A) The interactions between CED-1-CT and SRC-1 (WT, lack of SH2, SH3, or Tyrkc) were examined by yeast two-hybrid analyses. (B) Phosphorylation of CED-1-CT by SRC-1 was analyzed using phosphorylation staining and an anti-P-Tyr-100 antibody staining. CED-1-CT was fused with FLAG tag. * indicates phosphotyrosine bands. (C) ced-1 mRNA in control worms and those treated sta-2 RNAi was determined by qRT-PCR. Data were from three independent experiments. (D) The endogenous CED-1 was examined by immunoblot analysis in N2 treated with control and sta-2 RNAi. The graph shows the quantification of the protein level of CED-1. (E) The adult stage (48 hr post-L4) germ cell corpses were quantified in N2 treated with control or sta-2 RNAi. 15 adult worms were scored for each strain.(F) Schematic illustration of the mutation generated by CRISPR-Cas9 editing of the endogenous src-1 loci. The amino acids near the mutation sites are indicated. (G) Mass spectrum of a CED-1-CT peptide that was phosphorylated at tyrosine 89 (mean CED-1 full-length 1019 site). The peptide was identified from the phosphorylation of CED-1-CT by SRC-1 in vitro. (H, I) The interactions between CED-1-CT (WT or sites mutant: N962A, Y965F, and Y1019F) and TRIM-21 were examined by yeast two-hybrid analyses (H) and GST pull-down assays (I). (J) The endogenous CED-1 in ced-1::flag, ced-1(N962A)::flag and ced-1(Y1019F)::flag worms (grown in liquid medium at 20°C) treated with 0.2 mg/ml CHX (protein synthesis inhibitor) 0, 3, and 6 hr post-induction were detected by immunoblot analyses (β-actin loaded control). (K) The graph shows quantification of the 0, 3, and 6 hr post-CHX induction level of CED-1 in ced-1::flag, ced-1(N962A)::flag, and ced-1(Y1019F)::flag worms using ImageJ and analyzed by GraphPad Prism 8 and Excel. Data were from three independent experiments. (L) Ubiquitination of CED-1 was examined in worms carrying both ced-1::flag(WT, N962A or Y1019F) and ha::ubq-2. FLAG IP was performed, followed by detection of ubiquitination with anti-HA antibodies. (M) The graph shows quantification of the level of ubiquitination using ImageJ. The ratio of ubiquitin versus CED-1 was determined and normalized to onefold in WT. Data were from three independent experiments. The unpaired t-test was performed in this figure. *p<0.05. All bars indicate means and SEM.
-
Figure 4—figure supplement 1—source data 1
Related protein levels and poly-ubiquitination in indicated strains, mRNA levels of ced-1 and germ cell corpses in N2 treated with sta-2 RNAi, related proteins interactions and phosphorylation of CED-1-CT in vitro.
Including uncropped Western blot images and raw statistics.
- https://cdn.elifesciences.org/articles/76436/elife-76436-fig4-figsupp1-data1-v2.zip
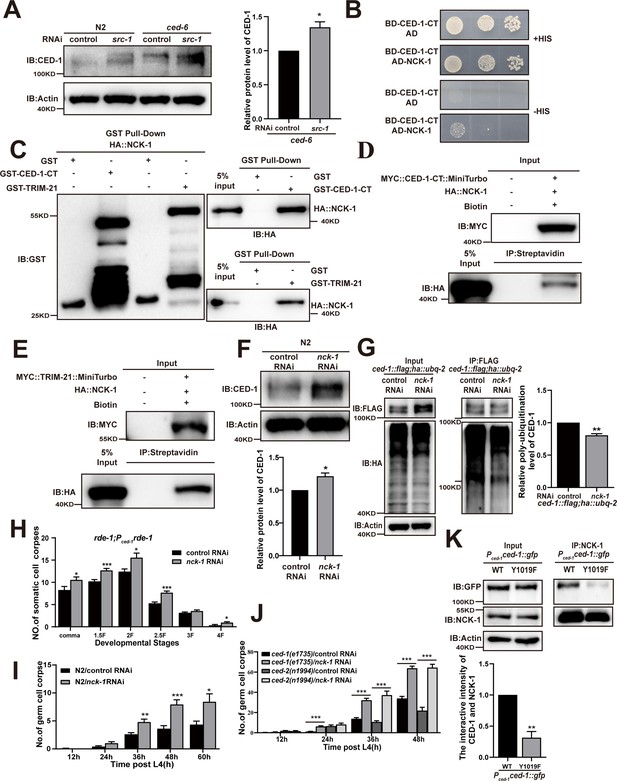
The adaptor NCK-1 is required for CED-1 degradation.
(A) The endogenous CED-1 was examined by immunoblot analysis in N2 and null alleles mutant ced-6(n1813) treated with control or src-1 RNAi. The graph shows the quantification of the level of CED-1 in ced-6(n1813) treated with control and src-1 RNAi. Data were from three independent experiments. (B–E) The interactions between CED-1-CT–NCK-1 were examined by yeast two-hybrid (Y2H) (B), CED-1-CT–NCK-1 and TRIM-21–NCK-1 levels were detected by GST pull-down assays (C), co-IP by 0.5 mM biotin in 293T cells (D, E). (F) The endogenous CED-1 was examined by immunoblot analysis in N2 treated with control or nck-1 RNAi. The graph shows quantification of the level of CED-1. (G) Ubiquitination of CED-1 was examined in worms carrying both ced-1::flag and ha::ubq-2 treated with control or nck-1 RNAi. FLAG IP was performed, followed by detection of ubiquitination with anti-HA antibodies. The graph shows quantification of the level of ubiquitination. The ratio of ubiquitin versus CED-1 was determined and normalized to onefold in the control. Data were from three independent experiments. (H–J) The embryonic or gonadal cell corpses were quantified in the indicated strains treated with control or nck-1 RNAi, the development stages of embryo cell corpses in tissue-specific expression strain rde-1; Pced-1rde-1 (H), the germ cell corpses with different stages post-L4 in N2 (I), and null alleles mutant ced-1(e1735) and null alleles mutant ced-2(n1994) (J). 15 adult worms or embryos were scored at each stage for each strain. (K) NCK-1 IP was performed, followed by detection of the interaction between CED-1 (WT, Y1019F) and NCK-1 in Pced-1ced-1::gfp (WT, Y1019F) worms with anti-GFP antibodies. The graph shows the protein level of CED-1/NCK-1. The ratio of CED-1 versus NCK-1 was determined and normalized to onefold in N2. Data were from three independent experiments. An unpaired t-test was performed in this figure. *p<0.05, **p<0.01, ***p<0.001. All bars indicate means and SEM.
-
Figure 5—source data 1
Related protein levels and poly-ubiquitination in indicated strains, cell corpses in indicated strains and related proteins interactions.
Including uncropped Western blot images and raw statistics.
- https://cdn.elifesciences.org/articles/76436/elife-76436-fig5-data1-v2.zip
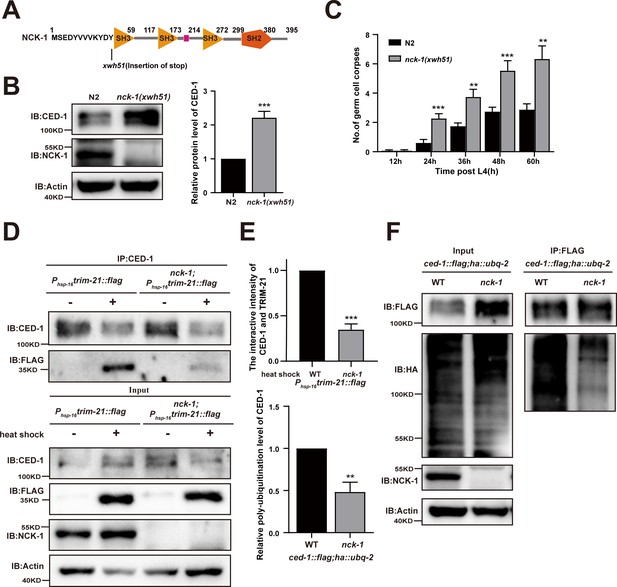
Role of NCK-1 in CED-1 degradation.
(A) Schematic illustration of the mutation generated by CRISPR-Cas9 editing of the endogenous nck-1 loci. The amino acids near the mutation sites are indicated. (B) The endogenous CED-1 was examined by immunoblot analysis in N2 and null alleles mutant nck-1(xwh51). The graph shows the quantification of the level of CED-1 in N2 and nck-1(xwh51). Data were from three independent experiments. (C) N2 and nck-1(xwh51) germ cell corpses were quantified in different adult stages (hr post L4). Fifteen adult worms were scored at each stage for each strain. (D, E) CED-1 IP was performed, followed by detection of interaction between CED-1 and TRIM-21. The interaction was observed in N2 and nck-1(xwh51). (E, above) The graph shows the quantification of the protein level of TRIM-21/CED-1. The ratio of TRIM-21 versus CED-1 was determined and normalized to onefold in N2. Data were from three independent experiments. (F) FLAG IP was performed, followed by detection of ubiquitination in WT and nck-1(xwh51) mutant carrying both ced-1::flag and ha::ubq-2 with anti-HA antibodies. (E, below) The graph shows the relative poly-ubiquitination of CED-1. The ratio of ubiquitin versus CED-1 was determined and normalized to onefold in WT. Data were from three independent experiments. The unpaired t-test was performed in this figure. **p<0.01, ***p<0.001. All bars indicate means and SEM.
-
Figure 5—figure supplement 1—source data 1
Related protein levels and poly-ubiquitination in indicated strains and germ cell corpses in nck-1 mutants.
