Rescue of Escherichia coli auxotrophy by de novo small proteins
Figures
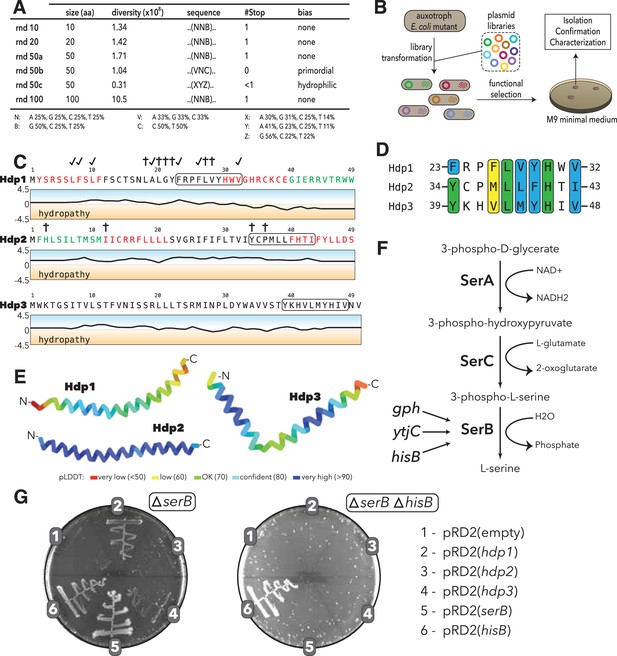
Experimental setup and sequence characteristics of the isolated small proteins.
(A) Libraries cloned into the expression vector pRD2. (B) Plasmid transformation into auxotrophic mutants and selection for rescue of auxotrophic mutants. (C) Hydropathy profiles of the three isolated small proteins (Hdp1-3). Colored amino acids denote residues that could (green) or could not (red) be removed while maintaining functionality. Loss-of-function mutations for Hdp1 and Hdp2 are indicated by daggers (†), and mutations that retain rescue activity are indicated by check marks; mutations are further described in Supplementary file 1b. The box spanning 10 amino acids denotes a potential region of similarity between the small proteins. (D) Sequence alignment showing the putative region of similarity shared between the three small proteins. Green coloring indicates identical amino acids, while blue and yellow coloring indicate strongly and weakly similar amino acids, respectively. (E) Structure prediction of Hdp1-3 obtained from AlphaFold (Jumper et al., 2021; Mirdita et al., 2022). Colors represent the per-residue prediction confidence level (pLDDT), based on the lDDT-cα metric (Mariani et al., 2013). (F) The three enzymes SerA, SerC, and SerB catalyze the last step in L-serine biosynthesis. Essentiality of SerB on minimal medium can be suppressed by overexpression of the phosphatases Gph, YtjC, or HisB. (G) Growth of E. coli ∆serB and ∆serB ∆hisB deletion strains carrying either the empty pRD2 plasmid or pRD2 encoding Hdp1-3, SerB, or HisB on M9 minimal medium supplemented with 50 μg/ml ampicillin, 0.2% glucose, and 1 mM IPTG. Pictures taken after 1-wk incubation at 37°C. Growth on M9-glucose indicates the ability of the cloned construct to rescue ∆serB auxotrophy.
-
Figure 1—source data 1
Random sequence libraries cloned into the expression vector pRD2.
- https://cdn.elifesciences.org/articles/78299/elife-78299-fig1-data1-v2.xlsx
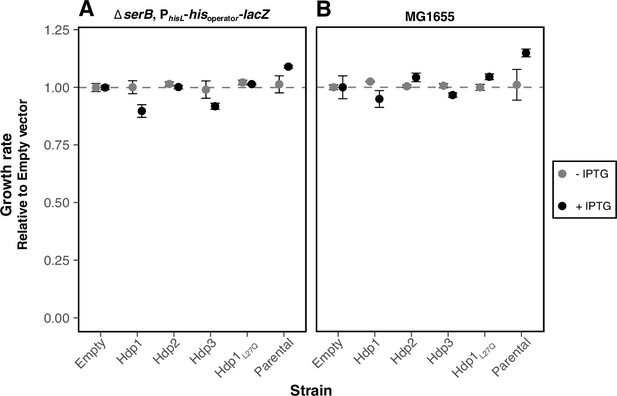
Growth rates of strains expressing Hdps relative to empty pRD2 control.
