Multiple ciliary localization signals control INPP5E ciliary targeting
Figures
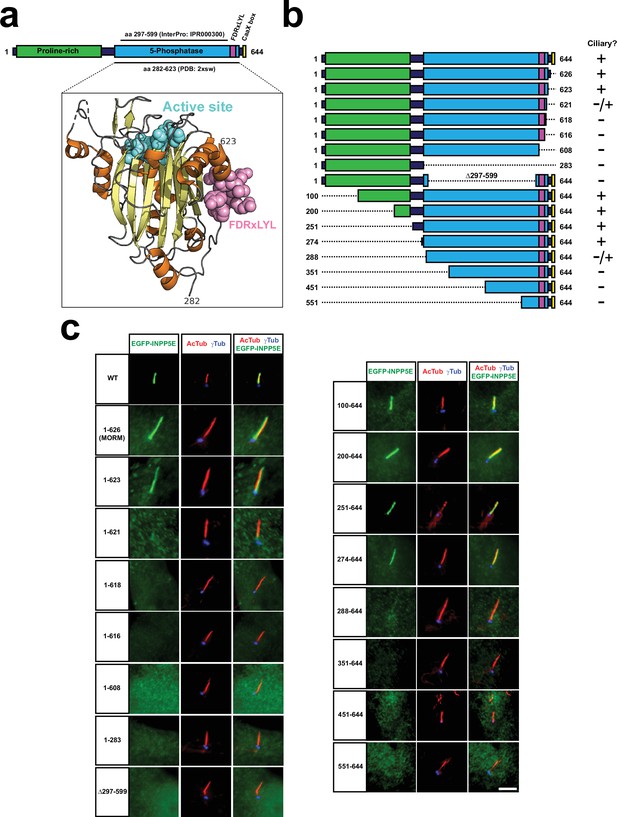
INPP5E catalytic domain encompasses FDRxLYL motif and is required for ciliary targeting.
(a) Top diagram represents full length human INPP5E protein sequence (aa 1–644). Depicted are the proline-rich region (aa 10–242, Uniprot), the previously reported ciliary localization signal (FDRxLYL, aa 609–615 Humbert et al., 2012), and the CaaX box driving farnesylation (aa 641–644). Also shown is the inositol polyphosphate 5-phosphatase catalytic domain, whose most conserved core corresponds to InterPro domain IPR000300 (aa 297–599), but which actually spans aa 282–623, as revealed by its crystal structure, available at the Protein Data Bank (PDB) and displayed below (PDB ID: 2xsw). Notice how FDRxLYL residues (in magenta above and below) are part of the catalytic domain, on whose surface they fold. The 3D structure also shows active site residues in cyan, alpha-helices in orange, and beta-strands in yellow (including the beta-sandwich at the domain’s core, and a small beta-hairpin near the active site). (b) Schematic representation of full length human INPP5E (1-644) and its deletion mutants used in (c), indicating on the right which ones localize to cilia. (c) Immunofluorescence images of cilia from hTERT-RPE1 cells transfected with the indicated EGFP-INPP5E constructs. Cells were stained with antibodies against acetylated α-tubulin (AcTub), γ-tubulin (γTub) and EGFP to detect the fusion proteins. Images are representative of at least two independent experiments per construct, with >30 transfected-cell cilia visualized per construct and experiment. Scale bar, 5 µm.
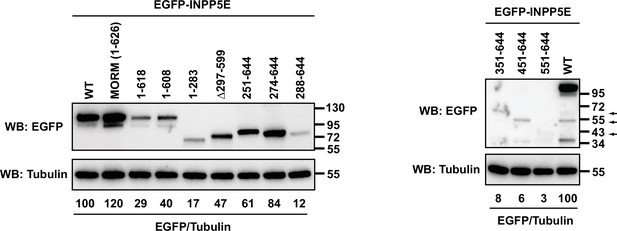
Expression levels of EGFP-INPP5E constructs from Figure 1.
The indicated constructs were expressed in HEK293T cells. Lysates were analyzed by SDS-PAGE and immunoblotting with the indicated antibodies. Molecular weight markers in kilodaltons (kDa) are shown on the right. Arrows indicate locations of the 351–644, 451–644, and 551–644 constructs, whose predicted molecular weights are 61, 50, and 39 kDa, respectively. The numbers under the tubulin blots are EGFP/Tubulin band intensity ratios, normalized so that WT equals 100%.
-
Figure 1—figure supplement 1—source data 1
Uncropped immunoblots from Figure 1—figure supplement 1.
Relevant bands are inside rectangles. See Figure 1—figure supplement 1 for more details.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig1-figsupp1-data1-v1.pdf
-
Figure 1—figure supplement 1—source data 2
Uncropped immunoblot from Figure 1—figure supplement 1 (WB: EGFP).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig1-figsupp1-data2-v1.tif
-
Figure 1—figure supplement 1—source data 3
Uncropped immunoblot from Figure 1—figure supplement 1 (WB: Tubulin).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig1-figsupp1-data3-v1.tif
-
Figure 1—figure supplement 1—source data 4
Uncropped immunoblot from Figure 1—figure supplement 1 (WB: EGFP).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig1-figsupp1-data4-v1.tif
-
Figure 1—figure supplement 1—source data 5
Uncropped immunoblot from Figure 1—figure supplement 1 (WB: Tubulin).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig1-figsupp1-data5-v1.tif
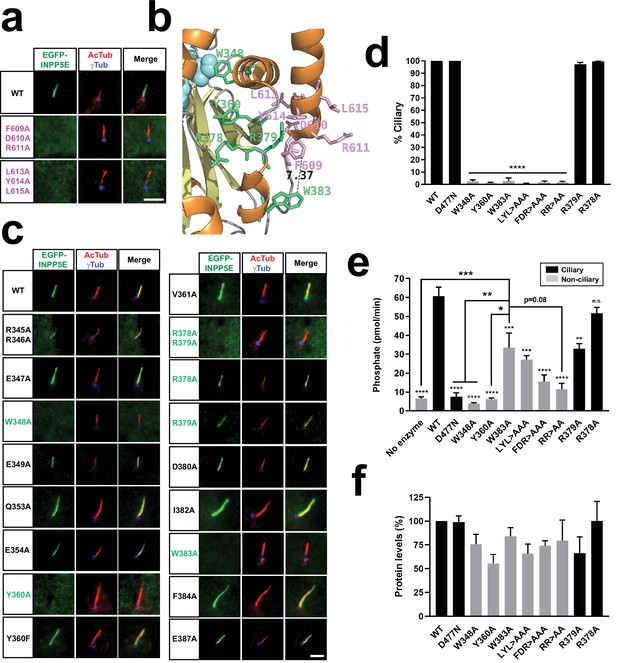
W383 and FDRxLYL motifs act as CLSs on the catalytic domain surface.
(a) Cilia localization of the indicated FDRxLYL mutants of EGFP-INPP5E was analyzed in hTERT-RPE1 cells as in Figure 1. Scale bars, 5 µm. (b) Magnification from INPP5E structure (PDB ID: 2xsw) showing the FDRxLYL motif residues (pink) and adjacent catalytic domain residues shown here to affect ciliary targeting (green). Distance between W383 and F609 is indicated in angstroms. Beta-sheets and alpha-helices are shown as yellow and orange ribbons, respectively. Notice active site region on top left (cyan). (c) Cilia localization of the indicated EGFP-INPP5E constructs was analyzed as in (a). In both cases, images are representative of at least two independent experiments per construct, with >30 transfected-cell cilia visualized per construct and experiment. Scale bars, 5 µm. (d) Percentage of positive cilia was quantitated for each of the indicated constructs. Data are mean ± SEM of n=3 independent experiments. Data were analyzed by one-way ANOVA followed by Tukey’s multiple comparisons tests. Significance relative to WT is shown as p<0.0001(****). LYL >AAA: L613A+Y614A+L615 A; FDR >AAA: F609A+D610A+R611 A; RR >AA: R378A+R379 A. (e) 5-phosphatase activity, expressed as picomoles of released inorganic phosphate per minute, was measured, using PI(4,5)P2 as substrate, in immunoprecipitates of HEK293T cells transfected with the indicated EGFP-INPP5E variants. Cilia-localized constructs shown as black columns, non-ciliary as grey. Data are mean ± SEM of n=9,9,5,4,3,5,3,3,2,3,3 independent experiments (from left to right). Data were analyzed by one-way ANOVA followed by Tukey’s multiple comparisons tests. Significance relative to WT is shown as small asterisks directly above each bar. Significance relative to W383A is shown as bigger asterisks as indicated. In all cases, significance is represented as: p<0.05(*), p<0.01(**), p<0.001(***), p<0.0001(****), or n.s. (not significant). (f) Protein levels in the immunoprecipitates used for the activity assays in (e). Western blot bands were quantitated and plotted as percentage of WT. Data are mean ± SEM of n=8,5,3,3,5,2,3,3,2,3 independent experiments (from left to right). One-way ANOVA revealed no significant differences.
-
Figure 2—source data 1
Source data for Figure 2d.
Data from n=3 independent experiments.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig2-data1-v1.xlsx
-
Figure 2—source data 2
Source data for Figure 2e.
Data from n=9 independent experiments.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig2-data2-v1.xlsx
-
Figure 2—source data 3
Source data for Figure 2f.
Data from n=8 independent experiments.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig2-data3-v1.xlsx
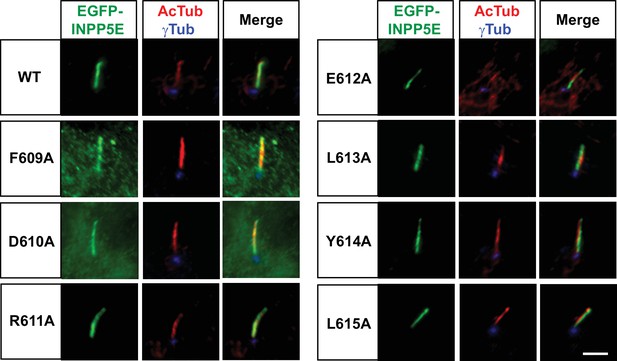
FDRELYL motif residues are not individually required for INPP5E cilia localization.
