Integrating analog and digital modes of gene expression at Arabidopsis FLC
Figures
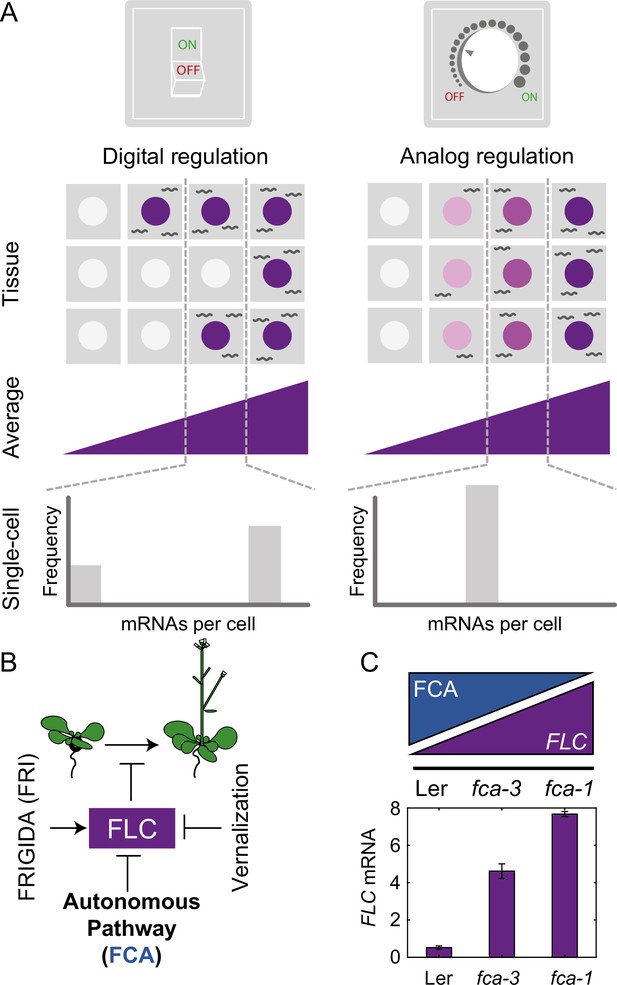
Schematic of digital and analog gene regulation.
(A) Digital regulation (left) corresponds to loci being in an ‘ON’ state (purple) or ‘OFF’ state (white), where we assume for simplicity that there is only one gene copy per cell. At the tissue level, moving from low to high average expression (columns left to right) is achieved by a change in the fraction of cells in each of the two states. This mode is distinct from analog regulation (right), where each cell has a graded expression level that roughly corresponds to the overall population average. (B) FLC represses the transition to flowering and is controlled by FRIGIDA, the autonomous pathway and the vernalization pathway (inducing digital epigenetic silencing in the cold). (C) Reduced FCA activity leads to higher cell population-level FLC expression in fca mutants. Wildtype Ler has lowest FLC expression, fca-3 intermediate and fca-1 highest. Expression is measured by qPCR relative to the house-keeping gene index (geometric mean of PP2A and UBC). Error bars show SEM of n = 3 biological replicates measured 7 d after sowing. Statistical tests: multiple comparisons following ANOVA (F-value = 214.62, p-value=2.6 · 10-6) with Tukey HSD post hoc tests for fca-3 – fca-1: p-value=0.00029; fca-3 – Ler: p-value=5.5 · 10-5; fca-1 – Ler: p-value<2.6 · 10-6 .

Characterization of fca alleles and FLC-Venus transgene.
(A) Flowering time measured in days from sowing until first appearance of a floral meristem (bolting). 12 plants were measured for each genotype. Boxplots show median and interquartile range (IQR), with data points outside median ±1.5 IQR shown as crosses. (B) Diagram showing positions of qPCR primers (red) to quantify spliced and unspliced endogenous FLC and transgenic FLC-Venus RNA. (C, D) FLC and FLC-Venus RNA measured by qPCR from whole seedlings, relative to UBC RNA. No RT (reverse transcription)-negative control in (D) shows no expression with same unspliced primers. Error bars show SEM of n = 2 biological replicates for fca-3 and n = 3 biological replicates for other genotypes, harvested 14 d after sowing.
