Dysregulated H19/Igf2 expression disrupts cardiac-placental axis during development of Silver-Russell syndrome-like mouse models
Figures
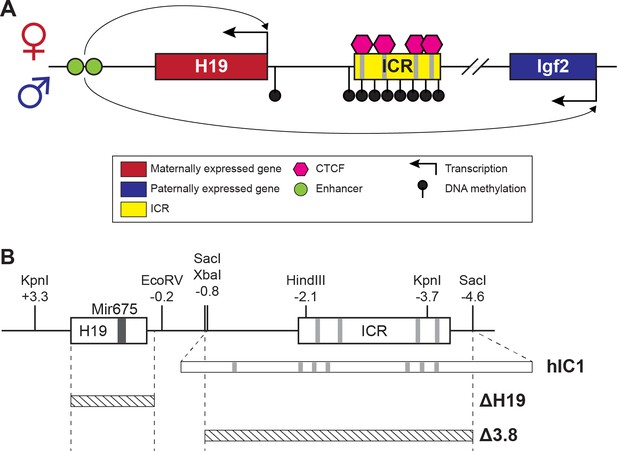
H19/Igf2 cluster and mouse models utilized in this study.
(A) A schematic representation of the wild-type H19/Igf2 cluster in mouse. The maternally expressed H19 and the paternally expressed Igf2 genes are shown in red and blue, respectively. Black lollipops on the paternal allele represent DNA methylation. The maternal imprinting control region (ICR) is bound to CTCF proteins (pink hexagons) at CTCF-binding sites, forming an insulator that blocks the maternal Igf2 promoter from the shared enhancers (green circles). These enhancers interact with the H19 promoter on the maternal allele and Igf2 promoter on the paternal allele. (B) Schematic of the mouse endogenous H19/Igf2 ICR, H19hIC1 (shortened as hIC1; Hur et al., 2016), H19Δ2.8kb-H19 (shortened as Δ H19), and H19 Δ3.8kb-5’H19 (shortened as Δ 3.8; Thorvaldsen et al., 2002; Thorvaldsen et al., 2006) alleles. Restriction site locations (kb) are relative to the H19 transcription start site. Gray lines on the ICR represent conserved CTCF-binding sequences.
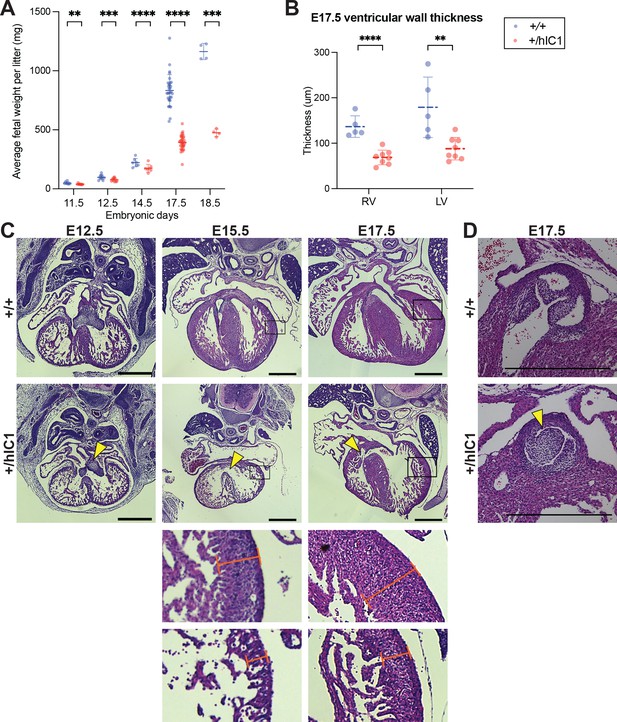
Growth anomalies and cardiac defects of +/hIC1 embryos.
(A) Fetal weight of the wild-type (blue) and +/hIC1 (red) embryos at E11.5, E12.5, E14.5, E17.5, and E18.5 (mean ± SD). Each data point represents an average weight of each genotype from one litter. 8 litters for E11.5, 10 litters for E12.5, 7 litters for E14.5, 31 litters for E17.5, and 4 litters for E18.5 are presented. (B) Quantification of ventricular wall thickness (µm), measured from E17.5 wild-type and +/hIC1 hearts (mean ± SD). Each data point represents an individual conceptus. Five wild-type and eight +/hIC1 embryos from four different litters were examined. (C) Representative cross-sections of wild-type and +/hIC1 embryonic hearts at E12.5, E15.5, and E17.5, stained with hematoxylin and eosin. All the hearts represented here are from female fetuses. Note the lack of fusion between atrioventricular (AV) cushions at E12.5, the ventricular septal defect (VSD) at E15.5, and E17.5 in +/hIC1 hearts (yellow arrowheads). The boxed regions of E15.5 and E17.5 images are enlarged at the bottom of the figure, to show where the ventricular wall thickness is measured. Scale bars = 500 µm. (D) A representative image of pulmonary valves in E17.5 wild-type and +/hIC1 hearts. The +/hIC1 right pulmonary cusp is enlarged (yellow arrowhead), and there is no cusp in the anterior position, in contrast to the tricuspid structure in the wild-type heart. Scale bars = 500 µm. Statistics used are (A) multiple paired t-test and (B) multiple unpaired t-test with **p<0.01, ***p<0.001, ****p<0.0001, and ns = not significant.