Ciliary targeting of EGFP-INPP5E wild type (WT) or the indicated FDRELYL motif mutants was assessed by immunofluorescence microscopy in transfected hTERT-RPE1 cells, which were stained with antibodies against acetylated α-tubulin (AcTub), γ-tubulin (γTub) and EGFP to detect the fusion proteins. Images are representative of n=2 independent experiments, with >30 transfected-cell cilia visualized per experiment. Scale bar, 5 µm.
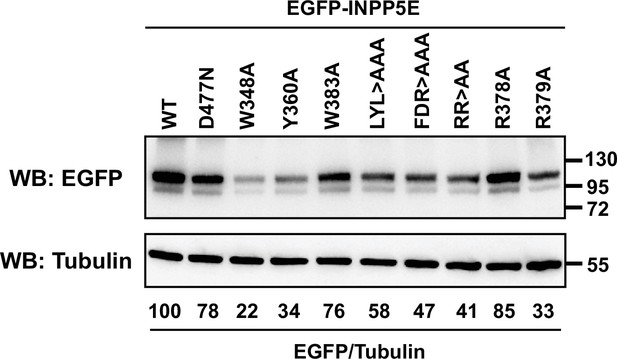
Expression levels of EGFP-INPP5E constructs from Figure 2.
The indicated constructs were expressed in HEK293T cells. Lysates were analyzed by SDS-PAGE and immunoblotting with the indicated antibodies. Molecular weight markers in kilodaltons are shown on the right. The numbers under the tubulin blots are EGFP/Tubulin band intensity ratios, normalized so that WT equals 100%.
-
Figure 2—figure supplement 2—source data 1
Uncropped immunoblots from Figure 2—figure supplement 2.
Relevant bands are inside rectangles. See Figure 2—figure supplement 2 for more details.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig2-figsupp2-data1-v1.pdf
-
Figure 2—figure supplement 2—source data 2
Uncropped immunoblot from Figure 2—figure supplement 2 (WB: EGFP).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig2-figsupp2-data2-v1.tif
-
Figure 2—figure supplement 2—source data 3
Uncropped immunoblot from Figure 2—figure supplement 2 (WB: Tubulin).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig2-figsupp2-data3-v1.tif
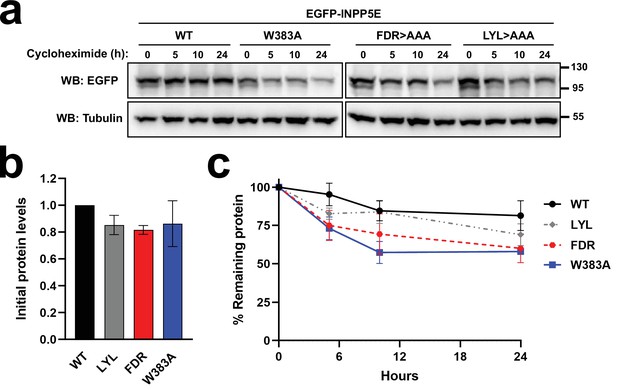
Stability of W383A and FDRxLYL mutants.
(a) The indicated EGFP-INPP5E forms were expressed in HEK293T cells for 24 hr, at which time 200 µg/ml cycloheximide was added for the indicated number of hours. Cell lysates were then analyzed by western blot with antibodies against EGFP and α-tubulin. Molecular weight markers in kDa shown on the right. FDR >AAA: F609A+D610A+R611 A; LYL >AAA: L613A+Y614A+L615 A. (b) Protein levels at the time of cycloheximide addition for the same EGFP-INPP5E forms as in (a). EGFP/Tubulin band intensity ratios were normalized to WT and plotted as mean ± SEM from n=3 independent experiments, including one in (a). One-way ANOVA revealed no significant differences. (c) Time course of protein levels after cycloheximide addition. Amounts are EGFP/Tubulin ratios for each EGFP-INPP5E form, normalized to the ratio at 0 hr. Data are mean ± SEM (n=3 independent experiments). Unpaired t-tests reveal non-significance except for WT versus W383A at 10 hr (p=0.048).
-
Figure 2—figure supplement 3—source data 1
Uncropped immunoblots from Figure 2—figure supplement 3a.
Relevant bands are inside rectangles. See Figure 2—figure supplement 3 for more details.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig2-figsupp3-data1-v1.pdf
-
Figure 2—figure supplement 3—source data 2
Uncropped immunoblot from Figure 2—figure supplement 2A (WB: EGFP for WT and W383A).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig2-figsupp3-data2-v1.tif
-
Figure 2—figure supplement 3—source data 3
Uncropped immunoblot from Figure 2—figure supplement 2A (WB: EGFP for FDR >AAA and LYL >AAA).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig2-figsupp3-data3-v1.tif
-
Figure 2—figure supplement 3—source data 4
Uncropped immunoblot from Figure 2—figure supplement 2A (WB: Tubulin for WT and W383A).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig2-figsupp3-data4-v1.tif
-
Figure 2—figure supplement 3—source data 5
Uncropped immunoblot from Figure 2—figure supplement 2A (WB: Tubulin for FDR >AAA and LYL >AAA).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig2-figsupp3-data5-v1.tif
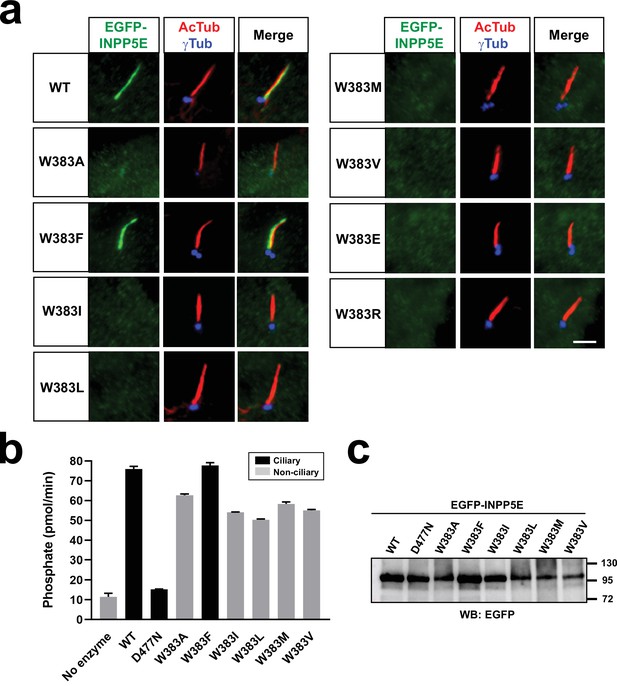
Effect of W383 mutations on ciliary targeting and activity.
(a) Cilia localization of the indicated EGFP-INPP5E variants was analyzed in hTERT-RPE1 cells as in Figures 1–2. Images are representative of n=2 independent experiments, with >30 transfected-cell cilia visualized per experiment. Scale bars, 5 µm. (b) 5-phosphatase activity, expressed as picomoles of released inorganic phosphate per minute, was measured, using PI(4,5)P2 as substrate, in immunoprecipitates of HEK293T cells transfected with the indicated EGFP-INPP5E variants. Cilia-localized constructs shown as black columns, non-ciliary as grey. Data are mean ± SEM of n=3 technical replicates. (c) Western blot of the anti-EGFP immunoprecipitates used for the activity assays in (b). Molecular weight markers are on the right (kDa).
-
Figure 2—figure supplement 4—source data 1
Uncropped immunoblot from Figure 2—figure supplement 4c.
Relevant bands are inside rectangle. See Figure 2—figure supplement 4 for more details.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig2-figsupp4-data1-v1.pdf
-
Figure 2—figure supplement 4—source data 2
Uncropped immunoblot from Figure 2—figure supplement 4c (WB: EGFP).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig2-figsupp4-data2-v1.zip
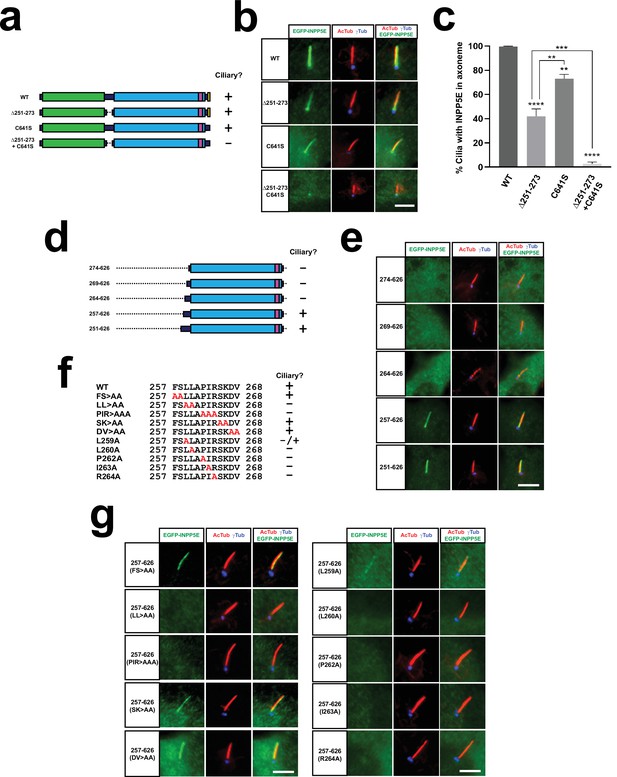
The LLxPIR motif is a novel CLS that cooperates with the CaaX box to mediate INPP5E ciliary targeting.
(a) Schema of full length human INPP5E and its mutants used in (b–c). Cilia localization of each mutant is indicated on the right. (b) Cilia localization of WT and indicated mutants was analyzed in hTERT-RPE1 cells as in Figures 1–2. Images are representative of n=3 independent experiments, with >30 transfected-cell cilia visualized per construct and experiment. Scale bar, 5 µm. (c) Quantitation of data from (b). The percentage of positive cilia in transfected cells is shown for the indicated EGFP-INPP5E constructs. Data are mean ± SEM of n=3 independent experiments. Data were analyzed by one-way ANOVA with post-hoc Tukey multiple comparisons tests. Statistical significance is depicted as p<0.01(**), p<0.001(***), or p<0.0001(****). Significance is shown relative to WT unless otherwise indicated. (d) Schema of INPP5E deletion mutants used to map the CLS within aa 251–273. None of these mutants contains the CaaX box (aa 641–644), so their ciliary targeting is strictly dependent on residues 251–273. Cilia localization of each mutant is indicated on the right. (e) Cilia localization of the mutants from (d) was analyzed in hTERT-RPE1 cells as in Figures 1—3. (f) Sequence of aa 257–268 in wild type INPP5E and indicated mutants, whose ciliary localization in shown on the right. (g) Ciliary targeting of INPP5E(257-626) containing the mutations from (f) was analyzed as in (e). In both cases, images are representative of n=2 independent experiments, with >30 transfected-cell cilia visualized per construct and experiment. Scale bars, 5 µm.
