Wolbachia action in the sperm produces developmentally deferred chromosome segregation defects during the Drosophila mid-blastula transition
Figures
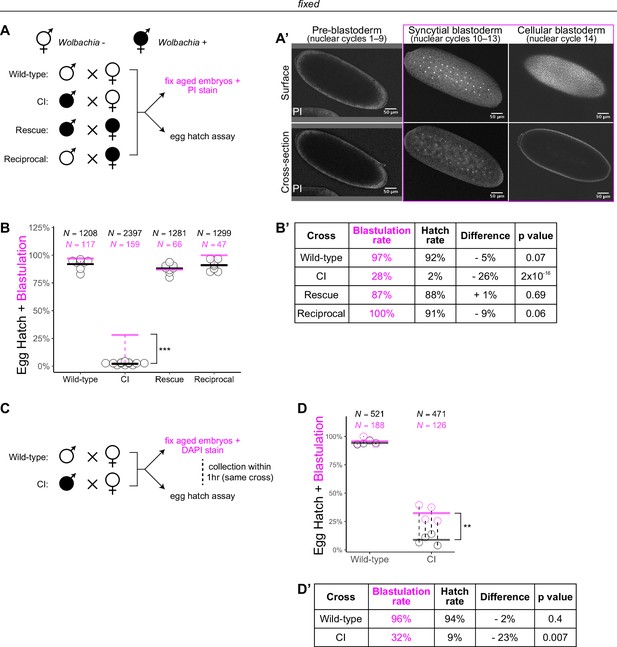
Wolbachia induces both early and late embryonic lethality.
(A) Wolbachia infection status is indicated by filled circles. Embryos were collected from each of the four crosses and either used for egg hatch assays or aged prior to fixing and staining DNA with propidium iodide (PI). (A’) Confocal imaging of PI-stained embryos allowed categorization of embryo stage as pre-blastoderm (cycles 2–9), syncytial blastoderm (cycles 10–13), or cellular blastoderms (cycle 14). Scale bars are 50 μm. *** = p = 2 x 10-16 χ-squared test. (B, B’) Comparison between blastulation rate (% of fixed embryos staged as progressing beyond cycle 9) and egg hatch rate between each of the four crosses. Each circle represents one biological replicate of an egg hatch assay. Black and magenta lines represent the average egg hatch rate and the blastulation rate, respectively. N values correspond to the total number of embryos scored. While the hatch rate from wild-type, rescue, and reciprocal crosses closely corresponded to the blastulation rate, the hatch rate from cytoplasmic incompatibility (CI) embryos was statistically significantly decreased compared to the blastulation rate. Displayed p values were determined by χ-squared tests. (C) Embryos were collected from wild-type and CI crosses and were used to either determine embryo stage (by 4',6-diamidino-2-phenylindole (DAPI) staining) or egg hatch percentage in paired assays (collections were from the same crosses within 1 hr of each other). (D, D’) Comparison between blastulation rate (% of fixed embryos staged as progressing beyond cycle 9) and egg hatch for each cross. Each circle represents a technical replicate of one egg hatch assay and one staging experiment. Dashed lines connect paired experiments. Black and magenta lines represent the average egg hatch rate and the blastulation rate, respectively. N values correspond to the total number of embryos scored. The difference between blastulation rate and hatch rate was statistically significant by a two-sided paired t-test. ** = p = 0.007. (D’). See also Figure 1—figure supplement 1.
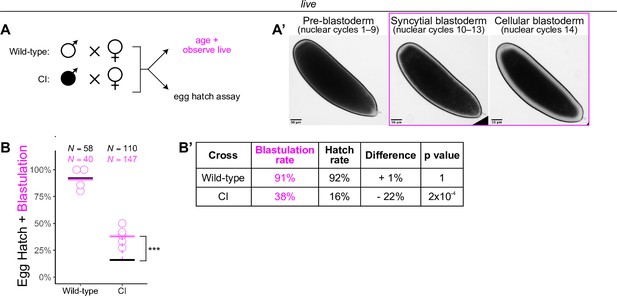
Wolbachia induces late embryonic lethality.
