Autoantibody discovery across monogenic, acquired, and COVID-19-associated autoimmunity with scalable PhIP-seq
Figures
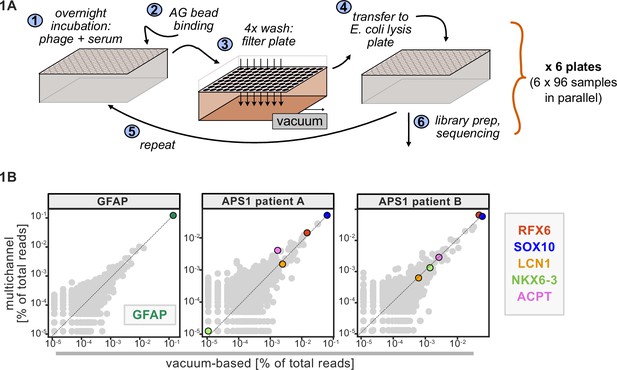
Advantages of and considerations motivating scaled phage-immunoprecipitation sequencing (PhIP-seq).
(A) Schematic of vacuum-based scaled PhIP-seq protocol, allowing for parallelized batches of 600–800 samples. (B) Comparison of moderate-throughput multichannel protocol data to high-throughput vacuum-based protocol data, with axes showing normalized read percentages. Controls include a commercial polyclonal anti-GFAP antibody (left), APS1 patient A with known and validated autoantibodies RFX6, SOX10, ACPT, and LCN1 (center), and APS1 patient B with the same known and validated autoantibodies as well as NKX6-3.
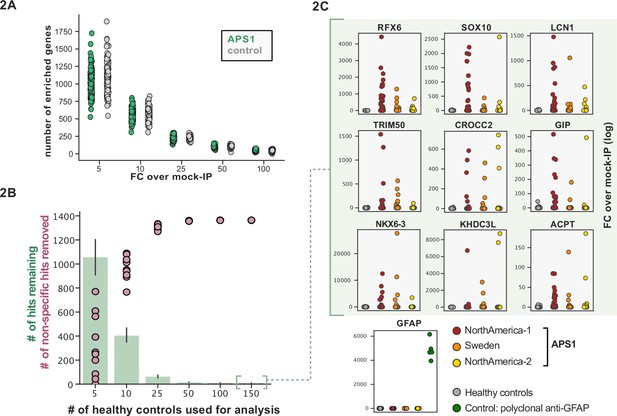
Application of scaled phage-immunoprecipitation sequencing to expanded APS1 and healthy control cohorts.
(A) Number of hits per sample reaching 5, 10, 25, 50, and 100-fold enrichment relative to mock-IP samples. Each dot represents a single APS1 patient (green) or non-APS1 control (gray). (B) When looking for disease-specific hits, increasing the number of healthy controls results in fewer apparent hits and is therefore critical. Shared hits are defined as gene-level signal (>10-fold change over mock-IP) which is shared among 10% of APS1 samples (n=128), present in fewer than 2% of healthy controls, and with at least one APS1 sample with a high signal (FC of 50<). Random downsampling was performed 10 times for each healthy control bin. (C) Nine gene-level hits are present in 10%< of a combined three-group APS1 cohort. North-America-1, n=62; Sweden, n=40; North-America-2, n=26. Anti-GFAP control antibody (n=5) indicates that results are consistent across plates and exhibit no well-to-well contamination.
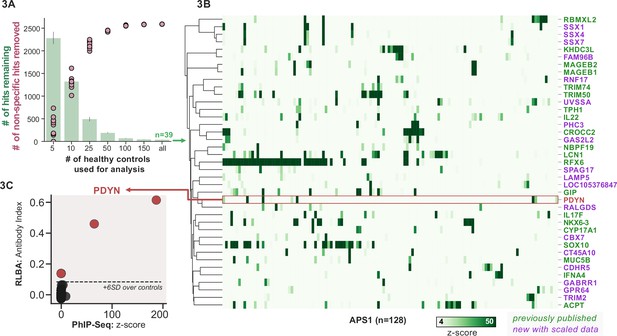
Replication and expansion of APS1 autoantigens across multiple cohorts using scaled phage-immunoprecipitation sequencing (PhIP-seq).