Including uncropped Western blot images and raw statistics.
- https://cdn.elifesciences.org/articles/76436/elife-76436-fig5-figsupp1-data1-v2.zip
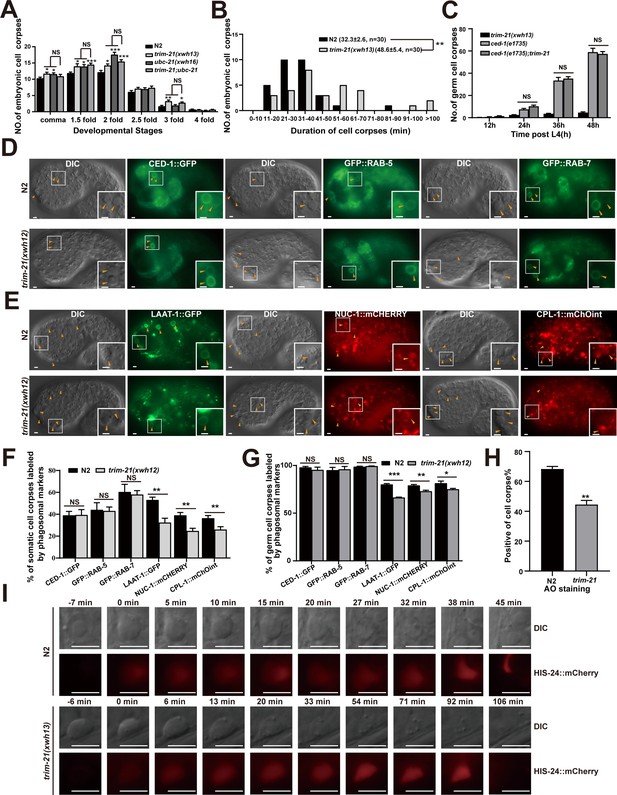
TRIM-21 acts in the CED-1 pathway to regulate phagosome maturation.
(A) Different stages of embryonic corpses were quantified (mean ± SEM) in the indicated mutants. Fifteen embryos were scored at each stage for each strain. (B) Four-dimensional microscopy analysis of cell corpse duration was performed in N2 and trim-21(xwh13). The persistence of 30 cell corpses from embryos was monitored. The mean duration (± SEM) is shown in parenthesis. (C) trim-21(xwh13), null alleles mutant ced-1(e1735) and double null alleles mutant trim-21(xwh13); ced-1(e1735) germ cell corpses were quantified in different adult stages (hr post L4). Fifteen adult worms were scored at each stage for each strain. (D, E) The cell corpse labeling by the phagosome markers CED-1::GFP, GFP::RAB-5, GFP::RAB-7 (D), LAAT-1::GFP, NUC-1::mCHERRY, and CPL-1::mChOint (E) in N2 and trim-21(xwh12) embryos were captured using Imager M2 (Zeiss). Bars, 2 µm. (F, G) The cell corpses positive for phagosome markers in N2 and trim-21(xwh12) embryos (F) and germlines (G) were quantified. At least 100 cell corpses were scored for each strain. Data were from three independent experiments. (H) The cell corpse labeled by 0.1 mg/ml acridine orange was quantified (mean ± SEM) in N2 and trim-21(xwh12) adult worms. Data were from three independent experiments. (I) Time-lapse chasing of button-like cell corpses in DIC and HIS-24::mCherry-positive phagolysosomes in N2 and trim-21 (xwh13) germlines. The time point that the HIS-24::mCherry ring was first detected on a cell corpses was set as 0 min. Bars, 5 µm. An unpaired t-test was performed in this figure. *p<0.05, **p<0.01, ***p<0.001, NS, no significance. All bars indicate means and SEM.
-
Figure 6—source data 1
Cell corpses and cell corpses duration in indicated strains, cell corpses labeled by phagosome markers and AO stainging in trim-21 mutants.
Including raw statistics.
- https://cdn.elifesciences.org/articles/76436/elife-76436-fig6-data1-v2.zip
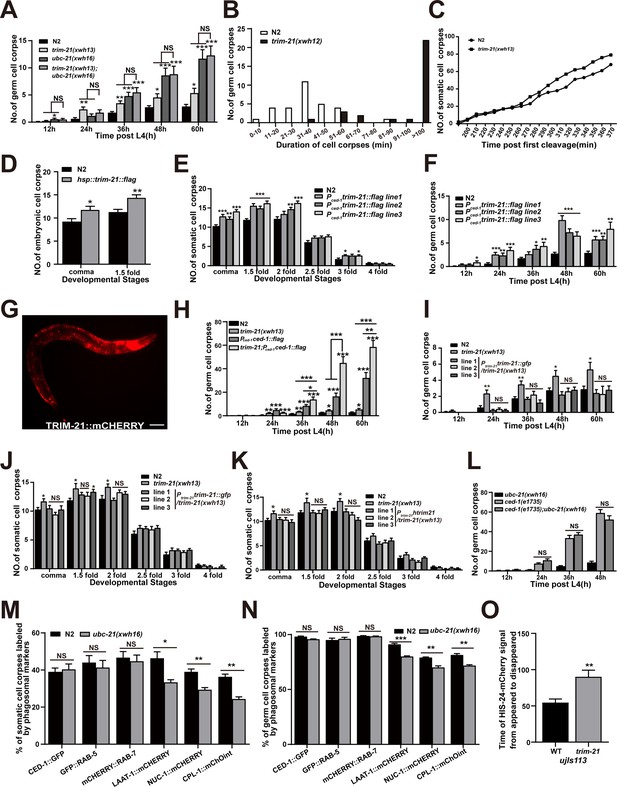
Loss of TRIM-21 and UBC-21 affect phagosome maturation.
(A) The different adult stages (hr post L4) of germ cell corpses were quantified in indicated mutants. Fifteen adult worms were scored at each stage for each strain. (B) Four-dimensional microscopy analysis of germ cell corpse duration was performed in N2 and trim-21(xwh12). The persistence of 30 germ cell corpses was monitored. (C) The number of embryo corpses was measured by four-dimensional microscopy analysis in N2 and trim-21(xwh13). Embryos (two cell stage) were isolated from adult worms and imaged every minute for 400 min. (D–F, H–L) The development of adult or embryo stages corpses was quantified in indicated strains. Fifteen adult worms or embryos were scored at each stage for each strain. (G) The TRIM-21::mCHERRY localization in L1 of xwhIs33(Ptrim-21trim-21::mcherry) worms was imaged by the mCherry channel used Imager M2 microscope. Bars, 50 µm. (M, N) The percentage of cell corpses labeled by phagosomal markers was quantified in N2 and ubc-21(xwh16) embryos (M) and germlines (N). Also, 100 apoptotic cells were counted, and this was repeated three times. (O) The persistence of five germ cell corpses labeled with HIS-24::mCherry was monitored. At least 100 cell corpses were scored for each strain. Data were from three independent experiments. An unpaired t-test was performed in this figure. *p<0.05, **p<0.01, ***p<0.001. All bars indicate means and SEM.
-
Figure 6—figure supplement 1—source data 1
Cell corpses and cell corpses duration in indicated strains, cell corpses labeled by phagosome markers in ubc-21 mutants and the persistence of cell corpses labeled with HIS-24::mCherry in trim-21 mutants.
Including raw statistics.
- https://cdn.elifesciences.org/articles/76436/elife-76436-fig6-figsupp1-data1-v2.zip
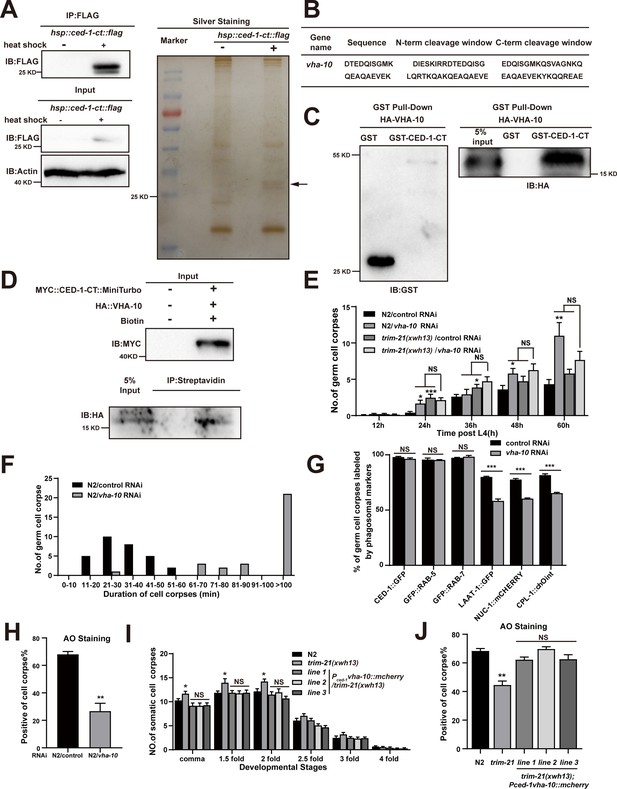
Excessive CED-1 binding to VHA-10 in trim-21 mutant worms negatively affects the maturation and acidification of cell corpse-containing phagosomes.