-
Figure 3—source data 1
Source data for Figure 3c.
Data from n=3 independent experiments.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig3-data1-v1.xlsx
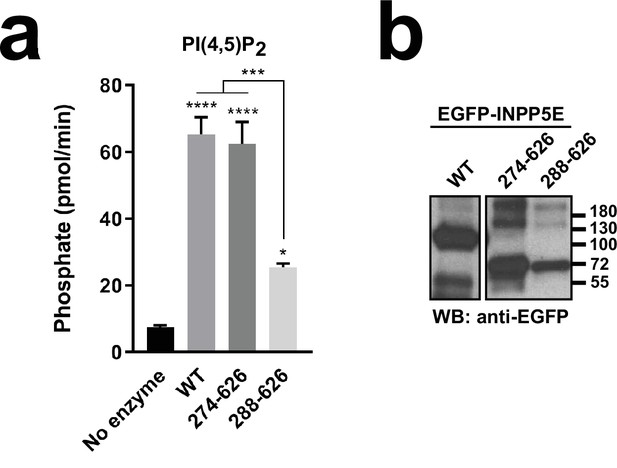
CaaX box and the CLS-containing residues 251–273 do not affect enzyme activity.
(a) Phosphatase activity, expressed as picomoles of released inorganic phosphate per minute, was measured in immunoprecipitates of HEK293T cells transfected with the indicated EGFP-INPP5E variants, or in control buffer (no enzyme). Activity was measured using PI(4,5)P2 as substrate. Data are shown as mean ± SEM of n=3 independent experiments. Data were analyzed by one-way ANOVA followed by Tukey’s multiple comparisons tests. Significance (relative to no enzyme unless otherwise indicated) shown as p<0.05(*), p<0.001(***), or p<0.0001(****). (b) Representative anti-EGFP immunoblot showing protein levels of EGFP-INPP5E variants in the immunoprecipitates used in (a). All three samples were run in the same SDS-PAGE gel and immunoblotted and detected in parallel. Molecular weight markers, in kDa, are shown on the right.
-
Figure 3—figure supplement 1—source data 1
Uncropped immunoblot from Figure 3—figure supplement 1b.
Relevant bands are inside rectangles. See Figure 3—figure supplement 1 for more details.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig3-figsupp1-data1-v1.pdf
-
Figure 3—figure supplement 1—source data 2
Uncropped immunoblot from Figure 3—figure supplement 1b (WB: EGFP).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig3-figsupp1-data2-v1.tif
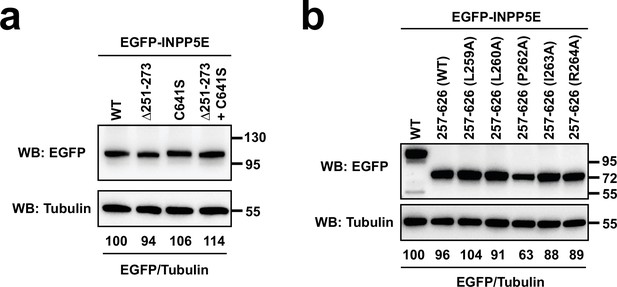
Expression levels of EGFP-INPP5E constructs from Figure 3.
(a) Constructs from Figure 3a–c were expressed in HEK293T cells. Lysates were analyzed by SDS-PAGE and immunoblotting with the indicated antibodies. Molecular weight markers in kilodaltons are shown on the right. The numbers under the tubulin blots are EGFP/Tubulin band intensity ratios, normalized so that WT equals 100%. (b) Same was done for the indicated constructs from Figure 3d–g.
-
Figure 3—figure supplement 2—source data 1
Uncropped immunoblots from Figure 3—figure supplement 2.
Relevant bands are inside rectangles. See Figure 3—figure supplement 2 for more details.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig3-figsupp2-data1-v1.pdf
-
Figure 3—figure supplement 2—source data 2
Uncropped immunoblot from Figure 3—figure supplement 2 (WB: EGFP).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig3-figsupp2-data2-v1.tif
-
Figure 3—figure supplement 2—source data 3
Uncropped immunoblot from Figure 3—figure supplement 2 (WB: Tubulin).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig3-figsupp2-data3-v1.zip
-
Figure 3—figure supplement 2—source data 4
Uncropped immunoblot from Figure 3—figure supplement 2 (WB: EGFP).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig3-figsupp2-data4-v1.tif
-
Figure 3—figure supplement 2—source data 5
Uncropped immunoblot from Figure 3—figure supplement 2 (WB: Tubulin).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig3-figsupp2-data5-v1.tif
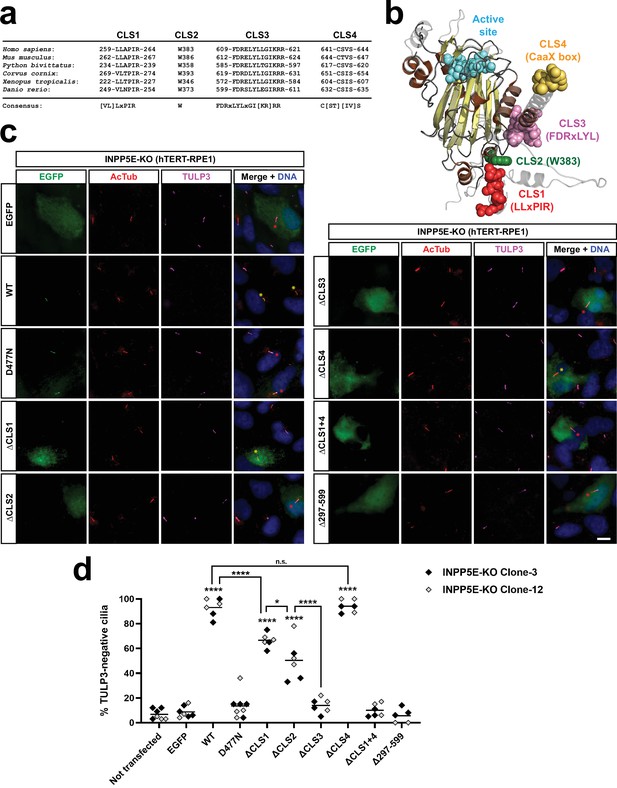
CLS1-4 are conserved ciliary localization signals affecting INPP5E function.
(a) CLS1-4 are highly evolutionarily conserved in vertebrates, including human (NP_063945.2), mouse (AAH80295.1), python (XP_007441606.1), crow (XP_039417670.1), toad (XP_002935265.1), and zebrafish (NP_001096089.2). Consensus sequences are shown below. (b) AlphaFold model of INPP5E 3D structure (AF-Q9NRR6-F1) depicting predicted locations of CLS1 (red), CLS2 (green), CLS3 (pink) and CLS4 (yellow). Active site in cyan. Beta-strands and alpha-helices in yellow and brown, respectively. Proline-rich N-terminal region (aa 1–200), predicted to be highly flexible, is not shown. CLS1 is probably also part of a flexible region, and its position in the AlphaFold model has a low confidence score (pLDDT). See Uniprot entry Q9NRR6 for more details. (c) Rescue assay assessing the ability of INPP5E or its mutants to lower the abnormally high TULP3 levels characteristic of INPP5E-KO cilia. The indicated constructs were transfected into INPP5E-KO RPE1 cells, generated via CRISPR-Cas9 (Figure 4—figure supplement 2). Cells were fixed and stained for EGFP, acetylated tubulin (AcTub), TULP3, and DNA (DAPI), as indicated. Scale bar, 10 µm. Note how untransfected INPP5E-KO cells have high ciliary TULP3 levels, as previously described. Transfected cell cilia are labeled with asterisks in the merge panels: yellow asterisks for rescued TULP3-negative cilia, and red asterisks for non-rescued TULP3-positive cilia. (d) Quantitation of the rescue experiment shown in (c). For each construct, the percentage of TULP3-negative transfected-cell cilia was counted. Data come from five independent experiments. Each point in the graph indicates an independent transfection. Between 12 and 39 transfected-cell cilia were counted per transfection (with exception of the highest data point in ΔCLS2, where only 9 cilia could be counted). Experiments were performed in parallel with two different INPP5E-KO clones (clones 3 and 12). Graph shows individual data points, color-coded by clone as indicated, and the overall median is indicated with a line. Two-way ANOVA revealed significant differences between constructs (p<0.0001) but no significant differences between the clones. All data were then analyzed by one-way ANOVA followed by Tukey tests. Significance is shown relative to EGFP unless otherwise indicated. p<0.0001 (****); p<0.05 (*); not significant (n.s.).
-
Figure 4—source data 1
Source data for Figure 4d.
Data from n=5 independent experiments.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig4-data1-v1.xlsx
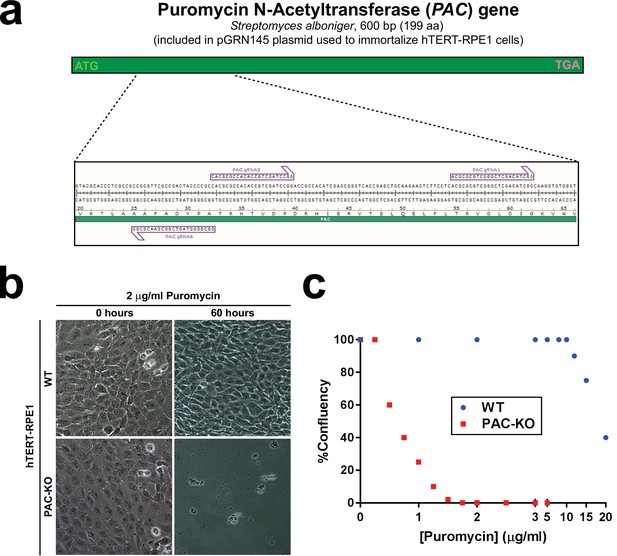
Generation of puromycin-sensitive hTERT-RPE1 cells by CRISPR/Cas9.