Relative exponential phase growth rates of E. coli strains expressing various Hdp constructs in either the (A) ∆sierB PhisL-hisoperator-lacZ reporter or (B) MG1655 wild-type genetic background. Growth rate measurements were performed at 37°C in LB medium supplemented with 50 µg/ml ampicillin ±1 mM IPTG. The values reported are relative to those measured for the empty plasmid strain with the corresponding genetic background and growth conditions and represent the mean of three biological replicates (with two technical replicates each); error bars represent the standard deviation. For each genetic background, ‘Parental’ strain is the corresponding E. coli strain without plasmid. Hdp1 and Hdp3 show a slight, but measurable fitness cost in rich medium. While the direct reason for this cost is unclear, it is likely to be associated with the strength of the rescue activity, rather than some general metabolic cost, since Hdp2, which confers the weakest rescue, is less costly than Hdp1 and Hdp3, and the L27Q mutation concomitantly abrogates the rescuing ability and cost of Hdp1.
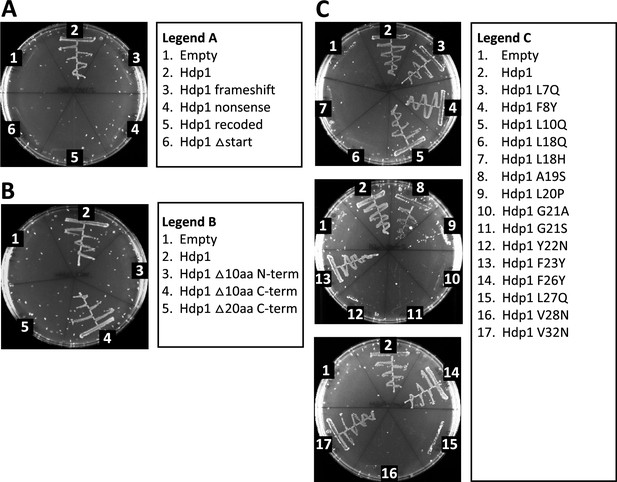
Characterization of Hdp1 mutations and truncations.
Growth of the E. coli ∆serB auxotrophic strain carrying either the empty pRD2 plasmid or pRD2 containing various mutations or truncations to the Hdp1 coding sequence on M9 minimal medium supplemented with 50 µg/ml ampicillin, 0.2% glucose, and 1 mM IPTG. Pictures taken after 1-wk incubation at 37°C. Growth on M9-glucose indicates ability of the cloned construct to rescue the ∆serB auxotroph. (A) Mutations characterizing Hdp1 as a protein-coding sequence. (B) Truncational analyses of Hdp1 function. (C) Functional analyses of Hdp1 mutations generated during random mutagenesis. For cloned insert sequences and a summary of random mutations, refer to Supplementary file 1f and b, respectively.
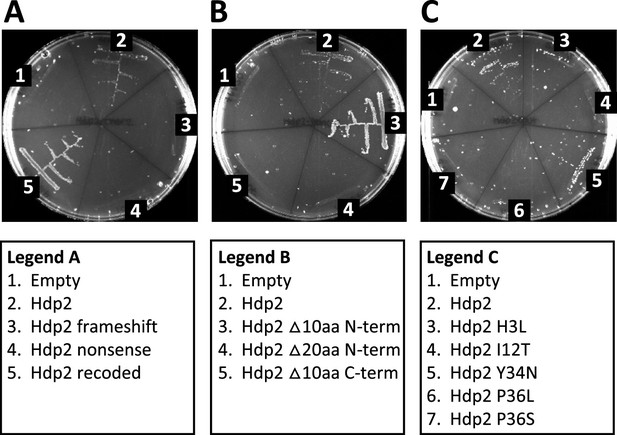
Characterization of Hdp2 mutations and truncations.
Growth of the E. coli ∆serB auxotrophic strain carrying either the empty pRD2 plasmid or pRD2 containing various mutations or truncations to the Hdp2 coding sequence on M9 minimal medium supplemented with 50 µg/ml ampicillin, 0.2% glucose, and 1 mM IPTG. Pictures taken after 1-wk incubation at 37°C. Growth on M9-glucose indicates ability of the cloned construct to rescue the ∆serB auxotroph. (A) Mutations characterizing Hdp2 as a protein-coding sequence. (B) Truncational analyses of Hdp2 function. (C) Functional analyses of Hdp2 mutations generated during random mutagenesis. For cloned insert sequences and a summary of random mutations, refer to Supplementary file 1f and b, respectively.

Hdp structure predictions using JPred4.
The consensus prediction of Hdp1-3 based on the secondary structure prediction server JPred4. Predicted helices are indicated with red and sheets are marked as green arrows. Confidence values corresponding to the confidence estimate of the prediction are shown; higher values indicate higher confidence.