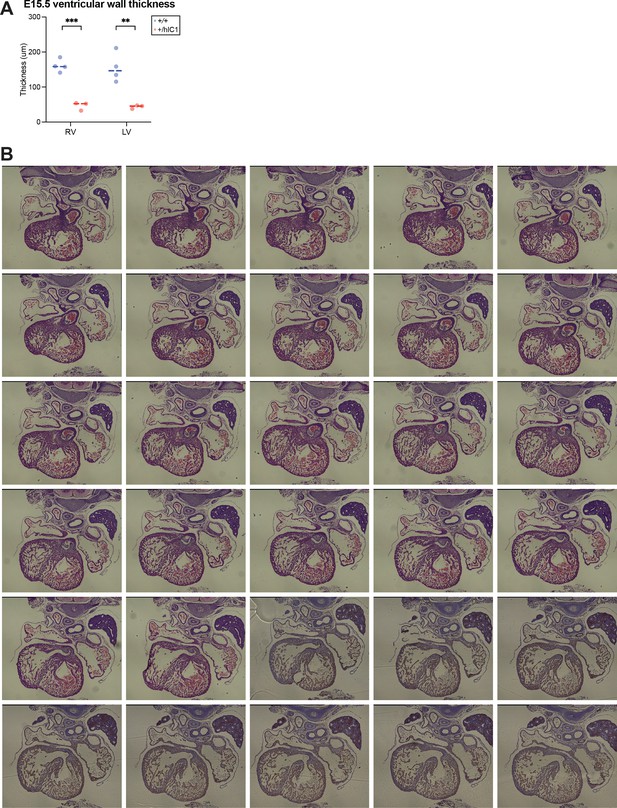
Supplementary data for anomalies observed in +/hIC1 hearts.
(A) Quantification of ventricular wall thickness (µm), measured from E15.5 wild-type and +/hIC1 hearts (mean ± SD). Four wild-type and three +/hIC1 embryos from two different litters were examined. Each data point represents an individual conceptus from different litters. Statistics used are multiple unpaired t-tests. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, and ns = not significant. (B) Sequential histological sections demonstrating the bicuspid pulmonary valve (BPV) phenotype.
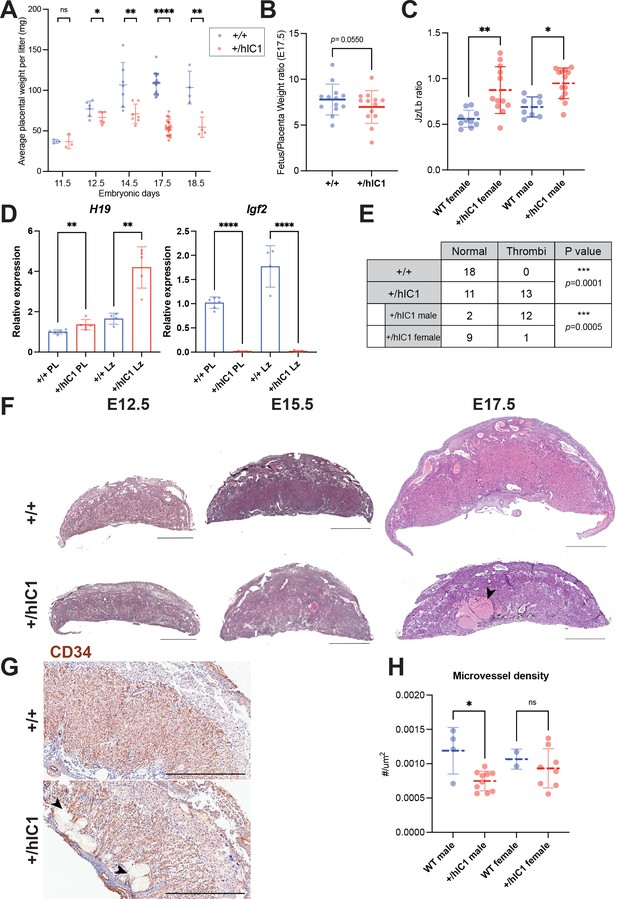
Placental defects of +/hIC1 embryos.