(A) The FLAG IP was performed on Phsp-16ced-1-ct::flag worms (heat shock for 1 hr at 33°C), followed by identification of proteins that interact with CED-1 in MS. The immunoblot analysis (left) and silver staining (right) results are shown. (B) The peptides were identified by MS analysis. (C, D) The interaction between CED-1-CT and VHA-10 was examined by GST pull-down assays (C) and co-IP assays in 293T cells treated with 0.5 mM biotin (D). (E) The different adult stages of germ cell corpses were quantified in N2 and trim-21(xwh13) treated with control or vha-10 RNAi. 15 adult worms were scored at each stage for each strain. (F) Four-dimensional microscopy analyses of 30 germ cell corpse duration were performed in N2 treated with control or vha-10 RNAi. (G) The germ cell corpses positive for phagosome markers in N2 treated with control or vha-10 RNAi were quantified. At least 100 cell corpses were scored for each strain. Data were from three independent experiments. (H) The cell corpse labeled by 0.1 mg/ml acridine orange was quantified in N2 treated with control or vha-10 RNAi adult worms. At least 100 cell corpses were scored for each strain. Data were from three independent experiments. (I) Embryonic cell corpses were quantified in the indicated strains. Fifteen embryos at different stages were scored for each strain. (J) Cell corpse labeled by 0.1 mg/ml acridine orange was quantified (mean ± SEM) in the indicated strains of adult worms. At least 100 cell corpses were scored for each strain. Data were from three independent experiments. An unpaired t-test was performed in this figure. *p<0.05, **p<0.01, ***p<0.001, NS, no significance. All bars indicate means and SEM.
-
Figure 7—source data 1
The immunoblot and silver staining in Phsp-16ced-1-ct::flag worms, related proteins interactions, cell corpses in indicated strains, cell corpses duration and cell corpses labeled by phagosome markers in N2 treated with vha-10 RNAi and AO staining in indicated strains.
Including uncropped Western blot images and raw statistics.
- https://cdn.elifesciences.org/articles/76436/elife-76436-fig7-data1-v2.zip
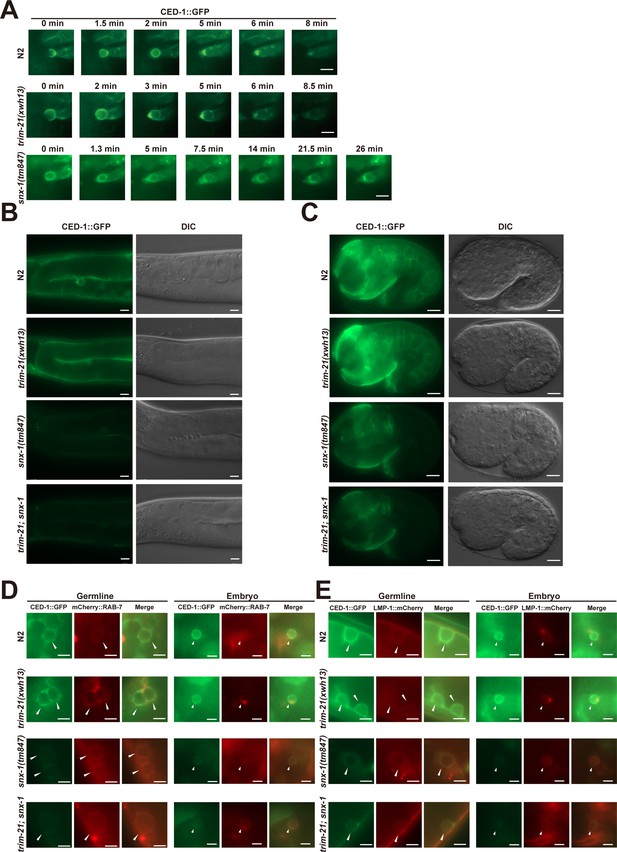
TRIM-21-mediated proteasome degradation of CED-1 independent of lysosomal degradation of CED-1 by loss of function of the retromer complex.
(A) Time-lapse monitoring of CED-1::GFP on phagosomes in N2, trim-21(xwh13), and snx-1(tm847) embryos. The point when the CED-1::GFP ring was first detected on the cell corpse was set as 0 min. Bars, 2 µm. (B, C) CED-1::GFP signals in germline (B) and 1.5-fold embryos (C) of N2, trim-21(xwh13), snx-1(tm847), and trim-21; snx-1 worms. Fluorescence images were obtained using equal exposure times. Bars, 10 µm in germline, 5 µm in embryo. (D) Localization of CED-1::GFP and mCherry::RAB-7 in germline or embryo phagosomal in N2, trim-21(xwh13), null alleles mutant snx-1(tm847) and trim-21; snx-1 double mutants. Fluorescence images were obtained using equal exposure times. Bars, 5 µm in germline, 2 µm in embryo. (E) Localization of CED-1::GFP and LMP-1::mCherry in germline or embryo phagolysosomal in N2, trim-21(xwh13), snx-1(tm847), and trim-21; snx-1 double mutants. Fluorescence images were obtained using equal exposure times. Bars, 5 µm in germline, 2 µm in embryo.
-
Figure 7—figure supplement 1—source data 1
Time-lapse mointoring of CED-1::GFP on phagosomes in N2, trim-21 and snx-1 embryos.
Including raw statistics.
- https://cdn.elifesciences.org/articles/76436/elife-76436-fig7-figsupp1-data1-v2.zip
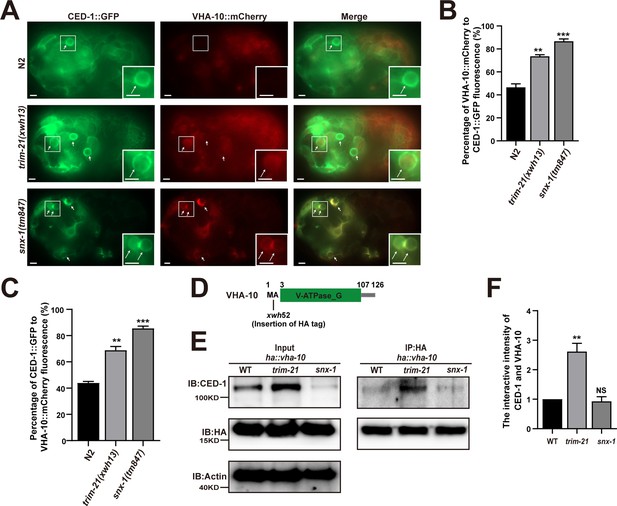
Excessive CED-1 in trim-21 mutant worms binds VHA-10.
(A) Co-localization of CED-1::GFP and VHA-10::mCherry in N2, trim-21(xwh13), and snx-1(tm847) mutant embryos. Arrows indicate cell corpses. Boxed regions are magnified (2×) in insets. Bars, 2 µm. (B, C) Graph of quantification of CED-1::GFP and VHA-10::mCherry co-localization on cell corpses in N2, trim-21(xwh13), and snx-1(tm847) mutant embryos. The percentage referred to the ratio of VHA-10::mCherry to CED-1::GFP (B). The percentage referred to the ratio of CED-1::GFP to VHA-10::mCherry (C). At least 100 cell corpses were scored for each strain. Data were from three independent experiments. (D) Schematic illustration of HA tag insertion generated by CRISPR-Cas9 editing of endogenous vha-10 loci. Amino acids near the insertion site are indicated. (E) HA IP was performed, followed by detection of the interaction between CED-1 and VHA-10 in WT, trim-21(xwh13), and null alleles mutant snx-1(tm847) carrying ha::vha-10 with anti-CED-1 antibody. The graph shows the protein level of CED-1/NCK-1. (F) Ratio of CED-1 versus VHA-10 was determined and normalized to onefold in N2. Data were from three independent experiments. An unpaired t-test was performed in this figure. **p<0.01, ***p<0.001. All bars indicate means and SEM.
-
Figure 7—figure supplement 2—source data 1
The percentage of VHA-10::mCherry and CED-1::GFP and the interaction between CED-1 and VHA-10 in N2, trim-21, snx-1 mutants.
Including uncropped Western blot images and raw statistics.
- https://cdn.elifesciences.org/articles/76436/elife-76436-fig7-figsupp2-data1-v2.zip
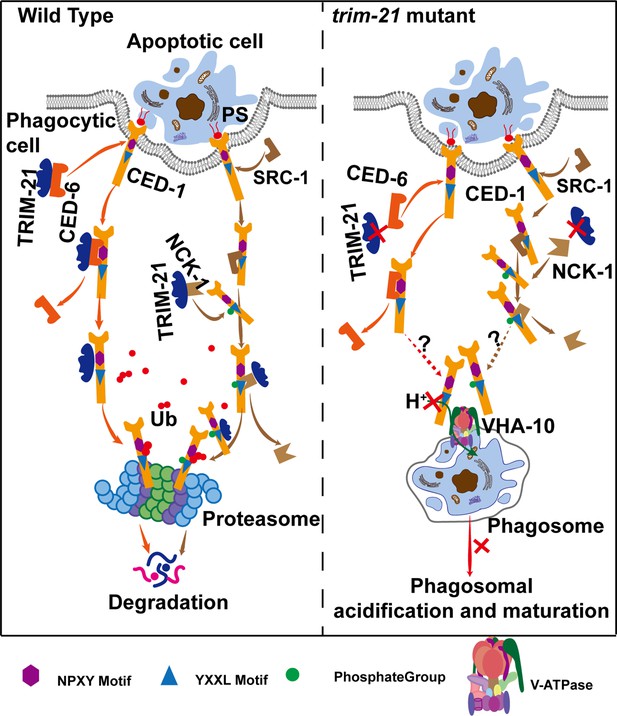
Model of TRIM-21-mediated CED-1 degradation through the proteasome pathway.