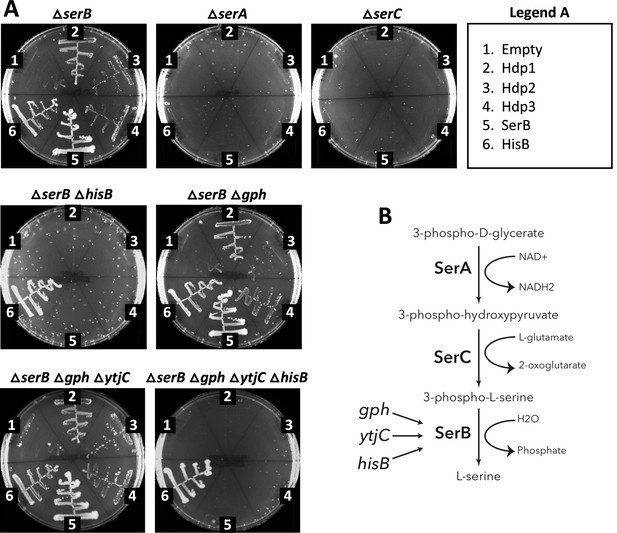
Hdp complementation of various deletion strains.
(A) Growth of various E. coli deletion strains carrying either the empty pRD2 plasmid or pRD2 encoding Hdp1-3, SerB, or HisB on M9 minimal medium supplemented with 50 µg/ml ampicillin, 0.2% glucose, and 1 mM IPTG. Pictures taken after 1-wk incubation at 37°C. Growth on M9-glucose indicates ability of the cloned construct to rescue the ∆serB auxotroph. (B) The last three steps of the serine biosynthetic pathway, showing the alternate phosphatases, Gph, YtjC, and HisB, that can complete serine biosynthesis in lieu of SerB, from main text Figure 1F.
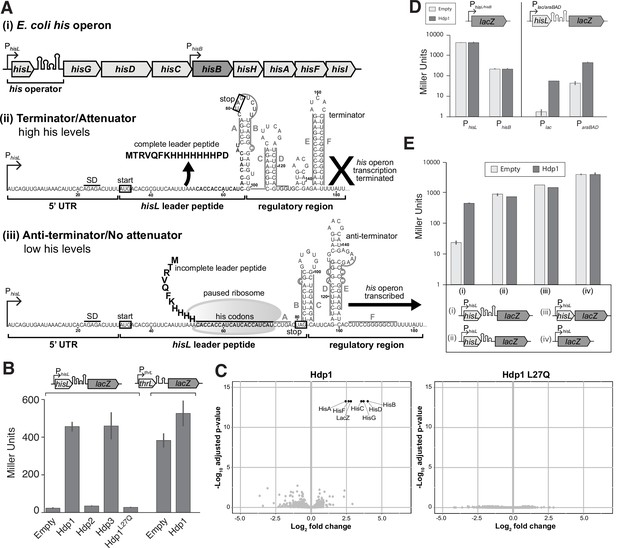
Overview of his operon regulation in E. coli and regulatory activity of the Hdps.
(A) (i) The regulatory region and structural genes of the his operon, including hisB (highlighted in dark gray), (ii) the his operator and RNA secondary structure under histidine-rich conditions, and (iii) the his operator and RNA secondary structure under histidine starvation. PhisL is the promoter; hisL is the leader peptide; SD is the Shine-Dalgarno sequence for hisL; the start and stop codons of the hisL coding region are boxed. Nucleotides are numbered from the transcription start site, +1. RNA secondary structures are adapted from Johnston et al., 1980; Kolter and Yanofsky, 1982. (B) β-Galactosidase activity (in Miller Units) of the strain carrying the full-length PhisL-hisoperator-lacZ reporter upon expression of Hdp1-3 and the Hdp1 L27Q mutant, and the strain containing a PthrL-throperator-lacZ reporter upon the expression of Hdp1. (C) ‘Volcano plots’ showing significant changes in protein abundance in the ΔserB mutant containing the full-length PhisL-hisoperator-lacZ reporter upon expression of Hdp1 or the Hdp1 L27Q nonfunctional mutant versus the empty plasmid control. An adjusted p-value cut-off of 0.01 and log2 fold change cut-off of 2 was used. Due to their small size, HisL and the Hdp variants were not recovered in these proteomics samples. Total proteome analysis was performed with at least three biological replicates for each strain. Some data points overlap. (D) β-Galactosidase activity of various lacZ reporter constructs under the control of different promoters upon expression of Hdp1 versus the empty plasmid control: lacZ transcriptional fusions under the control of the native hisL and hisB promoters (no his operator regulatory sequence) and the full-length hisoperator-lacZ reporter construct under the control of the IPTG-inducible Plac or arabinose-inducible ParaBAD promoters. (E) β-Galactosidase activity of various truncated PhisL-hisoperator-lacZ reporter constructs upon expression of Hdp1 versus the empty plasmid control. Refer to Supplementary file 1c for the nucleotide sequences of each lacZ reporter construct. For the β-galactosidase data presented in panels (B, D, E), the values reported represent the mean of three or more independent biological replicates; error bars represent the standard deviation.