(A) Embryos were collected from wild-type and cytoplasmic incompatibility (CI) crosses and either used for egg hatch assays or aged and observed live. (A’) Live observation of dechorionated embryos under a high-power dissecting scope enabled categorization of embryo stage as pre-blastoderm, syncytial blastoderm, or cellular blastoderms. Scale bars are 50 μm. (B, B’) Comparison between blastulation rate and egg hatch rate between wild-type and CI crosses. Each circle represents one round of live categorization (biological replicates). Black and magenta lines represent the egg hatch rate and the average blastulation rate, respectively. N values correspond to the total number of embryos scored. Displayed p values were determined by χ-squared tests (*** = 2 x 10-4). See also Figure 1.
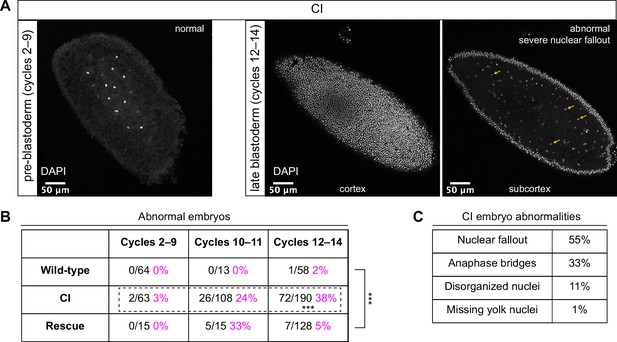
Cytoplasmic incompatibility (CI)-derived embryos proceed normally through pre-blastoderm divisions and then exhibit cellular defects during blastoderm divisions.
(A) Examples of fixed and DAPI-stained CI-derived embryos from pre-blastoderm (cycles 2–9) and late blastoderm (12–14) stages. While the CI-derived pre-blastoderm appears normal, the late blastoderm exhibits severe nuclear fallout (nuclei receded from the cortex and into subcortical regions). Arrows point to several examples of fallen out nuclei. Scale bars are 50 μm. (B) Comparison of the percentage of abnormal embryos from wild-type, CI, and rescue crosses during different stages of embryogenesis. While CI-derived embryos developed normally through cycles 2–9, they exhibited significantly increased abnormalities during cycles 10–14 and 12-14 (*** = p = 1.9 x 10-8 by two-sided Fisher's exact test). Abnormalities in cycles 12–14 were significantly reduced in embryos from rescue crosses (*** = p = 3.8 x 10-16 by two-sided Fisher's exact test). N values correspond to the number of embryos scored (biological replicates). (C) Classification of abnormalities observed in CI-derived embryos.
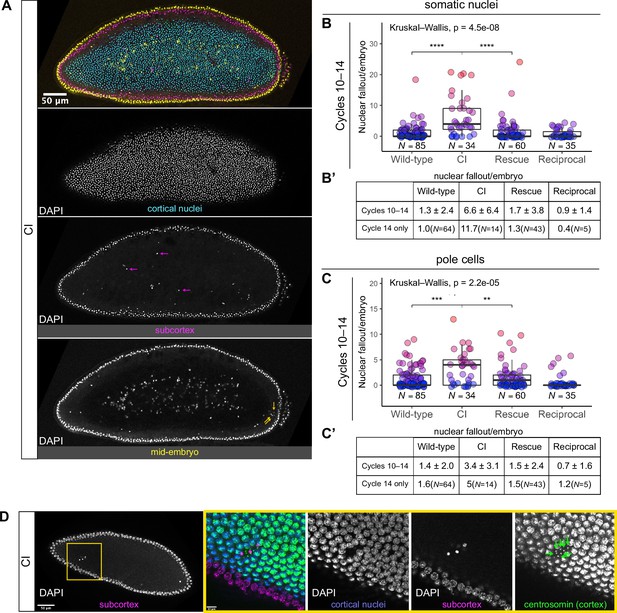
Developed cytoplasmic incompatibility (CI)-derived embryos exhibit increased rates of nuclear fallout.