After phagocytic receptors (CED-1) recognize apoptotic cells and cell corpse engulfment, TRIM-21 is recruited to the surfaces of phagosomes through two parallel pathways and ubiquitinates part of CED-1 for proteasomal degradation. One pathway occurs after the NPXY motif of CED-1 binds to the adaptor protein CED-6, and TRIM-21 released from CED-6 and ubiquitinates CED-1. The other occurs when a tyrosine residue in the YXXL motif of CED-1 is phosphorylated by tyrosine kinase SRC-1, and the phosphorylated CED-1 YXXL motif recruits an adaptor protein NCK-1 (containing the SH2 domain), followed by TRIM-21 being released from NCK-1 and ubiquitinating CED-1. In the absence of TRIM-21, part of CED-1 fails to be ubiquitinated for proteasomal degradation and accumulates in phagosomes to bind to VHA-10 and affect phagosomal acidification and maturation, thus resulting in cell corpse degradation defects. It is currently unknown how the excess CED-1 in trim-21 mutants specifically binds to VHA-10.
Tables
The CED-1 protein level, regulated by RNAi of C. elegans genes encoding E3 ubiquitin ligases.
| C. elegans E3 ubiquitin ligases (RNAi) | Endogenous level of CED-1 | ||
|---|---|---|---|
| 1st | 2nd | 3rd | |
| control | - | - | - |
| T09B4.10 | + | - | ++ |
| R10A10.2 | ++ | - | - |
| T24D1.3 | + | - | - |
| Y51F10.2 | ++ | - | - |
| F10G7.10 | ++ | - | + |
| C34F11.1 | ++ | - | - |
| M110.3 | + | - | - |
| D2089.2 | + | - | - |
| R06F6.2 | + | - | - |
| B0281.8 | ++ | ++ | + |
| ZK1240.1 | + | - | ++ |
| ZK1320.6 | + | - | - |
| F43C11.8 | + | - | - |
| ZK1240.9 | + | - | - |
| F45H7.6 | ++ | - | - |
| K01G5.1 | + | ++ | ++ |
| F40G9.12 | ++ | - | - |
| M88.3 | + | - | - |
| R05D3.4 | + | - | - |
| ZK637.14 | + | - | ++ |
| F43C11.7 | + | ++ | ++ |
| C09E7.5 | ++ | ++ | - |
| T02C1.2 | + | - | - |
| Y47D3A.22 | ++ | - | - |
| Y47D3B.11 | + | - | - |
| C09E7.9 | ++ | - | - |
| K12B6.8 | ++ | ++ | - |
| T08D2.4 | + | - | - |
| Y45G12B.2 | + | - | - |
| M142.6 | ++ | ++ | - |
| C32D5.10 | ++ | ++ | - |
| C36A4.8 | ++ | - | - |
The CED-1 protein level, regulated by RNAi of C. elegans genes encoding E2 ubiquitin-conjugating enzymes.
| C. elegans E2 ubiquitin-conjugating enzymes (RNAi) | Endogenous level of CED-1 | ||
|---|---|---|---|
| 1st | 2nd | 3rd | |
| control | - | - | - |
| ubc-1 | - | - | - |
| ubc-2 | - | - | + |
| ubc-3 | - | + | - |
| ubc-6 | - | + | - |
| ubc-7 | + | + | - |
| ubc-8 | - | + | - |
| ubc-9 | - | - | - |
| ubc-12 | - | - | - |
| ubc-13 | - | + | - |
| ubc-14 | - | - | - |
| ubc-15 | - | - | - |
| ubc-16 | + | - | - |
| ubc-17 | - | - | - |
| ubc-18 | - | - | + |
| ubc-19 | - | - | - |
| ubc-20 | - | - | - |
| ubc-21 | + | + | + |
| ubc-22 | - | - | - |
| ubc-23 | + | - | - |
| ubc-24 | - | - | + |
| ubc-25 | - | - | - |
| ubc-26 | - | - | + |
Cell corpse phenotypes in N2, ced-1(e1735), and overexpression of ced-1 or ced-1 site mutants in ced-1(e1735).
| Transgene | No. of somatic cell corpses (developmental stages) | |||||
|---|---|---|---|---|---|---|
| Comma | 1.5F | 2F | 2.5F | 3F | 4F | |
| N2 (-) | 9.73 ± 0.47 | 12.27 ± 0.48 | 11.40 ± 0.39 | 6.67 ± 0.29 | 2.6 ± 0.32 | 0.67 ± 0.12 |
| ced-1(e1735) | 21.67 ± 0.85 *** | 28.80 ± 0.79 *** | 34.73 ± 0.99 *** | 34.20 ± 1.45 *** | 31.20 ± 1.84 *** | 30.93 ± 1.23 *** |
| Pced-1ced-1 line 1 / ced-1(e1735) | 9.73 ± 0.36 NS | 12.47 ± 0.28 NS | 9.00 ± 0.30 NS | 8.20 ± 0.72 NS | 2.07 ± 0.43 NS | 0.93 ± 0.31 NS |
| Pced-1ced-1 line 2 / ced-1(e1735) | 9.47 ± 0.31 NS | 11.73 ± 0.31 NS | 11.00 ± 0.74 NS | 7.87 ± 0.53 NS | 2.53 ± 0.48 NS | 1.07 ± 0.18 NS |
| Pced-1ced-1 line 3 / ced-1(e1735) | 10.73 ± 0.31 NS | 11.87 ± 0.58 NS | 11.40 ± 0.78 NS | 6.40 ± 0.46 NS | 1.73 ± 0.33 NS | 0.53 ± 0.26 NS |
| Pced-1ced-1(N962A) line 1/ced-1(e1735) | 23.60 ± 0.51 *** | 32.00 ± 0.99 *** | 35.73 ± 0.71 *** | 31.53 ± 1.14*** | 30.87 ± 1.60*** | 30.93 ± 1.05*** |
| Pced-1ced-1(N962A) line 2/ced-1(e1735) | 23.93 ± 0.38 *** | 30.40 ± 1.12*** | 35.93 ± 1.19 *** | 33.47 ± 1.11 *** | 27.13 ± 0.80*** | 30.93 ± 1.46*** |
| Pced-1ced-1(N962A) line 3/ced-1(e1735) | 22.40 ± 0.49 *** | 32.40 ± 1.07 *** | 34.80 ± 0.99 *** | 33.07 ± 0.71*** | 32.00 ± 0.91*** | 32.00 ± 0.60*** |
| Pced-1ced-1(Y965F) line 1/ced-1(e1735) | 12.73 ± 0.89 ** | 16.67 ± 0.70 *** | 18.53 ± 1.61 *** | 12.67 ± 0.62*** | 11.00 ± 1.82 *** | 3.33 ± 0.50 *** |
| Pced-1ced-1(Y965F) line 2/ced-1(e1735) | 15.67 ± 0.50 *** | 18.20 ± 0.48 *** | 21.00 ± 0.80 *** | 11.27 ± 0.69 *** | 7.87 ± 0.52 *** | 2.73 ± 0.33 *** |
| Pced-1ced-1(Y965F) line 3/ced-1(e1735) | 12.07 ± 0.73 * | 15.00 ± 0.52 *** | 17.27 ± 1.54 ** | 11.33 ± 0.51 *** | 17.27 ± 1.54*** | 3.27 ± 0.47 *** |
| Pced-1ced-1(Y1019F) line 1/ced-1(e1735) | 16.47 ± 0.69 *** | 19.13 ± 0.70 *** | 20.27 ± 0.61 *** | 11.87 ± 0.63 *** | 10.07 ± 0.89*** | 3.07 ± 0.35 *** |
| Pced-1ced-1(Y1019F) line 2/ced-1(e1735) | 15.07 ± 0.43 *** | 16.60 ± 0.51 *** | 17.80 ± 0.54 *** | 12.13 ± 0.53*** | 5.53 ± 0.55 *** | 2.13 ± 0.21 *** |
| Pced-1ced-1(Y1019F) line 3/ced-1(e1735) | 14.33 ± 0.29 *** | 19.73 ± 0.49 *** | 20.53 ± 0.79 *** | 11.60 ± 0.57 *** | 8.20 ± 0.46 *** | 3.00 ± 0.37 *** |
-
At least 15 embryos were scored at each stage for each strain. *p<0.05, **p<0.01, ***p<0.001, NS, no significance.
-
Table 3—source data 1
The number of somatic different developmental stages cell corpses in indicated strains.
Including raw statistics.