(a) hTERT-RPE1 cells were originally immortalized with pGRN145, a plasmid expressing both human telomerase (hTERT) and puromycin N-acetyltransferase (PAC) from Streptomyces alboniger, a 199 aa enzyme that confers puromycin resistance (top). Three single guide RNAs were selected in the region encoding aa 25–62 of PAC (bottom). A pool of these gRNAs was then used for CRISPR/Cas9-mediated knockout of PAC gene in hTERT-RPE1 cells. (b) Phase contrast images of WT and PAC-KO hTERT-RPE1 cells right before (0 hr) or after treatment for 60 hr with 2 µg/ml puromycin. This treatment was sufficient to completely kill PAC-KO cells but did not noticeably affect WT cells. (c) Dose-response curves showing percentage confluence of WT or PAC-KO cells after 60 hr treatment with the indicated puromycin doses. Approximate IC50 values are 18 µg/ml for WT and 0.6 µg/ml for PAC-KO cells, a 30-fold difference.
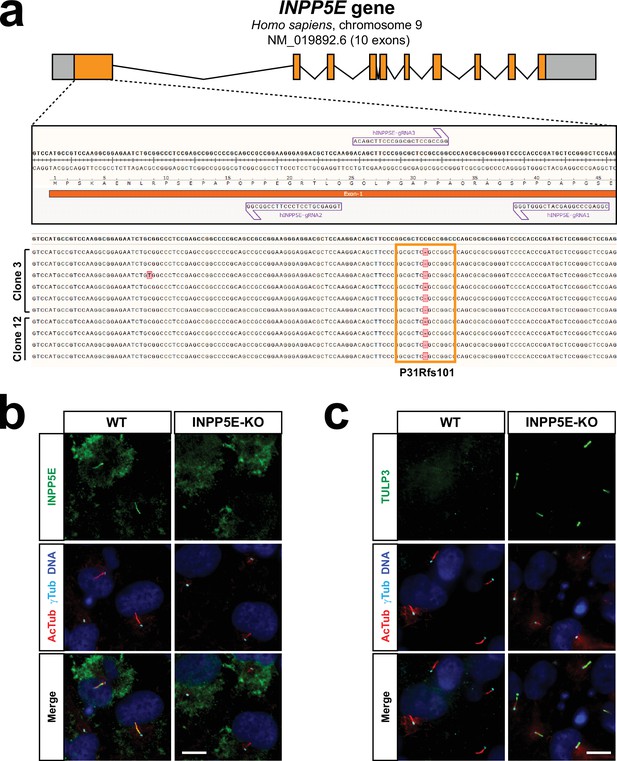
Generation of INPP5E-KO hTERT-RPE1 cells by CRISPR/Cas9.
(a) Top: diagram of human INPP5E gene depicting its 10 exons as boxes, with coding sequence in orange. Middle: the three indicated guide RNAs (arrows) were used to target INPP5E exon-1 in RPE-PS cells using CRISPR. Bottom: the genomic regions of interest from clones 3 and 12 were PCR-amplified, ligated into pJET1.2/blunt vector, transformed, and DNA from bacterial colonies sequenced (6 and 4 colonies for clones 3 and 12, respectively). All sequenced clones contained the same truncating mutation (P31Rfs101), which causes frameshift and only leaves intact the first 30 residues of INPP5E. (b) Immunofluorescence staining of clone-3 (INPP5E-KO) and its parental RPE-PS cells (WT) with the indicated antibodies. Ciliary staining of INPP5E is seen in WT but not INPP5E-KO cells, which only show non-specific staining outside cilia. Same results were obtained for clone-12 (not shown). (c) Immunofluorescence staining of clone-3 (INPP5E-KO) and its parental RPE-PS cells (WT) with the indicated antibodies. As previously reported, TULP3 strongly accumulates in INPP5E-KO cilia. Same results were obtained for clone-12 (not shown).
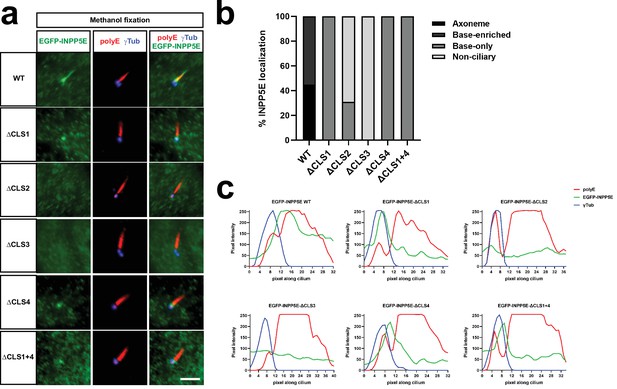
CLS1 and CLS4 mutants are only seen at the transition zone in methanol-fixed cells.
(a) The indicated EGFP-INPP5E constructs were expressed in hTERT-RPE1 cells, which were fixed with methanol as described in Materials and methods. Immunofluorescence was carried out with antibodies against EGFP, polyglutamylated alpha-tubulin (polyE), and gamma-tubulin (γTub), as indicated. Scale bar, 5 mm. (b) Percentage of cilia with the specified EGFP-INPP5E localization was quantified for each of the constructs in (a). Data are from an individual experiment. The number of transfected cell cilia counted for each construct was, from left to right: n=31,13,15,10,12,10. (c) Pixel intensity profiles of the images in (a). A line was drawn along the cilium using Fiji/ImageJ, and the Plot Profile function of this program was used to obtain the densitometric data for each channel. Note how EGFP-INPP5E WT, ΔCLS1, ΔCLS4 and ΔCLS1 +4 accumulate at the transition zone, distal from the basal body (γTub, blue) and proximal to the axoneme (polyE, red).
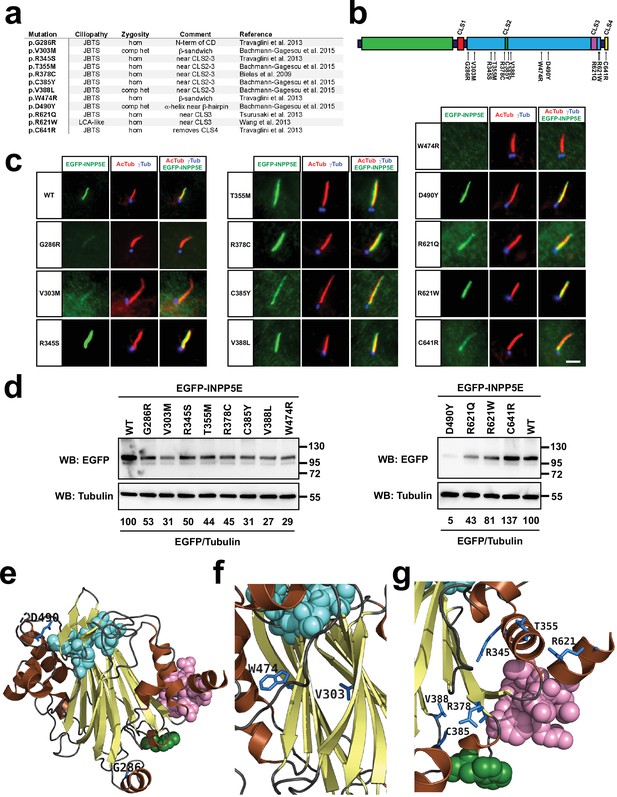
A subset of Joubert syndrome INPP5E mutations abolishes ciliary targeting.
(a) Table of INPP5E ciliopathy mutations analyzed here. JBTS: Joubert syndrome; LCA: Leber congenital amaurosis; hom: homozygous; comp het: compound heterozygous. (b) Schema of INPP5E protein sequence indicating the locations of the ciliopathy mutations from (c) relative to its four CLSs, its catalytic domain (cyan) and its N-terminal proline-rich region (green). (c) Ciliary localization of mutants from (a–b) was analyzed in hTERT-RPE1 cells as in Figures 1—3. Images are representative of at least two independent experiments per construct, with >30 transfected-cell cilia visualized per construct and experiment. Scale bar, 5 µm. (d) The mutants from (a–c) were expressed in HEK293T cells and their protein levels analyzed by SDS-PAGE and immunoblotting with anti-EGFP antibody, and anti-alpha tubulin as loading control. Molecular weight markers in kilodaltons are shown on the right. The numbers under the tubulin blots are EGFP/Tubulin band intensity ratios, normalized so that WT equals 100%. (e–g) 3D views of INPP5E catalytic domain (PDB ID: 2xsw) showing the ciliopathy-mutated residues from (a) in dark blue (other colors as in Figure 4b). (e) Full catalytic domain showing G286 (bottom) and D490 (top left). (f) closeup view of beta-sandwich showing W474 and V303. (g) closeup view of CLS2-3 region showing R345, T355, R378, C385, V388L, and R621.
-
Figure 5—source data 1
Uncropped immunoblots from Figure 5d.
Relevant bands are inside rectangles. See Figure 5 for more details.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig5-data1-v1.pdf
-
Figure 5—source data 2
Uncropped immunoblot from Figure 5d (WB: EGFP).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig5-data2-v1.tif
-
Figure 5—source data 3
Uncropped immunoblot from Figure 5d (WB: Tubulin).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig5-data3-v1.tif
-
Figure 5—source data 4
Uncropped immunoblot from Figure 5d (WB: EGFP).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig5-data4-v1.tif
-
Figure 5—source data 5
Uncropped immunoblot from Figure 5d (WB: Tubulin).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig5-data5-v1.zip
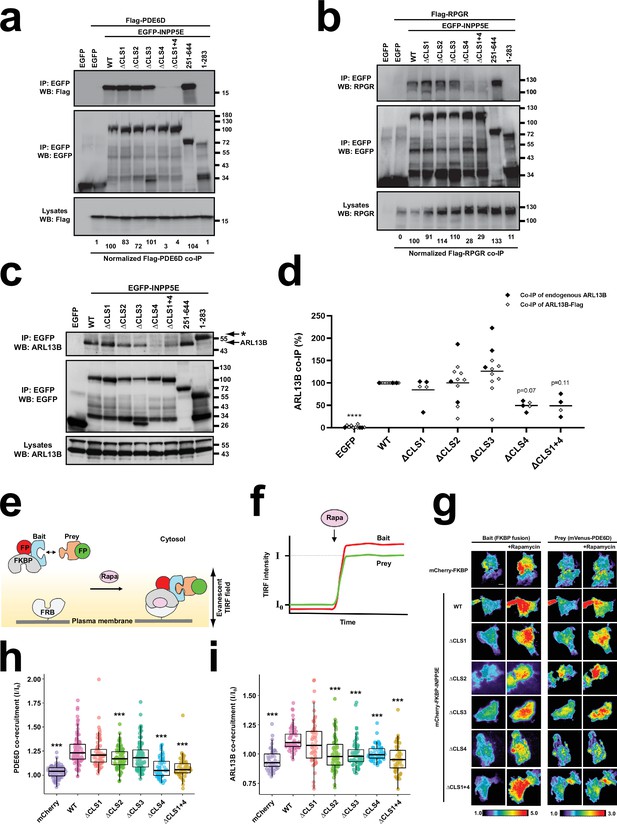
CLS4 promotes INPP5E binding to PDE6D, RPGR and ARL13B.