(A) Image of a CI-derived blastoderm exhibiting moderate nuclear fallout. Cortical nuclei (cyan) are on the surface of the embryo. Nuclei that have fallen out of the cortex can clearly be observed 5–10 μm beneath the cortex (subcortex, magenta) and at the mid-plane of the embryo (yellow). Magenta arrows point to examples of somatic nuclei that have fallen out. Yellow arrows point to examples of pole cells that have fallen out. Scale bar is 50 μm (B, B’) Comparison of somatic nuclear fallout in cycle 10–14 embryos from wild-type, CI, rescue, and reciprocal crosses. (B) Each dot represents the number of fallen nuclei per embryo. Asterisks indicate significance by Mann–Whitney tests (Wild-type to CI: p = 4.5 x 10-8; CI to rescue: p = 1.7 x 10-6). (B’) Averages and standard deviations are summarized. CI-derived embryos have significantly increased somatic nuclear fallout compared to wild-type- and rescue-derived embryos. N values correspond to the number of embryos scored (biological replicates). (C, C’) Comparison of pole cell nuclear fallout in cycle 10–14 embryos from wild-type, CI, rescue, and reciprocal crosses. (C) Each dot represents the number of fallen nuclei per embryo. Asterisks indicate significance by Mann–Whitney tests (Wild-type to CI: p = 6.8 x 10-4; CI to Rescue: p = 2.5 x 10-3). (C’) Averages and standard deviations are summarized. CI-derived embryos have significantly increased somatic nuclear fallout compared to wild-type- and rescue-derived embryos. N values correspond to the number of embryos scored (biological replicates). (D) CI-derived embryo stained with anti-centrosomin antibody to mark centrosomes and counterstained with DAPI. While cortical nuclei (blue) remain strongly associated with their centrosomes (green), nuclei that recede into the subcortex (magenta) detach from their centrosomes that are left at the cortex (green arrows). Yellow box indicates zoomed region. Scales bars are 50 and 10 μm for unzoomed and zoomed regions, respectively.
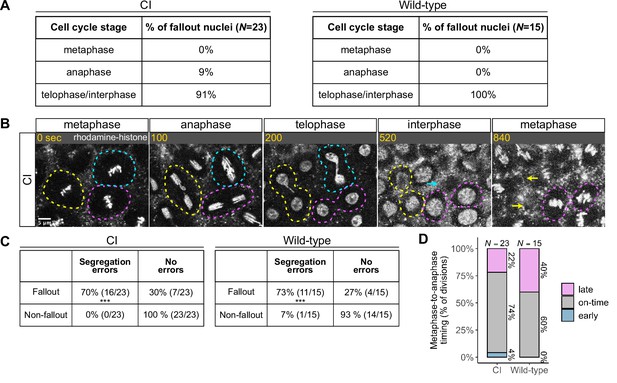
Chromosome segregation errors are the proximate cause of nuclear fallout in cytoplasmic incompatibility (CI)-derived embryos.
(A) Comparison of when in the cell cycle nuclei fallout in both CI- and wild-type-derived embryos. (B) Nuclei that fallout (yellow and blue circles) exhibit severely lagging chromosomes in the previous division, while nuclei that remain at the cortex (magenta circle) exhibit normal chromosome segregation. Scale bar is 5 μm, and time is written in seconds. See also Figure 4—video 1. (C) Chromosome segregation errors were significantly more frequent for nuclei destined to fallout compared to neighboring nuclei (non-fallout) in both CI- (*** = p = 3.4 × 10–6 by χ-squared test) and wild-type-derived embryos (*** = p = 8.0 × 10–4 by χ-squared test). (D) Comparison of metaphase-to-anaphase timing between nuclei destined to fallout and their neighbors that remain at the cortex in both CI- and wild-type-derived embryos. ‘Early’ = fallout nuclei enter anaphase before their neighbors. ‘On-time’ = fallout and neighboring nuclei enter anaphase simultaneously. ‘Late’ = fallout nuclei enter anaphase after their neighbors. N values correspond to the number of cells scored (biological replicates). See also Figure 4—figure supplement 1.