- https://cdn.elifesciences.org/articles/76436/elife-76436-table3-data1-v2.zip
Cell corpse phenotypes caused by RNAi of C. elegans genes encoding tyrosine kinases.
| C. elegans tyrosine kinases(RNAi) | No. of germ cell corpses(mean ± SEM) | C. elegans tyrosine kinases(RNAi) | No. of germ cell corpses(mean ± SEM) |
|---|---|---|---|
| Control | 2.667 ± 0.2425 | T06C10.6 | 2.478 ± 0.4484 |
| F49B2.5 | 4.459 ± 0.3435 | T13H10.1 | 4.854 ± 0.2828** |
| Y47G6A.5 | 3.735 ± 0.2873 | T25B9.4 | 4.577 ± 0.385 |
| Y48G1C.10 | 2.806 ± 0.2948 | W01B6.5 | 1.75 ± 0.3096 |
| C35E7.10 | 3.143 ± 0.5084 | Y4C6A.k | 2.75 ± 0.3708 |
| F22D6.1 | 2.529 ± 0.5363 | ZK593.9 | 1.188 ± 0.2453 |
| F23C8.7 | 2.235 ± 0.5391 | T25B9.5 | 2.063 ± 0.17 |
| F26E4.5 | 2.286 ± 0.3097 | W08D2.8 | 4.545 ± 0.2995 |
| F53G12.6 | 3.559 ± 0.3409 | Y69E1A.3 | 2.063 ± 0.2657 |
| F59A3.8 | 2.619 ± 0.4654 | F11E6.8 | 1.5 ± 0.2739 |
| T21G5.1 | 1.933 ± 0.4306 | T22B11.4 | 1.111 ± 0.1962 |
| ZC581.7 | 1.529 ± 0.2443 | Y116A8C.24 | 1.313 ± 0.2846 |
| W04G5.6 | 2.4 ± 0.3352 | T08G5.2 | 3.879 ± 0.3191 |
| C34F11.5 | 2.333 ± 0.2323 | M01B2.1 | 3.591 ± 0.3984 |
| F46F5.2 | 1.733 ± 0.3157 | T01G5.1 | 2.688 ± 0.3125 |
| M176.9 | 1.4 ± 0.3055 | C16D9.2 | 3.105 ± 0.4319 |
| R05H5.4 | 4.829 ± 0.4056** | C24G6.2 | 2.125 ± 0.482 |
| Y62F5A.10 | 2 ± 0.3086 | F40A3.5 | 1.789 ± 0.4811 |
| ZK622.1 | 4 ± 0.6249 | T10H9.2 | 4.05 ± 0.397 |
| C08H9.5 | 5.275 ± 0.4236*** | Y38H6C.20 | 1.842 ± 0.3356 |
| C08H9.8 | 3.826 ± 0.469 | F09G2.1 | 2 ± 0.2425 |
| M176.6 | 4.741 ± 0.4356* | B0302.1 | 4.571 ± 0.3864 |
| M176.7 | 1.579 ± 0.2791 | D1073.1 | 1.5 ± 0.2415 |
| R09D1.12 | 3.969 ± 0.4804 | M79.1 | 5.172 ± 0.4915** |
| R09D1.13 | 2.438 ± 0.3287 | F59F5.3 | 2 ± 0.3162 |
| ZK938.5 | 5.515 ± 0.4809*** | B0198.3 | 1.818 ± 0.3872 |
| B0252.1 | 2.04 ± 0.3628 | F54F7.5 | 2 ± 0.3208 |
| C01G6.8 | 3.063 ± 0.359 | C16B8.1 | 1.938 ± 0.335 |
| M03A1.1 | 3 ± 0.3291 | C25F6.4 | 3.192 ± 0.4039 |
| B0523.1 | 4.188 ± 0.4002 | F11D5.3 | 1.813 ± 0.2617 |
| F57B9.8 | 1.563 ± 0.3412 | F58A3.2 | 1.938 ± 0.2495 |
| W03A5.1 | 2.875 ± 0.2869 | F59F3.1 | 2.063 ± 0.359 |
| C15H7.3 | 4.293 ± 0.3479 | F59F3.5 | 3.375 ± 0.3146 |
| T17A3.8 | 4.273 ± 0.4661 | T14E8.1 | 2.188 ± 0.3788 |
| R151.1 | 3.103 ± 0.2595 | F08F1.1 | 2.438 ± 0.3158 |
| C01C7.1 | 2.438 ± 0.4741 | F09A5.2 | 2.625 ± 0.3637 |
| C18H7.4 | 5.093 ± 0.4046*** | ZK1067.1 | 4.9 ± 0.2969** |
| C25A8.5 | 2.7 ± 0.4872 | C30F8.4a | 3.313 ± 0.3619 |
| C55C3.4 | 2.85 ± 0.4881 | M142.1 | 3.313 ± 0.3502 |
| F01D4.3 | 4.188 ± 0.366 | Y55D5A.5a.2 | 1.188 ± 0.4002 |
| F22B3.8 | 8.643 ± 0.8517*** | T17A3.1 | 2.5 ± 0.3028 |
| K07F5.4 | 3.95 ± 0.3507 | W02A2.6 | 3.063 ± 0.193 |
| K09B11.5 | 5.532 ± 0.3231*** | Y50D4B.6 | 2.813 ± 0.3191 |
| R11E3.1 | 2.565 ± 0.4066 | T22B11.4 | 5.522 ± 0.6188*** |
| T04B2.2 | 5.892 ± 0.3948*** | Y92H12A.1 | 9.625 ± 0.6575*** |
| T06C10.3 | 2.579 ± 0.3182 |
-
At least 15 adult worms were scored for each RNAi treatment. *p<0.05, **p<0.01, ***p<0.001.
-
Table 4—source data 1
Germ cell corpses in N2 treated with tyrosine kinases RNAi.
Including raw statistics.
- https://cdn.elifesciences.org/articles/76436/elife-76436-table4-data1-v2.zip
Cell corpse phenotypes caused by RNAi of C. elegans genes encoding SH2 domain proteins.
| C. elegans SH2 domain proteins (RNAi) | No. of germ cell corpses(mean ± SEM) | ||
|---|---|---|---|
| 1st | 2nd | 3rd | |
| Control | 4.27 ± 0.44 | 3.47 ± 0.46 | 3.27 ± 0.55 |
| chin-1 | 5.53 ± 0.62 | 7.27 ± 0.69*** | 4.73 ± 0.64 |
| shc-1 | 4.93 ± 0.53 | 4.93 ± 0.64 | 5.33 ± 0.73* |
| csk-1 | 5.07 ± 0.71 | 6.67 ± 0.78** | 7.00 ± 2.20 |
| F39B2.5 | 4.07 ± 0.49 | 5.80 ± 0.82* | 5.73 ± 0.73 |
| sli-1 | 4.27 ± 0.68 | 5.80 ± 0.77* | 4.80 ± 0.66 |
| sem-5 | 8.40 ± 1.08** | ND | ND |
| rin-1 | 5.13 ± 0.80 | 5.33 ± 0.64* | 6.00 ± 0.90* |
| F13B12.6 | 2.47 ± 0.42 | ND | ND |
| vav-1 | 4.73 ± 0.65 | 7.67 ± 0.56*** | 4.13 ± 0.43 |
| gap-3 | 6.20 ± 0.85 | 5.60 ± 0.59* | 5.20 ± 0.76 |
| nck-1 | 6.80 ± 0.76** | 7.00 ± 1.02** | 6.53 ± 0.36*** |
| tns-1 | 5.53 ± 0.85 | ND | 5.13 ± 0.74 |
| C18A11.4 | 6.20 ± 0.98 | 4.07 ± 0.42 | 6.93 ± 0.94** |
| Y43C5B.2 | 5.87 ± 0.80 | 4.33 ± 0.68 | 4.20 ± 0.96 |
| K11E4.2 | 5.00 ± 0.78 | 4.13 ± 0.40 | 4.00 ± 0.56 |
| plc-3 | 2.67 ± 0.34 | 2.13 ± 0.48 | 2.13 ± 0.48 |
| sta-2 | 3.80 ± 0.50 | 9.67 ± 2.67* | 7.33 ± 0.87*** |
| Y116A8C.38 | 3.47 ± 0.46 | 4.40 ± 0.58 | 4.20 ± 0.55 |
| soem-1 | 5.33 ± 0.42 | 3.27 ± 0.49 | 5.60 ± 0.61* |
| Y52D5A.2 | 5.27 ± 0.66 | 4.93 ± 0.75 | 4.47 ± 0.55 |
| sta-1 | 5.53 ± 0.58 | 5.40 ± 0.65* | 4.27 ± 0.62 |
| aap-1 | 3.47 ± 0.56 | 3.40 ± 0.41 | 4.67 ± 0.71 |
| shc-2 | 4.67 ± 0.54 | 3.93 ± 0.46 | 5.33 ± 0.62* |
| Y37D8A.4 | 5.93 ± 0.79 | 5.20 ± 0.78 | 4.73 ± 68 |
| ptp-1 | 9.53 ± 1.05 | 7.26 ± 0.91 | 3.6 ± 0.58 |
| emb-5 | ND | ND | ND |
-
15 adult worms were scored for each RNAi treatment. Data were from three independent experiments. *p<0.05, **p<0.01, ***p<0.001, ND, no data.
-
Table 5—source data 1
Germ cell corpses in N2 treated with SH2 domain proteins RNAi.
Including raw statistics.
- https://cdn.elifesciences.org/articles/76436/elife-76436-table5-data1-v2.zip
The factors identified by liquid chromatography–tandem mass spectrometry (LC-MS/MS) to be associated with the CED-1 pathway in C. elegans.
| Gene names | Number of proteins | Peptides | Unique peptides | Sequence coverage (%) | Mol. weight (kDa) | Sequence length | Sequence coverage No HS (%) | Sequence coverage HS (%) | LFQ intensity No HSP | LFQ intensity HS |
|---|---|---|---|---|---|---|---|---|---|---|
| hsp-16.1 | 2 | 2 | 2 | 18.6 | 16.253 | 145 | 0 | 18.6 | 0 | 63691000 |
| vha-10 | 1 | 2 | 2 | 15.1 | 14.485 | 126 | 0 | 15.1 | 0 | 1.71E+08 |
| iffb-1 | 1 | 1 | 1 | 1 | 120.42 | 1074 | 0 | 1 | 0 | 0 |
| his-35 | 4 | 1 | 1 | 5.5 | 13.418 | 127 | 0 | 5.5 | 0 | 2.69E+08 |
| CELE_ T10C6.7 | 1 | 1 | 1 | 3.8 | 37.523 | 317 | 0 | 3.8 | 0 | 0 |
| rpn-8 | 1 | 1 | 1 | 2.5 | 40.687 | 362 | 0 | 2.5 | 0 | 0 |
| sumv-1 | 1 | 1 | 1 | 1.1 | 112.21 | 1024 | 0 | 1.1 | 0 | 0 |
| CELE_ F55H12.4 | 1 | 1 | 1 | 3.8 | 22.043 | 208 | 0 | 3.8 | 0 | 86366000 |
| pdi-6 | 1 | 1 | 1 | 3.4 | 47.727 | 440 | 0 | 3.4 | 0 | 19651000 |
| C17C3.3 | 1 | 1 | 1 | 3.5 | 35.904 | 316 | 0 | 3.5 | 0 | 0 |
| gmeb-3 | 1 | 1 | 1 | 6.6 | 42.769 | 376 | 0 | 6.6 | 0 | 34430000 |
| F38B2.4 | 1 | 1 | 1 | 4.3 | 22.597 | 210 | 0 | 4.3 | 0 | 46574000 |
| npr-10 | 2 | 1 | 1 | 3 | 40.576 | 362 | 0 | 3 | 0 | 0 |
| otub-2 | 2 | 1 | 1 | 2.4 | 56.275 | 499 | 0 | 2.4 | 0 | 0 |
| CELE_ Y62H9A.5 | 1 | 1 | 1 | 4.2 | 18.496 | 165 | 0 | 4.2 | 0 | 79508000 |
-
LFQ indicated the protein signal intensity after correction by LFQ algorithm. As a result of relative quantification of proteins, it is usually used for screening of differential proteins between different samples.