(A) Placental weight of the wild-type (blue) and +/hIC1 (red) samples at E11.5, E12.5, E14.5, E17.5, and E18.5 (mean ± SD). Each data point represents an average weight of each genotype from one litter. 3 litters for E11.5, 6 litters for E12.5, 7 litters for E14.5, 22 litters for E17.5, and 4 litters for E18.5 are presented. (B) Fetal to placental weight ratios in E17.5 wild-type and +/hIC1 samples (mean ± SD). Each data point represents the average F/P ratio of each genotype from one litter. 13 litters are presented. (C) Junctional zone (Jz) to labyrinth (Lb) ratio in E17.5 wild-type and +/hIC1 placentas (mean ± SD). (D) Relative total expression of H19 and Igf2 in E17.5 wild-type and +/hIC1 placentas and Lb samples (mean ± SD). (E) Number of wild-type, male, and female +/hIC1 placentas with thrombi observed. (F) Representative cross-sections of E12.5, E15.5, and E17.5 wild-type and +/hIC1 placentas stained with hematoxylin and eosin. All depicted E12.5 and E15.5 placentas are female. The E17.5 wild-type placenta is female, and +/hIC1 placenta is male. Black arrowhead indicates a large thrombus formed in the +/hIC1 Lb. Scale bars = 1 mm. (G) Representative images of CD34 immunostaining counterstained with hematoxylin on E17.5 wild-type female and +/hIC1 male placental sections. Black arrowheads indicate thrombi in the +/hIC1 Lb. Scale bars = 1 mm. (H) Quantification of the microvessel density in E17.5 wild-type and +/hIC1 placentas. 4 wild-type male, 10 +/hIC1 male, 2 wild-type female, 8 +/hIC1 female placentas from six litters were quantified. (C, D, and H) Each data point represents an individual conceptus from different litters. Statistics used are (A and B) multiple paired t-test, (C) one-way ANOVA with Tukey’s multiple comparisons test, (D and H) multiple unpaired t-test, (E) Fisher’s exact test. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, and ns = not significant.
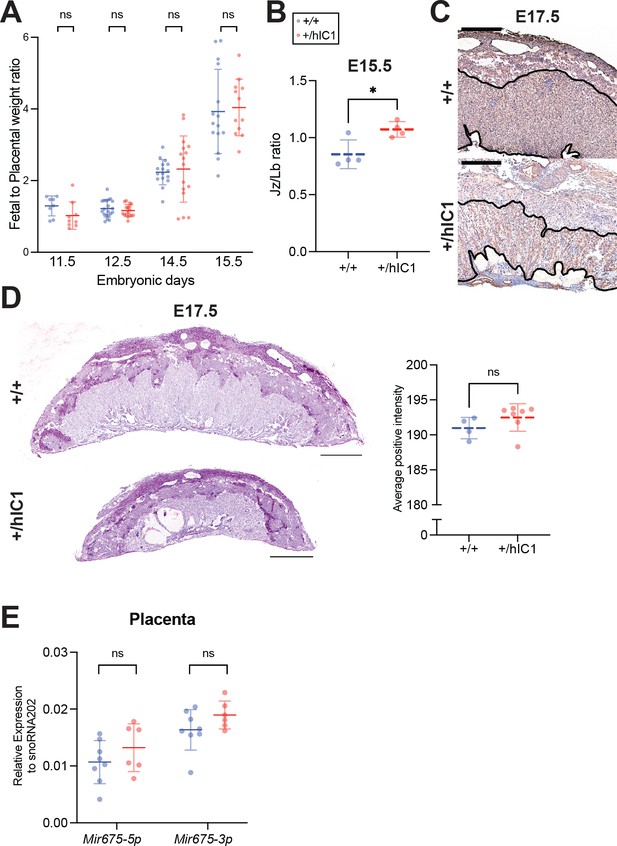
Supplementary data for anomalies observed in +/hIC1 placentas.
(A) Fetal to placental weight ratios of the wild-type (blue) and +/hIC1 (red) samples at E11.5, E12.5, E14.5, and E15.5. Each data point represents an average ratio of each genotype from one litter. (B) Junctional zone (Jz) to labyrinth (Lb) ratio in E17.5 wild-type and +/hIC1 placentas. (C) Example of Lb zones that were used to quantify the microvessel density in CD34 immunostained E17.5 wild-type and +/hIC1 placental sections from Figure 3G. Thrombi in +/hIC1 placentas were excluded. (D) Left: Representative images of Periodic acid-Schiff (PAS) staining counterstained with hematoxylin on E17.5 wild-type and +/hIC1 male placental sections. Right: Quantification of PAS stained images. (E) Relative total expression of Mir675-3p and Mir675-5p in E15.5 wild-type and +/hIC1 placentas (mean ± SD). (B, D, and E) Each data point represents an individual conceptus from different litters. Scale bars = 1 mm. Statistics used are (A) multiple paired t-test, (B, D, and E) multiple unpaired t-tests. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, and ns = not significant.