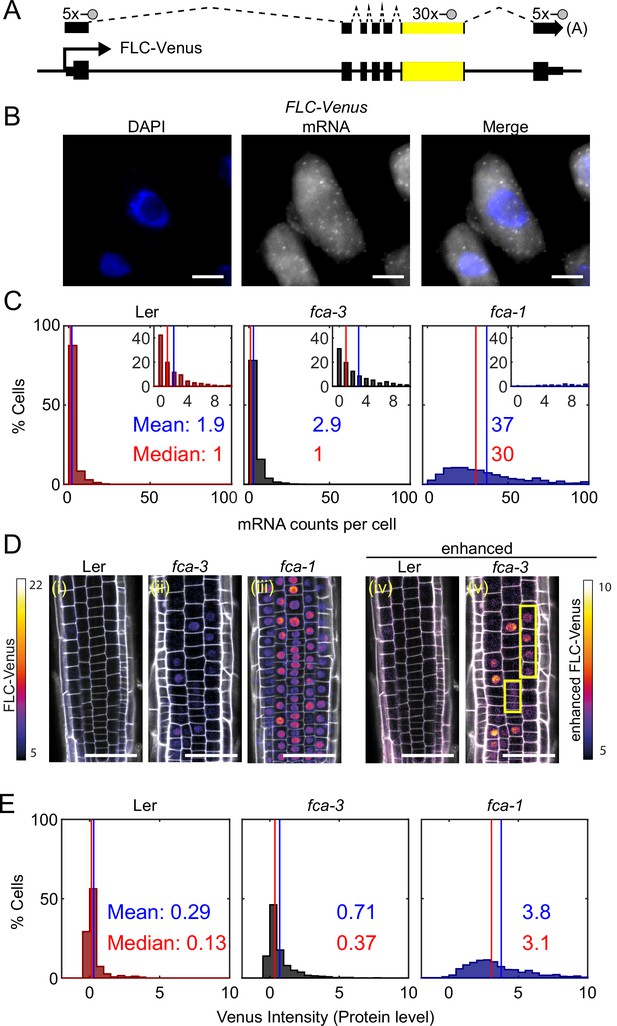
FLC expression per cell in fca mutants.
(A) Schematic diagram of FLC-Venus locus with transcript and exonic probe position indicated. A total of 40 probes were designed, 10 against the FLC sequence and 30 for the Venus sequence. (B) Detection of FLC-Venus transcripts in single cells. Representative images of isolated cells with DAPI staining (blue) and FLC-Venus mRNA (gray) obtained from Arabidopsis thaliana root squashes. Scale bar, 5 μm. (C) Histograms of single-molecule fluorescence in situ hybridization (smFISH) results for each genotype (Ler, fca-3, fca-1) showing the number of single-molecule FLC-Venus RNAs detected per cell. We show the distribution for Ler, but note that this is likely indistinguishable from background using our methods. The means and medians of the distributions are indicated in each panel. Insets show the same data for mRNA counts between 0 and 10 (where 10 is the 95th percentile of the fca-3 data). For Ler n = 853, fca-3 n = 1088, fca-1 n = 792; data from 7 d after sowing (two independent experiments). Statistical tests: three-way comparison with Kruskal–Wallis (, p-value=0) and pairwise comparisons with Wilcoxon rank sum tests with Bonferroni-adjusted p-values for fca-3 – fca-1: adj. p-valu =5.5 · 10-275; fca-3 – Ler: adj. p-value 1.2 · 10-9; fca-1 – Ler: adj. p-value=2.9 · 10-257. (D) Representative confocal images of roots for each genotype. FLC-Venus intensity indicated by color maps; gray shows the propidium iodide (PI) channel. Same settings were used for imaging and image presentation in (i–iii). Images in (iv) and (v) are the same as (i) and (ii), respectively, but adjusted to enhance the Venus signal by changing brightness and contrast (please note different scale of color map). Yellow boxes in (v) show short files of ON cells. Scale bar, 50 μm. (E) Histograms of FLC-Venus intensity per cell in each genotype. The means and medians of the distributions are indicated in each panel. For Ler n = 537 cells from 6 roots, fca-3 n = 1515 cells from 14 roots, fca-1 n = 1031 cells from 11 roots; data from 7 d after sowing (two independent experiments). Statistical tests: three-way comparison with Kruskal–Wallis (, p-value=0) and pairwise comparisons with Wilcoxon rank sum tests with Bonferroni-adjusted p-values for fca-3 – fca-1: adj. p-value=8.7 · 10-281; fca-3 – Ler: adj. p-value=1.7 · 10-32; fca-1 – Ler: adj. p-value=1.2 · 10-201.