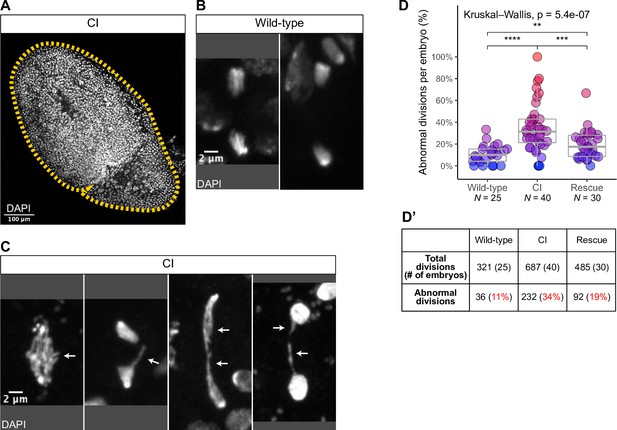
Cytoplasmic incompatibility (CI)-derived embryos exhibit increased rates of chromosome segregation errors and nuclear fallout.
(A) A whole CI-derived gastrulating embryo is outlined. Scale bars are 100 μm. Examples of divisions observed in wild-type- (B) and CI- (C) derived gastrulating embryoss. While divisions from wild-type-derived embryos are normal, divisions from CI-derived embryos exhibit a variety of lagging/bridging chromosome segregation errors (arrows). Scale bars are 2 μm. (D, D’) Comparison of chromosome segregation errors in wild-type-, CI-, and rescue-derived gastrulating embryos. Each dot represents one embryo (D), and a summation is presented in (D’). **** = p = 7.7 x 10-7; *** = p = 8.6 x 10-4; and ** = p = 0.009 by Mann—Whitney tests. N values correspond to the number of embryos scored (biological replicates). See also Figure 4.
Chromosome segregation immediately precedes nuclear fallout of cortical blastoderm nuclei.
A cytoplasmic incompatibility (CI)-derived embryo injected with rhodamine-labeled histone. Scale bar is 5 μm and time is in seconds. See also Figure 4.
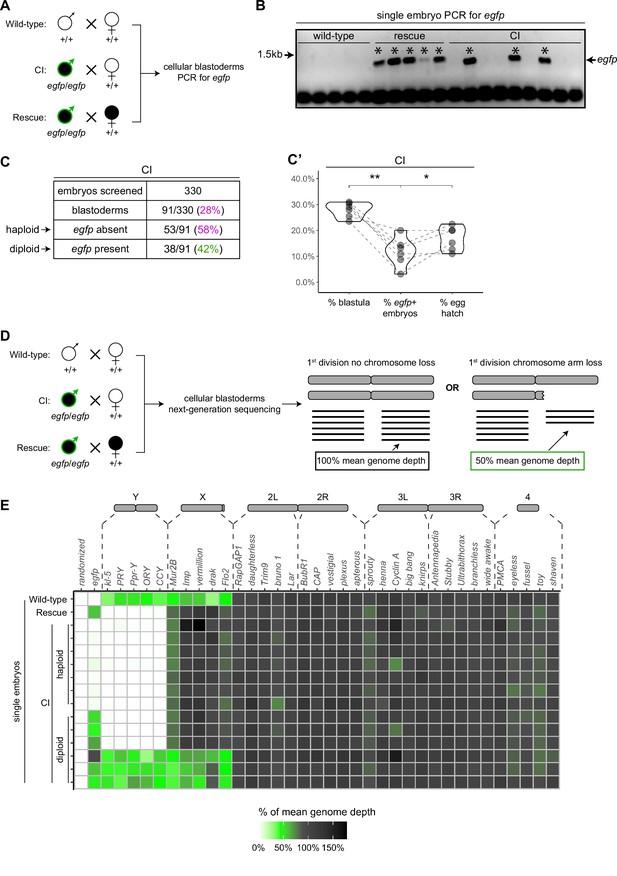
Late-stage cytoplasmic incompatibility (CI)-derived embryos are either haploid (maternal chromosome set) or diploid (both parental chromosome sets).