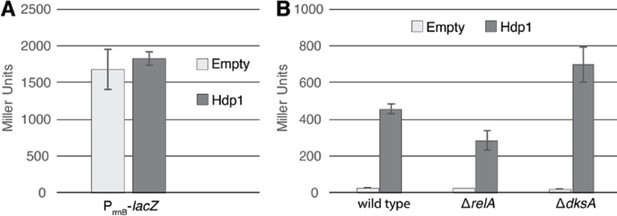
Hdp regulatory activity is independent of the stringent response.
(A) β-Galactosidase activity (in Miller Units) of the ∆serB strain carrying a lacZ transcriptional fusion under the control of the rrnB P1 promoter and accompanying regulatory sequence upon expression of Hdp1 versus the empty plasmid control. (B) β-Galactosidase activity (in Miller Units) of the ∆serB strain containing the full-length PhisL-hisoperator-lacZ reporter (denoted as ‘wild type’) and deletions of select stringent response genes upon expression of Hdp1 versus the empty plasmid control. The values reported represent the mean of three or more independent experimental replicates; error bars represent the standard deviation.
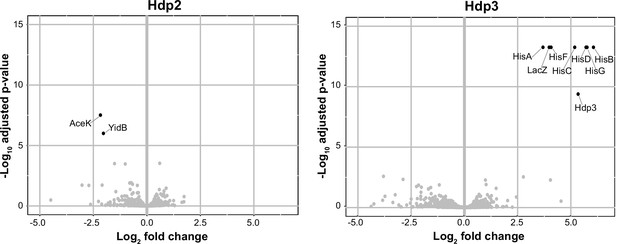
Protein abundance of strains expressing Hdps relative to the empty plasmid control strain.
‘Volcano plots’ showing significant changes in protein abundance in the ∆serB mutant containing the full-length PhisL-hisoperator-lacZ reporter upon expression of Hdp2 and Hdp3, versus the same strain expressing the empty plasmid control. An adjusted p-value cut-off of 0.01 and log2 fold change cut-off of 2 was used. Due to its small size, HisL and Hdp2 were not recovered in these proteomics samples; however, Hdp3 was detected. Some data points overlap. Total proteome analysis was performed on three biological replicates for each strain, grown in rich LB medium +1 mM IPTG.
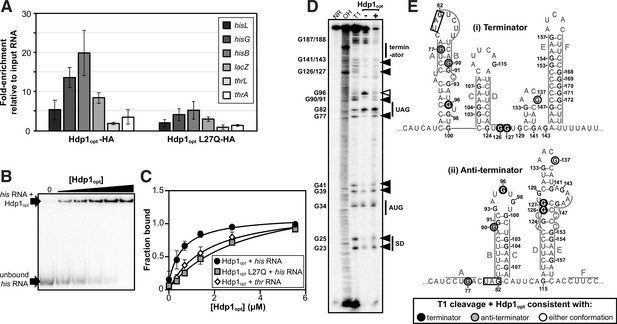
Characterization of Hdp1opt binding interactions with the his operator mRNA.
(A) Enrichment of select RNA transcripts in the HA-tagged Hdp1opt and Hdp1opt L27Q mutant pull-down samples, as quantified by RT-qPCR. thrL and thrA are nonspecific control RNAs. RT-qPCR data from pull-downs performed with untagged Hdp1opt was used as a normalization/background control. The data reported is the mean of three independent experimental replicates; error bars represent the standard deviation. (B) Representative electrophoretic mobility shift assay (EMSA) of the full-length his operator RNA in the presence of increasing concentrations of Hdp1opt (0–5.5 μM). Refer to Figure 3—figure supplement 3 for representative EMSA gels for all RNA and protein combinations assayed. All EMSAs were repeated independently three or more times. (C) Binding curves obtained from fitting the data quantified from the EMSAs to a standard 1:1 hyperbolic binding model. Each data point represents the mean of three or more independent experimental replicates; error bars represent the standard error. Some error bars fall within the boundaries of the markers. For the parameters used and the EMSA data fit to a binding model accounting for positive cooperativity, refer to Supplementary file 1d and e. (D) RNase T1 probing gel for the full-length his operator RNA in the absence and presence of Hdp1opt (0 and 5.5 μM, respectively). NR denotes RNA subject to no reaction, OH indicates partial alkaline hydrolysis, and T1 is an RNase T1 digest of the RNA under denaturing conditions used to map the RNA sequence. Numbering of G nucleotides is shown on the left. Arrows highlight changes in RNA cleavage in the presence of Hdp1opt: black arrows indicate nucleotides with increased cleavage and white arrows indicate reduced cleavage. Sequence and/or structure characteristics of the his operator RNA are also indicated on the right (i.e., the Shine-Dalgarno sequence [SD], start codon [AUG], and stop codon [UAG] of the hisL leader peptide coding sequence). This experiment was repeated three independent times with similar results; a representative gel is shown. The data from additional replicates can be found in Figure 3—figure supplement 4. (E) Nucleotides demonstrating a change in RNase T1 cleavage in the presence of Hdp1opt mapped to both the (i) terminator and (ii) anti-terminator conformations of the his operator RNA. The conformation consistent with cleavage in the presence of Hdp1opt is color-coded according to the legend; the hisL stop codon is boxed. Nucleotides are numbered from the transcription start site,+1.