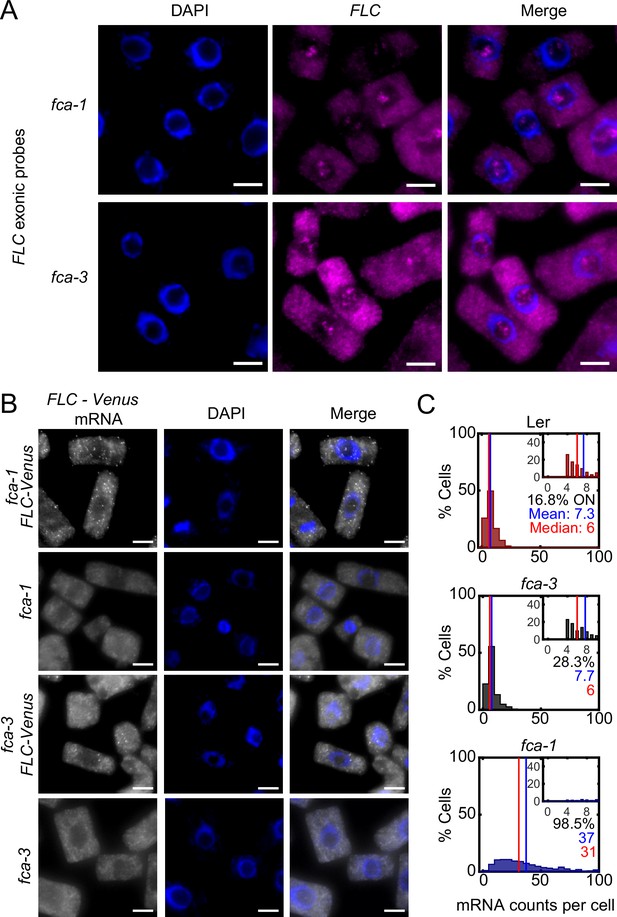
Single-molecule fluorescence in situ hybridization (smFISH) method for FLC-Venus imaging.
(A) Detection of nucleolar signal using FLC-specific probes (magenta) in Ler background fca-1 and fca-3 plants in Arabidopsis thaliana root cells (monolayer of cells derived from root squashes). The nucleus is stained with DAPI (blue). Scale bar, 5 μm. (B) FLC-Venus probes (gray) bind specifically to the transgenic FLC-Venus RNA and not to the endogenous FLC transcript. All data from seedlings 7 d after sowing. Scale bar, 5 μm. (C) The data of Figure 2C plotted for the three genotypes but showing only measurements in ‘ON cells.’ These are defined as cells with more than three FLC-Venus mRNA. The means and medians of the distributions are indicated in each panel, as well as the percentage of ON cells. For Ler n = 143, fca-3 n = 308, fca-1 n = 780, from 7 d post sowing. Statistical tests: three-way comparison with Kruskal–Wallis (, p-value=5.9 · 10-147) and pairwise comparisons with Wilcoxon rank sum tests with Bonferroni-adjusted p-values for fca-3 – fca-1: p-value=1.4 · 10-113; fca-3 – Ler: p-value=1; fca-1 – Ler: p-value=1.7 · 10-64. We note that the ‘ON cells’ in Ler come primarily from background signal.