(a) The indicated EGFP-INPP5E variants were coexpressed in HEK293T cells with Flag-PDE6D, as indicated. Lysates were immunoprecipitated with GFP-Trap beads and analyzed by Western blot with the indicated antibodies. Molecular weight markers in kilodaltons are shown on the right. Quantitation of Flag-PDE6D co-immunoprecipitation (co-IP), as percentage relative to WT, is shown at the bottom. Quantitations are normalized relative to both immunoprecipitated EGFP constructs and lysate amounts of Flag-PDE6D. (b) Same experiment as in (a) but Flag-RPGR was used instead of Flag-PDE6D. Quantitations at the bottom are normalized relative to immunoprecipitated EGFP constructs. (c) Coimmunoprecipitation of endogenous ARL13B with the indicated EGFP-INPP5E constructs in HEK293T cells. Asterisk points to EGFP-INPP5E(1-283) band. (d) Quantitation of ARL13B co-IP with the indicated EGFP-INPP5E constructs from n=11 independent experiments in HEK293T cells. Black and grey dots correspond, respectively, to experiments where endogenous ARL13B or exogenous ARL13B-Flag co-IP was assessed. Not all samples were present in all experiments (but EGFP, WT, ΔCLS2 and ΔCLS3 were). Two-way ANOVA revealed significant differences between constructs (p<0.0001) but no significant differences between using endogenous or exogenous ARL13B. All data were then analyzed by one-way ANOVA followed by Tukey tests. Significance is shown relative to WT: p<0.0001 (****). (e) Schema of chemically-inducible co-recruitment assay. Rapamycin (Rapa)-induced interaction between FKBP and FRB is used to quantitate binding of prey candidates to a bait. FKBP is fused to the prey along with a fluorescent protein, while FRB is tethered to inner leaflet of plasma membrane. Upon rapamycin addition, FKBP binds to FRB, bringing bait (red FP) and associated prey (green FP) to the plasma membrane. (f) Recruitment of bait and prey to plasma membrane can be sensitively detected by TIRF microscopy as an increased fluorescence signal. The ratio of final to initial TIRF intensity upon rapamycin addition (I/I0) for the prey provides a quantitative measure of prey’s co-recruitment to plasma membrane by bait, and hence of the prey-bait interaction. (g) TIRF microscopy images showing rapamycin-induced plasma membrane recruitment of bait constructs (left) and the corresponding co-recruitment of prey (mVenus-PDE6D, right). Intensity scales are depicted at bottom. Scale bar, 10 µm. (h) Normalized rapamycin-induced co-recruitment of mVenus-PDE6D (prey) by mCherry-FKBP-INPP5E (WT or indicated mutants), or by mCherry-FKBP (mCherry) as negative control. Individual measurements of n>50 cells per condition are shown. Box and whisker plots represent median, first and third quartiles, and 95% confidence intervals. Statistical significance relative to WT is shown as *** p<0.001 (unpaired Student’s t-tests). (i) Normalized rapamycin-induced co-recruitment of ARL13B-EYFP (prey) by mCherry-FKBP-INPP5E (WT or indicated mutants), or by mCherry-FKBP (mCherry) as negative control. Data acquisition, analysis and representation as in (h).
-
Figure 6—source data 1
Uncropped immunoblots from Figure 6.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig6-data1-v1.pdf
-
Figure 6—source data 2
Uncropped immunoblot from Figure 6a (IP: EGFP; WB: Flag).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig6-data2-v1.tif
-
Figure 6—source data 3
Uncropped immunoblot from Figure 6a (IP: EGFP; WB: EGFP).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig6-data3-v1.tif
-
Figure 6—source data 4
Uncropped immunoblot from Figure 6a (Lysates, WB: Flag).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig6-data4-v1.tif
-
Figure 6—source data 5
Uncropped immunoblot from Figure 6b (IP: EGFP; WB: RPGR).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig6-data5-v1.zip
-
Figure 6—source data 6
Uncropped immunoblot from Figure 6b (IP: EGFP; WB: EGFP).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig6-data6-v1.zip
-
Figure 6—source data 7
Uncropped immunoblot from Figure 6b (Lysates, WB: RPGR).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig6-data7-v1.zip
-
Figure 6—source data 8
Uncropped immunoblot from Figure 6c (IP: EGFP; WB: ARL13B).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig6-data8-v1.tif
-
Figure 6—source data 9
Uncropped immunoblot from Figure 6c (IP: EGFP; WB: EGFP).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig6-data9-v1.zip
-
Figure 6—source data 10
Uncropped immunoblot from Figure 6c (Lysates, WB: ARL13B).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig6-data10-v1.zip
-
Figure 6—source data 11
Source data from Figure 6d.
Data from n=11 experiments.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig6-data11-v1.xlsx
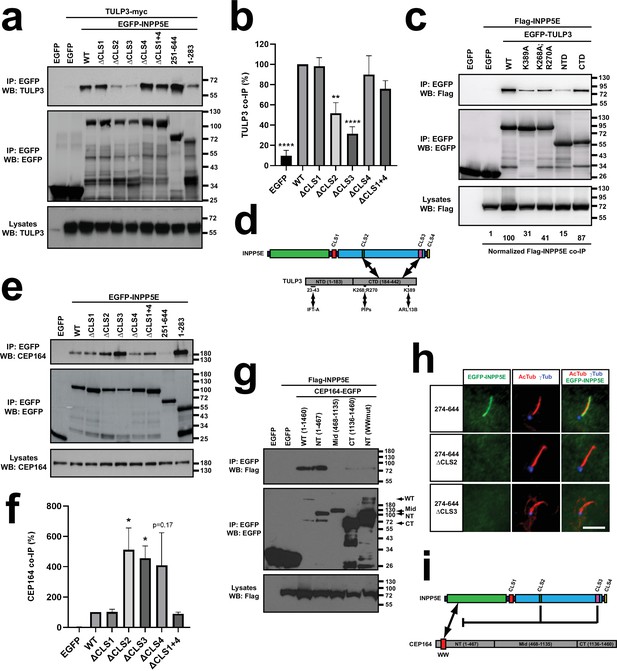
CLS2 and CLS3 regulate INPP5E binding to TULP3 and CEP164.
(a) The indicated EGFP-INPP5E variants were coexpressed in HEK293T cells with TULP3-myc as indicated. Lysates were immunoprecipitated with GFP-Trap beads and analyzed by Western blot with TULP3 and EGFP antibodies, as indicated. (b) Quantitation of TULP3-myc co-IP by the indicated EGFP-INPP5E constructs in HEK293T cells. Co-IP levels, expressed as percentage of WT, are normalized by the amounts of both immunoprecipitated EGFP-INPP5Es and TULP3-myc lysate levels. Data are mean ± SEM of n=7,7,4,7,7,4,4 independent experiments and were analyzed by one-way ANOVA followed by Tukey multiple comparisons tests. Significance relative to WT is shown as p<0.01(**) and p<0.0001(****). (c) The indicated EGFP-TULP3 variants were coexpressed in HEK293T cells with Flag-INPP5E as indicated. Lysates were immunoprecipitated with GFP-Trap beads and analyzed by Western blot with the indicated antibodies. NTD: N-terminal domain (aa 1–183); CTD: C-terminal Tubby domain (aa 184–442). Numbers at the bottom show quantitation of Flag-INPP5E co-IP as percentage of WT, normalized by both immunoprecipitated EGFP-TULP3, and by Flag-INPP5E lysate levels. (d) Schema of INPP5E-TULP3 interaction. On INPP5E’s side, the interaction mostly involves the catalytic domain, requiring CLS2 and CLS3. On TULP3’s side, the interaction occurs mostly through the CTD and is affected by the ARL13B-binding K389, and by the phosphoinositide (PIPs)-binding K268 and R270. (e) Lysates of HEK293T cells expressing the indicated EGFP-INPP5E variants were immunoprecipitated with GFP-Trap beads and the levels of endogenous CEP164 and exogenous EGFP were analyzed by Western blot as indicated. Molecular weight markers on the right. (f) Quantitation of endogenous CEP164 co-IP by the indicated EGFP-INPP5E constructs in HEK293T cells. Co-IP levels were calculated and plotted as in (b). Data are mean ± SEM of n=5,5,3,5,5,3,3 independent experiments and were analyzed by one-way ANOVA followed by Dunnett multiple comparisons tests relative to WT. Significance is shown as p<0.05(*). (g) Flag-INPP5E was coexpressed in HEK293T cells with the indicated CEP164-EGFP variants, including full length CEP164 (aa 1–1460), its N-terminal (NT, 1–467), middle (Mid, 468–1135) and C-terminal (CT, 1136–1460) regions, and NT carrying a mutated WW domain (WW: aa 56–89; mutation: Y73A+Y74 A). Arrows indicate the positions of these proteins. Lysates were immunoprecipitated with GFP-Trap beads and analyzed by Western blot with antibodies against Flag or EGFP, as indicated. Molecular weight markers are displayed on the right. (h) CLS2 and CLS3 are still required for INPP5E ciliary targeting in mutants unable to bind CEP164. Cilia localization was analyzed as in previous figures for the indicated EGFP-INPP5E variants, all of which lack aa 1–273 and hence cannot bind CEP164. Images are representative of n=2 independent experiments, with >30 transfected-cell cilia visualized per construct and experiment. Scale bar, 5 µm. (i) Schema summarizing results from (e–h). CEP164-NT is sufficient for INPP5E binding provided the WW domain is intact. On INPP5E’s side, the proline-rich N-terminal region (aa 1–283) is sufficient to interact with CEP164. Moreover, INPP5E(1-283), INPP5E(∆CLS2) and INPP5E(∆CLS3) mutants all bind CEP164 more intensely than INPP5E(WT), indicating that INPP5E’s C-terminal region downregulates CEP164 binding in a CLS2/3-dependent manner. This may or may not be necessary for INPP5E ciliary targeting, but it is clearly not sufficient, as shown by the data in (h).