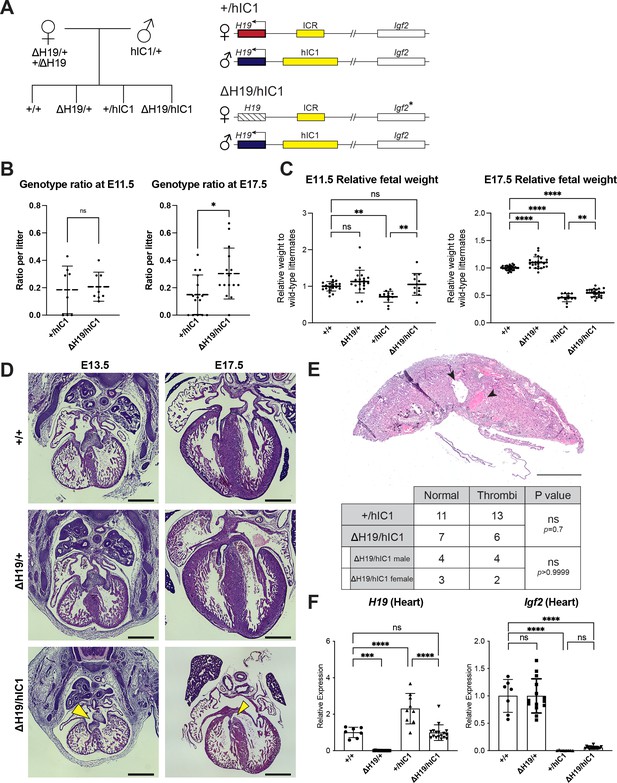
Normalizing H19 expression through the maternal deletion of H19.
(A) A schematic representation of the rescue breeding between △H19 heterozygous female and hIC1/+ male mice. △H19/hIC1 embryos are expected to express H19 only from the paternal allele and maternally express Igf2 in a tissue-specific manner. (B) Ratio of +/hIC1 and △H19/hIC1 embryos observed in E11.5 and E17.5 litters (mean ± SD). 8 E11.5 litters and 15 E17.5 litters with litter size larger than five pups were examined. Each data point represents one litter. (C) Relative fetal weights of wild-type, △H19/+, +/hIC1, and △H19/hIC1 embryos at E11.5 and E17.5, normalized to the average body weight of the wild-type littermates (mean ± SD). (D) Representative cross-sections of wild-type, △H19/+ and △H19/hIC1 embryonic hearts at E13.5 and E17.5, stained with hematoxylin and eosin. Note the cushion defect at E13.5 and the ventricular septal defect (VSD) at E17.5 in △H19/hIC1 hearts (yellow arrows). All E13.5 samples and E17.5 wild-type sample are male, and E17.5 △H19/+ and △H19/hIC1 samples are female. Scale bars = 500 µm. (E) Top: Representative cross-section of E17.5 △H19/hIC1 male placenta stained with hematoxylin and eosin. Black arrowheads indicate thrombi. Scale bar = 1 mm. Bottom: Number of the wild-type, male, and female +/hIC1 placentas with thrombi observed. (F) Relative total expression of H19 and Igf2 in E17.5 wild-type, △H19/+, +/hIC1, and △H19/hIC1 hearts (mean ± SD). (C and F) Each data point represents an individual conceptus from different litters. Statistics used are (B, C, and F) one-way ANOVA with Tukey’s multiple comparisons test and (E) Fisher’s exact test. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, and ns = not significant.
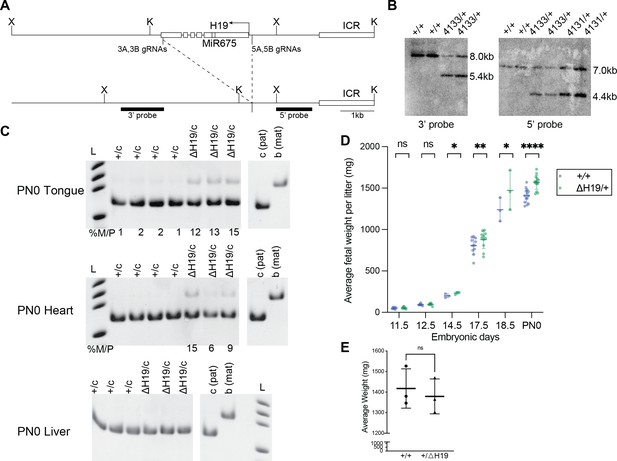
Characterization of ΔH19 allele.
(A) Targeting strategy to generate the ΔH19 allele. The endogenous H19 locus is shown with restriction sites, gRNA locations used to generate the deletion and binding sites for probes used in Southern blot analysis (thick lines). (B) Southern blot analysis of ΔH19 allele. Founder 4131 line was generated using gRNA pair A, and founder 4133 line was generated using gRNA pair B (Supplementary file 1). 3’ probe and 5’ probe were hybridized to XbaI- and KpnI-digested DNA, respectively. The sizes of the DNA fragments are shown on the right. (C) Allele-specific Igf2 expression in wild-type and ΔH19/+ neonatal tongue, heart, and liver analyzed by restriction fragment length polymorphism (RFLP). Ladder, genotypes and c (Mus castaneus, paternal) and b (C57BL/6, maternal) allele controls are indicated above each gel. Quantification of band densitometry is shown below each gel, with percent of maternal allele expression relative to paternal allele indicated. No expression from the maternal Igf2 allele was observed in liver. (D) Embryonic and neonatal body weight of the wild-type (blue) and ΔH19/+ (green) samples at E11.5, E12.5, E14.5, E17.5, E18.5, and PN0 (mean ± SD). 6 litters for E11.5, 6 litters for E12.5, 3 litters for E14.5, 11 litters for E17.5, 3 litters for E18.5, and 14 litters for PN0 are presented. (E) Body weight of +/ΔH19 neonates (mean ± SD). Three litters are presented. (D and E) Each data point represents an average weight of each genotype from one litter. Paired Student’s t-test; *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, and ns = not significant.
