Cellular and molecular dynamics in the lungs of neonatal and juvenile mice in response to E. coli
Figures
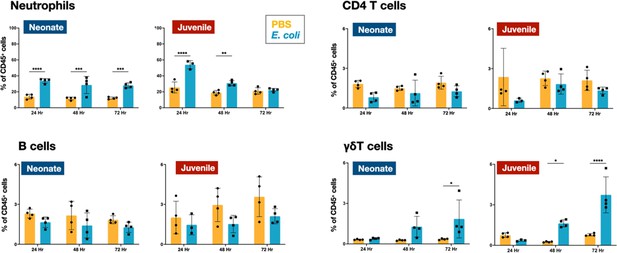
E. coli-induced changes in neutrophil and lymphocyte dynamics in neonatal and juvenile lungs.
Changes in the dynamics of Ly6g+ neutrophils, CD4+ T cells, B220+ B cells, and γδ+ T cells in neonatal and juvenile lungs at 24, 48, or 72 hr, post-E. coli challenge (PEC) as assessed by flow cytometry (see Figure 3—figure supplement 1 for gating) and expressed as the percentage of total CD45+ cells. Statistical differences were determined using two-way analysis of variance (ANOVA) with multiple comparisons. Error bars represent standard deviation of the mean. ns – not significant; *p < 0.05; **p < 0.005; ***p = 0.0001; ****p < 0.0001 (n = 4 group).
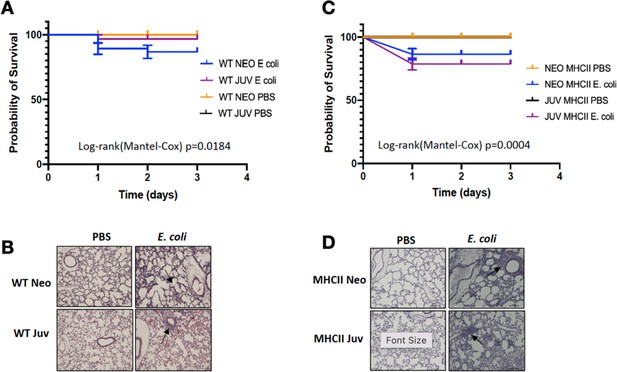
Survival analysis of WT and MHCII−/− neonatal and juvenile mice after challenge with E. coli.
(A) Mortality of WT neonates and juveniles at 24, 48, and 72 hr post-E. coli challenge with 2.4 × 106 CFUs or phosphate-buffered saline (PBS) treatment. Comparison of survival curves were determined using log-rank (Mantel–Cox) test (n = 24–47/group). (B) Representative examples of the histological appearance of the lungs from WT neonates and juveniles at 48 hr post-E. coli challenge or control PBS treatment (arrows point to inflammatory foci). (C) Mortality of MHCII−/− neonates and juveniles at 24, 48, and 72 hr post-E. coli challenge with 2.4 × 106 CFUs or PBS treatment. Comparison of survival curves were determined using log-rank (Mantel–Cox) test (n = 12–37/group). (D) Representative examples of the histological appearance of the lungs from MHCII−/− neonates and juveniles at 48 hr post-E. coli challenge or control PBS treatment (arrows point to inflammatory foci).
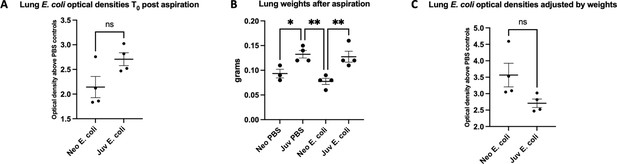
Similar bacterial burden between neonatal and juvenile mice at Time0 post-E. coli aspiration.
(A) Optical density at 600 nM wavelengths (OD600) from E. coli and phosphate-buffered saline (PBS)-treated mice at T0. (B) Lungs weights of neonatal and juvenile post-E. coli or PBS aspiration at T0 (*p < 0.5, **p < 0.007). (C) OD600 absorbance normalized for lung weights (n = 4 for each condition).