-
Figure 3—source data 1
Source data for Figure 3B and D.
- https://cdn.elifesciences.org/articles/78299/elife-78299-fig3-data1-v2.zip
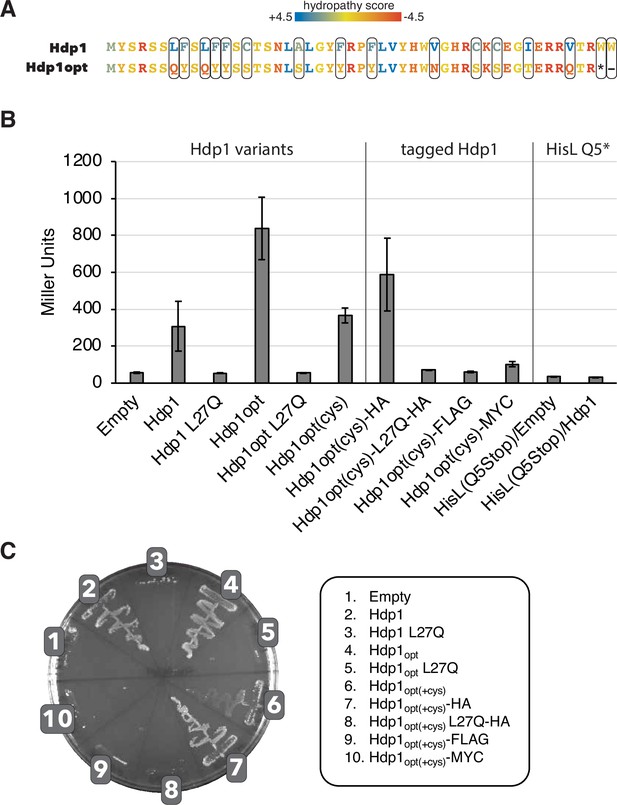
Characterization of Hdp1opt, the Hdp1 variant optimized for increased water solubility.
(A) Amino acid changes that generated the Hdp1opt variant with improved water solubility. Hdp1opt(+cys) is a prototype Hdp1opt variant that retains its cysteine residues. Colors represent the hydropathy score of each amino acid. The grand average of hydropathy (GRAVY) was reduced from –0.043 (Hdp1) to –1.46 (Hdp1opt). (B) β-Galactosidase activity (in Miller Units) of strains carrying the full-length PhisL-hisoperator-lacZ reporter upon expression of the originally isolated Hdp1 protein or the optimized Hdp1opt variants versus the empty plasmid control (left). The function of C-terminal-tagged versions of Hdp1opt(+cys) were also assessed for use in the in vivo co-immunoprecipitation assays (middle), as well as the regulatory activity of a PhisL-hisoperator-lacZ reporter containing a missense mutation in the hisL coding region (Q5stop, right). The values reported represent the mean of three or more independent experimental replicates; error bars represent the standard deviation. (C) Growth of the E. coli ∆serB auxotrophic strain carrying either the empty pRD2 plasmid or pRD2 containing the various Hdp1 constructs on M9 minimal medium supplemented with 50 µg/ml ampicillin, 0.2% glucose, and 1 mM IPTG. Pictures taken after 1-wk incubation at 37°C. Growth on M9-glucose indicates ability of the cloned construct to rescue the ∆serB auxotroph.