-
Figure 4—figure supplement 1—source data 1
Raw gel images for Figure 4—figure supplement 1B.
- https://cdn.elifesciences.org/articles/78754/elife-78754-fig4-figsupp1-data1-v1.zip
-
Figure 4—figure supplement 1—source data 2
Raw gel images for Figure 4—figure supplement 1C.
- https://cdn.elifesciences.org/articles/78754/elife-78754-fig4-figsupp1-data2-v1.zip
-
Figure 4—figure supplement 1—source data 3
Raw gel images for Figure 4—figure supplement 1C.
- https://cdn.elifesciences.org/articles/78754/elife-78754-fig4-figsupp1-data3-v1.zip
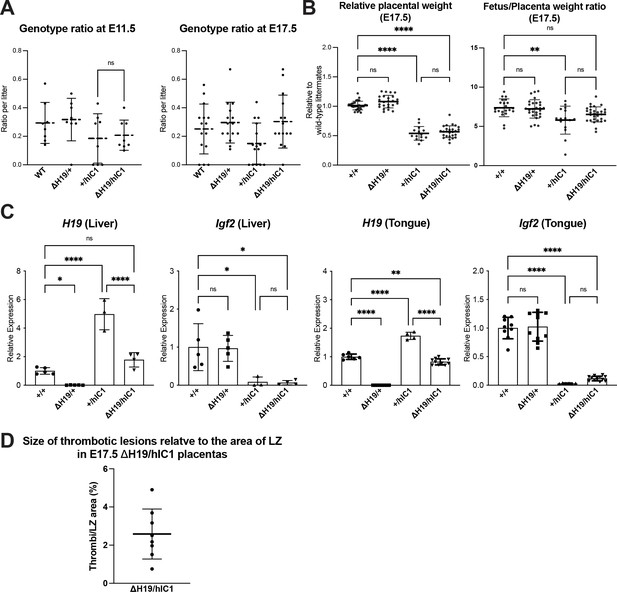
Supplementary data for rescue upon maternal ΔH19 transmission.
(A) Ratio of wild-type, ΔH19/+, +/hIC1, and ΔH19/hIC1 embryos observed in E11.5 and E17.5 litters (>5 pups; mean ± SD). Each data point represents one litter. (B) Left: Relative placental weights of E17.5 wild-type, ΔH19/+, +/hIC1, and ΔH19/hIC1 samples, normalized to the average placental weight of the wild-type littermates (mean ± SD). Right: Fetal to placental weight ratio in E17.5 wild-type, ΔH19/+, +/hIC1, and ΔH19/hIC1 samples (mean ± SD). (C) Relative total expression of H19 and Igf2 in E17.5 wild-type, ΔH19/+, +/hIC1, and ΔH19/hIC1 liver and tongue samples (mean ± SD). (D) % Area of thrombotic clots in ΔH19/hIC1 placentas relative to labyrinth zone. (B and C) Each data point represents an individual conceptus from different litters. Statistics used are (A) paired Student’s t-test and (B and C) one-way ANOVA with Tukey’s multiple comparisons test; *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, and ns = not significant.
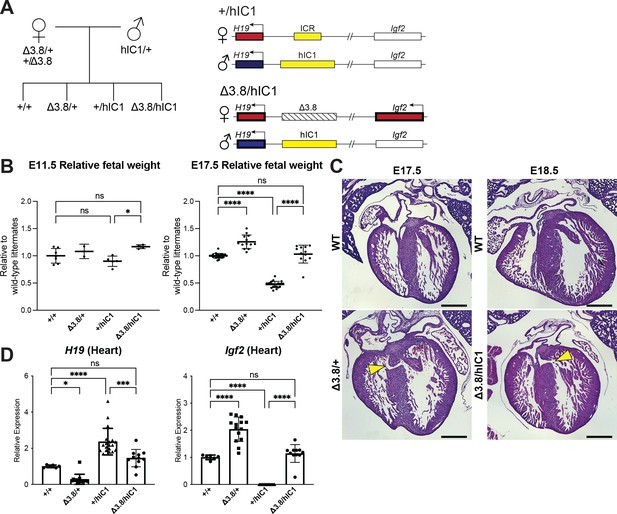
Restoring H19 and Igf2 expression utilizing maternal H19/Igf2 imprinting control region (ICR) deletion.