(A) Embryos were collected from wild-type crosses or CI and rescue crosses in which the father was homozygous for an egfp transgene. Embryos were staged live, and cellular blastoderms were selected for single embryo PCR analysis with primers recognizing egfp. (B) A representative gel showing detection of egfp (asterisks) in all rescue-derived embryos and in a mix of CI-derived embryos. No egfp is detected in the wild-type control. See also Figure 5—source data 1. (C) Summary of the percentage of screened CI-derived cellular blastoderms in which either egfp was absent (haploid) or egfp was present (diploid). N values correspond to the number of embryos scored (biological replicates). (C’) Comparison of the percentages of embryos that had reached at least cycle 10 (% blastula), had detectable egfp bands (% egfp + embryos) and a concomitant egg hatch (% egg hatch). Each dot represents one technical replicate, and lines connect values for the same experiment. The percentage of egfp + embryos (diploids) was more associated with the percentage of eggs that hatched (p = 0.045 by Mann–Whitney test) than with the percentage of blastoderms screened (p = 0.001 by Mann–Whitney test), suggesting haploid embryos do not hatch. (D) Embryos were collected from wild-type crosses or CI and rescue crosses in which the father was homozygous for an egfp transgene. Embryos were staged live, and cellular blastoderms were selected for single embryo sequencing. If chromosome arms were not lost during the first division, the mean depth of reads mapping to the chromosome arm should be near the mean depth of reads mapping to the genome (black box). If chromosome arms were lost during the first division, the mean depth of reads aligning to that chromosome arm should be 50% of the mean depth of reads aligning to the genome (green box). In haploids, maternal chromosome arm loss would result in no reads mapping to that chromosome arm. (E) Sequenced embryos were sorted as haploid or diploid based on the depth of reads mapping to egfp. Each box represents the mean depth of reads aligning to that gene divided by the mean depth of reads aligning to the whole genome (‘mean genome depth’). White = 0% of mean genome depth; green = 50% of mean genome depth; gray = 100% of mean genome depth; black = 150% of mean genome depth. Consistent with no partial chromosome/chromosome arm loss, genes across all chromosomes were present at 100% mean genome depth for both haploids and diploids (or 50% for X-linked genes when embryos are male). See also Figure 5—figure supplement 1. See also Figure 5—source data 2.
-
Figure 5—source data 1
Raw gel images for single embryo PCR assay egfp presence/absence.
- https://cdn.elifesciences.org/articles/81292/elife-81292-fig5-data1-v1.zip
-
Figure 5—source data 2
Normalized depth of reads aligning to coding sequences and egfp in wild-type-, cytoplasmic incompatibility (CI)-, and rescue-derived cellular blastoderms.
Numerical data used to create Figure 5E.
- https://cdn.elifesciences.org/articles/81292/elife-81292-fig5-data2-v1.xlsx
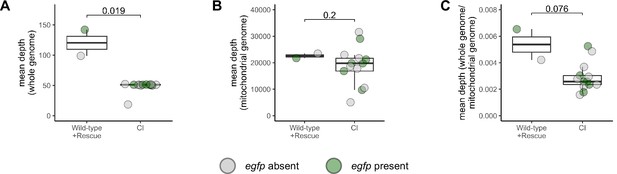
Depth of reads aligning to genome in cytoplasmic incompatibility (CI)-derived cellularized embryos is decreased compared to wild-type- and rescue-derived embryos.
Comparison of the mean depth of reads aligning to the reference genome (A), the mitochondrial genome (B), and the reference genome normalized to the mitochondrial genome (C) in wild-type-, CI-, and rescue-derived embryos. Each dot represents one embryo (biological replicate). Gray dots are embryos in which egfp was not detected (CI = haploids, wild-type = diploid). Green dots represent embryos in which egfp was detected (CI + rescue = diploids). Displayed p values were determined with Mann–Whitney tests. See also Figure 5.
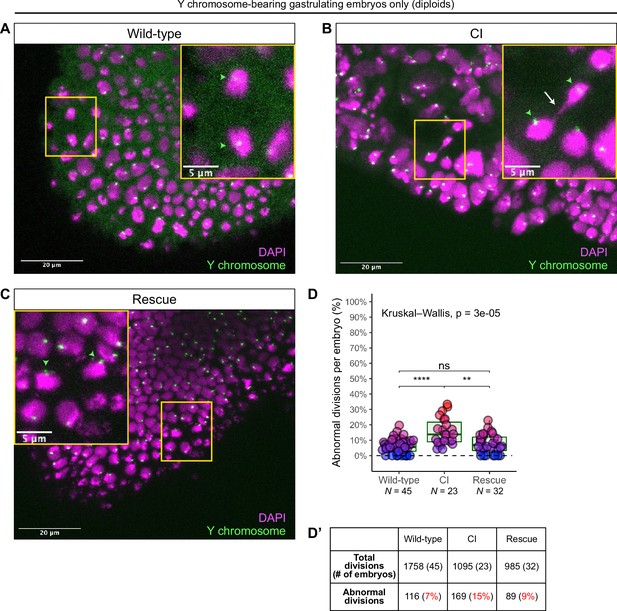
Diploid cytoplasmic incompatibility (CI)-derived gastrulating embryos that have escaped the first division defect exhibit increased chromosome segregation errors.