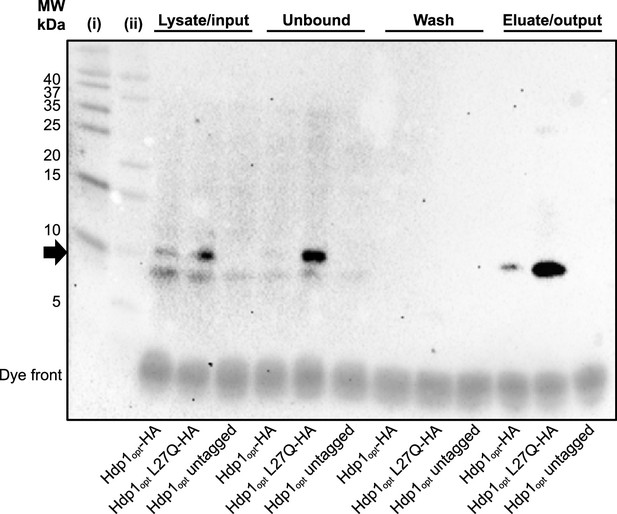
Western blot detection of HA-tagged Hdp1 variants following co-immunoprecipitation assays.
Protein fractions of the different steps of the co-immunoprecipitation experiments were separated on 16.5% Tris-Tricine polyacrylamide gels and the HA-tagged proteins of interest (~7.7 kDa, indicated by black arrow) were visualized via western blot using HRP-conjugated anti-HA mouse monoclonal antibody. Exposed blot image is merged with white light image of the membrane to show protein ladders for size reference: (i) Thermo Scientific PageRuler Prestained Protein Ladder (10–180 kDa); (ii) Bio-Rad Precision Plus Protein Dual Xtra (2–250 kDa). An equal volume of each sample/fraction (10 µl) was loaded onto the gel for quick confirmation of successful pull-downs; sample loading was not normalized to OD units and/or protein concentration, although experiments were performed using approximately the same amount of cells for each strain. Co-immunoprecipitation experiments were performed with three biological replicates for each strain; a representative western blot containing one set of biological replicates for each Hdp1opt variant is shown.
-
Figure 3—figure supplement 2—source data 1
Western blot containing protein fractions from the different steps of the co-immunoprecipitation experiments.
HA-tagged proteins of interested were detected using HRP-conjugated anti-HA mouse monoclonal antibody and Amersham ECL Prime Western Blotting Detection Reagent (Cytiva) and visualized using a Bio-Rad ChemiDoc MP System (Chemi Hi Sensitivity setting). Uncropped membrane from experimental replicate 1.
- https://cdn.elifesciences.org/articles/78299/elife-78299-fig3-figsupp2-data1-v2.zip
-
Figure 3—figure supplement 2—source data 2
Western blot containing protein fractions from the different steps of the co-immunoprecipitation experiments.
White light image of membrane to show protein ladders for size reference. Uncropped membrane from experimental replicate 1.
- https://cdn.elifesciences.org/articles/78299/elife-78299-fig3-figsupp2-data2-v2.zip
-
Figure 3—figure supplement 2—source data 3
Western blot containing protein fractions from the different steps of the co-immunoprecipitation experiments.
Merged image of membranes from Figure 3—figure supplement 2—source data 1 and Figure 3—figure supplement 2—source data 2 to show protein ladders for size reference alongside detected proteins of interest. Uncropped membrane from experimental replicate 1; included in Figure 3—figure supplement 2 as a representative western blot for the pull-down assays.
- https://cdn.elifesciences.org/articles/78299/elife-78299-fig3-figsupp2-data3-v2.zip
-
Figure 3—figure supplement 2—source data 4
Western blot containing protein fractions from the different steps of the co-immunoprecipitation experiments.
HA-tagged proteins of interested were detected using HRP-conjugated anti-HA mouse monoclonal antibody and Amersham ECL Prime Western Blotting Detection Reagent (Cytiva) and visualized using a Bio-Rad ChemiDoc MP System (Chemi Hi Sensitivity setting). Uncropped membrane from experimental replicate 2.
- https://cdn.elifesciences.org/articles/78299/elife-78299-fig3-figsupp2-data4-v2.zip
-
Figure 3—figure supplement 2—source data 5
Western blot containing protein fractions from the different steps of the co-immunoprecipitation experiments.
White light image of membrane to show protein ladders for size reference. Uncropped membrane from experimental replicate 2.
- https://cdn.elifesciences.org/articles/78299/elife-78299-fig3-figsupp2-data5-v2.zip
-
Figure 3—figure supplement 2—source data 6
Western blot containing protein fractions from the different steps of the co-immunoprecipitation experiments.
Merged image of membranes from Figure 3—figure supplement 2—source data 4 and Figure 3—figure supplement 2—source data 5 to show protein ladders for size reference alongside detected proteins of interest. Uncropped membrane from experimental replicate 2.