(A) A schematic representation of the offspring produced when △3.8 heterozygous female and hIC1/+ male mice are mated is depicted. △3.8/hIC1 embryos are expected to show activation of maternal Igf2 expression as well as paternal H19 expression. (B) Relative fetal weights of wild-type, △3.8/+, +/hIC1, and △3.8/hIC1 embryos at E11.5 and E17.5, normalized to the average body weight of wild-type littermates (mean ± SD). 2 E11.5 litters and 10 E17.5 litters are presented. (C) Representative cross-sections of E17.5 △3.8/+ and E18.5 △3.8/hIC1 embryonic hearts with ventricular septal defects (VSDs; yellow arrows), stained with hematoxylin and eosin. Sections from wild-type littermates are shown together for comparison. The E17.5 wild-type sample is male, and the rest are female. Scale bars = 500 µm. (D) Relative total expression of H19 and Igf2 in E17.5 wild-type, △3.8/+, +/hIC1, and △3.8/hIC1 hearts (mean ± SD). (B and D) Each data point represents an individual conceptus from different litters. One-way ANOVA with Tukey’s multiple comparisons test was used with *p<0.05, ***p<0.001, ****p<0.0001, and ns = not significant.
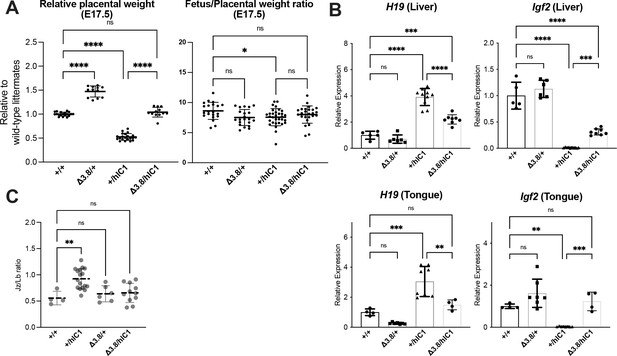
Supplementary data for rescue upon maternal Δ3.8 transmission.
(A) Left: Relative placental weights of E17.5 wild-type, Δ3.8/+, +/hIC1, and Δ3.8/hIC1 samples, normalized to the average placental weight of the wild-type littermates (mean ± SD). Right: Fetal to placental weight ratio in E17.5 wild-type, Δ3.8/+, +/hIC1, and Δ3.8/hIC1 samples (mean ± SD). (B) Relative total expression of H19 and Igf2 in E17.5 wild-type, Δ3.8/+, +/hIC1, and Δ3.8/hIC1 liver and tongue samples (mean ± SD). (C) Junctional zone (Jz) to labyrinth (Lb) ratio in E17.5 wild-type, Δ3.8/+, +/hIC1, and Δ3.8/hIC1 placentas. (A, B, and C) Each data point represents an individual conceptus from different litters. One-way ANOVA with Tukey’s multiple comparisons test; *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, and ns = not significant.
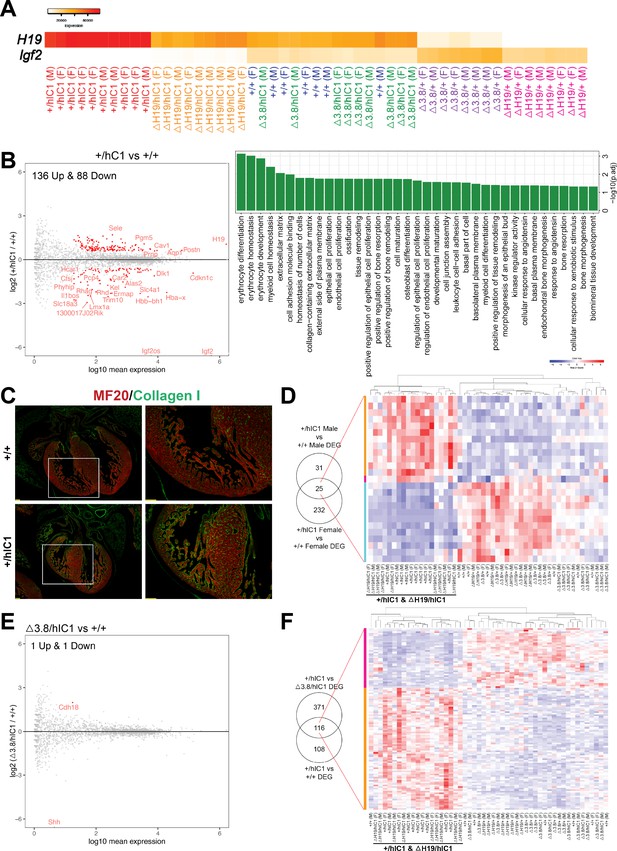
Transcriptomic analysis of E12.5 cardiac endothelial cells from wild-type and mutant embryos.