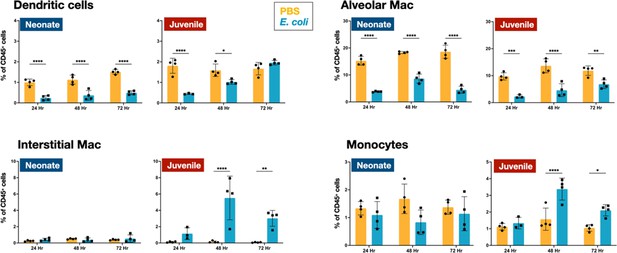
E. coli-induced changes in mononuclear cell dynamics in neonatal and juvenile lungs.
Changes in the dynamics of CD11c+SiglecF−F4/80−MHCIIhi dendritic cells, CD11c+F4/80+SiglecF+ alveolar macrophages, CD11b+F4/80+SiglecF− interstitial macrophages, and CD11b+Ly6c+MHCII+ monocytes in neonatal and juvenile lungs at 24, 48, or 72 hr, post-E. coli challenge (PEC) as assessed by flow cytometry (see Figure 3—figure supplement 1 for gating) and expressed as the percentage of total CD45+ cells. Statistical differences were determined using two-way analysis of variance (ANOVA) with multiple comparisons. Error bars represent standard deviation of the mean. ns – not significant; *p < 0.05; **p < 0.005; ***p = 0.0001; ****p < 0.0001 (n = 3 or 4/group).
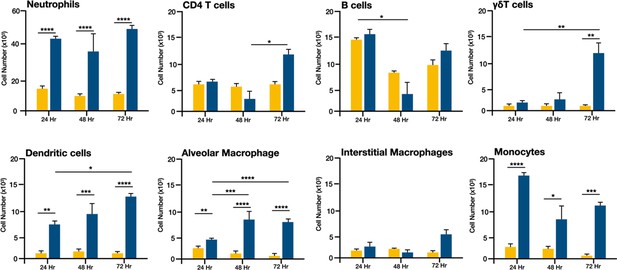
E. coli-induced changes in the number of immune cells in lungs from neonatal mice.
Changes in the numbers of Ly6g+ neutrophils, CD4+ T cells, B220+ B cells, γδ+ T cells, CD11c+SiglecF−F4/80−MHCIIhi dendritic cells, CD11c+F4/80+SiglecF+ alveolar macrophages, CD11b+F4/80+SiglecF− interstitial macrophages, and CD11b+Ly6c+MHCII+ monocytes in neonatal (PND 3–5) lungs at 24, 48, or 72 hr post-E. coli challenge (PEC) as assessed by flow cytometry (see Figure 1—figure supplement 1 for gating) and expressed as cell number (×103). Statistical differences were determined using two-way analysis of variance (ANOVA) with multiple comparisons. Error bars represent standard deviation of the mean. *p < 0.05; **p < 0.005; ***p = 0.0001; ****p < 0.0001 (n = 4/time point).
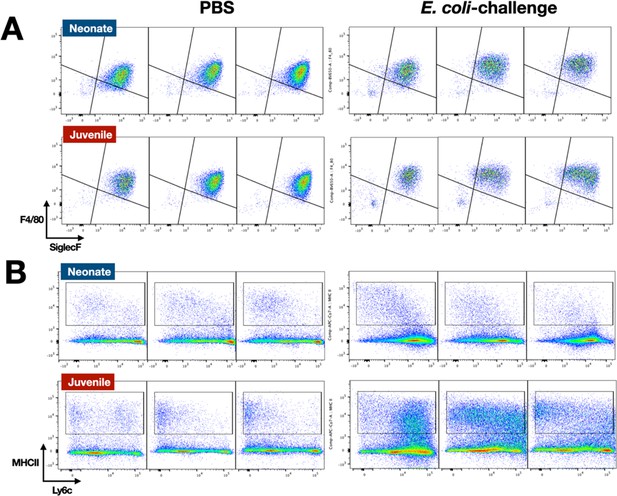
E. coli-induced changes in the surface phenotypes of alveolar macrophage and interstitial macrophage compartments.
(A) A representative flow cytometry analysis of F4/80+SiglecF+ alveolar macrophages from E. coli-challenged or phosphate-buffered saline (PBS)-treated neonatal and juvenile lungs at 24, 48, and 72 hr (see Figure 3—figure supplement 1 for gating). (B) A representative flow cytometry analysis of the CD11b+MHCII+Ly6cvariable interstitial macrophage compartment from E. coli-challenged or PBS-treated neonatal and juvenile lungs at 24, 48, and 72 hr (see Figure 3—figure supplement 1 for gating).
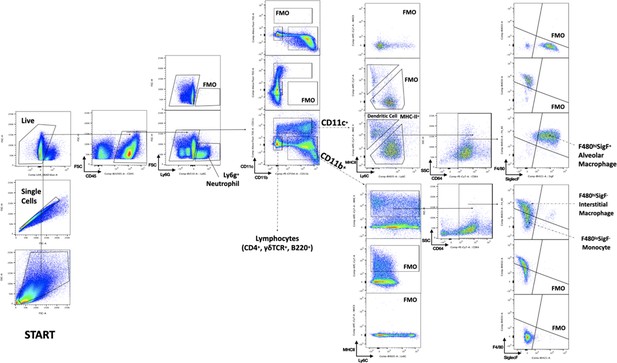
Gating strategy.