-
Figure 7—source data 1
Uncropped immunoblots from Figure 7.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-data1-v1.pdf
-
Figure 7—source data 2
Uncropped immunoblot from Figure 7a (IP: EGFP; WB: TULP3).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-data2-v1.tif
-
Figure 7—source data 3
Uncropped immunoblot from Figure 7a (IP: EGFP; WB: EGFP).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-data3-v1.zip
-
Figure 7—source data 4
Uncropped immunoblot from Figure 7a (Lysates, WB: TULP3).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-data4-v1.tif
-
Figure 7—source data 5
Uncropped immunoblot from Figure 7c (IP: EGFP; WB: Flag).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-data5-v1.zip
-
Figure 7—source data 6
Uncropped immunoblot from Figure 7c (IP: EGFP; WB: EGFP).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-data6-v1.zip
-
Figure 7—source data 7
Uncropped immunoblot from Figure 7c (Lysates, WB: Flag).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-data7-v1.tif
-
Figure 7—source data 8
Uncropped immunoblot from Figure 7e (IP: EGFP; WB: CEP164).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-data8-v1.tif
-
Figure 7—source data 9
Uncropped immunoblot from Figure 7e (IP: EGFP; WB: EGFP).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-data9-v1.tif
-
Figure 7—source data 10
Uncropped immunoblot from Figure 7e (Lysates, WB: CEP164).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-data10-v1.tif
-
Figure 7—source data 11
Uncropped immunoblot from Figure 7g (IP: EGFP; WB: Flag).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-data11-v1.tif
-
Figure 7—source data 12
Uncropped immunoblot from Figure 7g (IP: EGFP; WB: EGFP).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-data12-v1.tif
-
Figure 7—source data 13
Uncropped immunoblot from Figure 7g (Lysates, WB: Flag).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-data13-v1.tif
-
Figure 7—source data 14
Source data from Figure 7b.
Data from n=7 experiments.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-data14-v1.xlsx
-
Figure 7—source data 15
Source data from Figure 7f.
Data from n=5 experiments.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-data15-v1.xlsx
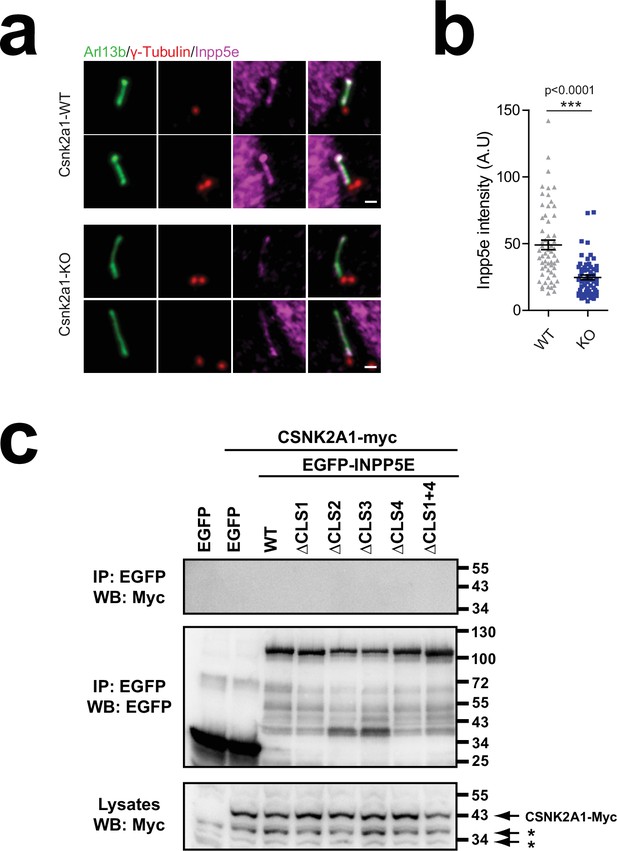
CSNK2A1 regulates INPP5E ciliary targeting without strongly interacting with it.
(a) Csnk2a1-WT and KO MEFs were stained with antibodies against Arl13b (green), gamma-tubulin (red), and Inpp5e (magenta). Scale bar, 1 µm. (b) Quantitation of Inpp5e ciliary intensity in Csnk2a1 WT and KO MEFs. Data are mean ± SEM. Statistical significance in Mann-Whitney non-parametric two-tailed test is shown. (c) The indicated EGFP-INPP5E variants were coexpressed in HEK293T cells with CSNK2A1-myc as indicated. Lysates were immunoprecipitated with GFP-Trap beads and analyzed by western blot with the indicated antibodies. No interaction was detected between EGFP-INPP5E and CSNK2A1-myc. Asterisks indicate non-specific bands. Molecular weight markers are shown on the right.
-
Figure 7—figure supplement 1—source data 1
Uncropped immunoblots from Figure 7—figure supplement 1c.
Relevant bands are inside rectangles. EGFP bands also visible on Myc immunoblot on the left. See Figure 7—figure supplement 1 for more details.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-figsupp1-data1-v1.pdf
-
Figure 7—figure supplement 1—source data 2
Uncropped immunoblot from Figure 7—figure supplement 1c (IP: EGFP; WB: Myc).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-figsupp1-data2-v1.zip
-
Figure 7—figure supplement 1—source data 3
Uncropped immunoblot from Figure 7—figure supplement 1c (IP: EGFP; WB: EGFP).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-figsupp1-data3-v1.zip
-
Figure 7—figure supplement 1—source data 4
Uncropped immunoblot from Figure 7—figure supplement 1c (Lysates; WB: Myc).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig7-figsupp1-data4-v1.zip
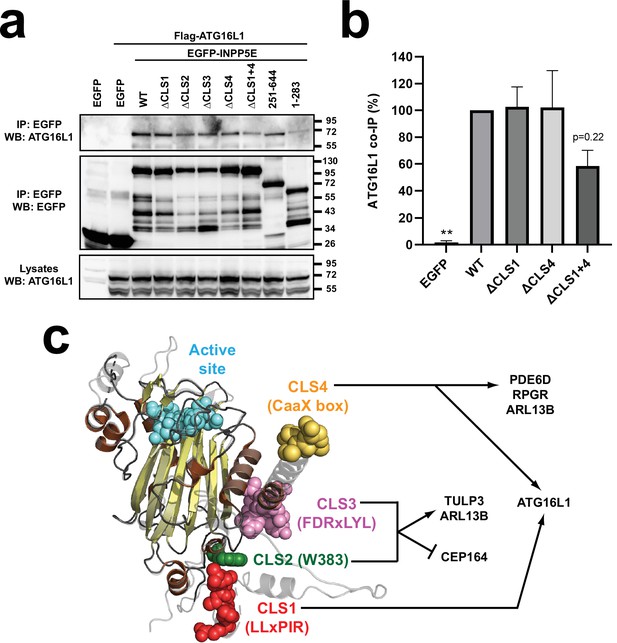
CLS1 and CLS4 jointly modulate the ATG16L1-INPP5E interaction.
(a) The indicated EGFP-INPP5E constructs were coexpressed in HEK293T cells with Flag-ATG16L1, as shown. Lysates were immunoprecipitated with GFP-Trap beads and analyzed by Western blot with the indicated antibodies. (b) Quantitation of Flag-ATG16L1 co-IP by the indicated EGFP-INPP5E constructs in HEK293T cells. Co-IP levels, expressed as percentage of WT, are normalized by the amounts of both immunoprecipitated EGFP-INPP5Es and Flag-ATG16L1 lysate levels. Data are mean ± SEM of n=3 independent experiments and were analyzed by one-way ANOVA followed by Dunnett tests relative to WT. Significance is shown as p<0.01(**). (c) Schema of INPP5E structure depicting CLS1-4 and the proteins through which they regulate INPP5E ciliary targeting, as shown herein.
-
Figure 8—source data 1
Uncropped immunoblots from Figure 8a.
Relevant bands are inside rectangles. See Figure 8 for more details.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig8-data1-v1.pdf
-
Figure 8—source data 2
Uncropped immunoblot from Figure 8a (IP: EGFP; WB: ATG16L1).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig8-data2-v1.zip
-
Figure 8—source data 3
Uncropped immunoblot from Figure 8a (IP: EGFP; WB: EGFP).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig8-data3-v1.zip
-
Figure 8—source data 4
Uncropped immunoblot from Figure 8a (Lysates, WB: ATG16L1).
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig8-data4-v1.zip
-
Figure 8—source data 5
Source data from Figure 8b.
Data from n=3 experiments.
- https://cdn.elifesciences.org/articles/78383/elife-78383-fig8-data5-v1.xlsx
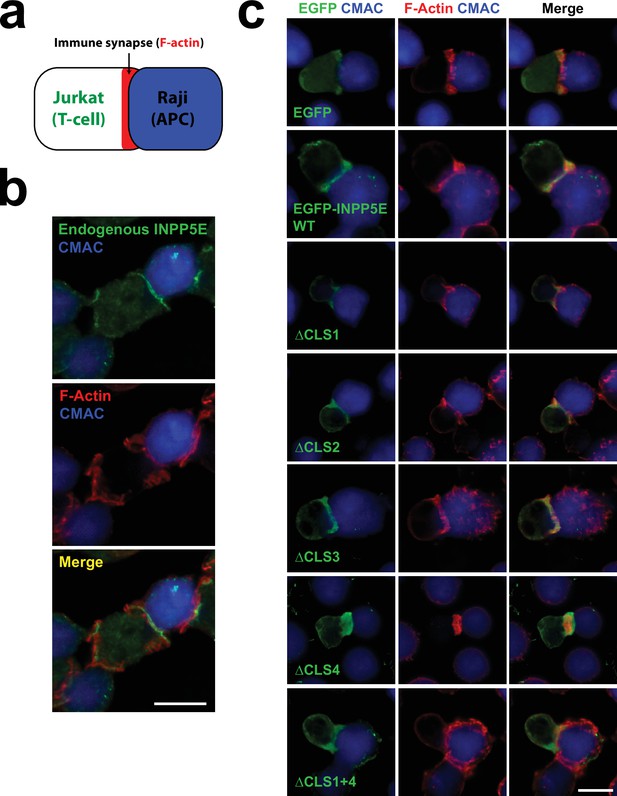
INPP5E targeting to the T-cell immune synapse is CLS-independent.