-
Table 6—source data 1
The factors identified by LC-MS/MS.
Including raw statistics.
- https://cdn.elifesciences.org/articles/76436/elife-76436-table6-data1-v2.zip
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Caenorhabditis elegans) | ced-1::flag(xwh17) I 2× | This paper | SNU19 | Figure 1 Figure 4—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhIs27[Pced-1ced-1::flag, sur-5::gfp] | This paper | SNU31 | Figure 1 ; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhIs28[Phsp-16trim-21::flag, sur-5::gfp] | This paper | SNU32 | Figure 1 Figure 3 Figure 3—figure supplement 1 Figure 5—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | trim-21(xwh12) II 2× | This paper | SNU12 | Figure 1 Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | trim-21(xwh13) II 6× | This paper | SNU13 | Figure 1 Figure 6 Figure 6—figure supplement 1 Figure 7; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | smIs34[Pced-1ced-1::gfp, rol-6(su1006)] | Dr. Chonglin Yang | CU1546 | Figure 1 |
| Strain, strain background (C. elegans) | smIs110[Pced-1ced-1DC::gfp] | Dr. Chonglin Yang (Chen et al., 2013) | N/A | Figure 1 |
| Strain, strain background (C. elegans) | bcIs39[Plim-7ced-1::gfp, lin-15(+)] | Dr. Chonglin Yang | MD701 | Figure 1 |
| Strain, strain background (C. elegans) | trim-21(xwh12); smIs34[Pced-1ced-1::gfp, rol-6(su1006)] | This paper | SNU33 | Figure 1 Figure 6 Figure 7—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | trim-21(xwh12); bcIs39[Plim-7ced-1::gfp, lin-15(+)] | This paper | SNU34 | Figure 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | trim-21(xwh12); smIs110[Pced-1ced-1DC::gfp] | This paper | SNU35 | Figure 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ced-1::flag(xwh17); ha::ubq-2(xwh20) | This paper | SNU22 | Figure 2 Figure 3 Figure 4 Figure 4—figure supplement 1 Figure 5 Figure 5—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ubc-21(xwh15) X 3× | This paper | SNU15 | Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ubc-21(xwh16) X 6× | This paper | SNU16 | Figure 6 Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ubc-21(xwh15); ced-1::flag(xwh17); ha::ubq-2(xwh20) | This paper | SNU36 | Figure 2; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | trim-21(xwh12); ced-1::flag(xwh17); ha::ubq-2(xwh20) | This paper | SNU37 | Figure 2; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | trim-21(xwh12); ced-1::flag(xwh17); ha::ubq-2-K48R(xwh23) | This paper | SNU25 | Figure 2; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | trim-21(xwh12); ced-1::flag(xwh17); ha::ubq-2-K63R(xwh24) | This paper | SNU26 | Figure 2; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhIs29[Pced-1trim-21::flag, sur-5::gfp] | This paper | SNU38 | Figure 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ced-6(xwh25); xwhIs29[Pced-1trim-21::flag, sur-5:: gfp] | This paper | SNU39 | Figure 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | smIs34[Pced-1ced-1::gfp, rol-6(su1006)]; xwhIs30[Pced-1trim-21::mcherry, rol-6(su1006)] | This paper | SNU43 | Figure 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | smIs34[Pced-1ced-1::gfp, rol-6(su1006)]; xwhIs31[Pced-1mcherry::ced-6, rol-6(su1006)] | This paper | SNU44 | Figure 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhIs31[Pced-1mcherry::ced-6, rol-6(su1006)]; xwhIs32[Pced-1trim-21::gfp, rol-6(su1006)] | This paper | SNU45 | Figure 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx34[Pced-1ced-1::flag, sur-5:: gfp]/ced-1(e1735) | This paper | SNU47 | Figure 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx35[Pced-1ced-1(N962A)::flag, sur-5:: gfp]/ced-1(e1735) | This paper | SNU48 | Figure 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ced-6(xwh25) III | This paper | SNU27 | Figure 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ced-6(xwh25); xwhIs28[Phsp-16trim-21::flag, sur-5::gfp] | This paper | SNU49 | Figure 3 Figure 3—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ubc-21(xwh16); trim-21(xwh13) | This paper | SNU17 | Figure 6 Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | src-1(xwh26); +/hT2 III | This paper | SNU28 | Figure 4; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | qxIs408[Pced-1gfp::rab-5] | Dr. Chonglin Yang (Chen et al., 2013) | N/A | Figure 6 |
| Strain, strain background (C. elegans) | trim-21(xwh12); qxIs408[Pced-1gfp::rab-5] | This paper | SNU50 | Figure 6; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | qxIs66[Pced-1gfp::rab-7] | Dr. Chonglin Yang (Liu et al., 2012) | N/A | Figure 6 |
| Strain, strain background (C. elegans) | trim-21(xwh12); qxIs66[Pced-1gfp::rab-7] | This paper | SNU51 | Figure 6; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | qxIs354[Pced-1laat-1::gfp] | Dr. Chonglin Yang (Liu et al., 2012) | N/A | Figure 6 |
| Strain, strain background (C. elegans) | trim-21(xwh12); qxIs354[Pced-1laat-1::gfp] | This paper | SNU52 | Figure 6; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | qxIs257[Pced-1nuc-1::mcherry] | Dr. Chonglin Yang (Chen et al., 2013) | N/A | Figure 6 |
| Strain, strain background (C. elegans) | trim-21(xwh12); qxIs257[Pced-1nuc-1::mcherry] | This paper | SNU53 | Figure 6; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | yqEx620[Pced-1cpl-1::mchoint] | Dr. Chonglin Yang (Xu et al., 2014) | N/A | Figure 6 |
| Strain, strain background (C. elegans) | trim-21(xwh12); yqEx620[Pced-1cpl-1::mchoint] | This paper | SNU54 | Figure 6; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ced-1(e1735); trim-21(xwh13) | This paper | SNU61 | Figure 6; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ced-1(e1735); ubc-21(xwh16) | This paper | SNU62 | Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhIs36[Phsp-16ced-1-ct::flag, sur-5::gfp] | This paper | SNU63 | Figure 7; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ced-1(N962A)::flag(xwh17) I | This paper | SNU73 | Figure 4—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ced-1(Y1019F)::flag(xwh17) I | This paper | SNU74 | Figure 4—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ced-1(N962A)::flag(xwh17); ha::ubq-2(xwh20) | This paper | SNU75 | Figure 4—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ced-1(N962A)::flag(xwh17); ha::ubq-2(xwh20) | This paper | SNU76 | Figure 4—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx37[Pced-1trim-21::flag, sur-5:: gfp] line1 | This paper | SNU64 | Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx38[Pced-1trim-21::flag, sur-5:: gfp] line2 | This paper | SNU65 | Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx39[Pced-1trim-21::flag, sur-5:: gfp] line3 | This paper | SNU66 | Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx40[Ptrim-21trim-21::gfp, sur-5:: gfp] line1/trim-21(xwh13) | This paper | SNU67 | Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx41[Ptrim-21trim-21::gfp, sur-5:: gfp] line2/trim-21(xwh13) | This paper | SNU68 | Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx42[Ptrim-21trim-21::gfp, sur-5:: gfp] line3/trim-21(xwh13) | This paper | SNU69 | Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx43[Pced-1vha-10::mcherry, sur-5:: gfp] line1/trim-21(xwh13) | This paper | SNU70 | Figure 7; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx44[Pced-1vha-10::mcherry, sur-5:: gfp] line2/trim-21(xwh13) | This paper | SNU71 | Figure 7; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx45[Pced-1vha-10::mcherry, sur-5:: gfp] line3/trim-21(xwh13) | This paper | SNU72 | Figure 7; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx46[Ptrim-21htrim21, sur-5:: gfp] line1/trim-21(xwh13) | This paper | SNU77 | Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx47[Ptrim-21htrim21, sur-5:: gfp] line2/trim-21(xwh13) | This paper | SNU78 | Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx48[Ptrim-21htrim21, sur-5:: gfp] line3/trim-21(xwh13) | This paper | SNU79 | Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhIs49[Pced-1rde-1, rol-6(su1006)]/rde-1 | This paper | SNU81 | Figure 5; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ujIs113 [Ppie-1H2B::mCherry, unc-119(+); Pnhr-2HIS-24::mCherry, unc-119(+)] | Dr. Chonglin Yang | JIM113 | Figure 6 |
| Strain, strain background (C. elegans) | trim-21(xwh13); ujIs113 [Ppie-1H2B::mCherry, unc-119(+); Pnhr-2HIS-24::mCherry, unc-119(+)] | This paper | SNU80 | Figure 6; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ubc-21(xwh15); smIs34[Pced-1ced-1::gfp, rol-6(su1006)] | This paper | SNU55 | Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ubc-21(xwh15); qxIs408[Pced-1gfp::rab-5] | This paper | SNU56 | Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | qxIs68[Pced-1mcherry::rab-7] | Dr. Chonglin Yang (Cheng et al., 2015) | N/A | Figure 6—figure supplement 1 |