(A) A gradient H19 and Igf2 expression levels are depicted in E12.5 wild-type, +/hIC1, △H19/+, △H19/hIC1, △3.8/+, and △3.8/hIC1 cardiac endothelial cells. M: male and F: female. (B) Left: Comparison between +/hIC1 and the wild-type samples with a volcano plot shows 224 differentially expressed genes (DEGs) between +/hIC1 and the wild-type samples. Right: Gene ontology (GO) pathways that are enriched for the 224 DEGs. (C) Immunofluorescence staining for MF20 (red) and collagen (green) on E17.5 wild-type and +/hIC1 hearts. Images on the right are enlarged from the boxed area of images on the left. Scale bars = 100 µm. (D) Expression pattern of 25 genes that are differentially expressed in both male and female +/hIC1 samples is compared to wild-type. (E) A volcano plot represents two DEGs between △3.8/hIC1 and the wild-type samples. (F) Expression pattern of 116 genes that are commonly differentially expressed in +/hIC1 compared to the wild-type and △3.8/hIC1 samples.
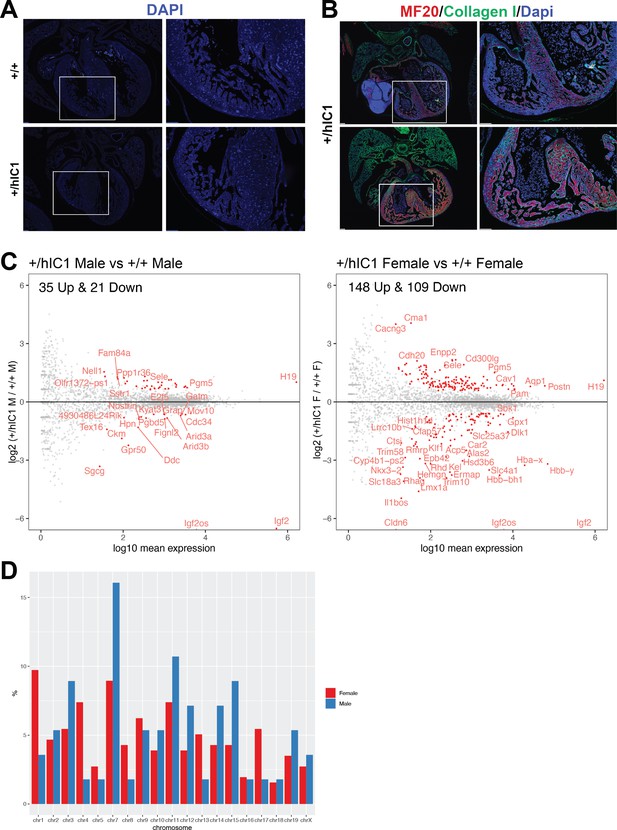
Supplementary data for differential gene expression of +/hIC1 hearts.
(A) DAPI (4′,6-diamidino-2-phenylindole; blue) on E17.5 wild-type and +/hIC1 hearts from Figure 6C. Images on the right are enlarged from the boxed area of images on the left. Scale bars = 100 µm. (B) Immunofluorescence staining for MF20 (red), collagen I (green), and DAPI (blue) on E17.5 +/hIC1 hearts. Images on the right are enlarged from the boxed area of images on the left. Scale bars = 100 µm. (C) Same-sex comparisons between +/hIC1 and the wild-type samples analyzed by RNA sequencing. (D) Chromosomal distribution of differentially expressed genes (DEGs) from comparisons between +/hIC1 and wild-type males and females.
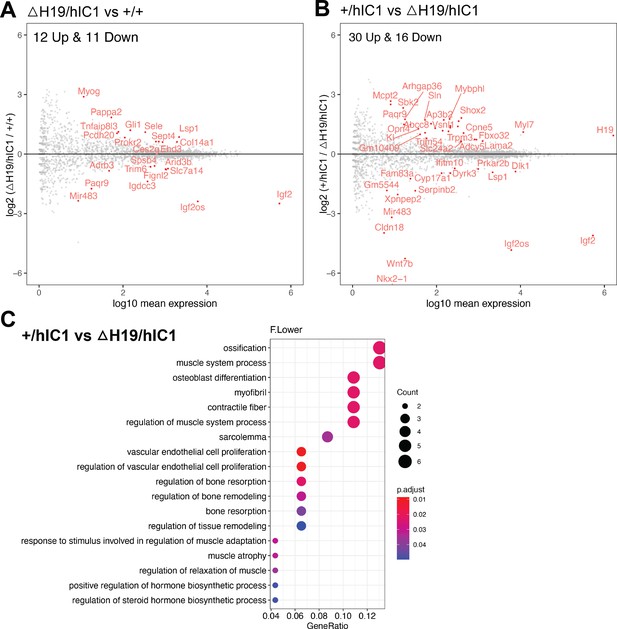
Further RNA-seq analysis involving ΔH19/hIC1 endothelial cells.
(A) Volcano plot depicting comparison of ΔH19/hIC1 and wild-type cardiac endothelial cells. (B) Volcano plot depicting comparison of +/hIC1 and ΔH19/hIC1 cardiac endothelial cells. (C) Gene ontology (GO) pathways that are enriched for 46 differentially expressed genes (DEGs) between +/hIC1 and ΔH19/hIC1 samples.