(a) Schema depicting an immune synapse (IS) between an antigen-presenting cell (APC) and a T-cell. Herein, Raji and Jurkat cells were used as APC and T-cells, respectively. F-actin is a marker well known to accumulate in productive immune synapses. (b) Jurkat cells were challenged with CMAC-labelled SEE-pulsed Raji cells to induce synaptic conjugate formation. Cells were fixed, permeabilized and stained with Alexa Fluor 546-conjugated phalloidin (F-actin, red) and anti-INPP5E antibody (green). CMAC (7-amino-4-chloromethylcoumarin) is shown in blue. Scale bar, 10 µm. (c) Jurkat cells expressing the indicated EGFP fusion proteins were challenged and stained as in (b), except that an anti-EGFP antibody (green) was used instead of anti-INPP5E. Scale bar, 10 µm. Cells in (b–c) were imaged by epifluorescence microscopy.
Tables
CLS-dependence of INPP5E protein-protein interactions.
Cilia localization and the indicated interactions are shown for each EGFP-INPP5E construct on the left column. For both localization and interactions, meaning of arrows is as follows: two upward green arrows (strong), one upward green arrow (moderate), one downward red arrow (low), and two downward red arrows (undetectable).
| INPP5E construct | Ciliary? | INPP5E interactors | |||||
|---|---|---|---|---|---|---|---|
| PDE6D | RPGR | ARL13B | TULP3 | CEP164 | ATG16L1 | ||
| WT | ↑↑ | ↑ | ↑ | ↑ | ↑ | ↑ | ↑ |
| ΔCLS1 | ↑ | ↑ | ↑ | ↑ | ↑ | ↑ | ↑ |
| ΔCLS2 | ↓↓ | ↑ | ↑ | ↑ | ↓ | ↑↑ | ↑ |
| ΔCLS3 | ↓↓ | ↑ | ↑ | ↑ | ↓ | ↑↑ | ↑ |
| ΔCLS4 | ↑ | ↓↓ | ↓↓ | ↓ | ↑ | ↑ | ↑ |
| ΔCLS1+4 | ↓↓ | ↓↓ | ↓↓ | ↓ | ↑ | ↑ | ↓ |
| CT (251-644) | ↑↑ | ↑ | ↑ | ↑ | ↑ | ↓↓ | ↑ |
| NT (1-283) | ↓↓ | ↓↓ | ↓↓ | ↓↓ | ↓ | ↑↑ | ↓ |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Cell line (Homo sapiens) | hTERT-RPE1 | ATCC | Cat # CRL-4000 | Derived from retinal pigmented epithelium |
| Cell line (Homo sapiens) | Puromycin-sensitive hTERT-RPE1 | This study | Puromycin Acetyltransferase (PAC)-KO cells | Also used in Gonçalves et al., 2021 |
| Cell line (Homo sapiens) | INPP5E-KO hTERT-RPE1 | This study | Clone 3 | Derived from puromycin-sensitive hTERT-RPE1 |
| Cell line (Homo sapiens) | INPP5E-KO hTERT-RPE1 | This study | Clone 12 | Derived from puromycin-sensitive hTERT-RPE1 |
| Cell line (Homo sapiens) | 293T | ATCC | Cat # CRL-3216 | Derived from human embryonic kidney |
| Cell line (Homo sapiens) | HeLa | ATCC | Cat # CCL-2 | Derived from cervical carcinoma |
| Cell line (Homo sapiens) | Jurkat, Clone E6-1 | ATCC | Cat # TIB-152 | T lymphoblasts from acute T cell leukemia |
| Cell line (Homo sapiens) | Raji | ATCC | Cat # CCL-86 | Lymphoblast-like cells from Burkitt’s lymphoma |
| Cell line (Mus musculus) | Csnk2a1-WT MEFs | Loukil et al., 2021 | N/A | Control mouse embryonic fibroblasts (MEFs) |
| Cell line (Mus musculus) | Csnk2a1-KO MEFs | Loukil et al., 2021 | N/A | Casein kinase 2 subunit alpha-null MEFs |
| Antibody | Anti-acetylated α-tubulin (mouse monoclonal) | Sigma-Aldrich (Merck) | Cat # T7451 (clone 6-11B-1) | IF: 1:10,000 |
| Antibody | Anti-α-tubulin (mouse monoclonal) | Proteintech | Cat # 66031–1-Ig | WB: 1:1000 |
| Antibody | Anti-γ-tubulin (mouse monoclonal) | Santa Cruz | Cat # sc-17787 | IF: 1:200 |
| Antibody | Anti-EGFP (mouse monoclonal) | Proteintech | Cat # 66002–1-Ig | WB: 1:1000 |
| Antibody | Anti-EGFP (mouse monoclonal) | Santa Cruz | Cat # sc-9996 | IF: 1:200 |
| Antibody | Anti-ARL13B (mouse monoclonal) | Proteintech | Cat # 66739–1-Ig | WB: 1:1000 |
| Antibody | Anti-Flag (mouse monoclonal) | Sigma-Aldrich (Merck) | Cat # F3165 (clone M2) | WB: 1:2000 |
| Antibody | Anti-polyglutamylated tubulin (mouse monoclonal) | Adipogen | Cat # GT335 | IF: 1:2000 |
| Antibody | Anti-EGFP (rabbit polyclonal) | Proteintech | Cat # 50430–2-AP | IF: 1:200 WB: 1:1000 |
| Antibody | Anti-RPGR (rabbit polyclonal) | Proteintech | Cat # 16891–1-AP | WB: 1:1000 |
| Antibody | Anti-TULP3 (rabbit polyclonal) | Proteintech | Cat # 13637–1-AP | IF: 1:750 WB: 1:2000 |
| Antibody | Anti-CEP164 (rabbit polyclonal) | Proteintech | Cat # 22227–1-AP | WB: 1:1000 |
| Antibody | Anti-ATG16L (rabbit polyclonal) | MBL | Cat # PM040 | WB: 1:1000 |
| Antibody | Anti-Myc (rabbit polyclonal) | Proteintech | Cat # 16286–1-AP | WB: 1:1000 |
| Antibody | Anti-INPP5E (rabbit polyclonal) | Proteintech | Cat # 17797–1-AP | IF: 1:100 |
| Antibody | Anti- γ-tubulin (goat polyclonal) | Santa Cruz | Cat # sc-7396 (discontinued) | IF: 1:200 |
| Antibody | Alexa Fluor 488 donkey anti-rabbit IgG (donkey polyclonal) | Thermo Fisher | Cat # A21206 | IF: 1:10,000 |
| Antibody | Alexa Fluor 555 donkey anti-mouse IgG (donkey polyclonal) | Thermo Fisher | Cat # A31570 | IF: 1:10,000 |
| Antibody | Alexa Fluor 647 donkey anti-goat IgG (donkey polyclonal) | Thermo Fisher | Cat # A21447 | IF: 1:10,000 |
| Antibody | Alexa Fluor 488 goat anti-mouse IgG2a (goat polyclonal) | Thermo Fisher | Cat # A21131 | IF: 1:10,000 |
| Antibody | Alexa Fluor 555 goat anti-mouse IgG1 (goat polyclonal) | Thermo Fisher | Cat # A21127 | IF: 1:10,000 |
| Antibody | Alexa Fluor 555 goat anti-mouse IgG2b (goat polyclonal) | Thermo Fisher | Cat # A21147 | IF: 1:10,000 |
| Antibody | Alexa Fluor 647 goat anti-mouse IgG2a (goat polyclonal) | Thermo Fisher | Cat # A21241 | IF: 1:10,000 |
| Antibody | HRP-conjugated goat anti-mouse IgG (goat polyclonal) | Thermo Fisher | Cat # A16072 | WB: 62 ng/ml |
| Antibody | HRP-conjugated goat anti-rabbit IgG (goat polyclonal) | Thermo Fisher | Cat # A16104 | WB: 62 ng/ml |
| Antibody | GFP-Trap_MA magnetic agarose beads (alpaca monoclonal) | Chromotek (Proteintech) | Cat # gtma-20 | IP: 10 µl slurry for 500 µl lysate (1:50) |
| Recombinant DNA reagent | EGFP-INPP5E | Jacoby et al., 2009 | Human INPP5E NM_019892.6 (644 amino acids) | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (MORM) | Jacoby et al., 2009 | 1–626 (MORM: Δ627–644) | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (D477N) | Garcia-Gonzalo et al., 2015 | D477N | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (1-623) | This study | 1–623 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (1-621) | This study | 1–621 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (1-618) | This study | 1–618 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (1-616) | This study | 1–616 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (1-608) | This study | 1–608 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (1-283) | This study | 1–283 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (Δ297–599) | This study | Δ297–599 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (100-644) | This study | 100–644 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (200-644) | This study | 200–644 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (251-644) | This study | 251–644 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (274-644) | This study | 274–644 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (288-644) | This study | 288–644 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (351-644) | This study | 351–644 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (451-644) | This study | 451–644 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (551-644) | This study | 551–644 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (FDR609AAA) | This study | F609A+D610A+R611 A (aka ΔCLS3 or FDR >AAA) | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (LYL613AAA) | This study | L613A+Y614A+L615 A (aka LYL >AAA) | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (F609A) | This study | F609A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (D610A) | This study | D610A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (R611A) | This study | R611A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (E612A) | This study | E612A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (L613A) | This study | L613A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (Y614A) | This study | Y614A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (L615A) | This study | L615A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (R345A+R346 A) | This study | R345A+R346 A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (E347A) | This study | E347A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (W348A) | This study | W348A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (E349A) | This study | E349A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (Q353A) | This study | Q353A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (E354A) | This study | E354A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (Y360A) | This study | Y360A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (Y360F) | This study | Y360F | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (V361A) | This study | V361A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (R378A+R379 A) | This study | R378A+R379 A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (R378A) | This study | R378A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (R379A) | This study | R379A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (D380A) | This study | D380A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (I382A) | This study | I382A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (W383A) | This study | W383A (aka ΔCLS2) | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (W383F) | This study | W383F | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (W383I) | This