- https://cdn.elifesciences.org/articles/78299/elife-78299-fig3-figsupp2-data6-v2.zip
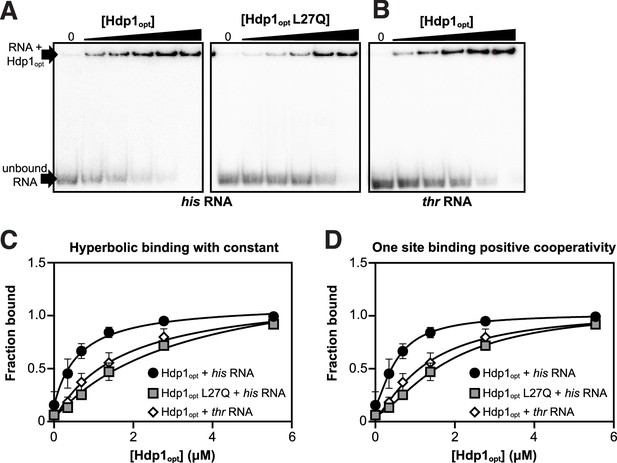
Electrophoretic mobility shift assay (EMSA) gels and binding curves from fitting of different binding models to the EMSA data.
(A) Electrophoretic mobility shift assays (EMSAs) of the full-length (A) his operator RNA or (B) thr operator RNA in the presence of increasing concentrations (0–5.5 μM) of Hdp1opt or the Hdp1opt L27Q mutant (his RNA only). EMSAs were repeated independently three or more times for each protein with similar results; representative gels are shown. (C) Binding curve and parameters for the EMSA data fitted to a standard 1:1 hyperbolic binding model (shown in Figure 3C). (D) Binding curve and parameters for the EMSA data fitted to a model accounting for positive cooperativity. Each data point represents the mean of three or more independent experimental replicates; error bars represent the standard error. Some error bars fall within the boundaries of the markers. For the fraction bound data as quantified and calculated from the EMSA gels and the parameters used in the binding curve calculations, refer to Supplementary file 1d and e, respectively. Curve fitting was performed with Prism 9 (GraphPad Software); see ‘Materials and methods’ for the equations used.
-
Figure 3—figure supplement 3—source data 1
Electrophoretic mobility shift assay (EMSA) of the full-length his operator RNA in the presence of increasing concentrations of Hdp1opt.
Uncropped gel from experimental replicate 1.
- https://cdn.elifesciences.org/articles/78299/elife-78299-fig3-figsupp3-data1-v2.zip
-
Figure 3—figure supplement 3—source data 2
Electrophoretic mobility shift assay (EMSA) of the full-length his operator RNA in the presence of increasing concentrations of Hdp1opt.
Uncropped gel from experimental replicates 2 and 3. Experimental replicate 2 is included in the main text figure as a representative EMSA gel image.
- https://cdn.elifesciences.org/articles/78299/elife-78299-fig3-figsupp3-data2-v2.zip
-
Figure 3—figure supplement 3—source data 3
Electrophoretic mobility shift assay (EMSA) of the full-length his operator RNA in the presence of increasing concentrations of Hdp1opt (left) or the Hdp1opt L27Q mutant (right).
Uncropped gel from Hdp1opt experimental replicate 4 and Hdp1opt L27Q experimental replicate 1. Hdp1opt L27Q experimental replicate 1 is included in Figure 3—figure supplement 3 as a representative EMSA gel image.
- https://cdn.elifesciences.org/articles/78299/elife-78299-fig3-figsupp3-data3-v2.zip
-
Figure 3—figure supplement 3—source data 4
Electrophoretic mobility shift assay (EMSA) of the full-length his operator RNA in the presence of increasing concentrations of the Hdp1opt L27Q mutant.
Uncropped gel from experimental replicates 2 and 3.
- https://cdn.elifesciences.org/articles/78299/elife-78299-fig3-figsupp3-data4-v2.zip
-
Figure 3—figure supplement 3—source data 5
Electrophoretic mobility shift assay (EMSA) of the full-length thr operator RNA in the presence of increasing concentrations of Hdp1opt.
Uncropped gel from experimental replicate 1; included in Figure 3—figure supplement 3 as a representative EMSA gel image. Unlabeled experiment on the left of the gel was not used for quantification or analysis due to the gel ripping.
- https://cdn.elifesciences.org/articles/78299/elife-78299-fig3-figsupp3-data5-v2.zip
-
Figure 3—figure supplement 3—source data 6
Electrophoretic mobility shift assay (EMSA) of the full-length thr operator RNA in the presence of increasing concentrations of Hdp1opt.
Uncropped gel from experimental replicates 2 and 3. ‘X’ denotes a well in which a reaction was loaded in the incorrect order.