| Strain, strain background (C. elegans) | ubc-21(xwh15); qxIs68[Pced-1mcherry::rab-7] | This paper | SNU57 | Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | qxIs352[Pced-1laat-1::mcherry] | Dr. Chonglin Yang (Liu et al., 2012) | N/A | Figure 6—figure supplement 1 |
| Strain, strain background (C. elegans) | ubc-21(xwh15); qxIs352[Pced-1laat-1::mcherry] | This paper | SNU58 | Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ubc-21(xwh15); qxIs257[Pced-1nuc-1::mcherry] | This paper | SNU59 | Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ubc-21(xwh15); yqEx620[Pced-1cpl-1::mchoint] | This paper | SNU60 | Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | nck-1(xwh51) | This paper | SNU83 | Figure 5—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ced-6(xwh25); smIs34[Pced-1ced-1::gfp, rol-6(su1006)]; xwhIs30[Pced-1trim-21::mcherry, rol-6(su1006)] | This paper | SNU85 | Figure 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ced-6(xwh25); xwhIs32[Pced-1trim-21::gfp, rol-6(su1006)] | This paper | SNU86 | Figure 3—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhIs53[Pced-1ced-1::gfp, Podr-1:: rfp] | This paper | SNU87 | Figure 5; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhIs54[Pced-1ced-1(Y1019F)::gfp, rol-6(su1006)] | This paper | SNU88 | Figure 5; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | trim-21(xwh13); xwhIs27[Pced-1ced-1::flag, sur-5::gfp] | This paper | SNU90 | Figure 6—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | snx-1(tm847) | Dr. Chonglin Yang (Chen et al., 2010) | N/A | Figure 6 |
| Strain, strain background (C. elegans) | qxIs58[Pced-1lmp-1::mcherry] | Dr. Chonglin Yang (Sasaki et al., 2013) | N/A | Figure 7—figure supplement 1 |
| Strain, strain background (C. elegans) | snx-1(tm847); smIs34[Pced-1ced-1::gfp, rol-6(su1006)] | This paper | SNU91 | Figure 7—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | trim-21(xwh13); smIs34[Pced-1ced-1::gfp, rol-6(su1006)] | This paper | SNU92 | Figure 7—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | trim-21(xwh13); snx-1(tm847); smIs34[Pced-1ced-1::gfp, rol-6(su1006)] | This paper | SNU93 | Figure 7—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | smIs34[Pced-1ced-1::gfp, rol-6(su1006)]; qxIs68[Pced-1mcherry::rab-7] | This paper | SNU94 | Figure 7—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | trim-21(xwh13); smIs34[Pced-1ced-1::gfp, rol-6(su1006)]; qxIs68[Pced-1mcherry::rab-7] | This paper | SNU95 | Figure 7—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | snx-1(tm847); smIs34[Pced-1ced-1::gfp, rol-6(su1006)]; qxIs68[Pced-1mcherry::rab-7] | This paper | SNU96 | Figure 7—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | trim-21(xwh13); snx-1(tm847); smIs34[Pced-1ced-1::gfp, rol-6(su1006)]; qxIs68[Pced-1mcherry::rab-7] | This paper | SNU97 | Figure 7—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | smIs34[Pced-1ced-1::gfp, rol-6(su1006)]; qxIs58[Pced-1lmp-1::mcherry] | This paper | SNU98 | Figure 7—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | trim-21(xwh13); smIs34[Pced-1ced-1::gfp, rol-6(su1006)]; qxIs58[Pced-1lmp-1::mcherry] | This paper | SNU99 | Figure 7—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | snx-1(tm847); smIs34[Pced-1ced-1::gfp, rol-6(su1006)]; qxIs58[Pced-1lmp-1::mcherry] | This paper | SNU100 | Figure 7—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | trim-21(xwh13); snx-1(tm847); smIs34[Pced-1ced-1::gfp, rol-6(su1006)]; qxIs58[Pced-1lmp-1::mcherry] | This paper | SNU101 | Figure 7—figure supplement 1; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | trim-21(xwh13); xwhIs53[Pced-1ced-1::gfp, Podr-1:: rfp] | This paper | SNU105 | Figure 7—figure supplement 2; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | snx-1(tm847); xwhIs53[Pced-1ced-1::gfp, Podr-1:: rfp] | This paper | SNU106 | Figure 7—figure supplement 2; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx55[Pced-1vha-10::mcherry/ xwhIs53[Pced-1ced-1::gfp, Podr-1:: rfp]] | This paper | SNU102 | Figure 7—figure supplement 2; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx56[Pced-1vha-10::mcherry/ trim-21(xwh13); xwhIs53[Pced-1ced-1::gfp, Podr-1:: rfp]] | This paper | SNU103 | Figure 7—figure supplement 2; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx57[Pced-1vha-10::mcherry/ snx-1(tm847); xwhIs53[Pced-1ced-1::gfp, Podr-1:: rfp]] | This paper | SNU104 | Figure 7—figure supplement 2; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | ha::vha-10(xwh52) | This paper | SNU89 | Figure 7—figure supplement 2; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | trim-21(xwh13); ha::vha-10(xwh52) | This paper | SNU107 | Figure 7—figure supplement 2; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | snx-1(tm847); ha::vha-10(xwh52) | This paper | SNU108 | Figure 7—figure supplement 2; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx58[Pced-1ced-1, sur-5:: gfp] line2/ced-1(e1735) | This paper | SNU109 | Table 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx59[Pced-1ced-1, sur-5:: gfp] line1/ced-1(e1735) | This paper | SNU110 | Table 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx60[Pced-1ced-1, sur-5:: gfp] line3/ced-1(e1735) | This paper | SNU111 | Table 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx61[Pced-1ced-1(N962A), sur-5:: gfp] line2/ced-1(e1735) | This paper | SNU112 | Table 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx62[Pced-1ced-1(N962A), sur-5:: gfp] line1/ced-1(e1735) | This paper | SNU113 | Table 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx63[Pced-1ced-1(N962A), sur-5:: gfp] line3/ced-1(e1735) | This paper | SNU114 | Table 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx64[Pced-1ced-1(Y965F), sur-5:: gfp] line2/ced-1(e1735) | This paper | SNU115 | Table 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx65[Pced-1ced-1(Y965F), sur-5:: gfp] line1/ced-1(e1735) | This paper | SNU116 | Table 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx66[Pced-1ced-1(Y965F), sur-5:: gfp] line3/ced-1(e1735) | This paper | SNU117 | Table 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx67[Pced-1ced-1(Y1019F), sur-5:: gfp] line2/ced-1(e1735) | This paper | SNU118 | Table 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx68[Pced-1ced-1(Y1019F), sur-5:: gfp] line1/ced-1(e1735) | This paper | SNU119 | Table 3; Available from the Xiao Lab |
| Strain, strain background (C. elegans) | xwhEx69[Pced-1ced-1(Y1019F), sur-5:: gfp] line3/ced-1(e1735) | This paper | SNU120 | Table 3; Available from the Xiao Lab |
| Strain, strain background (Escherichia coli) | OP50 | Dr. Chonglin Yang (Chen et al., 2010) | N/A | |
| Strain, strain background (E. coli) | DH5α | TaKaRa | Cat# 9057 | |
| Strain, strain background (E. coli) | HT115 | Dr. Chonglin Yang (Chen et al., 2010) | N/A | |
| Strain, strain background (E. coli) | BL21 | TaKaRa | Cat# 9126 | |
| Strain, strain background (E. coli) | BL21(DE3) | Solarbio | Cat# C1400 | |
| Strain, strain background (Saccharomyces cerevisiae) | Y2HGold | Clontech | Cat# 630498 | |
| Antibody | Anti-CED-1 (rabbit polyclonal) | Dr. Chonglin Yang (Chen et al., 2010) | N/A | IB(1:1000) Available from the Xiao Lab |
| Antibody | Anti-β-actin (mouse polyclonal) | This paper | N/A | IB(1:1000) Available from the Xiao Lab |
| Antibody | Anti-FLAG tag (DYKDDDDK) (rabbit polyclonal) | Sigma-Aldrich | Cat# SAB1306078 | IB(1:1000) |
| Antibody | Anti-Ub(P4D1) IgG1 (mouse monoclonal) | Santa Cruz Biotechnology | Cat# SC-8017; RRID:AB_628423 | IB(1:500) |
| Antibody | Anti-GST (mouse monoclonal) | Engibody | Cat# AT0027 | IB(1:1000) |
| Antibody | Anti-GST (mouse polyclonal) | This paper | N/A | IB(1:1000) Available from the Xiao Lab |
| Antibody | Anti-CED-1 (mouse polyclonal) | This paper | N/A | IB(1:1000) Available from the Xiao Lab |
| Antibody | Anti-FLAG M2 antibody (mouse monoclonal) | Sigma-Aldrich | Cat# F1804; RRID:AB_262044 | IB(1:1000) |
| Antibody | Anti-C-MYC-antibody (rabbit polyclonal) | Sigma-Aldrich | Cat# SAB4301136 | IB(1:1000) |
| Antibody | Anti-GFP (rabbit polyclonal) | Engibody | Cat# 1598 | IB(1:1000) |
| Antibody | Anti-GFP (mouse polyclonal) | This paper | N/A | IB(1:1000) Available from the Xiao Lab |
| Antibody | Anti-HA-Tag(C29F4) (rabbit monoclonal) | Cell Signaling Technology | Cat# 3724; RRID:AB_1549585 | IB(1:1000) |
| Antibody | Anti-CED-6 (rabbit polyclonal) | Dr. Chonglin Yang | N/A | IB(1:1000) Available from the Xiao Lab |
| Antibody | Anti-CED-6 (mouse polyclonal) | This paper | N/A | IB(1:1000) Available from the Xiao Lab |
| Antibody | Anti-P-tyrosine (P-Tyr-100) (mouse monoclonal) | Cell Signaling Technology | Cat# 9411; RRID:AB_331228 | IB(1:1000) |
| Antibody | Anti-NCK-1 (mouse polyclonal) | This paper | N/A | IB(1:1000) Available from the Xiao Lab |
| Antibody | Peroxidase-conjugated AffiniPure Goat Anti-Rabbit IgG(H+L) | Jackson ImmunoResearch | Cat# 111-035-003; RRID:AB_2313567 | IB(1:10,000) |