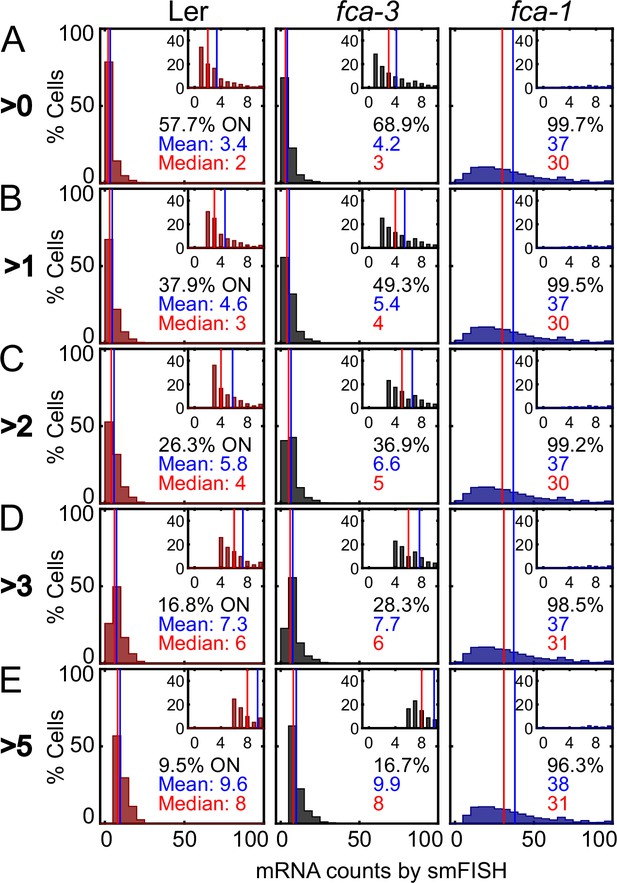
Threshold for ON/OFF state of cells in single-molecule fluorescence in situ hybridization (smFISH) experiments.
Figure 2—figure supplement 1C replotted for different thresholds 0 (A), >1 (B), >2 (C), >3 (D) and >5 (E) mRNAs. Red number indicates the median of the ON cells, while the blue number indicates the mean, and the black number is the percentage of cells that satisfy the threshold condition. We note that the ‘ON cells’ in Ler come primarily from background signal and the consistently lower percentage of such cells compared to fca-3 reflects this.
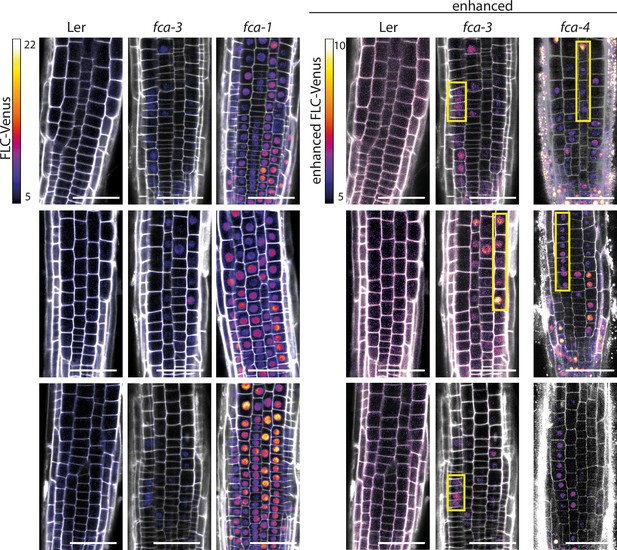
FLC-Venus imaging in fca alleles – root replicates and fca-4.
Representative images of FLC-Venus signal in four genotypes (Ler, fca-3, fca-1, fca-4) from 7 d post sowing. FLC-Venus intensity indicated by color maps; gray shows the propidium iodide (PI) channel. Same settings were used for imaging and image presentation in left three columns. ‘Enhanced’ images (three columns on right) are the same respective images for Ler and fca-3, adjusted to enhance FLC-Venus signal by changing brightness and contrast (please note different scale of color map). fca-4 images were acquired on a different microscope and were enhanced to a comparable level as in Ler and fca-3 (enhanced). Yellow boxes show files of ON cells in fca-3 and fca-4. Scale bar, 50 μm.

FLC-Venus imaging in fca mutants and wildtype, in young leaf tissue.
(A) Three representative images of FLC-Venus signal in the three genotypes (Ler, fca-3, fca-1) from 9 d post sowing. FLC-Venus intensity indicated by color map; gray shows the cell wall dye (SR2200). Scale bar, 20 μm. (B) Histograms of FLC-Venus intensity per cell in each genotype. The cell numbers (n), means, and medians of the distributions are indicated in each panel. Data from 9 d after sowing (two experiments). Statistical tests: three-way comparison with Kruskal–Wallis (X2(2) = 292.27, p-value=3.4 · 10-64) and pairwise comparisons with Wilcoxon rank sum tests with Bonferroni-adjusted p-values for fca-3 – fca-1: adj. p-value=2.7 · 10-42; fca-3 – Ler: adj. p-value=3.2 · 10-22; fca-1 – Ler: adj. p-value=1.4 · 10-40.