Gastrulating embryos from wild-type (A), CI (B), and rescue (C) crosses. Embryos are hybridized with probes that specifically recognize the D. simulans Y chromosome (green arrowheads) to select for diploidy (both maternal and paternal chromosome sets present) and counterstained with DAPI (magenta). Yellow boxes indicate zoomed in regions. Scale bars are 20 and 5 μm for unzoomed and zoomed images, respectively. (A) Diploid wild-type-derived embryos exhibit relatively normal chromosome segregation. (B) Diploid CI-derived embryos have elevated rates of bridging and lagging chromosomes (arrow). (C) Diploid rescue-derived embryos exhibit relatively normal chromosome segregation. (D) Comparison of the percentage of chromosome segregation errors observed in diploid wild-type-, CI-, and rescue-derived embryos. Each dot represents one embryo. N values correspond to the number of embryos scored (biological replicates). Asterisks indicate significance by Mann–Whitney tests. Wild-type to CI: p = 6.7 x 10-6; CI to rescue: p = 1.1 x 10-3. ns = 0.26. (D’) Summary of chromosome segregation errors in wild-type-, CI-, and rescue-derived embryos. See also Figure 6—figure supplements 1 and 2.
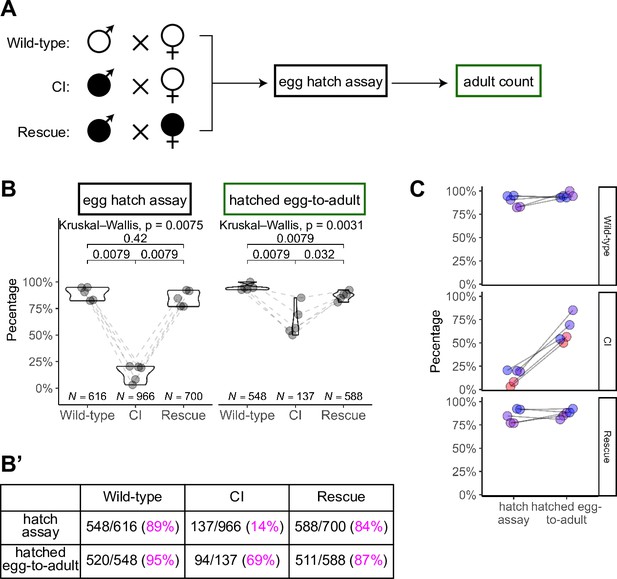
Hatched eggs from cytoplasmic incompatibility (CI) crosses exhibit significantly increased lethality prior to eclosion than those from wild-type or rescue crosses.
(A) Diagram of the ‘egg-to-adult’ experiment. Eggs from egg hatch assays were placed in new collection bottles. Adults were collected from hatched eggs and counted for as long as flies were eclosing. (B, B’) Comparison of egg hatch rates and hatched egg-to-adult rates. Hatched eggs from CI crosses had significantly reduced rates of developing into adults, suggesting the existence of a CI-induced lethal phase during larval development. Maternally supplied Wolbachia rescues this larval lethality. Each dot represents the rates from one collection (biological replicate). Lines connect experiments performed simultaneously. Unless otherwise indicated, p values displayed were determined with Mann–Whitney tests. (C) Comparison of the strengths of early CI defects (as judged by hatching) and late CI defects (as judged by the hatched egg-to-adult rates). In general, higher lethality in hatching correlated to higher lethality during egg-to-adult development. Each dot represents one technical replicate. Lines connect the egg hatch rate and the hatched egg-to-adult rate for each experiment. N values correspond to the total number of embryos/hatched eggs scored. See also Figure 6.