| Antibody | Peroxidase-conjugated AffiniPure Goat Anti-Mouse IgG(H+L) | Jackson ImmunoResearch | Cat# 115-035-003; RRID:AB_10015289 | IB(1:10,000) |
| Commercial assay or kit | Anti-FLAG M2 Affinity Gel | Sigma-Aldrich | Cat# A2220; RRID:AB_10063035 | |
| Commercial assay or kit | Anti-FLAG M2 Magnetic Beads | Sigma-Aldrich | Cat# M8823; RRID:AB_2637089 | |
| Commercial assay or kit | Pierce Streptavidin Magnetic Beads | Thermo Fisher Scientific | Cat# 88817 | |
| Commercial assay or kit | Ni-NTA Superflow | QIAGEN | Cat# 1018611 | |
| Commercial assay or kit | Glutathione Sepharose 4B | GE Healthcare | Cat# 17-0756 | |
| Commercial assay or kit | Glutathione High Capacity Magnetic Agarose Beads | Sigma-Aldrich | Cat# G0924 | |
| Commercial assay or kit | PureProteome Protein A/G Mix Magnetic Beads | Millipore | Cat# LSKMAGAG | |
| Commercial assay or kit | Pierce Anti-HA Magnetic Beads | Thermo Scientific | Cat# 88837 | |
| Chemical compound, drug | Glutathione, reduced | VWR AMRESCO | Cat# 0399 | |
| Chemical compound, drug | Cycloheximide (CHX) | INALCO SPA, Milan, Italy | Cat# 1758-9310 | |
| Chemical compound, drug | MG-132 | Selleck | Cat# S2619 | |
| Chemical compound, drug | Imidazole | Millipore | Cat# 288-32-4 | |
| Chemical compound, drug | TRIzol Reagent | Ambion | Cat# 15596018 | |
| Commercial assay or kit | Pro-Q Diamond Phosphoprotein Gel Stain | Invitrogen | Cat# P33301 | |
| Commercial assay or kit | HiScript III RT SuperMix for qPCR (+gDNA wiper) | Vazyme | Cat# R323 | |
| Commercial assay or kit | ChamQ Universal SYBR qPCR Master Mix | Vazyme | Cat# Q711 | |
| Recombinant DNA reagent | pGBKT7-ced-1-ct | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGBKT7-ubc-21 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGBKT7-ced-1-nt | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGBKT7-ced-1-ct-A | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGBKT7-ced-1-ct-B | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGBKT7-ced-1-ct-C | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGBKT7-ced-1(Y965F)-ct | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGBKT7-ced-1(Y1019F)-ct | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGBKT7-ced-1(N962A)-ct | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGADT7-trim-21 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGADT7-f43c11.7 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGADT7-k01g5.1 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGADT7-trim-21-ΔRING | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGADT7-trim-21-ΔBBOX | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGADT7-trim-21-ΔCC | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGADT7-trim-21-nt | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGADT7-trim-21-ct | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGADT7-ced-6-PTB | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGADT7-src-1 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGADT7-src-1(1-261aa) | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGADT7-src-1(229-533aa) | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGADT7-src-1-ΔSH2 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGADT7-src-1-ΔSH3 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGADT7-src-1-ΔTyrkc | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGADT7-nck-1 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGEX-KG-ced-1-ct | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGEX-KG-trim-21 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGEX-KG-ubc-21 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGEX-KG-ced-6 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pGEX-KG-htrim21 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pET28a-ced-1-ct-flag | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pET28a-myc-trim-21 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pET28a-ha-ubq-2 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pET28a-ha-ubq-2(K48R) | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pET28a-ha-ubq-2(K63R) | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pET28a-uba-1 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pET28a-myc-ubc-21 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pET28a-ha-ced-6 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pET28a-ced-1(Y965F)-ct-flag | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pET28a-ced-1(Y1019F)-ct-flag | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pET28a-ced-1(N962A)-ct-flag | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pET28a-myc-src-1 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pET28a-ha-vha-10 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pET28a-myc-trim-21-ΔRING | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pET28a-myc-trim-21-ΔBBOX | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pET28a-myc-trim-21-ΔCC | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pET28a-ha-nck-1 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pET28a-ha-megf10-ct | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pcDNA3.1-myc-src-1 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pcDNA3.1-ha-ced-1-ct-miniturbo | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pcDNA3.1-myc-ced-1-ct-miniturbo | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pcDNA3.1-ha-nck-1 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pcDNA3.1-myc-trim-21-miniturbo | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pPD49.26-Pced-1ced-1-flag | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pPD49.78-trim-21-flag | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pPD49.83-trim-21-flag | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pPD49.26-Pced-1trim-21-flag | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pPD49.26-Pced-1mcherry-ced-6 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pPD49.26-Pced-1trim-21-gfp | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pPD49.26-Pced-1ced-1(N962A)-flag | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pPD49.78-ced-1-ct-flag | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pPD49.83-ced-1-ct-flag | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pPD49.26-Pced-1vha-10-mcherry | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pPD95.77-Ptrim-21trim-21-mcherry | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pPD95.77-Ptrim-21trim-21-gfp | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pPD95.77-Ptrim-21trim21 | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pPD49.26-Pced-1ced-1-gfp | This paper | N/A | Available from the Xiao Lab |
| Recombinant DNA reagent | pPD49.26-Pced-1ced-1(Y1019F)-gfp | This paper | N/A | Available from the Xiao Lab |
| Sequence-based reagent | trim-21 sgRNA targeting sequence | This paper | N/A | GACTTCTCAAGTGAGGAGGATGG |
| Sequence-based reagent | ubc-21 sgRNA targeting sequence | This paper | N/A | TCGCATTGGCACGGGTCACACGG |
| Sequence-based reagent | ced-1-flag sgRNA targeting sequence | This paper | N/A | TGCGAACAAAAAACGTGCTCAGG |
| Sequence-based reagent | ha-ubq-2 sgRNA targeting sequence | This paper | N/A | AATCTTCGTCAAGACTCTGACGG |
| Sequence-based reagent | ced-1(N962A)-flag sgRNA targeting sequence | This paper | N/A | GGCCGAGAATTCCAGAATCCCCT |
| Sequence-based reagent | ced-1(Y1019F)-flag sgRNA targeting sequence | This paper | N/A | CCCAGACGACTACGCCTCCCTGG |
| Sequence-based reagent | src-1 sgRNA targeting sequence | This paper | N/A | GCGATCGGGAGGCAGTGATATGG |
| Sequence-based reagent | ha-ubq-2(K48R) sgRNA targeting sequence | This paper | N/A | AATTTCAGGAAAGCAACTCGAGG |
| Sequence-based reagent | ha-ubq-2(K63R) sgRNA targeting sequence | This paper | N/A | TTGGTGCTCCGTCTTCGTGGAGG |
| Sequence-based reagent | ced-6 sgRNA targeting sequence | This paper | N/A | GTCGGTGGAAATAATATTAATGG |
| Sequence-based reagent | nck-1 sgRNA targeting sequence | This paper | N/A | ATACGATTATTTAGCACAAGAGG |
| Sequence-based reagent | ha-vha-10 sgRNA targeting sequence | This paper | N/A | CAGTACCGAAAACCTTAAAATGG |
| Sequence-based reagent | QPCR, tbg-1, forward | This paper | N/A | cgtcatcagcctggtagaaca |
| Sequence-based reagent | QPCR, tbg-1, reverse | This paper | N/A | tgatgactgtccacgttgga |
| Sequence-based reagent | QPCR, ced-1, forward | This paper | N/A | ggatggactggaaaacattgtg |
| Sequence-based reagent | QPCR, ced-1, reverse | This paper | N/A | cggattcgcattgacattgg |
| Software, algorithm | SMART | EMBL | http://smart.embl-heidelberg.de/ | |
| Software, algorithm | ImageJ | NIH | https://imagej.nih.gov/ij/download.html | |
| Software, algorithm | GraphPad Prism 8 | GraphPad Software | https://www.graphpad.com/scientific-software/prism/ | |
| Software, algorithm | ClustalW2 | EMBL-EBI | https://www.ebi.ac.uk/Tools/msa/clustalw2/ | |
| Software, algorithm | ZEN 2 pro | ZEISS | https://www.zeiss.com/microscopy/int/products/microscope-software/zen.html |