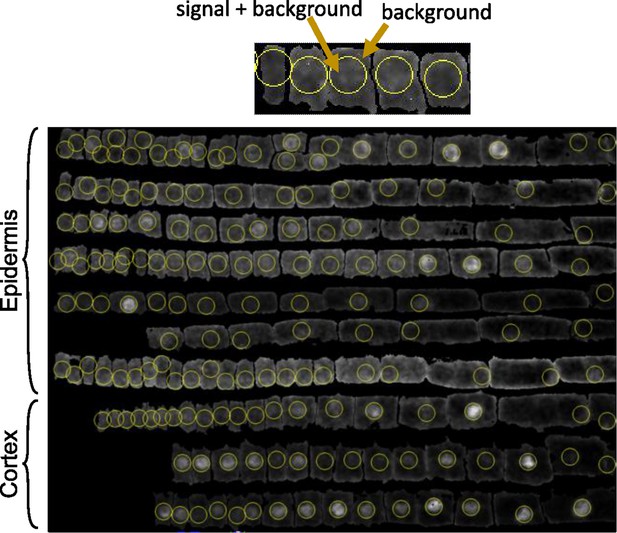
Segmentation and quantification method for FLC-Venus protein intensity per nucleus.
Custom visualization and segmentation software was used to manually correct automatic segmentations and annotate cells according to cell file. Top: diagram explaining quantification of FLC-Venus intensity. In ON cells, the sphere contains the nucleus and therefore the FLC-Venus signal as well as the background signal. In OFF cells, the sphere will contain only background signal. The mean intensity outside the sphere was used as the background signal, and this was subtracted from the sphere intensity. Bottom: cell files from a single root separated according to tissue type demonstrating the result of the nucleus finding algorithm. Longitudinal section shown. In each cell, the sphere of constant radius that maximized fluorescent intensity inside the sphere was determined. For each cell, the position of the sphere is shown, corresponding to the nucleus in ON cells.
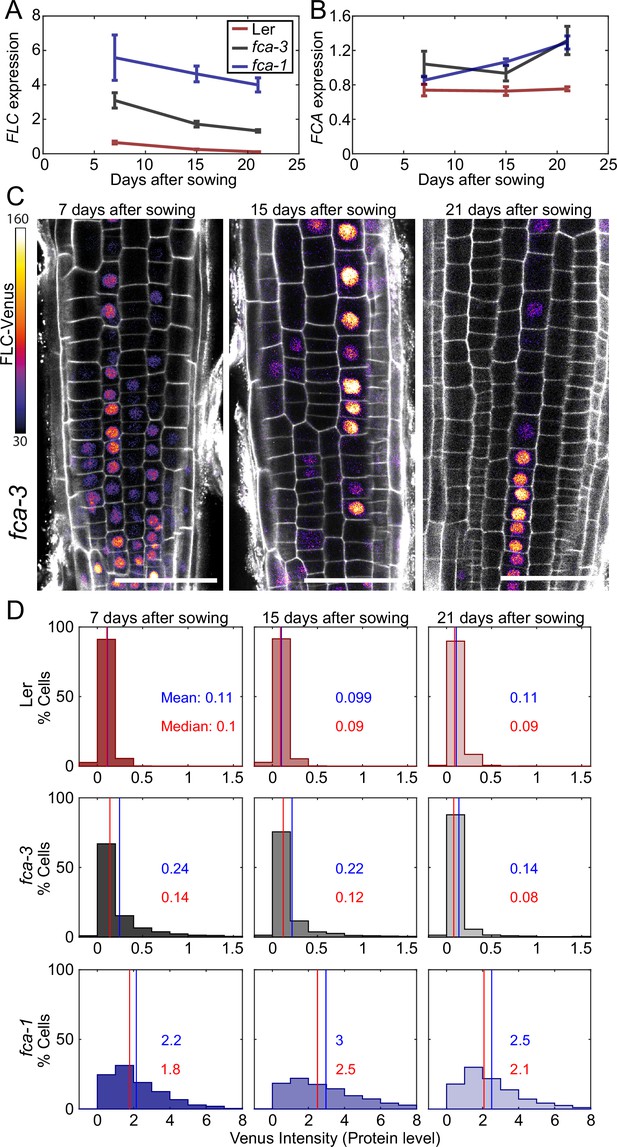
Experimental observation of gradual FLC silencing.