(A) Increasing the number of healthy controls results in fewer apparent hits and is therefore critical. Shared hits are defined as gene-level signal (>10-fold change over mock-IP) which is shared among 4%< of APS1 samples (n=128), present in fewer than 2% of healthy controls, and with at least one APS1 sample with a high signal (FC of 50<). Random downsampling was performed 10 times for each healthy control bin. (B) 39 candidate hits present in 4%< of the APS1 cohort. (C) Rare, novel anti-PDYN autoantibodies validate at whole-protein level, with PhIP-seq and whole-protein RLBA data showing good concordance.
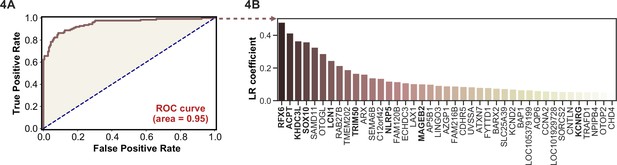
Logistic regression of phage-immunoprecipitation sequencing data enables APS1 disease prediction.
(A) Receiver operating characteristic (ROC) curve for prediction of APS1 versus control disease status. (B) The highest logistic regression (LR) coefficients include known antigens RFX6, KHDC3L, and others.
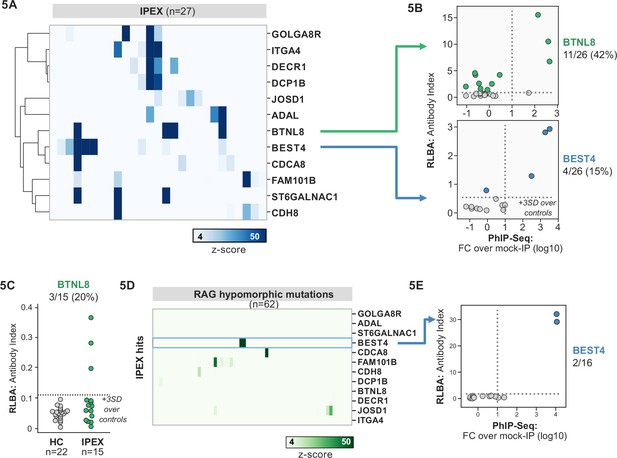
Phage-immunoprecipitation sequencing (PhIP-seq) screening in IPEX and RAG1/2 deficiency reveals novel, intestinally expressed autoantigens BEST4 and BTNL8.
(A) PhIP-seq heatmap of most frequent shared antigens among IPEX, with color indicating z-score relative to a cohort of non-IPEX controls. (B) Radioligand binding assay for BTNL8 reveals additional anti-BTNL8 positive IPEX patients (top). Radioligand binding assay for BEST4 autoantibodies correlates well with PhIP-seq data (bottom). (C) Discovery of additional anti-BTNL8 positive individuals in an independent IPEX cohort (n=15) by radioligand binding assay; dotted line indicates mean of healthy controls +3 SD. (D) PhIP-seq screen of patients with hypomorphic mutations in RAG1/2 reveals two patients with anti-BEST4 signal. (E) Orthogonal radioligand binding assay validation of anti-BEST4 antibodies in both PhIP-seq anti-BEST4 positive patients.