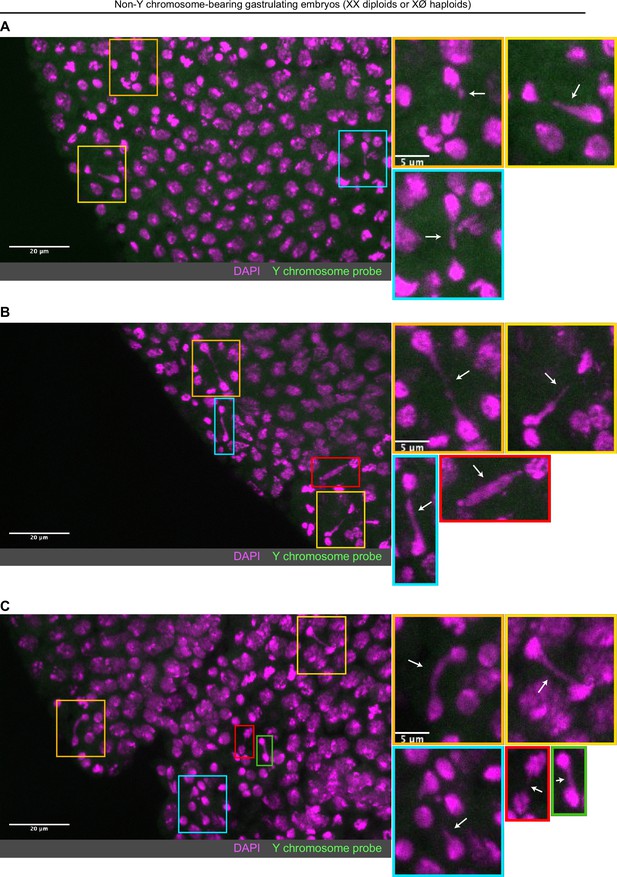
Non-Y chromosome-containing gastrulating embryos exhibit extensive chromosome segregation errorss.
(A–C) Examples of non-Y chromosome-containing gastrulating embryos (haploid or diploid) with extensive chromosome segregation errors. No probes targeting the Y chromosome (green) were detected in these embryos. Embryos were counterstained with DAPI (magenta). The extent of errors observed in these embryos is not observed in Y chromosome-containing diploid embryos, suggesting these embryos may be haploid and more strongly experience the second set of cytoplasmic incompatibility (CI)-induced defects. Boxes indicate zoomed in regions. Scale bars are 20 and 5 μm for unzoomed and zoomed images, respectively. See also Figure 6.
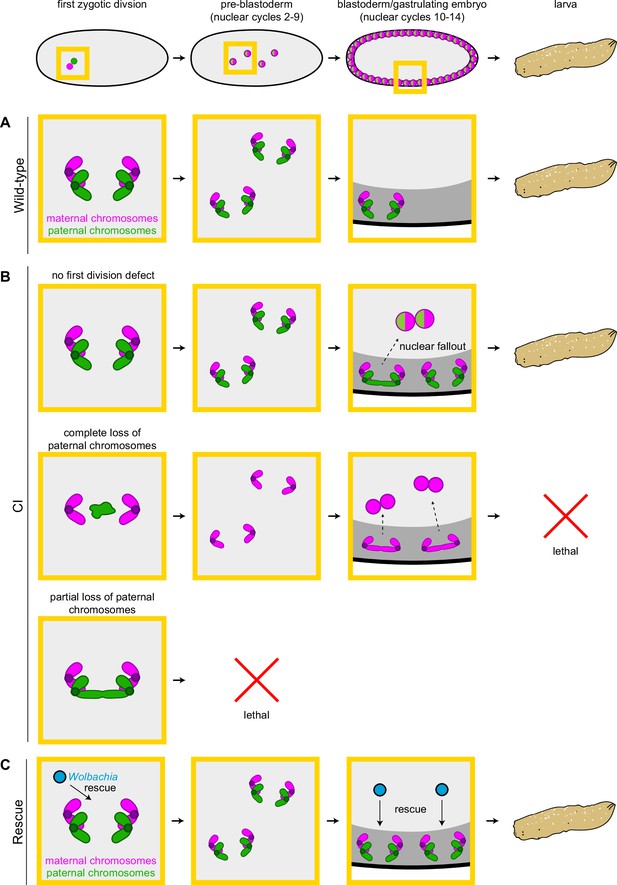
Cytoplasmic incompatibility (CI) induces independent first division and mid-blastula transition chromosome segregation errors.