The gating strategy is illustrated using a juvenile 72 hr phosphate-buffered saline (PBS)-treated sample. Neutrophils were identified first among total CD45+ cells by Ly6g expression. Remaining cells were segregated by expression of CD11c or CD11b. Populations in the CD11c+ compartment include MHC-IIhi dendritic cells and F480+SiglecF+ alveolar macrophages. Cells within the CD11b+ compartment include two F480loSiglecF− monocyte populations with variable MHC-II expression, and MHC-IIhiF480hiSiglecF− interstitial macrophages.
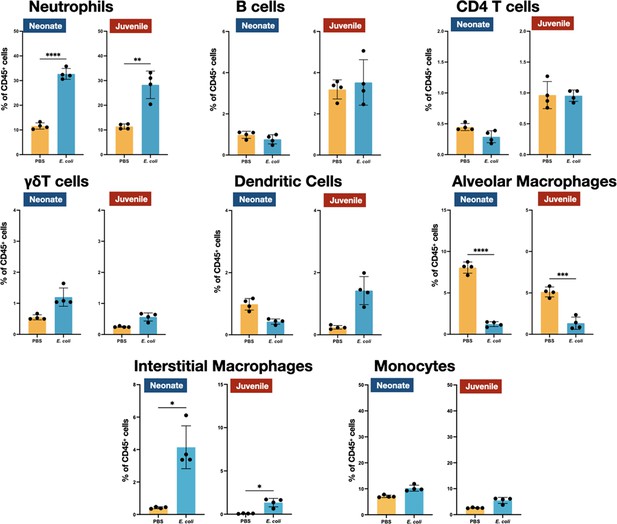
E. coli-induced changes in the proportions of neutrophils, lymphocytes, and mononuclear cells in the lungs of MHC−/− neonates and juveniles at 48 hr post-challenge with E. coli.
Changes in neutrophils, CD4 T cells, B cells, γδT cells, dendritic cells, alveolar macrophages, interstitial macrophages, and monocytes in neonatal and juvenile lungs at 48 hr post-E. coli challenge (PEC) as assessed by flow cytometry (see Figure 3—figure supplement 1 for gating) and expressed as the percentage of total CD45+ cells. Statistical differences were determined using two-way analysis of variance (ANOVA) with multiple comparisons. Error bars represent standard deviation of the mean. ns – not significant; *p < 0.05; **p < 0.005; ***p = 0.0001; ****p < 0.0001 (n = 3 or 4/group).
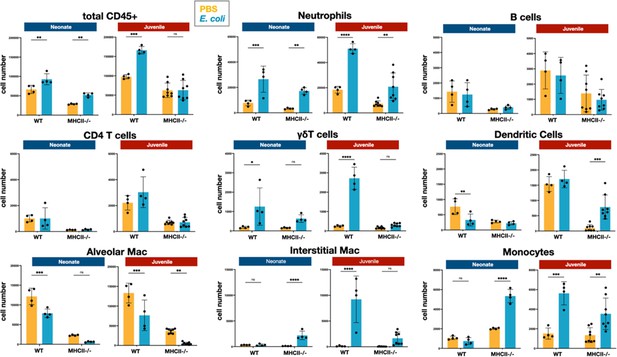
E. coli-induced changes in the numbers of neutrophils, lymphocytes, and mononuclear cells in the lungs of WT and MHC−/− neonates and juveniles at 48 hr post-challenge with E. coli.
Changes in the numbers of total CD45+ cells, neutrophils, CD4 T cells, B cells, γδT cells, dendritic cells, alveolar macrophages, interstitial macrophages, and monocytes in neonatal and juvenile lungs at 48 hr post-E. coli challenge (PEC) as assessed by flow cytometry (see Figure 3—figure supplement 1 for gating). Statistical differences were determined using two-way analysis of variance (ANOVA) with Holm–Šídák post hoc test for multiple comparisons. Error bars represent standard deviation of the mean. ns – not significant; *p < 0.05; **p < 0.005; ***p = 0.0001; ****p < 0.0001 (n = 4–8/group).