(A) Timeseries of FLC expression in fca mutant alleles transformed with the FLC-Venus construct. Expression is measured by qPCR in whole seedlings relative to the house-keeping gene index (geometric mean of PP2A and UBC). Error bars show SEM of n = 3 biological replicates. Statistical tests: samples were excluded as outliers based on Grubbs’ test with alpha = 0.05. Linear regression on timeseries for each genotype. Slope for fca-3 = −0.13, p-value=; slope for fca-1 = −0.11, p-value=0.12; slope for Ler = −0.038, p-value=. (B) Timeseries of FCA expression, otherwise as in (A). Statistical tests: samples were excluded as outliers based on Grubbs’ test with alpha = 0.05. Linear regression on timeseries for each genotype. Slope for fca-3 = 0.021, p-value=0.22; slope for fca-1 = 0.032, p-value=; slope for Ler = , p-value=0.84. (C) Representative images of fca-3 roots by confocal microscopy. FLC-Venus intensity indicated by color map; gray shows the PI channel. Same settings were used for imaging and image presentation. Scale bar, 50 μm. (D) Histograms of FLC-Venus intensity per cell at each timepoint. The means and medians of the distributions are indicated in each panel. For Ler: 7 d, n = 1121 cells from 10 roots (three independent experiments); 15 d, n = 1311 cells from 9 roots (two independent experiments); 21 d, n = 1679 cells from 12 roots (three independent experiments). For fca-3: 7 d, n = 2875 cells from 24 roots (three independent experiments); 15 d, n = 3553 cells from 23 roots (three independent experiments); 21 d, n = 3663 cells from 21 roots (three independent experiments). For fca-1: 7 d, n = 1022 cells from 9 roots (three independent experiments); 15 d, n = 1770 cells from 12 roots (three independent experiments); 21 d, n = 2124 cells from 12 roots (three independent experiments). Statistical tests: linear regression on timeseries for each genotype. Slope for fca-3 = −0.0077, p-value=; slope for fca-1 = 0.018, p-value=0.00064; slope for Ler = -0.00015, p-value=0.44.

Degradation rate of FLC mRNA and protein levels.
(A) Histogram of FLC-Venus mRNAs per cell measured by single-molecule fluorescence in situ hybridization (smFISH) after 4 hr or 6 hr of ActD treatment (top) or mock DMSO treatment (bottom). Intronic PP2A probes were used to confirm the effect of ActD on transcription. Mean mRNA number per cell, µ, is indicated in each panel. Half-life t1/2 of 5 hr calculated as in Ietswaart et al., 2017 using the formula , with , where is the degradation rate and are the mean RNA counts after 4 and 6 hr, respectively. For 0 hr, n = 1554; for ActD 4 hr, n = 820; for ActD 6 hr, n = 784; for DMSO 4 hr, n = 759; for DMSO 6 hr, n = 776, in seedlings 7 d after sowing. Three independent experiments were performed and combined. (B) Analysis of FLC-Venus protein stability with cycloheximide (CHX) treatment. 7-day-old seedlings of fca-1 plants carrying FLC-Venus reporter were subjected to liquid MS solution supplemented with 100 μM CHX for 0, 1.5, 3, 6, 12, or 24 hr. Blot shows representative replicate. Nontreatment control was also included (right-most well). Arrow indicates FLC-Venus. Ponceau staining was used as processing controls. (C) Quantification of the FLC-Venus from blot shown in (B) and additional replicates. Combined data from n = 3 biological replicates are shown with exponential decay profile fitted to the first three timepoints (half-life = 1.45 hr).

FLC and FCA expression in whole seedlings over time .
(A) Timeseries of FLC expression in fca mutant alleles. Expression is measured by qPCR relative to the house-keeping gene index (geometric mean of PP2A and UBC). Error bars show SEM of n = 3 biological replicates. FLC 7d timepoint is repeated from Figure 1C. Statistical tests: linear regression on timeseries for each genotype. Slope for fca-3 = −0.15, p-value=0.0063; slope for fca-1 = −0.091, p-value=0.0054; slope for Ler = −0.023, p-value=0.0030. (B) Timeseries of FCA expression, otherwise as in (A). Statistical tests: linear regression on timeseries for each genotype. Slope for fca-3 = −0.028, p-value=0.043; slope for fca-1 = 0.043, p-value=0.031; slope for Ler = −0.00445, p-value=0.56.