(A) During the first zygotic division in wild-type derived embryos, paternal (green) and maternal (magenta) chromosomes segregate normally. Chromosome segregation occurs normally during pre-blastoderm, blastula, and post-cellularization divisions. Embryos hatch. (B, top row) In CI-derived embryos, if there are no segregation defects during the first division, embryos develop as diploids containing full maternal and paternal chromosome sets. Pre-blastoderm divisions proceed normally. However, during blastoderm stages, CI induces a second set of defects, which cause chromosome segregation errors and subsequent nuclear fallout (dashed arrow). Chromosome segregation errors continue during gastrulation. These defects occur at moderate frequencies and embryos hatch. (B, middle row) If the paternal chromosomes are completely excluded during the first division, embryos develop as haploids from only the maternal chromosome set. Pre-blastoderm divisions proceed normally, followed by increased chromosome segregation errors and nuclear fallout during blastoderm divisions. Chromosome segregation errors continue during gastrulation. Perhaps due to CI being strong in haploid embryos (Bonneau et al., 2018), this second set of CI-induced defects is more severe, and embryos fail to hatch, due to their haploidy. (B, bottom row) If the paternal chromosomes are partially lost during the first divisions, embryos arrest due to severe aneuploidy. (C) Maternally supplied Wolbachia (blue circles) rescue both the first division defects and the late-stage defects independently.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Wolbachia riverside) | Wolbachia (wRi) | O’Neill and Karr, 1990 | ||
| Genetic reagent (Drosophila simulans) | Infected | Serbus et al., 2015 | Infected with wRi | |
| Genetic reagent (Drosophila simulans) | Uninfected | Casper-Lindley et al., 2011 | Generated by tetracycline-curing wRi-infected stocks | |
| Genetic reagent (Drosophila simulans) | egfp uninfected | Holtzman et al., 2010 | NDSS: 14021-0251.275 | Obtained from the National Drosophila Species Stock Center (NDSS) |
| Genetic reagent (Drosophila simulans) | egfp Wolbachia-infected | This paper | Generated by crossing Wolbachia from infected stock into the egfp uninfected stock | |
| Sequence-based reagent | egfp 5′ | Cruachem | PCR primers | ATCAAGCTTGTGA GCAAGGGCGAGGAGC |
| Sequence-based reagent | egfp 3′ | Cruachem | PCR primers | ACCTCGAGCTACTT GTACAGCTCGTCCATGC |
| Sequence-based reagent | Y-chr-488 | Ferree and Barbash, 2009 | FISH probe | (AAT-AAA-C)4 conjugated to Alexa488, synthesized by Integrated DNA technologies |
| Peptide, recombinant protein | Rhodamine-labeled histone | This paper | Recombinant rhodamine-histone fusion protein | |
| Antibody | Anti-centrosomin (Rabbit polyclonal) | Megraw et al., 1999 | (1:200) | |
| Antibody | Anti-rabbit-488 (Goat polyclonal) | Thermo Fisher | Cat# A-11008 | (1:1000) |
| Gene (Drosophila simulans) | D. simulans reference genome | Clark et al., 2007 | WUGSC mosaic 1.0/droSim1 assembly | UCSC Genome Browser |
| Gene (Wolbachia riverside) | wRi reference genome | Klasson et al., 2009 | CP001391.1 | GenBank |
| Gene (p-egfp plasmid) | egfp | Addgene | ||
| Software, algorithm | BWA-MEM2 | Md et al., 2019 | Version 2.2.1 | |
| Software, algorithm | Picard tools | Broad Institute | Version 2.27.1 | |
| Software, algorithm | BEDTools | Quinlan and Hall, 2010 | Version 2.26.0 | |
| Software, algorithm | Leica Application Suite Advanced Fluorescence | Leica | ||
| Software, algorithm | R | R Core Team | Version 4.0.5 | |
| Software, algorithm | ggplot2 | Wickham, 2016 | ||
| Other | DAPI in Vectashield | Vector Laboratories | Cat# H-1200 | DNA stain (1.5 µg/ml) in mounting media |
| Other | Halocarbon oil | Millipore Sigma | Cat# H8898 | For mounting embryos for live imaging |