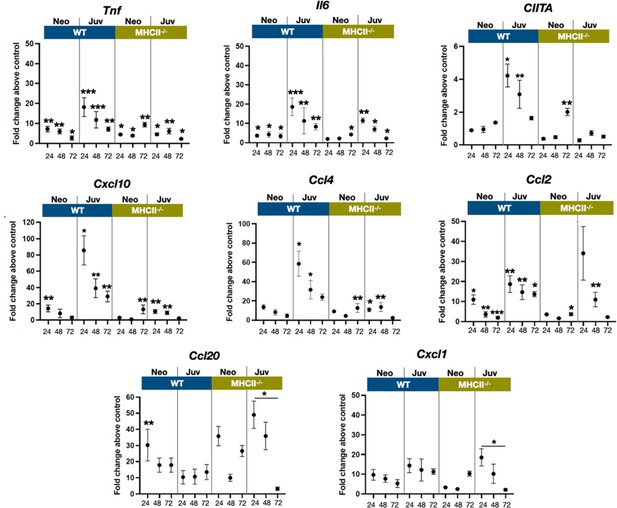
E. coli-induced changes in the expression dynamics of select regulatory, cytokine, and chemokine genes in WT and MHC−/− neonatal and juvenile lungs.
Quantitative RT-PCR analysis of total RNA isolated from the lungs of WT and MHCII−/− neonates and juveniles at 24, 48, and 72 hr post-E. coli challenge for changes in expression of class II major histocompatibility complex, transactivator (Ciita), tumor necrosis factor (Tnf), interleukin 6 (Il6), C-X-C motif chemokine ligand 10 (Cxcl10), C-C motif chemokine ligand 4 (Ccl4), C-C motif chemokine ligand 2 (Ccl2), C-C motif chemokine ligand 20 (Ccl20), and C-X-C motif chemokine ligand 1 (Cxcl1). Data present as fold change over the value for the corresponding age/genetic background phosphate-buffered saline (PBS) controls. Points represent means of the values from 3 to 5 animals per group (error bars represent standard error of the mean). Statistical significance between groups were determined using one-way analysis of variance (ANOVA) with multiple comparisons. *p < 0.01, **p < 0.004, ***p < 0.001.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Mus musculus) | B6;129S2-H2dlAb1-Ea/J | Jackson Laboratory | Strain #003584 | |
| Antibody | B220/AF488 (Rat monoclonal) | Biolegend | Cat# 103228 | IgG2a (clone RA3-6B2) used at 1/100 |
| Antibody | CD25/PE (Rat polyclonal) | BD | Cat# 561065 | IgG1 used at 1/50 |
| Antibody | CD64/PEcy7 (Mouse monoclonal) | Biolegend | Cat# 139314 | X54-5/7 used at 1/100 |
| Antibody | CD11b/PE-CF594 (Rat monoclonal) | BD | Cat# 562317 | M1/70 IgG2b used at 1/50 |
| Antibody | CD4/BB700 (Rat monoclonal) | BD | Cat# 566408 | RM4-5 IgG2a used at 1/50 |
| Antibody | CD11c/APC-R700 (Hamster monoclonal) | BD | Cat# 565872 | N418 IgG2 used at 1/40 |
| Antibody | PD-1/APC-AF647 (Rat monoclonal) | Biolegend | Cat# 135209 | 29F-1A12 IgG2a used at 1/30 |
| Antibody | MHCII/APC-Cy7 (Rat monoclonal) | Biolegend | Cat# 107627 | M5/114.15.2 IgG2b used at 1/50 |
| Antibody | SigF/BV421 (Rat monoclonal) | BD | Cat# 562681 | E50-2440 IgG2a used at 1/50 |
| Antibody | Ly6G/BV510 (Rat monoclonal) | BD | Cat# 740157 | 1A8 IgG2a used at 1/50 |
| Antibody | Ly6C/BV605 (Rat monoclonal) | BD | Cat# 128036 | HK1.4 IgG2c used at 1/100 |
| Antibody | F4/80/BV650 (Rat monoclonal) | BD | Cat# 743282 | T45.2342 IgG2a used at 1/20 |
| Antibody | CD3/BV711 (Hamster monoclonal) | BD | Cat# 563123 | 145-2C11 IgG1 used at 1/40 |
| Antibody | ITCRgd/BV786/SB780 (Hamster monoclonal) | Invitrogen | Cat# 78-5711-82 | GL3 IgG used at 1/30 |
| Antibody | Cd45/BUV395 (Rat monoclonal) | BD | Cat# 564279 | 30-F11 IgG2b used at 1/50 |
| Antibody | CD8/BUV737 (Rat monoclonal) | BD | Cat# 564297 | 5306.7 IgG2a used at 1/40 |