study | W383I | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (W383L) | This study | W383L | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (W383M) | This study | W383M | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (W383V) | This study | W383V | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (W383E) | This study | W383E | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (W383R) | This study | W383R | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (F384A) | This study | F384A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (E387A) | This study | E387A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (Δ251–273) | This study | Δ251–273 (aka ΔCLS1) | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (C641S) | This study | C641S (aka ΔCLS4) | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (Δ251–273+C641 S) | This study | Δ251–273+C641 S (aka ΔCLS1+4) | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (288-626) | This study | 288–626 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (274-626) | This study | 274–626 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (269-626) | This study | 269–626 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (264-626) | This study | 264–626 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (257-626) | This study | 257–626 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (251-626) | This study | 251–626 | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (257-626) +FS257AA | This study | (257-626)+F257A+S258 A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (257-626) +LL259AA | This study | (257-626)+L259A+L260 A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (257-626) +PIR262AAA | This study | (257-626)+P262A+I263A+R264 A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (257-626) +SK265AA | This study | (257-626)+S265A+K266 A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (257-626) +DV267AA | This study | (257-626)+D267A+V268 A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (257-626) +L259 A | This study | (257-626)+L259 A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (257-626) +L260 A | This study | (257-626)+L260 A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (257-626) +P262 A | This study | (257-626)+P262 A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (257-626) +I263 A | This study | (257-626)+I263 A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (257-626) +R264 A | This study | (257-626)+R264 A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (G286R) | This study | G286R | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (V303M) | This study | V303M | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (R345S) | This study | R345S | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (T355M) | This study | T355M | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (R378C) | This study | R378C | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (C385Y) | This study | C385Y | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (V388L) | This study | V388L | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (W474R) | This study | W474R | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (D490Y) | This study | D490Y | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (R621Q) | This study | R621Q | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (R621W) | This study | R621W | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (C641R) | This study | C641R | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (274-644) + ΔCLS2 | This study | (274-644)+W383 A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-INPP5E (274-644) + ΔCLS3 | This study | (274-644)+F609A+D610A+R611 A | XhoI-KpnI cloning into pEGFP-C1 |
| Recombinant DNA reagent | Flag-INPP5E | This study | Human INPP5E NM_019892.6 (644 amino acids) | EcoRI-KpnI cloning into pFlag-CMV4 |
| Recombinant DNA reagent | mVenus-PDE6D | This study | Human PDE6D NM_002601.4 (150 amino acids) | Cloned into p-mVenus-C1 |
| Recombinant DNA reagent | Flag-PDE6D | This study | Human PDE6D NM_002601.4 (150 amino acids) | mVenus-PDE6D cassette from eponymous plasmid excised with AgeI-EcoRI and replaced by Flag-PDE6D |
| Recombinant DNA reagent | pcDNA3.1(+)-N-DYK-RPGR | GenScript | Human RPGR NM_000328.3 (815 amino acids) | Plasmid, expresses Flag-RPGR (aka DYK-RPGR) |
| Recombinant DNA reagent | ARL13B-EYFP | This study | Human ARL13B NM_001174150.2 (428 amino acids) | Cloned into pEYFP-C1 |
| Recombinant DNA reagent | ARL13B-EGFP | This study | Human ARL13B NM_001174150.2 (428 amino acids) | XhoI-BamHI cloning into pEGFP-N1 |
| Recombinant DNA reagent | ARL13B-Flag | This study | Human ARL13B NM_001174150.2 (428 amino acids) | EGFP in ARL13B-EGFP was swapped by Flag using AgeI-NotI and pre-annealed Flag-encoding primers. |
| Recombinant DNA reagent | pcDNA3.1-TULP3-myc-his | Barbeito and Garcia-Gonzalo, 2021 | Human TULP3 NP_003315.2 (442 amino acids) | XhoI-BamHI into pcDNA3.1-myc-his(-)C |
| Recombinant DNA reagent | pG-LAP1-TULP3 | Mukhopadhyay et al., 2010 | Human TULP3 NP_003315.2 (442 amino acids) | Expresses EGFP-Stag-TULP3 (LAP-TULP3) |
| Recombinant DNA reagent | pG-LAP1-TULP3-KR | Mukhopadhyay et al., 2010 | TULP3 (K268A+R270 A) | Phosphoinositide binding-defective mutant |
| Recombinant DNA reagent | EGFP-TULP3 (NTD) | This study | TULP3 (1–183) | KpnI-BamHI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-TULP3 (CTD) | This study | TULP3 (184–442) (aka Tubby domain) | KpnI-BamHI cloning into pEGFP-C1 |
| Recombinant DNA reagent | EGFP-TULP3 (K389A) | This study | Human TULP3 NP_003315.2 (442 amino acids) | KpnI-BamHI cloning into pEGFP-C1 |
| Recombinant DNA reagent | CEP164-EGFP | Addgene | Addgene plasmid; RRID:Addgene_41149 | Human CEP164 NM_014956.5 (1460 amino acids) |
| Recombinant DNA reagent | CEP164-EGFP (1-467) | This study | CEP164-NT (aka N-term in Cajanek and Nigg, 2014) | CEP164-NT replaces full length in CEP164-EGFP (EcoRI-KpnI) |
| Recombinant DNA reagent | CEP164-EGFP (1-467)+WWmut | This study | (1-467) +Y73A+Y74 A | Mutant of WW domain |
| Recombinant DNA reagent | CEP164-EGFP (468–1135) | This study | CEP164-Mid (aka M-part in Cajanek and Nigg, 2014) | CEP164-Mid replaces full length in CEP164-EGFP (EcoRI-KpnI) |
| Recombinant DNA reagent | CEP164-EGFP (1136–1460) | This study | CEP164-CT (aka C-term in Cajanek and Nigg, 2014) | CEP164-CT replaces full length in CEP164-EGFP (EcoRI-KpnI) |
| Recombinant DNA reagent | pMRX-IP-SECFP-hATG16A1 | Addgene | Addgene plasmid; RRID:Addgene_58994 | Human ATG16L1 NP_060444.3 (588 amino acids) |
| Recombinant DNA reagent | Flag-ATG16L1 | This study | Human ATG16L1 NP_060444.3 (588 amino acids) | EcoRI insert from pMRX-IP-SECFP-hATG16A1 transferred to pFlagCMV4 |
| Recombinant DNA reagent | CSNK2A1-myc | This study | Mouse CSNK2A1 NP_031814.2 (391 amino acids) | XhoI-BamHI into pcDNA3.1-myc-his(-)C |
| Recombinant DNA reagent | FRB-CFP-CaaX | Roy et al., 2020 Biorxiv | FRB* domain of human mTOR (as in RRID:Addgene_20148) | CaaX box of K-Ras targets FRB-CFP to inner leaflet of plasma membrane |
| Recombinant DNA reagent | mCherry-FKBP | This study | Human FKBP1A (aka FKBP12) NP_000792.1 (108 amino acids) | Cloned into p-mCherry-C1 |
| Recombinant DNA reagent | mCherry-FKBP-INPP5E(WT) | This study | Human INPP5E NM_019892.6 (644 amino acids) | Cloned into mCherry-FKBP |
| Recombinant DNA reagent | mCherry-FKBP-INPP5E(ΔCLS1) | This study | INPP5E Δ251–273 | Cloned into mCherry-FKBP |
| Recombinant DNA reagent | mCherry-FKBP-INPP5E(ΔCLS2) | This study | INPP5E W383A | Cloned into mCherry-FKBP |
| Recombinant DNA reagent | mCherry-FKBP-INPP5E(ΔCLS3) | This study | INPP5E F609A+D610A+R611 A | Cloned into mCherry-FKBP |
| Recombinant DNA reagent | mCherry-FKBP-INPP5E(ΔCLS4) | This study | INPP5E C641S | Cloned into mCherry-FKBP |
| Recombinant DNA reagent | mCherry-FKBP-INPP5E (ΔCLS1+4) | This study | INPP5E Δ251–273+C641 S | Cloned into mCherry-FKBP |
| Recombinant DNA reagent | pSpCas9-sgPAC1 | This study | PAC gRNA#1 plasmid | PAC gRNA#1: ACGCGCGUCGGGCTCGACAUCGG |
| Recombinant DNA reagent | pSpCas9-sgPAC3 | This study | PAC gRNA#3 plasmid | PAC gRNA#3: CACGCGCCACACCGUCGAUCCGG |
| Recombinant DNA reagent | pSpCas9-sgPAC6 | This study | PAC gRNA#3 plasmid | PAC gRNA#6: GGCGGGGUAGUCGGCGAACGCGG |
| Recombinant DNA reagent | pSpCas9-hINPP5E-gRNA1 | This study | INPP5E gRNA#1 plasmid | INPP5E gRNA#1: CGGAGCCCGGAGCAUCGGGUGGG |
| Recombinant DNA reagent | pSpCas9-hINPP5E-gRNA2 | This study | INPP5E gRNA#2 plasmid | INPP5E gRNA#2: UGGAGCGUCCUCCCUUCCGGCGG |
| Recombinant DNA reagent | pSpCas9-hINPP5E-gRNA3 | This study | INPP5E gRNA#3 plasmid | INPP5E gRNA#3: ACAGCUUCCCGGCGCUCCGCCGG |
| Commercial assay or kit | Malachite Green Assay Kit | Echelon Biosciences (Tebu-Bio) | Cat # K-1500 | For measurement of phosphate release in activity assays. |
| Chemical compound, drug | PtdIns(4,5)P2-diC8 | Echelon Biosciences (Tebu-Bio) | Cat # P-4508 | Activity assays: 120 µM |
| Chemical compound, drug | n-octyl-β-D-glucopyranoside | Alfa Aesar | Cat # J67390.03 | Activity assays: 0.1% |
| Chemical compound, drug | Alexa Fluor 546 Phalloidin | Thermo Fisher | Cat # A22283 | IF: 1:100 |
| Software, algorithm | GraphPad Prism 9.4.0 | GraphPad Software Inc | RRID:SCR_002798 | https://www.graphpad.com/ |
| Software, algorithm | Fiji (Image J) | Schmidt et al., 2012 | RRID:SCR_002285 | http://imagej.net/Fiji |