- https://cdn.elifesciences.org/articles/78299/elife-78299-fig3-figsupp3-data6-v2.zip
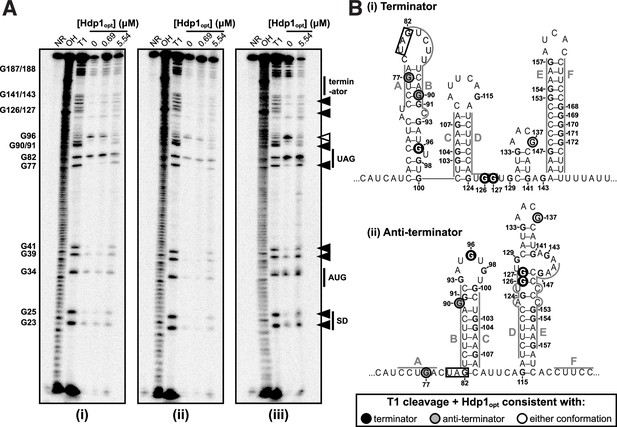
RNase T1 probing experimental replicates.
(A) RNase T1 probing gels for the full-length his operator RNA in the absence and presence of Hdp1opt (0, 0.69, and 5.5 μM). NR denotes RNA subject to no reaction, OH indicates partial alkaline hydrolysis, and T1 is an RNase T1 digest of the RNA under denaturing conditions used to map the RNA sequence. Numbering of G nucleotides is shown on the left. Arrows highlight changes in RNA cleavage in the presence of Hdp1opt: black arrows indicate nucleotides with increased cleavage and white arrows indicate reduced cleavage. Sequence and/or structure characteristics of the his operator RNA are also indicated on the right (i.e., the Shine-Dalgarno sequence [SD], start codon [AUG], and stop codon [UAG] of the hisL leader peptide coding sequence). This experiment was repeated three independent times with similar results: (i) probing reactions incubated with 0.05U RNase T1 for 5 min, (ii) probing reactions incubated with 0.01U RNase T1 for 5 min, and (iii) probing reactions incubated with 0.05U RNase T1 for 10 min. The reactions from (iii) are included in main text Figure 3D as a representative RNase T1 probing gel image. (B) Nucleotides demonstrating a change in RNase T1 cleavage in the presence of Hdp1opt mapped to both the (i) terminator and (i) anti-terminator conformations of the his operator RNA. The conformation consistent with cleavage in the presence of Hdp1opt is color-coded according to the legend; the hisL stop codon is boxed. Nucleotides are numbered from the transcription start site, +1. Included for reference from main text Figure 3E.
-
Figure 3—figure supplement 4—source data 1
Uncropped RNase T1 probing gel for the full-length his operator RNA in the absence and presence of Hdp1opt (0, 0.69, and 5.5 μM).
NR denotes RNA subject to no reaction, OH indicates partial alkaline hydrolysis, and T1 is an RNase T1 digest of the RNA under denaturing conditions used to map the RNA sequence. (i) Probing reactions incubated with 0.05U T1 RNase for 5 min. (ii) Probing reactions incubated with 0.01U T1 RNase for 5 min. (iii) Probing reactions incubated with 0.05U T1 RNase for 10 min. Numbering of G nucleotides is shown on the left. Arrows highlight changes in RNA cleavage in the presence of Hdp1opt: black arrows indicate nucleotides with increased cleavage and white arrows indicate reduced cleavage. Sequence and/or structure characteristics of the his operator RNA are also indicated on the right (i.e., the Shine-Dalgarno sequence [SD], start codon [AUG], and stop codon [UAG] of the hisL leader peptide coding sequence). The reactions from (iii) are included in the main text figure as a representative T1 RNase probing gel image; all replicates are in included in Figure 3—figure supplement 4. All reaction sets were performed with independent RNA and protein dilutions.
- https://cdn.elifesciences.org/articles/78299/elife-78299-fig3-figsupp4-data1-v2.zip
Additional files
-
Supplementary file 1
Supplementary Tables 1a-g.
(a) KEIO deletion strains screened with the random sequence libraries. (b) Summary of Hdp1-2 single mutants obtained via random mutagenesis. (c) Escherichia coli K-12 strains used in this study. (d) Fraction bound data as quantified and calculated from the electrophoretic mobility shift assay (EMSA) gels. (e) Parameters from fitting of different binding models to the EMSA data. (f) Plasmids used in this study. (g) Oligonucleotides used in this study.
- https://cdn.elifesciences.org/articles/78299/elife-78299-supp1-v2.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/78299/elife-78299-mdarchecklist1-v2.docx