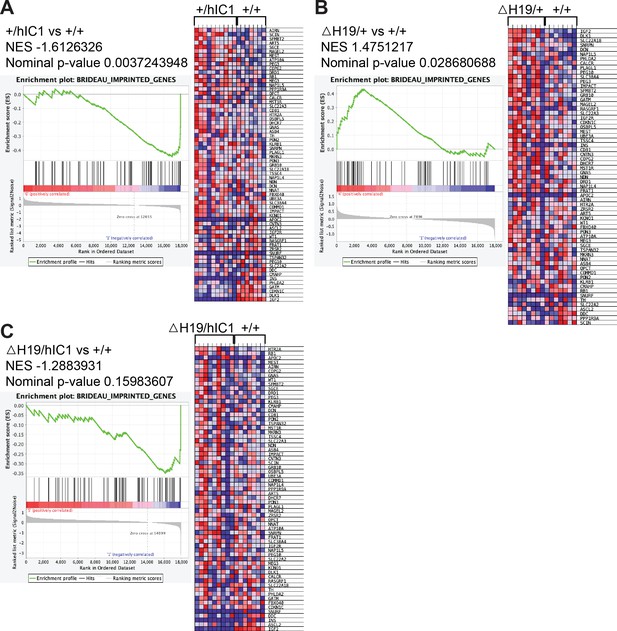
Enrichment tests for BRIDEAU_IMPRINTED_GENES using gene set enrichment analysis (GSEA).
Normalized enrichment score (NES), nominal p-value, enrichment plot, and expression heatmap of the analyzed gene set comparing (A) +/hIC1 and wild-type, (B) ΔH19/+ and wild-type, (C) ΔH19/hIC1 and wild-type cardiac endothelial cells.
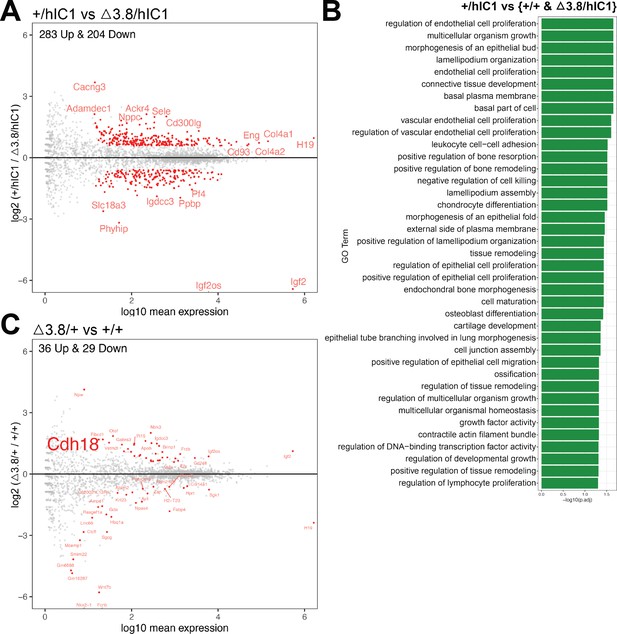
Further RNA-seq analysis utilizing Δ3.8/+ and Δ3.8/hIC1 endothelial cells.
(A) Volcano plot depicting comparison of +/hIC1 and Δ3.8/hIC1 cardiac endothelial cells. (B) Gene ontology (GO) pathways that are enriched for 116 differentially expressed genes (DEGs) that are commonly differentially expressed in +/hIC1 compared to wild-type and Δ3.8/hIC1 samples. (C) Volcano plot depicting comparison of Δ3.8/+ and wild-type cardiac endothelial cells.

Enrichment tests for HALLMARK_HYPOXIA, GROSS_HYPOXIA_VIA_HIF1A_DN using gene set enrichment analysis (GSEA).
Normalized enrichment score (NES), nominal p-value, enrichment plot, and expression heatmap of the (A) HALLMARK_HYPOXIA, (B) GROSS_HYPOXIA_VIA_HIF1A_DN gene set comparing +/hIC1 and wild-type cardiac endothelial cells.

Tetraploid aggregation experiment.
(A) Schematic description of tetraploid aggregation experiment. (B) Level of hIC1 and GFP in embryonic and placental gDNA from hIC1 aggregated samples. (C) Embryonic and placental weight of E17.5 aggregated samples. (D) Representative cross-sections of E17.5 4N wild-type placentas with +/hIC1 embryos stained with hematoxylin and eosin. Black arrowhead indicates a large thrombus formed in the +/hIC1 labyrinth. (E) Representative cross-sections of E17.5 4N wild-type placentas with +/hIC1 embryos stained with hematoxylin and eosin. (F) (G) (D, E) Scale bars = 1mm. (B, C, G) Each data point represents an individual conceptus from different aggregated litters. Statistics used are (B, C, G) multiple unpaired t-test. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, ns = not significant.
Additional files
-
Supplementary file 1
Primers and PCR conditions utilized in this study.
- https://cdn.elifesciences.org/articles/78754/elife-78754-supp1-v1.docx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/78754/elife-78754-mdarchecklist1-v1.docx