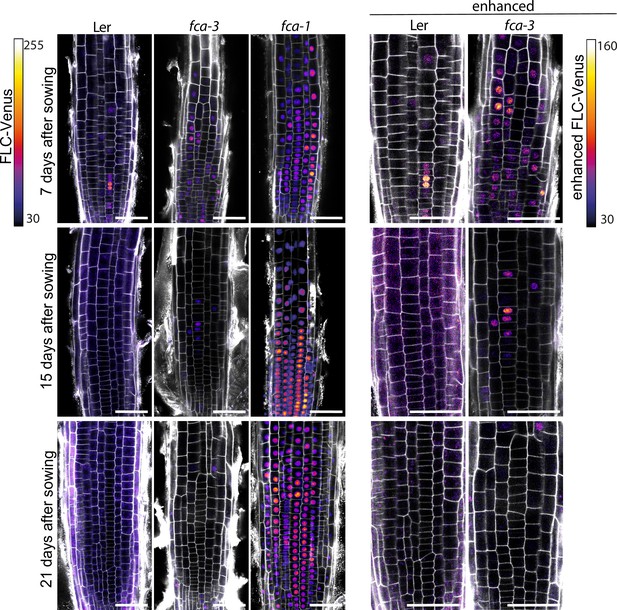
FLC-Venus time-course replicates in fca alleles.
Additional representative images of FLC-Venus signal in the three genotypes (Ler, fca-3, fca-1) at the three timepoints. FLC-Venus intensity is indicated by color maps; gray shows the PI channel. Same settings were used for imaging and image presentation in left three columns. ‘Enhanced’ images (two columns on right) are a zoomed-in area of the same respective images, adjusted to enhance FLC-Venus signal to similar level as fca-1 by changing brightness and contrast (please note different scale of color map). Scale bar, 50 μm.
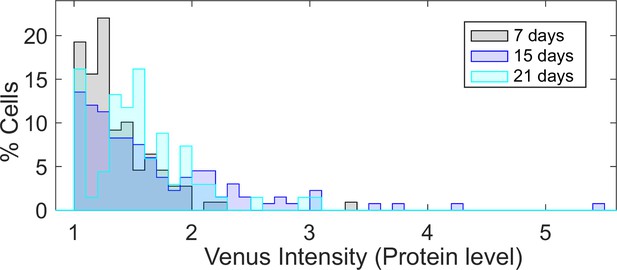
Experimental intensity of FLC-Venus in ON cells does not change over time in fca-3.
Histogram of ON cell tail in fca-3 at three timepoints normalized by the total number of cells in each. Only cells with FLC-Venus intensity above 1 are shown. For 7 d n = 109 cells from 17 roots, 15 d n = 133 cells from 14 roots, 21 d n = 68 cells from 6 roots. Data combined from three independent experiments.
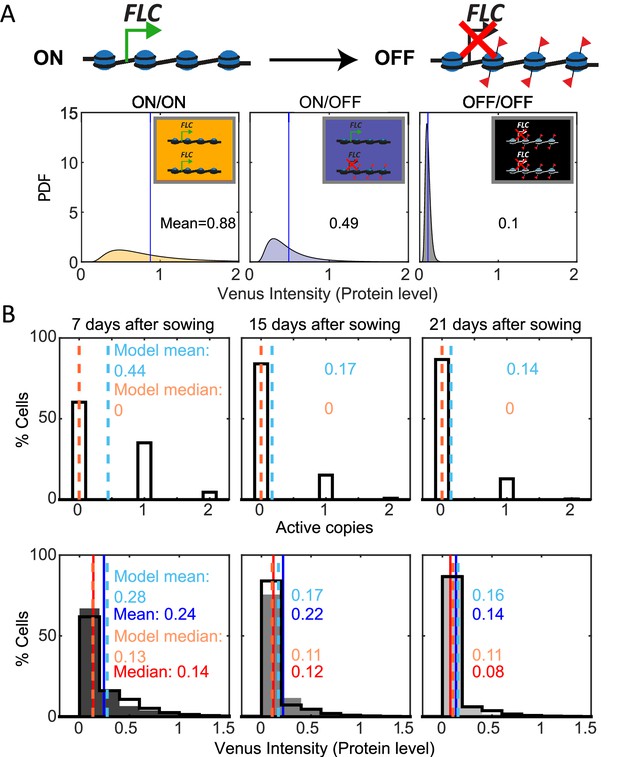
Mathematical model captures FLC regulation.