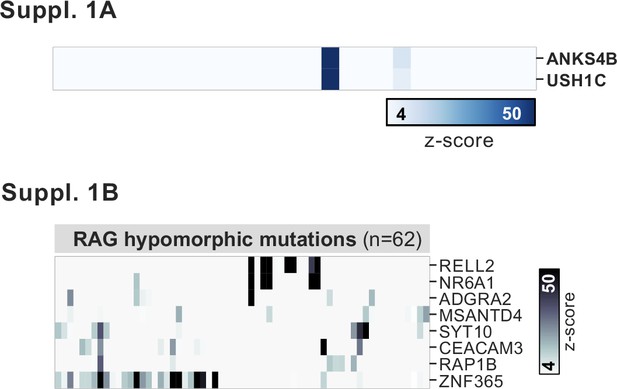
PhIP-seq application to a cohort of patients with hypomorphic RAG1/2 mutations.
(A) Phage-immunoprecipitation sequencing has low detection sensitivity for known antigens USH1C and ANKS4B, but those patients with positive signal exhibit the previously reported coupled signal for both antigens. (B) Additional, shared putative antigens within the cohort of RAG1/2-deficient patients (n=62).
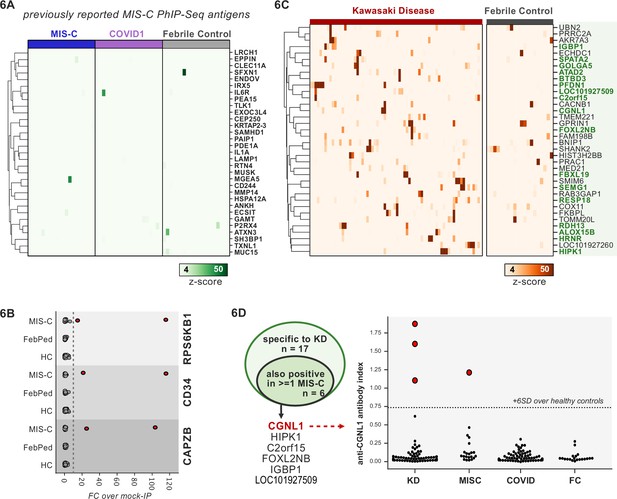
Phage-immunoprecipitation sequencing (PhIP-seq) screening of multisystem inflammatory syndrome in children (MIS-C) and Kawasaki disease (KD) cohorts.
(A) Heatmap of signal for putative hits from Gruber et al., 2020, among MIS-C, adult COVID-19 controls, and pediatric febrile controls (each n=20). (B) Only rare, shared PhIP-seq signals were found among n=20 MIS-C patients. (C) Heatmap of putative antigens in a cohort of n=70 KD patients. Hits that are specific to KD and are not found among n=20 febrile controls, are highlighted in green. (D) A small number of rare putative antigens are shared between KD and MIS-C (left), with radioligand binding assay confirmation of antibody reactivity to whole protein form of CGNL1 in three KD patients and one MIS-C patient (right).
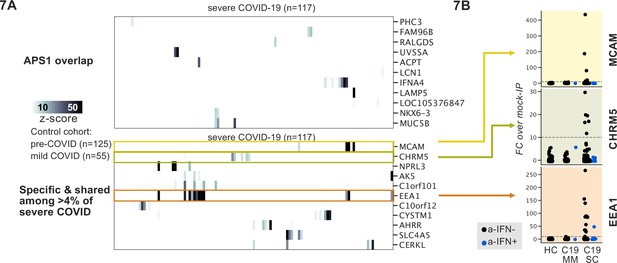
Phage-immunoprecipitation sequencing screening in severe forms of COVID-19 reveals putative novel autoantigens, including EEA1.
(A) Screening of patients with severe COVID-19 pneumonia shows little overlap with APS1 but enables discovery of possible novel disease-associated autoantigens including EEA1. (B) Putative novel antigens EEA1, CHRM5, and MCAM are primarily found in anti-IFN-negative patients, suggesting the possibility of other frequent, independent disease-associated antibodies in severe COVID-19.
Additional files
-
Supplementary file 1
Clinical metadata.
- https://cdn.elifesciences.org/articles/78550/elife-78550-supp1-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/78550/elife-78550-transrepform1-v2.pdf