(A) Diagram of mathematical model. Individual FLC gene copies can be ON or OFF, such that a cell can be in one of three states depending on the combination of ON and OFF gene copies within it (ON/ON, ON/OFF, OFF/OFF). Venus intensity (corresponding to amount of protein) within a cell was sampled from the distributions shown (described in ‘Materials and methods’ section), depending on the cell state. The means of the distributions are indicated in each panel. (B) Histograms of active FLC copies per cell (top) and Venus intensity per cell (bottom) at the indicated timepoints. Model histograms are plotted with black lines around empty bars and experimental data is shown as filled gray histograms with no outline. The means and medians of the distributions are indicated in each panel for the data (solid lines and matching color text) and the model (dashed lines and matching color text). Model simulated 1000 fca-3 cell files and histograms shown exclude bottom four cells of each file as cells near the QC are also not included in the imaging.
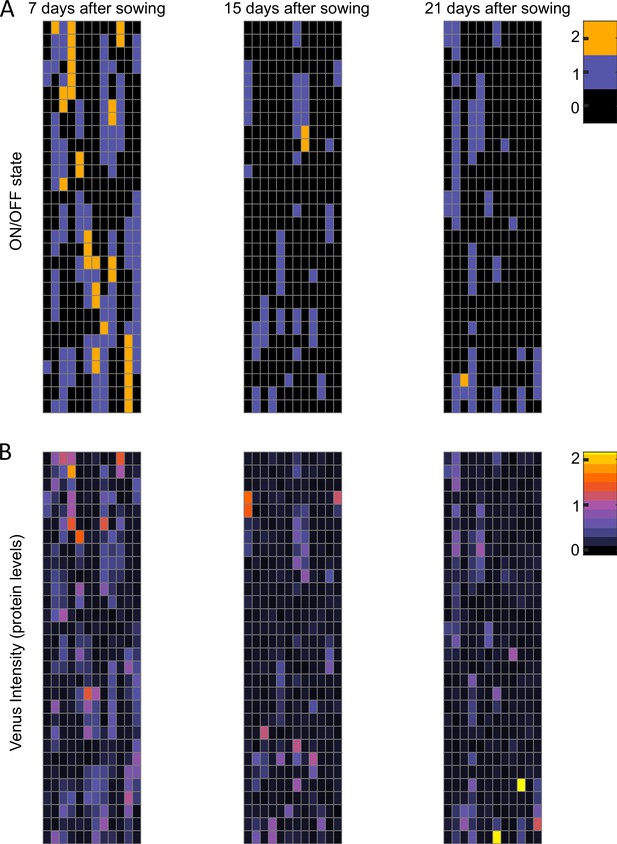
Modeling FLC regulation in clonal cell files in the root.
Representative model output for fca-3 root, showing 12 simulated cell files of length 30 cells each, at different timepoints, matching sampling times. (A) The state of the loci (both copies ON – orange, one copy ON – purple, both copies OFF – black). (B) FLC-Venus intensity (proxy for protein concentration) per cell, for the same cells.
Additional files
-
Supplementary file 1
Mathematical model parameters.
- https://cdn.elifesciences.org/articles/79743/elife-79743-supp1-v1.xlsx
-
Supplementary file 2
Primers used in this study.
- https://cdn.elifesciences.org/articles/79743/elife-79743-supp2-v1.xlsx
-
Supplementary file 3
smFISH probe sequences used to detect FLC Venus transcripts.
These probes were labeled with Quasar570.
- https://cdn.elifesciences.org/articles/79743/elife-79743-supp3-v1.xlsx
-
Supplementary file 4
smFISH probe sequences from Duncan et al., 2016 (A) used to detect unspliced PP2A transcripts and (B) used to detect FLC sense spliced.
Both probe sets were labeled with Quasar670.
- https://cdn.elifesciences.org/articles/79743/elife-79743-supp4-v1.xlsx
-
Supplementary file 5
Raw data from this study.
- https://cdn.elifesciences.org/articles/79743/elife-79743-supp5-v1.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/79743/elife-79743-mdarchecklist1-v1.pdf





